Introduction
Previous studies have suggested that alteration of
oxidative stress and thiol-redox are initiating and propagating
forces in the pathogenesis of bleomycin (BLM)-induced pulmonary
fibrosis (1,2). BLM is an effective chemotherapeutic
agent used primarily in the treatment of lymphoma, squamous cell
carcinoma, testicular cancer and malignant pleural effusion
(3). The antineoplastic effect of
BLM is hypothesized to be due to the formation of a BLM-oxygen
complex, which binds to the DNA and cleaves the
phosphodiester-deoxyribose backbone (4). However, the effective clinical use of
BLM in chemotherapy is limited as a result of the development of
dose- and time-dependent interstitial pneumonitis, which eventually
progresses to interstitial pulmonary fibrosis (5,6).
Oxidative stress in the cell is caused by an
imbalance between the formation of free radicals and their removal
by enzymatic and non-enzymatic anti-oxidant molecules (7). It was hypothesized that redox cycling
of the iron-BLM complex catalyzes the formation of reactive oxygen
species (ROS), which leads to an imbalance between ROS and the
available anti-oxidant defenses, principally glutathione (8). The generated ROS damage lung cells
and activate inflammatory cells, which accumulate in the lower
airways and produce further ROS, leading to oxidative stress. The
nuclear transcription factor, NF-E2-related transcription factor 2
(Nrf2), is a major sensor of oxidative stress in the cell (9). Under oxidative stress, Nrf2 is
phosphorylated and translocated into the nucleus, where it
activates the transcription of anti-oxidant and detoxifying genes
by binding to the anti-oxidant response elements (AREs) in their
regulatory regions (9).
A previous study investigating Kruppel-like factor 9
(Klf9) genes binding to the sequences of transcription factors in
order to regulate the oxidative stress response, suggested that
Klf9 promoter regions contained several conserved AREs (10), which are binding sites for Nrf2, a
major regulator of anti-oxidant defense in the cell (9,11).
Klf9 is a ubiquitously expressed member of the Sp1 C2H2-type zinc
finger family of transcription factors (12). The expression of Klf9 can be
increased by several stress-inducing agents, including the
proteasomal inhibitor, bortezomib, and the histone deacetylase
inhibitor, panobinostat, and Klf9 in turn mediates their
cytotoxicity (13). Therefore, the
present study aimed to investigate whether Klf9 was involved in the
pathogenesis of BLM-induced pulmonary toxicity in human pulmonary
fibroblasts (HPF), which was also analyzed in the Klf9 knockout
mouse model.
Materials and methods
Cell lines and reagents
HPF cells, isolated from human lung tissue, were
obtained from Promocell (Heidelberg, Germany). The cells were
cultured in Dulbecco's modified Eagle's medium (Sigma-Aldrich, St.
Louis, MO, USA), supplemented with 10% fetal calf serum (Gibco-BRL,
Gaithersburg, MD, USA), 2 mM glutamine (Gibco-BRL), and
penicillin-streptomycin antibiotics (Gibco-BRL). Hydrogen peroxide,
sulforaphane, BLM, 2′-7′-dichlorofluorescin diacetate (DCFH-DA),
actinomycin D and cycloheximide were purchased from
Sigma-Aldrich.
Determination of cell viability and
measurement of ROS
The HPF cells were seeded in a 96-well tissue
culture plate, at a density of ~1.25×104 cells/well, for
24 h. The media was discarded and the cells were treated with
various concentrations (25, 50, 100 or 200 µM) of BLM in
serum-free media for 24 h. The levels of intracellular ROS
generation were measured using a well-characterized probe, DCFH-DA.
DCFH-DA is hydrolyzed by esterases to DCFH, which is trapped within
the cell. This non-fluorescent molecule is subsequently oxidized to
fluorescent DCF by the action of cellular oxidants. The cells
treated with either BLM or standard media for 6 h were subjected to
DCFH staining and the fluorescence was determined at 485 nm
excitation and 520 nm emission, using a microplate reader
(FLUOstar; BMG Labtech, Inc., Durham, NC, USA).
Protein expression levels of
antioxidants
The harvested control and BLM-treated (50 or 100
µM) HPF cells were washed with ice-cold phosphate-buffered
saline and were lysed with radioimmunoprecipitation buffer (Cell
Signaling Technology, Inc., Danvers, MA, USA). The lysates were
centrifuged at 10,000 × g for 10 min at 4°C, and the protein
content of the supernatant was determined by the Bradford method
(14). The proteins (10–20
µg) were loaded and separated by 10% SDS-PAGE and were
blotted onto a polyvinylidene fluoride membrane (Thermo Fisher
Scientific, Inc., Frederick, MD, USA). The membranes were blocked
with 5% non-fat dry milk powder solution for 1 h at room
temperature prior to overnight incubation with the following
primary antibodies: Catalase (CAT; rabbit polyclonal; cat. no.
ab87529; 1:500), glutathione reductase (GR; rabbit polyclonal; cat.
no. ab137513; 1:500), super oxide dismutase (SOD; rabbit
polyclonal; cat. no. ab16831; 1:2,000) and thioredoxin reductase 2
(TR-2; rabbit polyclonal; cat. no. ab71262; 1:500), all purchased
from Abcam (Cambridge, MA, USA), and Nrf2 (rabbit polyclonal; cat.
no. sc-722; 1:500) and Klf9 (rabbit polyclonal; cat. no. sc-28195;
1:500) were purchased from Santa Cruz Biotechnology, Inc. (Santa
Cruz, CA, USA) at 4°C. Following rinsing of the membranes with
Tris-buffered saline with Tween® 20, (Sigma-Aldrich),
they were incubated with secondary antibody (goat anti-rabbit;
polyclonal; cat. no. ab97200; 1:2,000; Abcam) for 1 h at room
temperature and the bands were made visible by enhanced
chemiluminescence using the SuperSignal West Femto substrate
(Thermo Fisher Scientific, Inc.), visualized using the Alpha
Innotech FluorChem HD2 Imaging system (Alpha Innotech Co., San
Leandro, CA, USA). All antibodies, with the exception of α-tubulin
(rabbit polyclonal; cat. no. ab4074; 1:1,000; Abcam), were used at
a dilution of 1:250, and the α-tubulin antibody was used at a
dilution of 1:1,000.
Gene expression analysis
Reverse transcription-quantitative polymerase chain
reaction (RT-qPCR) was performed to analyze the gene expression
status using gene-specific primers with the following sequences:
Forward: 5′-ACAGTGGCTGTGGGAAAGTC-3′ and reverse:
5′-AACTGCTTTTCCCCAGTGTG-3′ for KLF9; and forward:
5′-AGTGGATCTGCCAACTACTC-3′ and reverse:
5′-CATCTACAAACGGGAATGTCTG-3′ for Nrf2. The total RNA was extracted
from retinal cells using the RNA Nanoprep kit (Agilent
Technologies, Santa Clara, CA, USA) and was reverse transcribed
using the Reverse Transcription system (Promega Corporation,
Madison, WI, USA) to synthesize cDNA. RT-qPCR was performed on the
Rotor-Gene Q Cycler (Qiagen, Valencia, CA, USA) using the SYBR
GREEN PCR Master mix (Qiagen). The data were analyzed using the
2−ΔΔCT method and amplification was performed for 2 min
at 50°C and 10 min at 95°C, followed by 40 cycles of 95°C for 10
sec, 60°C for 10 sec, and 72°C for 30 sec. For each gene, the
relative expression levels were calculated by comparison with a
standard curve, following normalization to the expression of the
housekeeping gene β-actin selected as a control.
Animal experiment
The animal experiments were performed, according to
the ethical guidelines and were approved by the animal Ethical
Committee of The First Hospital of Jilin University (Jilin, China).
Thirty female KLF9 knockout mice and age-matched wild-type female
mice (C57BL/6J; 14–16 weeks-old) were purchased from the Chinese
Academy of Medical Science (Beijing, China). They were anesthetized
and subjected to intratracheal instillation of saline or 2.5 mg/kg
BLM sulfate saline solution (Sigma-Aldrich). The mice were
euthanized via cervical dislocation 10 days following instillation
and the content of 8-oxo-20-deoxy-guanosine (8-OHdG), a marker of
oxidative DNA damage, was determined in the DNA isolated from the
lungs of the treated animals.
Histopathological examination
At 19 days following intratracheal administration of
BLM, the mice were euthanized and the lungs were inflated and fixed
with 4% paraformaldehyde. The lungs were surgically resected and
embedded in paraffin wax. Lung sections (5 mm) were cut and stained
with hematoxylin and eosin. The stained sections were examined with
a light microscope (Eclipse E100-LED; Nikon, Tokyo, Japan).
Statistical analysis
Data are presented as the mean ± standard deviation
of triplicate experiments. Statistical significance was determined
by one-way analysis of variance, followed by Tukey's multiple
comparison tests. Analyses were conducted using Prism 5.0 (GraphPad
Software, San Diego, CA, USA). P<0.05 was considered to indicate
a statistically significant difference.
Results
Effects of BLM treatment
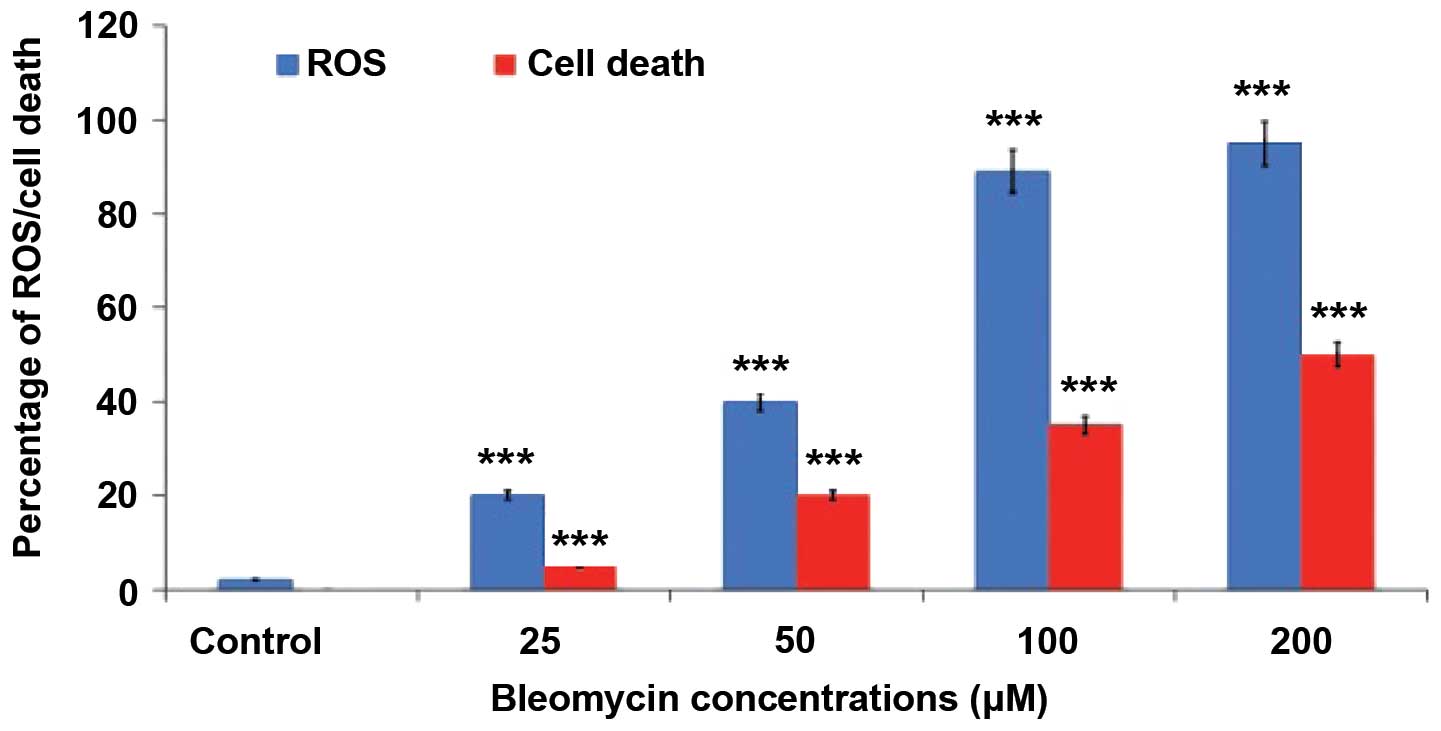
A dose-dependent increase in the production of ROS
and cell death were observed in BLM-treated cells compared with the
normal untreated cells (Fig. 1).
The present study next investigated the levels of the anti-oxidant
proteins, CAT, SOD, GR and TR-2, at 24 h in BLM-treated and control
HPFs. The protein expression levels of Nrf2 and Klf9 were markedly
increased at concentrations of BLM >50 µM (Fig. 2). However, with the exception of
Nrf2 and Klf9, all other antioxidant proteins demonstrated a
dose-dependent decrease in the expression levels (Fig. 2). To understand the mechanisms of
the Klf9-mediated response to oxidative stress, the gene expression
levels of Klf9 and Nrf2 were determined in the BLM treated cells.
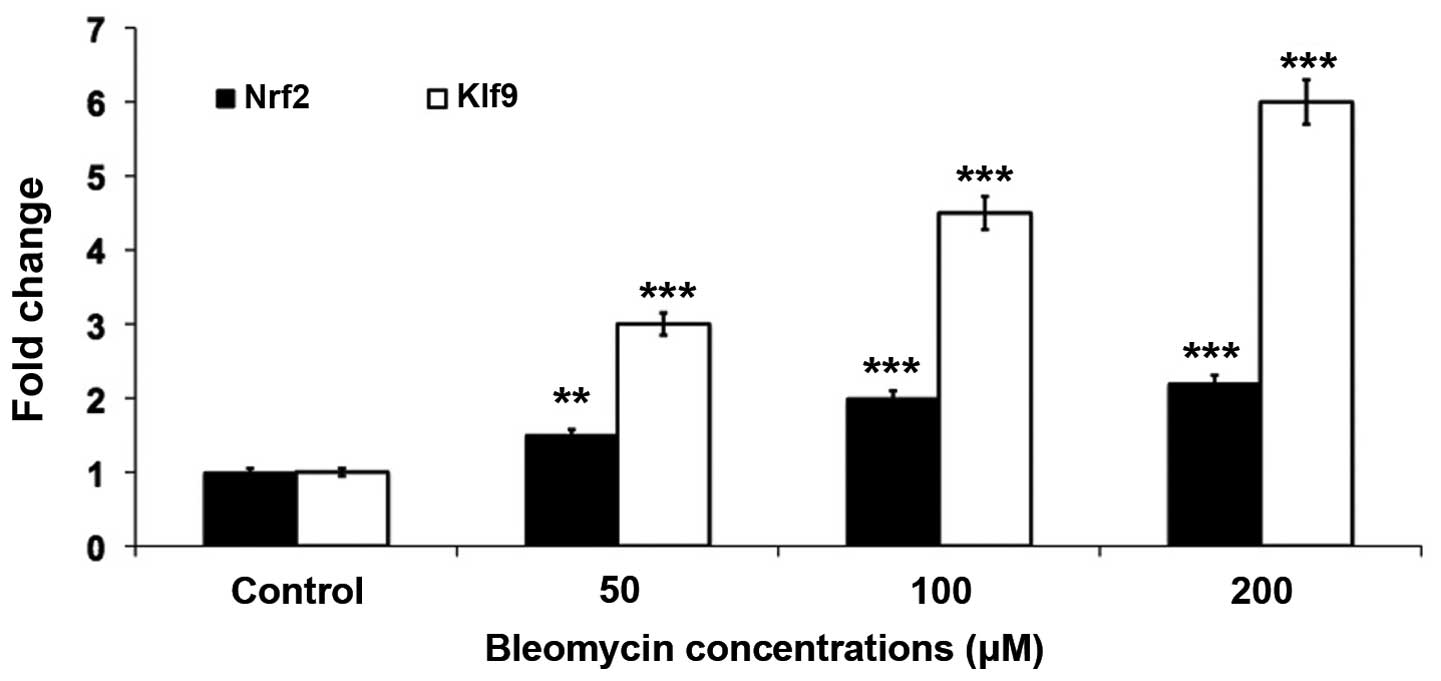
Similarly, gene expression analysis by RT-qPCR revealed that
oxidative stress induced the mRNA expression levels of Nrf2 and
Klf9 (Fig. 3). It was revealed
that a 4- and 6-fold increase in the gene expression levels of Nrf2
and Klf9, respectively, occurred in the BLM-treated cells.
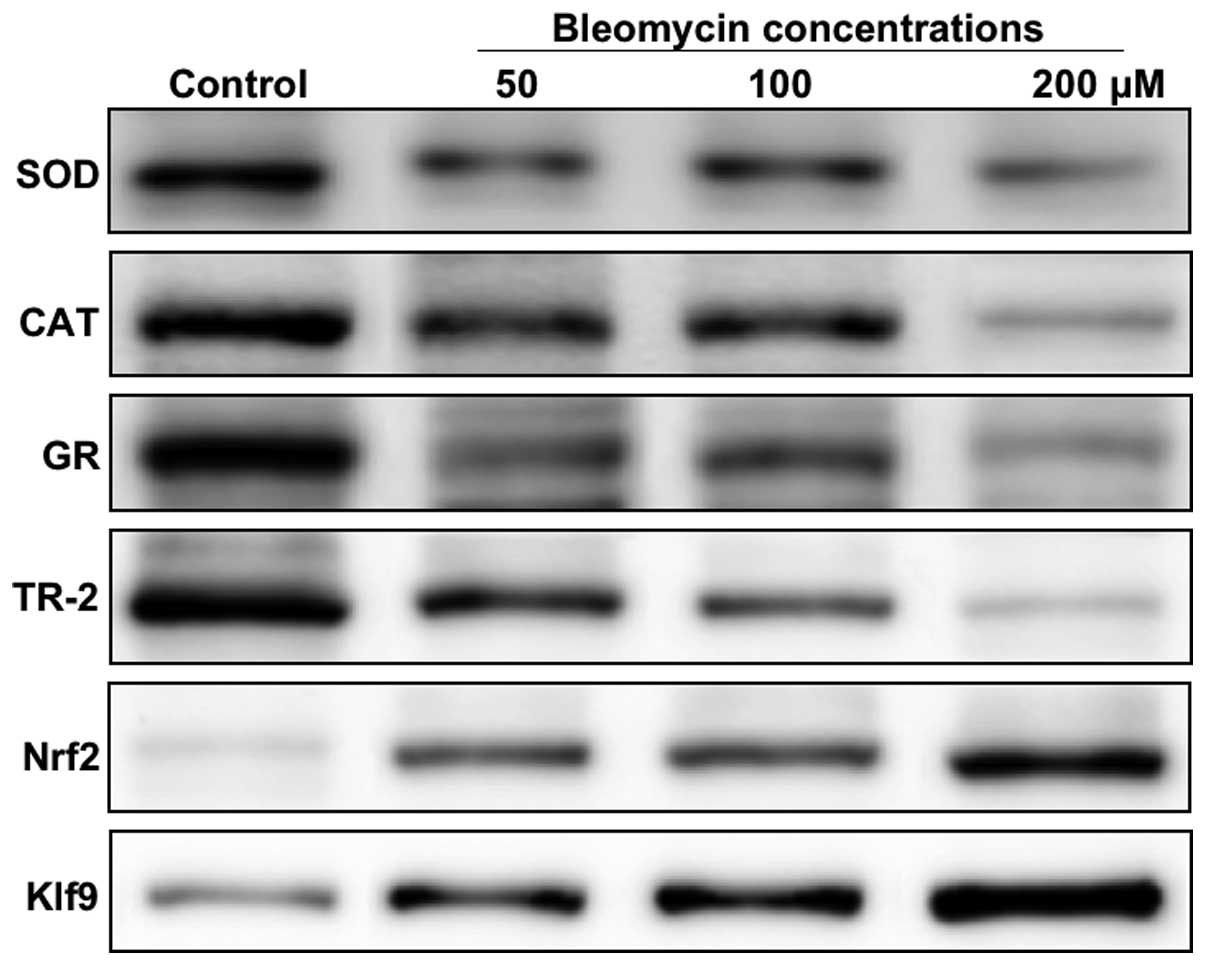 | Figure 2Western blotting was performed to
determine the protein expression levels of antioxidant proteins,
including SOD, CAT, GR, TR-2, Nrf2 and Klf9, in HPF cells treated
with different concentrations (50, 100 or 200 µM) of
bleomycin. SOD, superoxide dismutase; CAT, catalase; GR,
glutathione reductase, TR-2, thioredoxine reductase 2; Nrf2,
NF-E2-related transcription factor 2; Klf9, Kruppel-like factor
9. |
Effect of Klf9 deficiency on DNA damage
and lung fibrosis
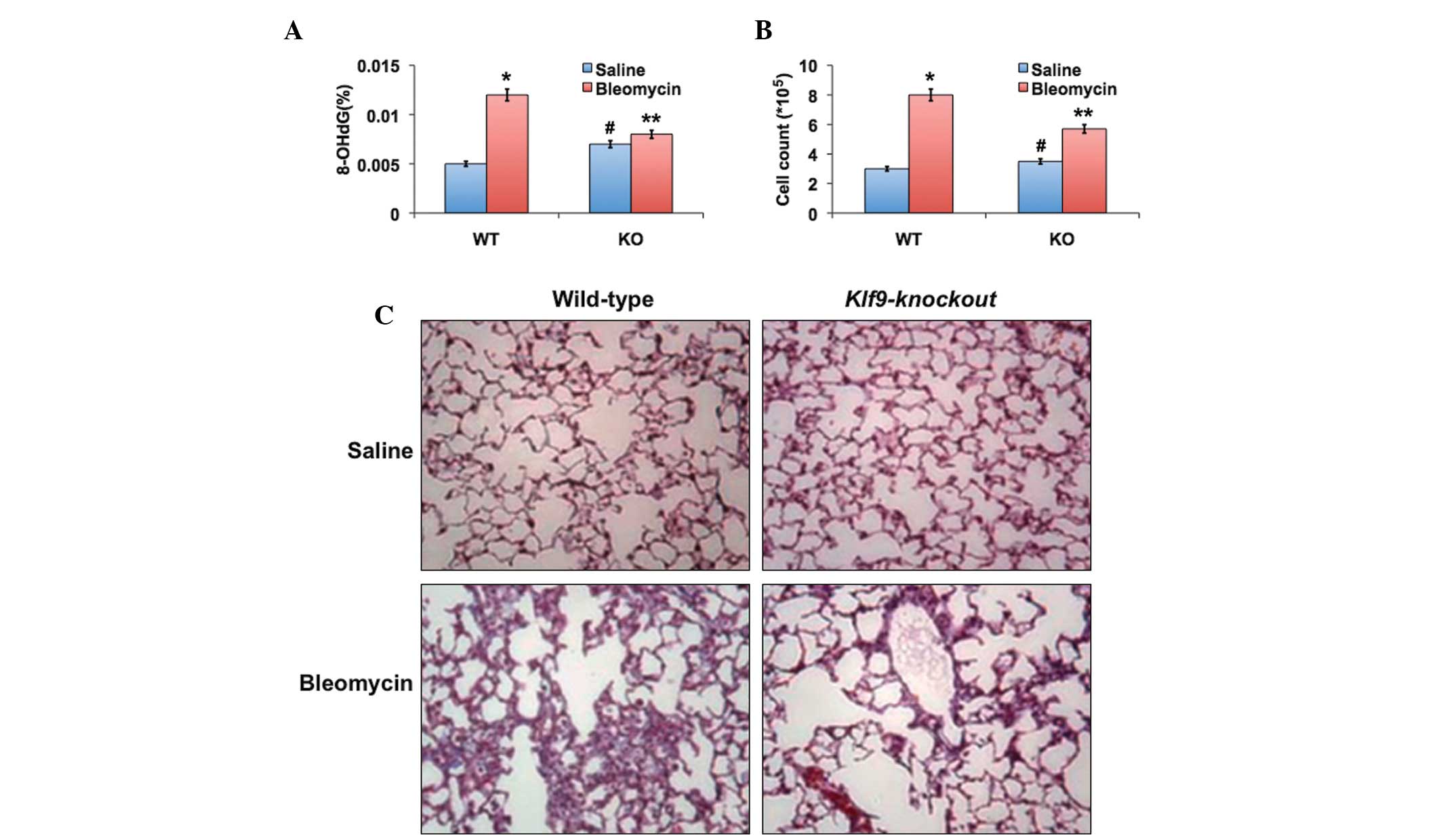
BLM treatment induced a statistically significant
increase in the levels of 8-OHdG, a marker of oxidative DNA damage,
in the lungs of wild-type mice (P=0.0003); however, it revealed no
significant effect on the levels of 8-OHdG in the lungs of Klf9
knockout mice (Fig. 4A). A second
cohort of saline- or BLM-treated mice were analyzed 19 days
following treatment for the accumulation of cells in the
bronchoalveolar lavage fluid (BALF) and for the accumulation of
collagen, each of which are markers of lung fibrosis. A significant
increase in the number of total BALF cells was observed in all
animals treated with BLM (Fig.
4B). However, this increase was significantly lower in the Klf9
knockout mice compared with the wild-type counterparts (P<0.05).
Fibrosis was observed in the histological lung sections stained
with hematoxylin and eosin (Fig.
4C). No sign of fibrosis was detected in the wild-type or Klf9
knockout mice treated with saline solution. Treatment with BLM led
to pulmonary fibrosis in the wild-type and knockout animals,
although the degree of fibrosis was significantly higher in the
wild-type mice compared with the knockout counterparts (P=0.038).
Therefore, Klf9 deficiency provided resistance to BLM-induced
oxidative stress and pulmonary fibrosis in mice.
Discussion
Due to the wide range of toxicities (pulmonary
toxicity) associated with BLM (antineoplastic drug), clinical use
is yet to be effective. Although several mechanisms have been
hypothesized to explain BLM-induced pulmonary toxicity, there is
clear evidence that ROS contribute to the pathogenesis of
BLM-induced lung injury through several pathways (15–17).
Pulmonary fibrosis is a chronic multifactorial disease, where
oxidative stress is important in disease progression (18). In the present study, the
transcription factor, Klf9, was assessed as a regulator of
intracellular ROS in BLM-induced pulmonary toxicity. Klf9 is a
ubiquitously expressed protein with a relatively understudied
function. A significant, concentration-dependent increase in the
total ROS was observed in the present study, indicating an increase
in oxidative stress in response to treatment with BLM (Fig. 1). These data are in agreement with
other previous studies, which have reported an increase in the
production of ROS upon treatment with BLM (2,19).
Antioxidant enzymes are involved in the detoxification or
scavenging of oxidative free radicals in the cells. These results
suggested that upon treatment with BLM, there is a significant
reduction in the major antioxidant proteins, CAT, SOD, GR and TR-2,
which are hypothesized to be an effective defense system in the
lung. This indicated the excess generation of ROS (Fig. 2). This data is consistent with
previous studies indicating that a decrease in antioxidant activity
following BLM treatment indicated the formation of ROS, which, in
turn, caused further damage to lungs (20–22).
However, an increase in the expression levels of Nrf2 (23) and Klf9 upon treatment with BLM has
also been reported (24).
The chemotherapeutic agent, BLM, has been
demonstrated to cause oxidative lung injuries in mice, and
BLM-treated mice are considered to be a classical model of
idiopathic pulmonary fibrosis (25). Currently, only a few genes involved
in the control of ROS have been implicated in the pathogenesis of
pulmonary fibrosis in mouse knockout models (18), including Nrf2 as a sole
transcription factor among these genes (12). Furthermore, Nrf2 was activated and
its targets, including Nqo1, were upregulated in mice treated with
BLM (23). Therefore, the
functional involvement of Klf9, and potentially its targets, in the
pathogenesis of pulmonary fibrosis uncovers a regulatory network of
this disease and may offer targets and/or prognostic markers. The
present investigation using Klf9 knockout mice clearly elucidated
the reduced risk of ROS formation in lungs of BLM-treated mice.
Since Klf9 depletion also suppressed oxidative stress, Klf9
downregulation may promote cancer progression and, potentially,
resistance to the treatment. In conclusion, the data suggested that
Klf9 is upregulated by Nrf2 under conditions of excessive oxidative
stress, therefore, suggesting unique functions and modalities of
the action of Nrf2 in the cell. These results are consistent with
studies regarding the prognostic value of the expression of Klf9 in
cancer and specifically Nrf2-regulated expression of Klf9.
Investigations focusing on the involvement of Nrf2 and Klf9 are
important for determining future therapeutic targets for
BLM-induced pulmonary toxicity.
References
|
1
|
Karam H, Hurbain-Kosmath I and Housset B:
Antioxidant activity in alveolar epithelial type 2 cells of rats
during the development of bleomycin injury. Cell Biol Toxicol.
14:13–22. 1998. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Patel RB, Kotha SR, Sauers LA, Malireddy
S, Gurney TO, Gupta NN, Elton TS, Magalang UJ, Marsh CB, Haley BE,
et al: Thiol-redox antioxidants protect against lung vascular
endothelial cytoskeletal alterations caused by pulmonary fibrosis
inducer, bleomycin: Comparison between classical thiol-protectant,
N-acetyl-L-cysteine and novel thiol antioxidant,
N,N′-bis-2-mercaptoethyl isophthalamide. Toxicol Mech Methods.
22:383–396. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Lazo JS and Chabner BA: Cancer
chemotherapy and biotherapy: principles and practice. Chabner BA
and Longo DL: Philadelphia: Lippincott-Raven; pp. 379–394. 1996
|
|
4
|
Hecht SM: Bleomycin: New perspectives on
the mechanism of action. J Nat Prod. 63:158–168. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Chandler DB: Possible mechanisms of
bleomycin-induced fibrosis. Clin Chest Med. 11:21–30.
1990.PubMed/NCBI
|
|
6
|
Keane MP, Belperio JA, Arenberg DA,
Burdick MD, Xu ZJ, Xue YY and Strieter RM: IFN-gamma-inducible
protein-10 attenuates bleomycin-induced pulmonary fibrosis via
inhibition of angiogenesis. J Immunol. 163:5686–5692.
1999.PubMed/NCBI
|
|
7
|
Finkel T and Holbrook NJ: Oxidants,
oxidative stress and the biology of ageing. Nature. 408:239–247.
2000. View
Article : Google Scholar : PubMed/NCBI
|
|
8
|
Cross CE, Warren D, Gerriets JE, Wilson
DW, Halliwell B and Last JA: Deferoxamine injection does not affect
bleomycin induced lung fibrosis in rats. J Lab Clin Med.
106:433–438. 1985.PubMed/NCBI
|
|
9
|
Itoh K, Wakabayashi N, Katoh Y, Ishii T,
Igarashi K, Engel JD and Yamamoto M: Keap1 represses nuclear
activation of antioxidant responsive elements by Nrf2 through
binding to the amino-terminal Neh2 domain. Genes Dev. 13:76–86.
1999. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Wasserman WW and Fahl WE: Functional
antioxidant responsive elements. Proc Natl Acad Sci USA.
94:5361–5366. 1997. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Itoh K, Chiba T, Takahashi S, Ishii T,
Igarashi K, Katoh Y, Oyake T, Hayashi N, Satoh K, Hatayama I, et
al: An Nrf2/small Maf heterodimer mediates the induction of phase
II detoxifying enzyme genes through antioxidant response elements.
Biochem Biophys Res Commun. 236:313–322. 1997. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Kikuchi N, Ishii Y, Morishima Y, Yageta Y,
Haraguchi N, Itoh K, Yamamoto M and Hizawa N: Nrf2 protects against
pulmonary fibrosis by regulating the lung oxidant level and Th1/Th2
balance. Respir Res. 11:312010. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Mannava S, Zhuang D, Nair JR, Bansal R,
Wawrzyniak JA, Zucker SN, Fink EE, Moparthy KC, Hu Q, Liu S, et al:
KLF9 is a novel transcriptional regulator of bortezomib- and
LBH589-induced apoptosis in multiple myeloma cells. Blood.
119:1450–1458. 2012. View Article : Google Scholar :
|
|
14
|
Bradford MM: A rapid and sensitive method
for the quantitation of microgram quantities of protein utilizing
the principle of protein-dye binding. Anal Biochem. 72:248–254.
1976. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Serrano-Mollar A, Closa D, Prats N, Blesa
S, Martinez-Losa M, Cortijo J, Estrela JM, Morcillo EJ and Bulbena
O: In vivo antioxidant treatment protects against bleomycin-induced
lung damage in rats. Br J Pharmacol. 138:1037–1048. 2003.
View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Mastruzzo C, Crimi N and Vancheri C: Role
of oxidative stress in pulmonary fi brosis. Monaldi Arch Chest Dis.
57:173–176. 2002.
|
|
17
|
Mata M, Ruiz A, Cerda M, Martinez-Losa M,
Cortijo J, Santangelo F, Serrano-Mollar A, Llombart-Bosch A and
Morcillo EJ: Oral N-acetylcysteine reduces bleomycininduced lung
damage and mucin Muc5ac expression in rats. Eur Respir J.
22:900–905. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Cheresh P, Kim SJ, Tulasiram S and Kamp
DW: Oxidative stress and pulmonary fibrosis. Biochim Biophys Acta.
1832:1028–1040. 2013. View Article : Google Scholar :
|
|
19
|
Sayed-Ahmed MM, Mansour HH, Gharib OA and
Hafez HF: Acetyl-L-carnitine modulates bleomycin-induced oxidative
stress and energy depletion in lung tissues. J Egypt Natl Canc
Inst. 16:237–243. 2004.
|
|
20
|
Ali EN and Mansour SZ: Boswellic acids
extract attenuates pulmonary fi brosis induced by bleomycin and
oxidative stress from gamma irradiation in rats. Chin Med.
6:362011. View Article : Google Scholar
|
|
21
|
Iraz M, Erdogan H, Kotuk M, Yagmurca M,
Kilic T, Ermis H, Fadillioğlu E and Yildirim Z: Ginkgo biloba
inhibits bleo-mycin-induced lung fi brosis in rats. Pharmacol Res.
53:310–316. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Erdogan H, Fadillioglu E, Kotuk M, Iraz M,
Tasdemir S, Oztas Y and Yildirim Z: Effects of Ginkgo biloba on
plasma oxidant injury induced by bleomycin in rats. Toxicol Ind
Health. 22:47–52. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Cho HY, Reddy SP, Yamamoto M and
Kleeberger SR: The transcription factor NRF2 protects against
pulmonary fibrosis. FASEB J. 18:1258–1260. 2004.PubMed/NCBI
|
|
24
|
Bieker JJ: Krüppel-like factors: Three
fingers in many pies. J Biol Chem. 276:34355–34358. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Mouratis MA and Aidinis V: Modeling
pulmonary fibrosis with bleomycin. Curr Opin Pulm Med. 17:355–361.
2011. View Article : Google Scholar : PubMed/NCBI
|