Introduction
Epithelial ovarian cancer (EOC) causes the highest
rates of mortality among genital tract malignancies in women. It is
most often diagnosed at the advanced stage of peritoneal
carcinomatosis with a poor prognosis (5-year survival rate 30–35%).
Screening and early detection could probably reduce the mortality
rate.
HE4 was originally identified in the epithelium of
the distal epididymis using northern blot analysis and in
situ transcript hybridization in 1991 (1,2). The
HE4 gene resides on human chromosome 20q12-13.1, a region that
harbors a locus of 14 genes encoding protein domains that have
homology with whey acidic protein (WAP) (3). Two functions attributed to this family
of proteins are the regulation of proinflammatory mediators and
anti-bacterial or anti-fungal activity (4,5). There
is a growing body of evidence demonstrating the tumor-promoting
roles of WAP domain family members (6,7). Among
these WAP genes is secretory leukocyte protease inhibitor (SLPI),
which is also overexpressed in ovarian cancer (6,8).
Hoskins et al (8) reported
that SLPI stimulated ovarian cancer invasion, modulated in part by
its serine protease inhibitory activity. Significantly, comparative
genomic hybridization studies have shown that 20q13 is among the
most frequently amplified chromosomal regions in ovarian cancers
(9–11). HE4 contains WAP domains (12). Based on structural and sequence
similarities with those from other WAP family members, it was
suggested that the protein may exert antiprotease activity. LeBleu
et al (13) identified HE4
as a protease inhibitor using mouse models of renal fibrosis
disease for the first time.
Expression of the HE4 gene is highly restricted in
normal human tissues, being largely limited to the epithelium of
the respiratory tracts, oral and reproductive tracts, and HE4 is
not expressed in normal ovarian surface epithelium (14,15).
HE4 has been reported to be upregulated in several types of cancers
including those of the ovary (16),
endometrium (17), lung (18,19),
breast (7,15), stomach and pancreas (20). The expression of HE4 in ovarian
cancer was initially reported in 1999 (16), and it was subsequently cleared as a
new biological marker of ovarian cancer in 2003 (21). Since then it has been shown to be
potentially useful for remission monitoring (22–24)
and has been approved by the US Food and Drug Administration (FDA)
for that use. Current studies are focusing on the clinical
application of HE4 in EOC as a biomarker, and the diagnostic and
predictive value of HE4 in EOC have been confirmed (25,26).
Several studies have aimed to establish a role for HE4 in cell
proliferation using overexpression and knockdown analyses (27–29),
although the results are controversial. Notably, few studies have
been carried out on the predictive value of HE4 on platinum
resistance and the molecular mechanism of chemoresistance mediated
by HE4.
In the present study, we focused on the effects of
the recombinant HE4 protein on cell proliferation and carboplatin
resistance in SKOV-3 cells, with the aim of providing a theoretical
foundation for HE4 to be used as a predictor for tumor growth
potential and resistance to platinum-based chemotherapy in EOC.
Materials and methods
Chemicals and reagents
The recombinant HE4 protein, which was expressed and
purified in eukaryotic cells, was purchased from Sino Biological,
Inc. (Beijing, China; cat. 12609-H08H). Carboplatin and dimethyl
sulfoxide (DMSO) were purchased from Sigma (St. Louis, MO, USA).
McCoy’s 5A Modified Medium, fetal bovine serum (FBS) and TRIzol
reagent were purchased from Invitrogen (Carlsbad, CA, USA). The
SYBR-Green PCR Master Mix kit was purchased from Toyobo (Osaka,
Japan). The Cell Counting Kit-8 (CCK-8) was purchased from Dojindo
(Kumamoto, Japan).
Cell cultures and treatments
The ovarian cancer cell line SKOV-3 was purchased
from the Cell Culture Collection of Shanghai (Shanghai, China) and
was propagated in McCoy’s 5A Modified Medium with 10% FBS. The cell
cultures were maintained in an incubator at 37°C in a humidified
atmosphere with 95% air and 5% CO2.
CCK-8 assay
SKOV-3 cells were seeded into 96-well plates
(2×103 cells/well) and cultured with serum-free McCoy’s
5A Modified Medium for 24 h. After treatment with the HE4 protein
at different concentrations (0.083, 0.2 or 1 μg/ml) or isometric
serum-free medium for 48 h, 10 μl of the tetrazolium substrate was
added to each well (starting volume of culture media, 100 μl). The
plates were incubated at 37°C for 2 h, and a microplate reader was
used to determine the absorbance of each group at 450 nm. For each
group, three wells were used to calculate the average absorbance
value. Finally, curves or column graphs were drawn to compare the
cell proliferation between the groups.
SKOV-3 cells were seeded into 96-well plates
(5×103 cells/well) and cultured in McCoy’s 5A Modified
Medium with 10% FBS for 48 h. After exposure to carboplatin (10,
25, 50, 100, 200 and 400 μg/ml) or isometric DMSO for different
duration (24, 48 and 72 h), 10 μl of the tetrazolium substrate was
added to each well (starting volume of culture media, 100 μl). The
plates were incubated at 37°C for 2 h, and a microplate reader was
used to determine the absorbance of each group at 450 nm. For each
group, 3 wells were used to calculate the average absorbance value.
The growth inhibitory rate and the IC50 value were
evaluated using SPSS statistical software (version 17.0) (SPSS,
Inc., Chicago, IL, USA).
Colony formation assay
SKOV-3 cells were seeded into 6-well plates
(1×103 cells/well) and were cultured in serum-free
medium that contained the recombinant HE4 protein at different
concentrations (0.083, 0.2 and 1 μg/ml) or isometric serum-free
medium. After 2 days, the cells were cultured in medium with 10%
FBS and the HE4 protein at the concentrations mentioned above; the
medium was replaced every 2 days. Two weeks later, the total number
of colonies in each plate was determined and analyzed using ImageJ
software (National Institutes of Health, Bethesda, MD, USA) version
1.31v. Each experiment was repeated three times for each group, and
the results were subjected to statistical analysis.
Flow cytometry (FCM) for analysis of the
cell cycle
SKOV-3 cells were seeded into 6-well plates
(12×104 cells/well) and were cultured in McCoy’s 5A
Modified Medium with 0.5% FBS for 24 h. After treatment with the
recombinant HE4 protein (0.2 μg/ml) or isometric serum-free medium
for 24 h, the cells were collected and fixed with 2 ml of 70%
ethanol at 4°C for 2 h. Then, the cells were washed with 1×
phosphate-buffered solution (PBS) and incubated for 15 min at room
temperature in 1× PBS containing 100 μg/ml RNase A and 50 μg/ml
propidium iodide (PI). DNA content and cell cycle analyses were
performed using FACScan (Becton-Dickinson, Franklin Lakes, NJ,
USA). Each experiment was repeated three times for each group, and
the results were subjected to statistical analysis.
FCM for analysis of apoptosis
SKOV-3 cells were plated into 6-well plates
(15×104 cells/well) and were cultured in McCoy’s 5A
Modified Medium with 0.5% FBS for 24 h. After being incubated with
the recombinant HE4 protein (0.2 μg/ml) or isometric serum-free
medium for 24 h, the cells were treated with carboplatin (50 μg/ml)
or isometric DMSO for 24 h. The cells were then collected, fixed
and washed with 1× PBS and were incubated for 15 min at room
temperature in 1× PBS containing 100 μg/ml RNase A and 50 μg/ml PI.
DNA content and cell cycle analyses were performed using FACScan.
Based on PI staining, the hypodiploid peak (sub-G1 peak) on the DNA
histogram was considered to be an indicator of apoptosis. We
therefore quantified the sub-G1 peak area. Each experiment was
repeated three times for each group, and the results were subjected
to statistical analysis.
Western blotting
The SKOV-3 cells were prepared as previously
described in apoptosis assay. Briefly, cells were seeded into
6-well plates (15×104 cells/well) and were cultured in
McCoy’s 5A Modified Medium with 0.5% FBS for 24 h. After being
treated with the HE4 protein (0.2 μg/ml) or isometric serum-free
medium for 24 h, carboplatin (50 μg/ml) or isometric DMSO was
added, respectively. The cells were harvested at 48 h
post-treatment. Proteins were separated by SDS-PAGE and transferred
to nitrocellulose membranes. Expression of the target proteins was
determined using the following primary antibodies: Bax (1:200) and
Bcl-2 (1:200) (both from Cell Signaling Technology, USA). The
protein expression levels were visualized using the enhanced
chemiluminescence method. The intensity of each protein band was
quantified using ImageJ 1.31v and normalized relative to the actin
protein expression level.
Microarray profiling and data
analysis
For microarray analysis, the SKOV-3 cell samples
were treated with the recombinant HE4 protein (0.2 μg/ml) or
isometric serum-free medium after starvation. At 12 h
post-treatment, the cells were rinsed once with ice-cold PBS and
lysed, and total RNA was isolated using TRIzol reagent. The two
samples were sent to Gene Tech, Ltd. (Shanghai, China) for
microarray hybridization and detection. We used the PrimeView Human
Gene Expression Array, which covers >36,000 transcripts,
variants and expressed sequence tags. All the samples were
normalized and summarized using the robust multichip analysis (RMA)
normalization method, which includes background correction,
normalization and calculation of the expression values. The data
analysis was performed using Partek Genomics Suite 6.6 software.
Genes were filtered on the basis of call, fold-change and P-value.
All genes with a signal more than ±1.5-fold and P-value <0.05
were chosen to be statistically altered by HE4. A Gene Ontology
(GO) term enrichment analysis was performed using DAVID version 6.7
(http://david.abcc.ncifcrf.gov/). The
Kyoto Encyclopedia of Genes and Genomes (KEGG) was used to identify
the biological pathways in which the differentially expressed genes
are involved.
Real-time quantitative polymerase chain
reaction
Template cDNA was synthesized from total RNA that
was isolated from the SKOV-3 cell samples. All the PCR reactions
were performed using the SYBR-Green PCR Master Mix kit. Briefly,
each PCR reaction contained 1× Master Mix, 1 μl of the diluted
cDNA, and 250 nM forward and reverse primers. PCR was performed
over 40 cycles (95°C for 20 sec and 60°C for 30 sec) following an
initial 3-min enzyme activation step at 95°C. The primers that were
used in the present study for PCR validation are listed in Table I. Each experiment was repeated three
times for each group, and β2-microglobulin (β2-MG) was used as an
endogenous control (reference) gene.
 | Table IPrimers used for quantitative
real-time PCR. |
Table I
Primers used for quantitative
real-time PCR.
| GenBank accession
no. | Gene | Forward primer | Reverse primer |
|---|
| NM_001039667 | ANGPTL4 |
5′-GACCAAGGGGCATGGAGCTT-3′ |
5′-CAGGGGACCTACACACAACAGCA-3′ |
| NM_053056 | CCND1 |
5′-GGATGCTGGAGGTCTGCG-3′ |
5′-GGAGTTGTCGGTGTAGATGC-3′ |
| NM_001145966 | MKI67 |
5′-ACGAGACGCCTGGTTACTATC-3′ |
5′-GCTCATCAATAACAGACCCATTTAC-3′ |
| NM_001191 | BCL2L1 |
5′-GCAGGTATTGGTGAGTCGGATCGC-3′ |
5′-CACAAAAGTATCCCAGCCGCCG-3′ |
| β2MG |
5′-ATGAGTATGCCTGCCGTGTGAAC-3′ |
5′-TGTGGAGCAACCTGCTCAGATAC-3′ |
Statistical analysis
SPSS version 17.0 (SPSS, Inc., Chicago, IL, USA)
software was used for the statistical analysis. The data are
expressed as means ± standard deviation (SD). The statistical
significance of the differences between the groups was evaluated
using one-way analysis of variance (ANOVA), followed by a least
significant difference (LSD) test. P<0.05 was considered to
indicate a statistically significant result.
Results
Validation of the recombinant protein
product
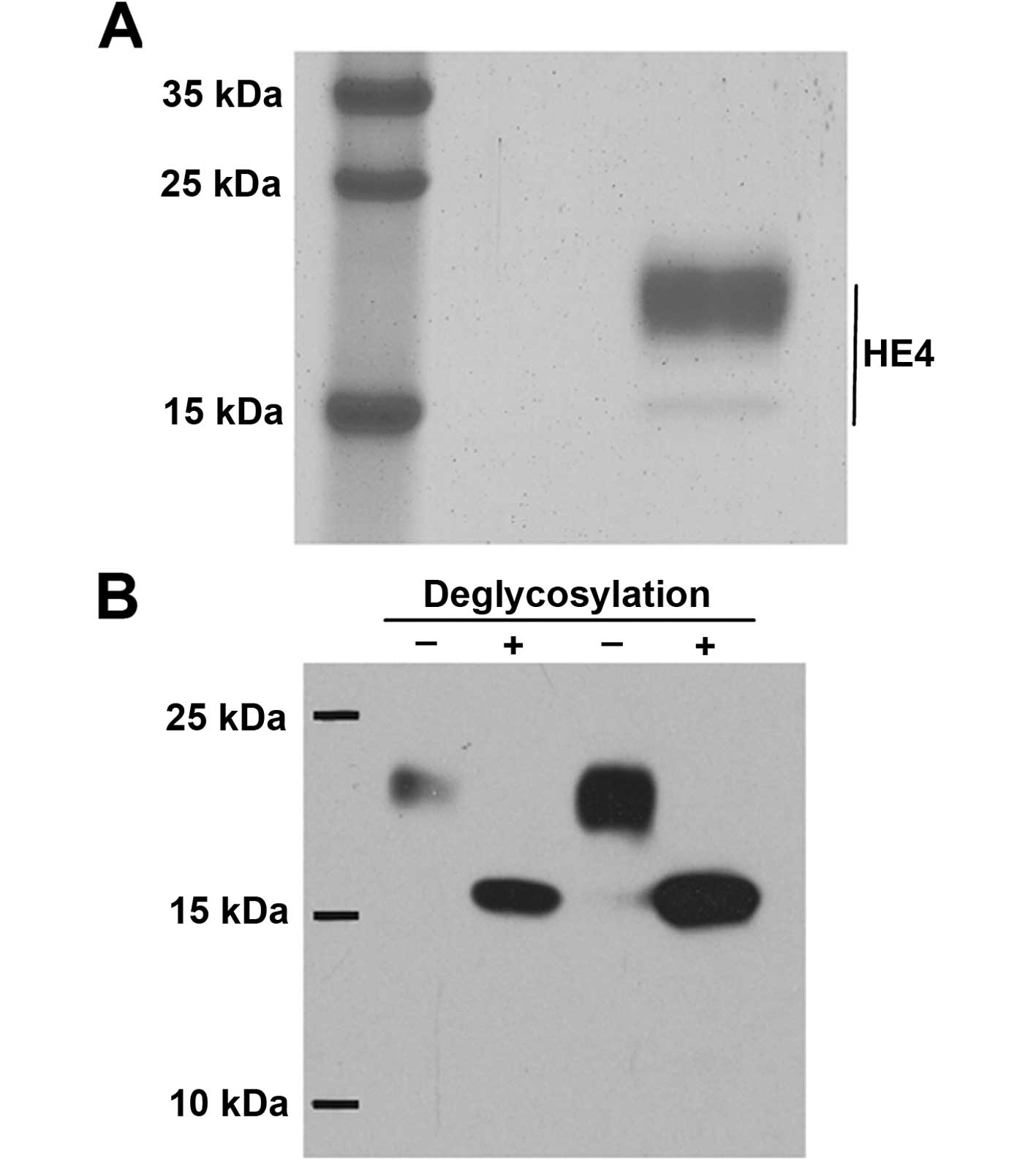
The recombinant HE4 protein product was validated
before the experiments were initiated. SDS-PAGE, stained with
Coomassie blue dye, revealed that the molecular mass was ~20 kDa
(Fig. 1A), which was consistent
with previous reports (13). Next,
glycation modification was tested, and the recombinant protein was
treated with a deglycosylation enzyme (PNGase), which indicated
that the migration of the protein was concentrated at the 15-kDa
position on SDS-PAGE (Fig. 1B).
Therefore, the protein could be expressed and could undergo
glycosylation modification, which is consistent with the
characteristics of HE4 as a glycoprotein. We further confirmed that
the recombinant protein was HE4, according to protein mass
spectrometry (data not shown).
According to the manufacturer’s instructions, the
initial concentration of the recombinant HE4 protein was 250 μg/ml.
Based on several in vitro trials involving recombinant
proteins (30) and the results of
preliminary experiments, the HE4 original solution was diluted
250-fold (1 μg/ml), 1,250-fold (0.2 μg/ml) and 3,000-fold (0.083
μg/ml), respectively, for follow-up experiments.
Recombinant HE4 protein stimulates cell
proliferation
Uncontrolled proliferation is a feature of malignant
cancer that potentially contributes to cancer progression. To
ascertain the effects of the recombinant HE4 protein on cell
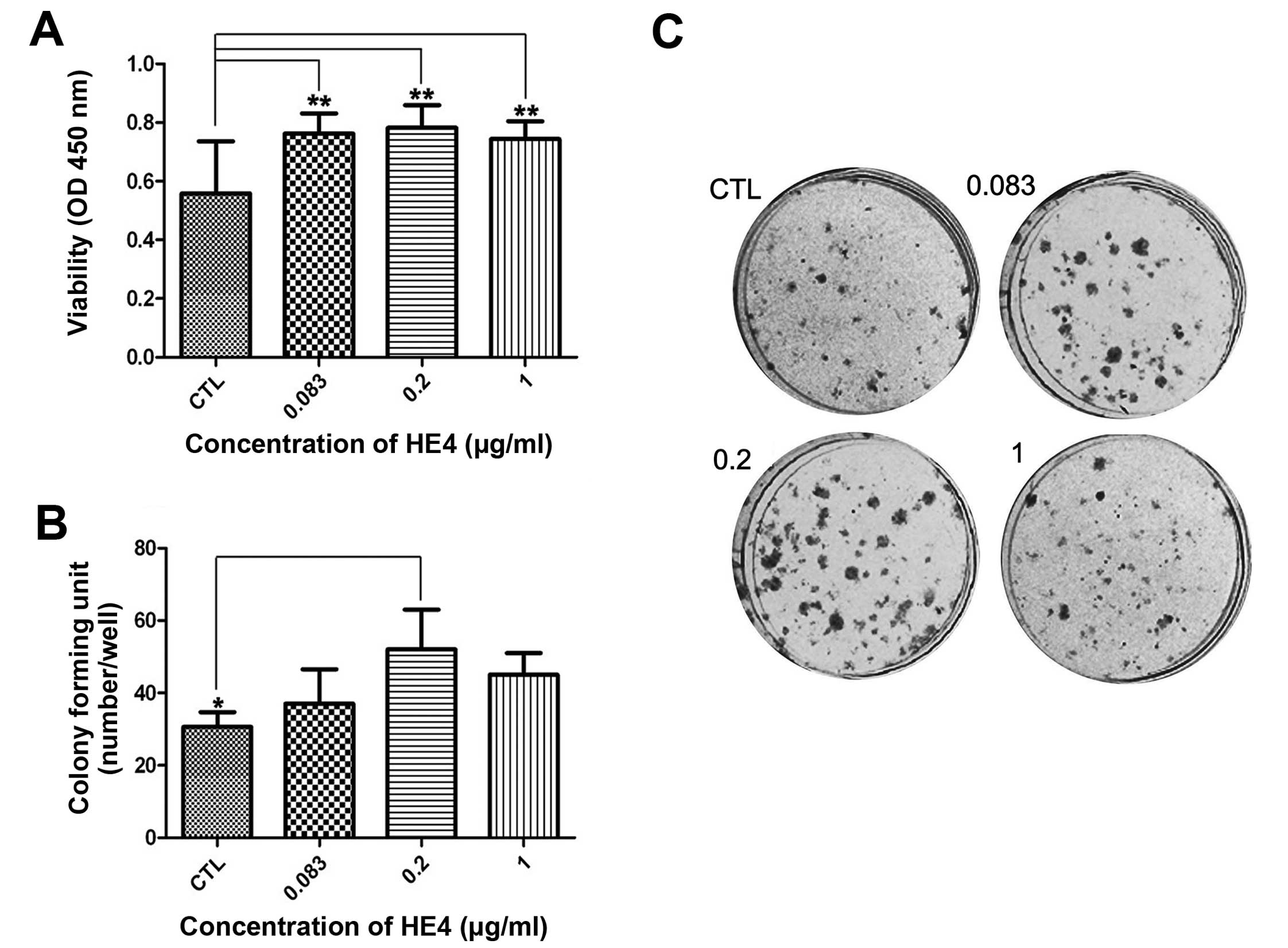
proliferation, we performed a CCK-8 assay using SKOV-3 cells. As
shown in Fig. 2A, compared with the
control cells, the HE4-treated cells exhibited enhanced viability
at 48 h. It is notable that the highest viability level was found
in cells exposed to 0.2 μg/ml HE4 protein (P<0.01).
Next, we carried out colony formation assays. As
shown in Fig. 2B and C, adding the
recombinant HE4 protein (0.2 μg/ml) to SKOV-3 cells resulted in the
formation of significantly more colonies compared with the control
cells (P<0.05).
These results indicated that the HE4 protein
promoted the proliferation of SKOV-3 cells, and the optimal
concentration of HE4 protein used in the CCK-8 and colony formation
assays was 0.2 μg/ml.
Recombinant HE4 protein promotes cell
cycle progression
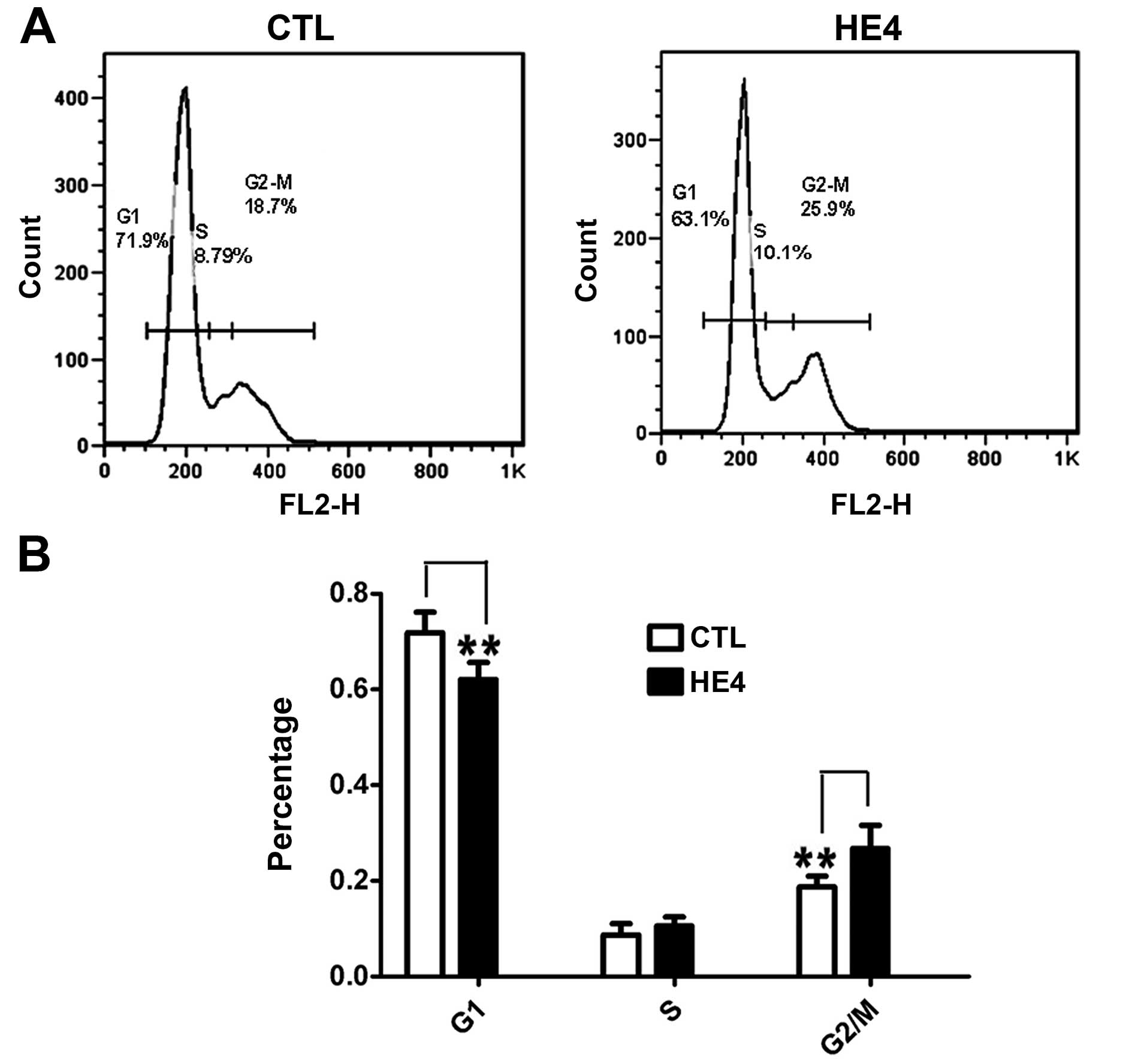
We examined the effects of HE4 on cell cycle
distribution using SKOV-3 cells. DNA content and cell cycle
distribution were determined by FCM. As shown in Fig. 3A and B, a significantly increased
number of cells in the G2/M phase (P<0.01) and a decreased
number of cells in the G0/G1 phase (P<0.01) were observed in the
HE4 protein-treated group. This result confirmed that the HE4
protein promotes cell cycle progression.
Recombinant HE4 protein represses
carboplatin-induced apoptosis
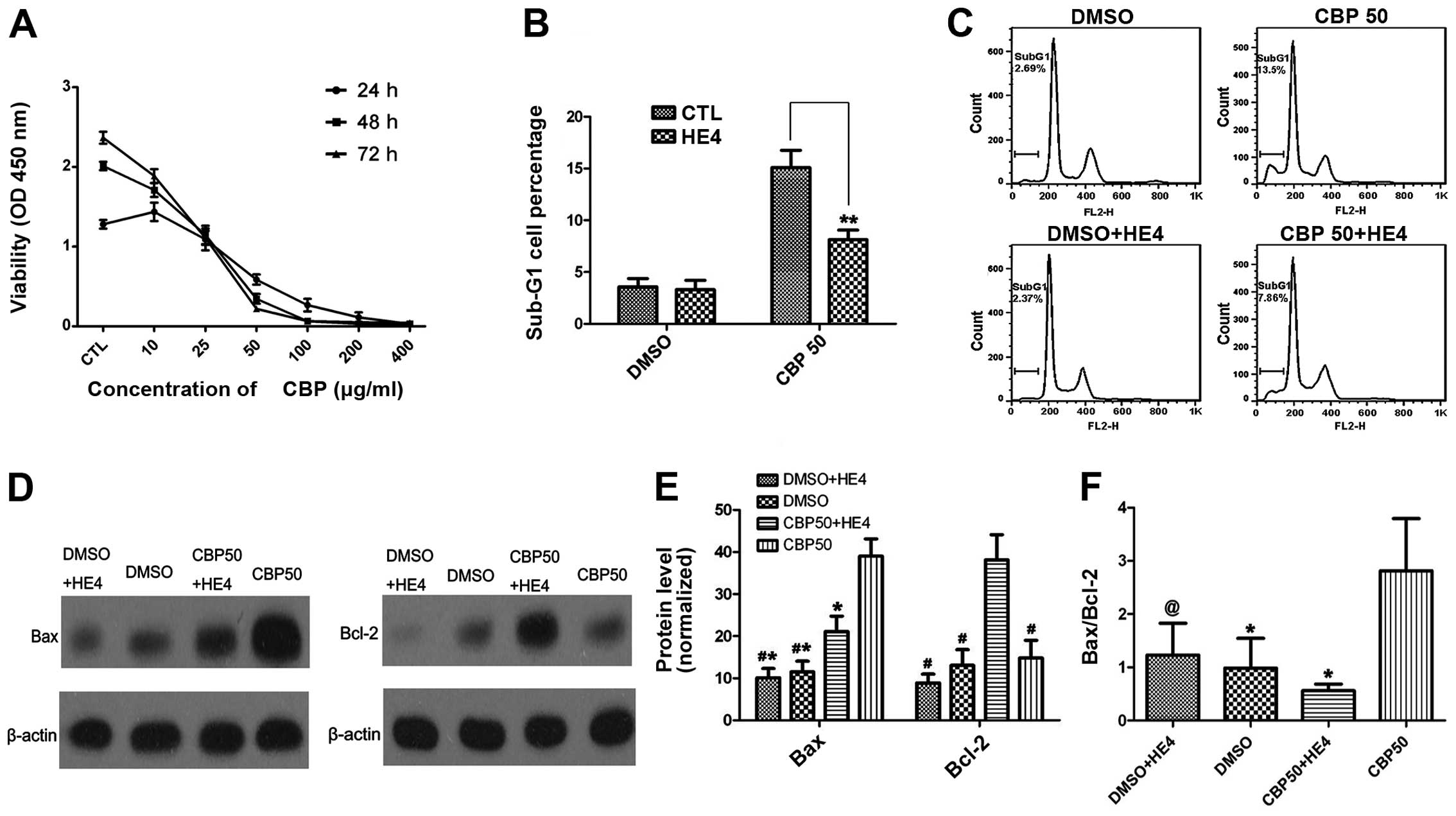
We firstly performed a cell proliferation assay to
determine the viability of cells exposed to carboplatin (0–400
μg/ml) and the growth inhibitory effects of carboplatin on SKOV-3
cells were evaluated. As shown in Fig.
4A, the SKOV-3 cells were relatively sensitive to carboplatin.
Based on an analysis of the cell growth inhibition rate performed
using SPSS statistical software (version 17.0), the IC50
value was 27.6±1.053 μg/ml.
Next, we tested whether the HE4 protein could
attenuate carboplatin-induced apoptosis. Considering that HE4 may
mediate the protective effect against apoptosis, we performed
preliminary experiments for analysis of apoptosis with different
concentrations of carboplatin (30 and 50 μg/ml) combined with HE4
(0, 0.2 μg/ml), and the optimal concentration of carboplatin was
determined to be 50 μg/ml (data not shown). In follow-up
experiments, SKOV-3 cells were treated with the combination of
carboplatin (50 μg/ml) with HE4 protein (0.2 μg/ml) or with
carboplatin alone (50 μg/ml). As shown in Fig. 4B and C, the percentage of cells in
the sub-G1 phase was decreased in the group treated with
carboplatin combined with HE4, compared with the group treated with
carboplatin alone (P<0.01), indicating that the HE4 protein
attenuated carboplatin-induced apoptosis.
We further assessed markers of apoptosis (Bax and
Bcl-2) by western blotting in order to confirm the role of HE4 in
apoptosis (Fig. 4D and E). Our
results indicated that the expression of the anti-apoptotic protein
Bcl-2 was markedly increased in the group treated with carboplatin
combined with HE4, compared with the group treated with carboplatin
alone (P<0.01). In contrast to Bcl-2, the expression of the
pro-apoptotic protein Bax was markedly reduced in the
combination-treatment group compared with the group treated with
carboplatin alone (P<0.01). HE4 markedly decreased the Bax/Bcl-2
ratio (P<0.01) (Fig. 4F).
Gene expression profile of the SKOV-3
cell line
To identify genes altered by HE4 in SKOV-3 cells,
mRNA expression in these cells incubated in the absence and
presence of HE4 protein (0.2 μg/ml, 12 h) was analyzed using
PrimeView Human Gene Expression Array. The HE4 treatment
significantly (P<0.05) altered the expression of 387 genes in
the SKOV-3 cells (236 upregulated and 151 downregulated) (Table II).
 | Table IIOverview of the relevant
differentially expressed genes following treatment with recombinant
HE4 protein (0.2 μg/ml) using microarray analyses. |
Table II
Overview of the relevant
differentially expressed genes following treatment with recombinant
HE4 protein (0.2 μg/ml) using microarray analyses.
| RefSeq ID | Gene title | Gene symbol | Fold-change |
|---|
| NM_006289 | Talin 1 | TLN1 | 2.57297 |
| NM_006281 | Serine/threonine
kinase 3 | STK3 | 2.24695 |
| NM_002526 | 5′-Nucleotidase,
ecto (CD73) | NT5E | 2.05391 |
| NM_001039535 | Spindle and
kinetochore associated complex subunit 1 | SKA1 | 2.03459 |
| NM_022897 | RAN binding protein
17 | RANBP17 | 2.0061 |
| NM_016107 | Zinc finger RNA
binding protein | ZFR | 2.00455 |
| NM_004759 | Mitogen-activated
protein kinase-activated protein kinase 2 | MAPKAPK2 | 1.98081 |
| NM_006041 | Heparan sulfate
(glucosamine) 3-O-sulfotransferase 3B1 | HS3ST3B1 | 1.96157 |
| NM_001145966 | Antigen identified
by monoclonal antibody Ki-67 | MKI67 | 1.95639 |
| NM_016343 | Centromere protein
F, 350/400 kDa (mitosin) | CENPF | 1.9259 |
| NM_001142650 | Heterogeneous
nuclear ribonucleoprotein L-like | HNRPLL | 1.92383 |
| NM_001032283 | Thymopoietin | TMPO | 1.91932 |
| NM_000916 | Oxytocin
receptor | OXTR | 1.90076 |
| NM_144508 | Cancer
susceptibility candidate 5 | CASC5 | 1.8975 |
| NM_004412 | tRNA aspartic acid
methyltransferase 1 | TRDMT1 | 1.88795 |
| NR_002976 | Small nucleolar RNA
host gene 12 (non-protein coding) | SNHG12 | 1.8766 |
| NM_152562 | Cell division cycle
associated 2 | CDCA2 | 1.87068 |
| NM_144643 | Sodium channel and
clathrin linker 1 | SCLT1 | 1.8599 |
| NM_001032283 | Thymopoietin | TMPO | 1.85677 |
| NM_174931 | Coiled-coil domain
containing 75 | CCDC75 | 1.84617 |
| NM_001161429 | RAN binding protein
3-like | RANBP3L | 1.82972 |
| NM_020772 | Nuclear fragile X
mental retardation protein interacting protein 2 | NUFIP2 | 1.81732 |
| NM_001164239 | DEAH
(Asp-Glu-Ala-His) box polypeptide 16 | DHX16 | 1.8158 |
| NM_000346 | SRY (sex
determining region Y)-box 9 | SOX9 | 1.78461 |
| NM_000222 | v-Kit
Hardy-Zuckerman 4 feline sarcoma viral oncogene homolog | KIT | 1.66719 |
| NM_006785 | Mucosa associated
lymphoid tissue lymphoma translocation gene 1 | MALT1 | 1.54842 |
| NM_001191 | BCL2-like 1 | BCL2L1 | 1.54115 |
| NM_053056 | Cyclin D1 | CCND1 | 1.53673 |
| NM_004958 | Mechanistic target
of rapamycin (serine/threonine kinase) | MTOR | 1.50036 |
| NM_000051 | Ataxia
telangiectasia mutated | ATM | −1.52656 |
| NM_004052 | BCL2/adenovirus E1B
19 kDa interacting protein 3 | BNIP3 | −1.73822 |
| NR_027183 | Hypothetical
LOC729678 | LOC729678 | −1.80157 |
| NM_001161520 | Component of
oligomeric golgi complex 5 | COG5 | −1.80739 |
| NM_019058 |
DNA-damage-inducible transcript 4 | DDIT4 | −1.81411 |
| NM_032932 | RAB11 family
interacting protein 4 (class II) | RAB11FIP4 | −1.81574 |
| NM_022783 | DEP domain
containing 6 | DEPDC6 | −1.82032 |
| NM_000096 | Ceruloplasmin
(ferroxidase) | CP | −1.8246 |
| NM_017429 | β-carotene
15,15′-monooxygenase 1 | BCMO1 | −1.82869 |
| NM_001085486 | Selenoprotein P,
plasma, 1 | SEPP1 | −1.83777 |
| NM_001164586 | Immunoglobulin-like
and fibronectin type III domain containing 1 | IGFN1 | −1.83843 |
| NM_005622 | Acyl-CoA synthetase
medium-chain family member 3 | ACSM3 | −1.84541 |
| NM_002133 | Heme oxygenase
(decycling) 1 | HMOX1 | −1.85928 |
| NM_004586 | Ribosomal protein
S6 kinase, 90 kDa, polypeptide 3 | RPS6KA3 | −1.88341 |
| NM_145238 | Zinc finger and
SCAN domain containing 20 | ZSCAN20 | −2.02897 |
| NM_005622 | Acyl-CoA synthetase
medium-chain family member 3 | ACSM3 | −2.09509 |
| NM_004864 | Growth
differentiation factor 15 | GDF15 | −2.12152 |
| NM_003325 | HIR histone cell
cycle regulation defective homolog A | HIRA | −2.6158 |
| NM_001039667 | Angiopoietin-like
4 | ANGPTL4 | −2.65737 |
| NM_002612 | Pyruvate
dehydrogenase kinase, isozyme 4 | PDK4 | −3.32334 |
Identification and analysis of
differentially expressed genes
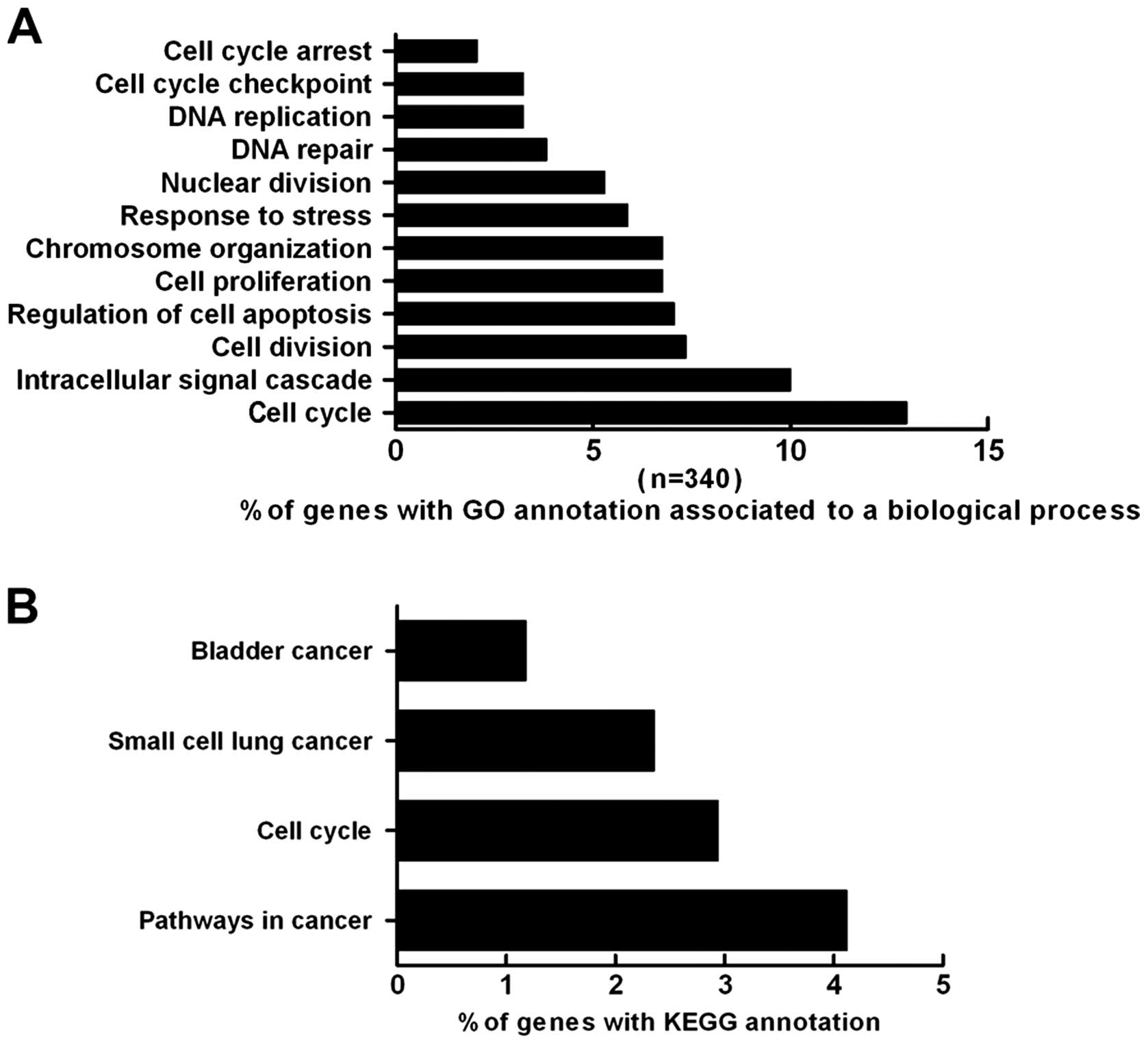
The GO project is an international system of
classification in which the major biological processes, cellular
components or molecular functions of genes and their products are
described using a controlled vocabulary GO terms (31). The GO annotation of our data set
indicated that the differentially expressed genes in the SKOV-3
cells after treatment with HE4 were involved in many processes,
such as cell cycle regulation (12.9%), signal transduction (10%),
cell proliferation (6.8%), apoptosis (7.1%), DNA repair (3.8%) and
the stress response (5.9%) (Fig.
5A).
The KEGG database was utilized to determine the
associations between genes and pathways. The present analysis
revealed that the genes altered by HE4 are involved in several
pathways. Overall, 4.12% of the genes are involved in cancer
pathways, and 2.94% of the genes are associated with the cell
cycle. Additionally, 1.18 and 2.35% of the genes are associated
with bladder cancer and small-cell lung cancer pathways,
respectively (Fig. 5B).
Quantitative real-time PCR
validation
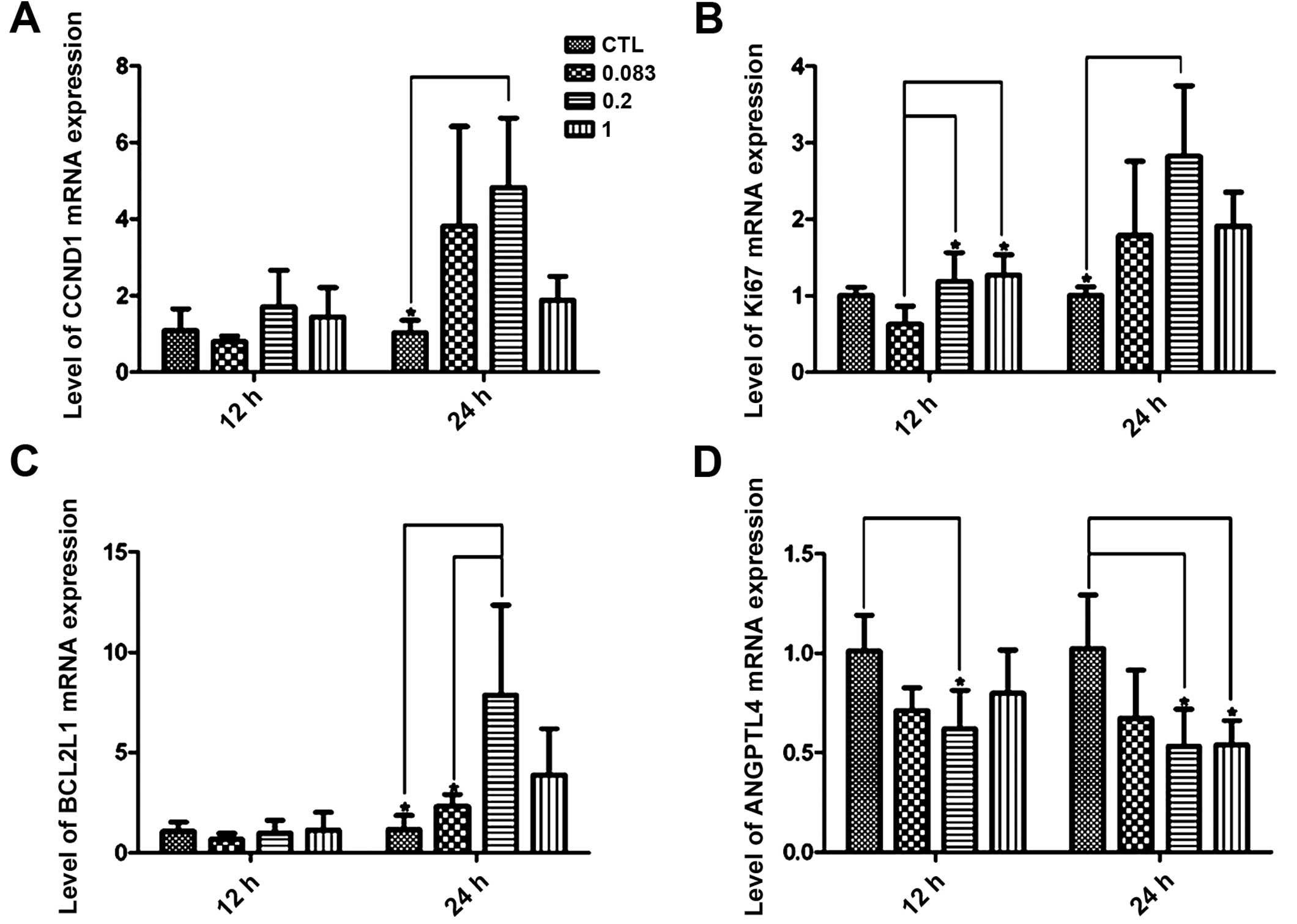
We next determined the differential expression of
four genes, CCND1, KI67, BCL2L1 and ANGPTL4, using quantitative
real-time PCR with independent RNA samples to confirm the validity
of the microarray results. In the majority of the samples, the
results were significantly correlated with the microarray data
(Table II and Fig. 6), and it was notable that the
relatively low concentrations of HE4 (0.2 μg/ml) significantly
altered the mRNA expression of all the four genes at the 24 h time
point (P<0.05).
Discussion
An increasing number of investigations validate the
diagnostic and predictive value of HE4 in EOC. A previous study by
our group (32) showed that the
HE4-positive staining was found more frequently in the epithelium
of serous ovarian carcinomas, compared with borderline and benign
tumors by immunohistochemical methods, and a positive correlation
between expression of HE4 protein and clinical stage was suggested.
It is being accepted recently that HE4 may play a critical role in
promoting the malignancy of ovarian cancer cells as well. Lu et
al (33) showed that
proliferation was significantly inhibited in a SKOV-3 stable strain
with silenced HE4, suggested that HE4 enhanced proliferation by
activating the EGFR-MAPK pathway. Moore et al (27) showed that HE4 overexpression
promoted xenograft tumor growth in a mouse model for ovarian
cancer. However, other reports have presented opposing views. Gao
et al (28) reported that
the upregulation of HE4 led to the significantly reduction in the
number of colonies in ovarian cancer cell lines SKOV-3 and ES-2.
Kong et al (29) suggested
that HE4 protein plays a protective role in SKOV-3 cells by
inhibiting cell proliferation. The present study indicated that
recombinant HE4 protein enhanced cell viability and promoted
accumulation of cells in the G2/M phase. The findings confirm the
proliferation-promoting activity of HE4. Contrary to results
obtained in our cell model, Kong et al (29) showed that neither conditioned medium
containing HE4 nor recombinant HE4 protein had an effect on
proliferation in SKOV-3 cells. It should be noted that our research
team performed serum starvation on cells 24 h before the HE4
treatment. Serum starvation is accepted as a method to synchronize
cells, and it can help to reduce errors among different groups to
some extent.
Apart from differential diagnosis and prediction of
recurrence and overall survival in EOC (25,26),
application of HE4 for other purposes such as a predictor of
platinum resistance has not been extensively investigated. Angioli
et al (25) suggested that
evaluating the serum HE4 values during first-line chemotherapy
could predict chemotherapy response in EOC patients. Hynninen et
al (34) showed that assessment
of serum HE4 could improve the reliability of response evaluation
during chemotherapy for serous EOC compared with CA125. To date,
there have been few studies on the role of HE4 in enhancing the
resistance to drug chemotherapy in ovarian cancer cells. Moore
et al (27) showed that HE4
overexpression induced resistance to cisplatin in a mouse model for
ovarian cancer. Previous experiments conducted in our laboratory
showed that the expression of HE4 antigen was significantly higher
in the drug-resistant group, and the expression of HE4 and the
pathological stage were both independent risk factors for drug
resistance in EOC (unpublished data). In the present study, we
showed that HE4 could attenuate apoptosis induced by carboplatin
through decreasing the mitochondrial Bax/Bcl-2 ratio; as HE4
markedly increased Bcl-2 expression while inhibiting Bax
expression. This finding provides new insight into the role of HE4
in carboplatin resistance in ovarian cancer cells.
Microarray analysis of SKOV-3 genes altered by HE4
identified 236 genes as upregulated and 151 genes as downregulated.
The significantly and consistently upregulated genes were genes
involved in cell cycle regulation and proliferation mainly MTOR,
CCND1, KIT and KI67, thus indicating cell proliferation to be
crucially targeted by HE4 protein. This suitably agrees with our
findings that HE4 protein is very significant in promoting the
proliferation of ovarian cancer cells. Among those within the
downregulated dataset include genes involved in several aspects of
the DNA damage response such as positive regulation of apoptosis,
ATM and BNIP3. This result is consistent with our finding that HE4
protein attenuated carboplatin-induced apoptosis.
We also performed quantitative real-time PCR to
analyze the expression of four genes, CCND1, KI67, BCL2L1 and
ANGPTL4, to confirm the validity of the microarray results and the
results were significantly correlated with the microarray data. We
observed that after 24 h, the mRNA expression of all four genes was
significantly altered in the cells treated with a relatively low
concentration of the HE4 protein (0.2 μg/ml), and the concentration
was consistent with the optimal concentration in the CCK-8 and
colony formation assays. We chose these four genes since the
expression of their mRNA differed clearly from those of the control
group and they were related to cell cycle regulation, proliferation
and apoptosis based on Gene Ontology (GO) terms. It is important
that the elevated expression of BCL2L1 (also known as Bcl-xL), an
essential anti-apoptotic regulator, was confirmed by quantitative
real-time PCR. We also demonstrated that HE4 protein regulated the
expression of the Bcl-2 family members Bax and Bcl-2 using western
blotting. We speculate that regulation of Bcl-2 family members is a
downstream event that occurs after HE4 treatment in SKOV-3 cells.
Further biological experiments are required to elucidate the exact
roles of HE4 in attenuating carboplatin-induced apoptosis.
Unfortunately, the current analysis did not reveal any information
concerning signaling pathways such as EGFR-MAPK, which were
reported in recent articles on HE4 (29,33).
We hypothesize that HE4 may initiate signaling within the cell by
binding to a receptor protein, altering the mRNA levels of critical
target genes associated with cell proliferation and apoptosis; and
then inducing biological effects, including enhanced proliferation
and resistance to apoptosis induced by carboplatin. Additional
research is needed to confirm this hypothesis.
In summary, the present study indicated that the HE4
protein played a promotive role in the proliferation and resistance
to carboplatin-induced apoptosis in the ovarian cancer cell line
SKOV-3. An analysis of the function and regulation of the candidate
genes screened by microarray will help to determine underlying
signaling pathways and target genes coordinated in the cellular
response to HE4. Our findings also provide a theoretical foundation
for HE4 to be used as a predictor for tumor growth potential and
resistance to platinum-based chemotherapy in EOC.
Acknowledgements
This study was supported by the National Natural
Science Foundation of China (nos. 81172491 and 81101527), the Ph.D.
Programs Foundation of Ministry of Education of China (nos.
20112104110016 and 20112104120019), and the Shengjing Free
Researcher Project (no. 201303).
References
|
1
|
Kirchhoff C, Habben I, Ivell R and Krull
N: A major human epididymis-specific cDNA encodes a protein with
sequence homology to extracellular proteinase inhibitors. Biol
Reprod. 45:350–357. 1991. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Kirchhoff C: Molecular characterization of
epididymal proteins. Rev Reprod. 3:86–95. 1998. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Clauss A, Lilja H and Lundwall A: A locus
on human chromosome 20 contains several genes expressing protease
inhibitor domains with homology to whey acidic protein. Biochem J.
368:233–242. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Williams SE, Brown TI, Roghanian A and
Sallenave JM: SLPI and elafin: one glove, many fingers. Clin Sci.
110:21–35. 2006. View Article : Google Scholar
|
|
5
|
Moreau T, Baranger K, Dadé S,
Dallet-Choisy S, Guyot N and Zani ML: Multifaceted roles of human
elafin and secretory leukocyte proteinase inhibitor (SLPI), two
serine protease inhibitors of the chelonianin family. Biochimie.
90:284–295. 2008. View Article : Google Scholar
|
|
6
|
Tanner MM, Grenman S, Koul A, et al:
Frequent amplification of chromosomal region 20q12-q13 in ovarian
cancer. Clin Cancer Res. 6:1833–1839. 2000.PubMed/NCBI
|
|
7
|
Larramendy ML, Lushnikova T, Björkqvist
AM, et al: Comparative genomic hybridization reveals complex
genetic changes in primary breast cancer tumors and their cell
lines. Cancer Genet Cytogenet. 119:132–138. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Hoskins E, Rodriguez-Canales J, Hewitt SM,
et al: Paracrine SLPI secretion upregulates MMP-9 transcription and
secretion in ovarian cancer cells. Gynecol Oncol. 122:656–662.
2011. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Sonoda G, Palazzo J, du Manoir S, et al:
Comparative genomic hybridization detects frequent
overrepresentation of chromosomal material from 3q26, 8q24, and
20q13 in human ovarian carcinomas. Genes Chromosomes Cancer.
20:320–328. 1997. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Kiechle M, Jacobsen A, Schwarz-Boeger U,
Hedderich J, Pfisterer J and Arnold N: Comparative genomic
hybridization detects genetic imbalances in primary ovarian
carcinomas as correlated with grade of differentiation. Cancer.
91:534–540. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Iwabuchi H, Sakamoto M, Sakunaga H, et al:
Genetic analysis of benign, low-grade, and high-grade ovarian
tumors. Cancer Res. 55:6172–6180. 1995.PubMed/NCBI
|
|
12
|
Idoji Y, Watanabe Y, Yamashita A, et al:
In silico study of whey-acidic-protein domain containing oral
protease inhibitors. Int J Mol Med. 21:461–468. 2008.PubMed/NCBI
|
|
13
|
LeBleu VS, Teng Y, O’Connell JT, et al:
Identification of human epididymis protein-4 as a
fibroblast-derived mediator of fibrosis. Nat Med. 19:227–231. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Drapkin R, von Horsten HH, Lin Y, et al:
Human epididymis protein 4 (HE4) is a secreted glycoprotein that is
overexpressed by serous and endometrioid ovarian carcinomas. Cancer
Res. 65:2162–2169. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Galgano MT, Hampton GM and Frierson HF Jr:
Comprehensive analysis of HE4 expression in normal and malignant
human tissues. Mod Pathol. 19:847–853. 2006.PubMed/NCBI
|
|
16
|
Schummer M, Ng WV, Bumgarner RE, et al:
Comparative hybridization of an array of 21,500 ovarian cDNAs for
the discovery of genes overexpressed in ovarian carcinomas. Gene.
238:375–385. 1999. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Zanotti L, Bignotti E, Calza S, et al:
Human epididymis protein 4 as a serum marker for diagnosis of
endometrial carcinoma and prediction of clinical outcome. Clin Chem
Lab Med. 50:2189–2198. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Tokuishi K, Yamashita S, Ohbo K and
Kawahara K: Splice variant HE4-V3 expression is associated with
favorable prognosis in pulmonary adenocarcinoma. Tumour Biol.
33:103–109. 2012. View Article : Google Scholar
|
|
19
|
Iwahori K, Suzuki H, Kishi Y, et al: Serum
HE4 as a diagnostic and prognostic marker for lung cancer. Tumour
Biol. 33:1141–1149. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
O’Neal RL, Nam KT, LaFleur BJ, et al:
Human epididymis protein 4 is up-regulated in gastric and
pancreatic adenocarcinomas. Hum Pathol. 44:734–742. 2013.
View Article : Google Scholar
|
|
21
|
Urban N, Thorpe J, Karlan BY, et al:
Interpretation of single and serial measures of HE4 and CA125 in
asymptomatic women at high risk for ovarian cancer. Cancer
Epidemiol Biomarkers Prev. 21:2087–2094. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Havrilesky LJ, Whitehead CM, Rubatt JM, et
al: Evaluation of biomarker panels for early stage ovarian cancer
detection and monitoring for disease recurrence. Gynecol Oncol.
110:374–382. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Anastasi E, Marchei GG, Viggiani V,
Gennarini G, Frati L and Reale MG: HE4: a new potential early
biomarker for the recurrence of ovarian cancer. Tumour Biol.
31:113–119. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Schummer M, Drescher C, Forrest R, et al:
Evaluation of ovarian cancer remission markers HE4, MMP7 and
Mesothelin by comparison to the established marker CA125. Gynecol
Oncol. 125:65–69. 2012. View Article : Google Scholar :
|
|
25
|
Angioli R, Capriglione S, Aloisi A, et al:
Can HE4 predict platinum response during first-line chemotherapy in
ovarian cancer? Tumour Biol. 35:7009–7015. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Braicu EI, Chekerov R, Richter R, et al:
HE4 expression in plasma correlates with surgical outcome and
overall survival in patients with first ovarian cancer relapse. Ann
Surg Oncol. 21:955–962. 2014. View Article : Google Scholar
|
|
27
|
Moore RG, Hill EK, Horan T, et al: HE4
(WFDC2) gene over-expression promotes ovarian tumor growth. Sci
Rep. 4:35742014. View Article : Google Scholar
|
|
28
|
Gao L, Cheng HY, Dong L, et al: The role
of HE4 in ovarian cancer: inhibiting tumour cell proliferation and
metastasis. J Int Med Res. 39:1645–1660. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Kong X, Chang X, Cheng H, Ma R, Ye X and
Cui H: Human epididymis protein 4 inhibits proliferation of human
ovarian cancer cells via the mitogen-activated protein kinase and
phosphoinositide 3-kinase/AKT pathways. Int J Gynecol Cancer.
24:427–436. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Xiao Q, Qu K, Wang C, et al: HDGF-related
protein-3 is required for anchorage independent survival and
chemoresistance in hepatocellular carcinomas. Gut. 62:440–451.
2013. View Article : Google Scholar
|
|
31
|
Camon E, Magrane M, Barrell D, et al: The
Gene Ontology Annotation (GOA) Database: sharing knowledge in
Uniprot with Gene Ontology. Nucleic Acids Res. 32:D262–D266. 2004.
View Article : Google Scholar :
|
|
32
|
Zhuang H, Gao J, Hu Z, Liu J, Liu D and
Lin B: Co-expression of Lewis y antigen with human epididymis
protein 4 in ovarian epithelial carcinoma. PLoS One. 8:e689942013.
View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Lu R, Sun X, Xiao R, Zhou L, Gao X and Guo
L: Human epididymis protein 4 (HE4) plays a key role in ovarian
cancer cell adhesion and motility. Biochem Biophys Res Commun.
419:274–280. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Hynninen J, Auranen A, Dean K, Lavonius M,
Carpen O, Perheentupa A, Seppänen M and Grénman S: Serum HE4
profile during primary chemotherapy of epithelial ovarian cancer.
Int J Gynecol Cancer. 21:1573–1578. 2011. View Article : Google Scholar : PubMed/NCBI
|