Introduction
Torque Teno virus (TTV) is a small, non-enveloped,
single-stranded circular DNA virus that was first classified as a
member of the family circoviridae, genus Anellovirus. TTV
was first discovered in 1997 in Japan from the serum of a patient
with acute post-transfusion hepatitis of unknown etiology (1). In addition to humans, TTV can also be
detected from non-human primates (2–4), domestic
animals (porcine, avian, bovine and ovine) (5,6), companion
animals (feline and canine) (7), wild
animals (wild boar and camels) (8,9) and marine
animals (sea lions and sea turtle) (10,11). Thus,
in 2009, TTV was classified by the International Committee on
Taxonomy of Viruses into the family Anelloviridae, which contains 9
genera (from Alphatorquevirus to
Zetatorquevirus).
TTV is widespread worldwide and ≥80% of healthy
adults have persistent viraemia (12).
A number of previous studies have reported that the prevalence of
TTV in children was 10–40% (13,14). The
infection and replication mechanisms and the pathogenicity of TTV
remain unknown.
The genome of TTV has a range of 2.1–3.9 kb in
length and contains 3 or 4 overlapping open reading frames (ORFs),
as well as a short stretch of untranslated region with high GC
content. Although TTV is a DNA virus, the TTV sequence exhibits a
wide range of sequence divergence, particularly in human isolates.
Based on the genetic diversity, TTV strains have been classified
into 5 distinct phylogenetic groups (groups 1–5) (15). Additionally, certain novel TTV
variants, which are widely distributed in China (16), have not yet been classified into
genomic groups. Group 1 is represented by the TA278 strain of
genotype 1 and group 2 is represented by the PMV isolate (17). Group 3 is composed of 11 genotypes and
includes SANBAN, TUS01 and TYM9 isolates (18,19).
Genogroups 1 and 3 are the most widespread, followed by 4 (KC009)
and 5 (CT39), while genogroup 2 viruses are less common (20).
In the present study, in order to characterize the
infection status of human TTV in China, serum samples were
investigated and collected from hospitalized patients with
cardiovascular disease, tumor or gastroenteritis, using ELISA and
polymerase chain reaction (PCR) assays.
Materials and methods
Samples
A total of 378 blood specimens were collected from
the hospitalized patients, aged from 19 to 89 years, during the
period between August and December in 2012, in Jiangxi, China. All
378 patients suffered from cardiovascular disease (171/378), tumor
(192/378) or gastroenteritis (15/378) at the sampling time. No
patients had a history of transfusions. Samples were centrifuged at
1,000 × g for 20 min, and supernatant serum aliquots were collected
and stored at −80°C until the testing was performed.
ELISA detection
TTV antigen (Ag) was determined using a commercial
ELISA kit (Huitebi Technology Development Co., Ltd., Beijing,
China) according to the manufacturer's instructions. The ELISA
plate was coated with purified TTV antibody to capture the TTV
virus from the detected serum and screened by the horseradish
peroxidase-tagged TTV antibody (catalogue number, 2R109). The kit
contained positive and negative controls, and the cut-off values
for the results assay were determined based on 0.15 plus the mean
optical density 450 values of the negative control samples.
DNA extraction
Viral DNA was extracted from 100 µl serum samples
using the MiniBEST Viral RNA/DNA Extraction kit version 4.0
(Takara, Dalian, China), according to the manufacturer's
instructions.
PCR detection of TTV DNA
All the positive samples detected by ELISA were
tested via nested PCR using specific primers corresponding to the 5
TTV genogroups (groups 1–5) (Table I),
and the reference sequences were as follows: Group 1: TA278
(accession no. AB017610) (21), group
2: PMV (AF261761) (22), group 3:
TUS01 (AB017613) (23), group 4: KC009
(24) (AB038621) and group 5: CT39F
(AB064604) (25). All PCR reactions
were carried out as follows: 32 cycles of denaturation at 94°C for
40 sec with an additional 7 min in the first cycle, annealing at
55°C for 40 sec, extension at 72°C for 70 sec and with an
additional 7 min in the last cycle. The amplification products were
excised from 1% agarose gels containing ethidium bromide (0.5
µg/ml), purified with the AxyPrep DNA Gel Extraction kit (Axygen
Biotechnology Co., Ltd., Silicon Valley, CA, USA), cloned into the
pMD-18T vector (Takara) and sequenced (Takara).
 | Table I.Primers employed in the analysis of
Torque Teno virus. |
Table I.
Primers employed in the analysis of
Torque Teno virus.
| Group | Polarity | Nucleotide
position | Nucleotide
sequence |
|---|
| 1 (TA278) | Sense-1 | 867–886 | 5′-ATG TGG CGA AAA
TAC TGT CA-3′ |
|
| Antisense-1 | 1903–1923 | 5′-CAT GTT GCC TTC
TCC TCT GTC-3′ |
|
| Sense-2 | 1102–1125 | 5′-ATT AAT ACC ATG
CCT CCT TTT CTA-3′ |
|
| Antisense-2 | 1903–1923 | 5′-CAT GTT GCC TTC
TCC TCT GTC-3′ |
| 2 (PMV) | Sense-1 | 107–126 | 5′-TGA GTT TTC CAC
GCC CGT CC-3′ |
|
| Antisense-1 | 1424–1433 | 5′-ATC CGG CGG TTA
TAC CAG TA-3′ |
|
| Sense-2 | 473–492 | 5′-CAG TGT GGC GGC
TCG TGT TG-3′ |
|
| Antisense-2 | 1364–1384 | 5′-TTG TGG TGA GCA
GAA CGG AAA-3′ |
| 3 (TUS01) | Sense-1 | 322–341 | 5′-GAA GGC ACC TGC
CAT GAG CT-3′ |
|
| Antisense-1 | 1936–1956 | 5′-GCC TTT GCC CTT
GTC CAT TAG-3′ |
|
| Sense-2 | 477–496 | 5′-TGC CTG CTA CCT
CTT CGC CT-3′ |
|
| Antisense-2 | 1358–1378 | 5′- AGC AGA ACG GAT
ACC GCA AGT-3′ |
| 4 (KC009) | Sense-1 | 137–158 | 5′-AGG CCA ATG AGG
ATC TTC TAC G-3′ |
|
| Antisense-1 | 982–1001 | 5′-GGG AGG GAA GTC
GTC CAT GT-3′ |
|
| Sense-2 | 360–381 | 5′-TTA AAA AAT TCC
CCC GCT CTG T-3′ |
|
| Antisense-2 | 918–937 | 5′-AGG GGC ATC CAT
CCT GTA AT-3′ |
| 5 (CT39) | Sense-1 | 106–125 | 5′-GAG TTT ATG CTG
CCC GTC CG-3′ |
|
| Antisense-1 | 722–742 | 5′-TCC ACC TCC TCC
GCC TCC TTA-3′ |
|
| Sense-2 | 191–210 | 5′-CGC AGT CAA GGG
GCA ATT CG-3′ |
|
| Antisense-2 | 671–690 | 5′-TCT AGG CCA TCG
TCT GCG AA-3′ |
Phylogenetic analysis
The sequences of the TTV isolates in the present
study were analyzed using the MegAlign software (DNAStar Inc.,
Madison, WI, USA). Phylogenetic trees were constructed by the
alignment of the TTV isolates in the study and the referenced
strains (GenBank number and source of regions are shown in Fig. 1). They were evaluated using the
neighbor-joining method with 1,000 bootstrap replicates in a
heuristic search with the Molecular Evolutionary Genetics Analysis
program (MEGA, version 4.0; Oxford University Press, New York, NY,
USA).
Results
Detection of TTV by ELISA
A total of 378 patient serum specimens were
collected from different departments (171 with cardiovascular
disease, 192 with tumor and 15 with gastroenteritis) and were
divided into 3 groups according to age. TTV was detected by the
human ELISA detection kit following the manufacturer's instructions
and 64 specimens were positive for TTV. The prevalence of TTV was
14.0% (24/171), 18.8% (36/192) and 26.7% (4/15) in cardiovascular,
tumor and gastroenteritis patients, respectively. The patients aged
<30 years had a higher prevalence in cardiovascular (50.0%, 3/6)
and tumor patients (33.3%, 1/3) and TTV in males, 20.2% (36/178),
was more common compared with female patients, 14.0% (28/200)
(Table II).
 | Table II.ELISA detection result of Torque Teno
virus in cardiovascular, tumor and gastroenteritis patients. |
Table II.
ELISA detection result of Torque Teno
virus in cardiovascular, tumor and gastroenteritis patients.
|
|
| Infection rate |
|
|---|
|
|
|
|
|
|---|
| Patients | Age, years | Male | Female | Total | Total rate for
patients |
|---|
| Cardiovascular
disease (n=171) | ≤30 | 33.3 (1/3) | 66.7 (2/3) | 50.0 (3/6) | 14.0 (24/171) |
|
| 31–60 | 18.5 (5/27) | 12.0 (3/25) | 15.4 (8/52) |
|
|
| ≥60 | 20.0 (10/50) | 4.7 (3/63) | 11.5 (13/113) |
|
| Tumor (n=192) | ≤30 | 100.0 (1/1) | 0.0 (0/2) | 33.3 (1/3) | 18.8 (36/192) |
|
| 31–60 | 4.7 (2/43) | 15.7 (11/70) | 11.5 (13/113) |
|
|
| ≥60 | 30.9 (13/42) | 26.5 (9/34) | 28.9 (22/76) |
|
| Gastroenteritis
(n=15) | ≤30 | 0.0 (0/0) | 0.0 (0/0) | 0.0 (0/0) | 26.7 (4/15) |
|
| 31–60 | 33.3 (2/6) | 0.0 (0/1) | 28.6 (2/7) |
|
|
| ≥60 | 33.3 (2/6) | 0.0 (0/2) | 25.0 (2/8) |
|
| Total rate for
gender | No. | 20.2 (36/178) | 14.0 (28/200) | No. |
|
PCR detection of TTV DNA
According to the result of ELISA, 64 patients (24
with cardiovascular disease, 36 with tumors and 4 with
gastroenteritis) were detected to be carrying TTV. In order to know
the most prominent genogroup, 64 serum specimens were analyzed by
nest-PCR with group-specific primer sets. The results showed that
the positive samples were 48 among the 64 samples (56.3%), and in
the 24 cardiovascular disease patients, the infection rate of TTV
groups 1–5 were 20.8% (5/24), 0.0% (0/24), 16.7% (4/24), 20.8%
(5/24) and 8.3% (2/24), respectively (Table III). A total of 4 patients had
co-infection of groups 3 and 4, or groups 3 and 5, or groups 1 and
5, or groups 1, 3 and 4 (Table IV).
The prevalence in the 36 tumor patients were 22.2% (8/36), 0.0%
(0/36), 13.9% (5/36), 41.2% (15/36) and 19.4% (7/36), respectively.
Among the tumor patients, 2 were co-infected with 3 groups (groups
1, 4 and 5), and 10 were co-infected with 2 groups (groups 1 and 3,
groups 1 and 4, groups 1 and 5, groups 3 and 4 or groups 4 and 5).
However, in gastroenteritis patients, groups 4 and 5 were detected
with the prevalence of 50.0% (2/4) (Table
IV).
 | Table III.Frequency of Torque Teno virus
detected by nest-polymerase chain reaction assay in cardiovascular,
tumor and gastroenteritis patients. |
Table III.
Frequency of Torque Teno virus
detected by nest-polymerase chain reaction assay in cardiovascular,
tumor and gastroenteritis patients.
| Patients | Group 1 | Group 2 | Group 3 | Group 4 | Group 5 |
|---|
| Cardiovascular
disease (n=24) | 20.8 (5/24) | 0.0 (0/24) | 16.7 (4/24) | 20.8 (5/24) | 8.3 (2/24) |
| Tumor (n=36) | 22.2 (8/36) | 0.0 (0/36) | 13.9 (5/36) | 41.7 (15/36) | 19.4
(7/36) |
| Gastroenteritis
(n=4) | 0.0 (0/4) | 0.0 (0/4) | 0.0 (0/4) | 50.0 (2/4) | 50.0 (2/4) |
| Total (n=64) | 21.7 (13/60) | 0.0 (0/60) | 15.0 (9/60) | 34.4 (22/64) |
17.2 (11/64) |
 | Table IV.Torque Teno virus infection from
different groups in cardiovascular, tumor and gastroenteritis
patients. |
Table IV.
Torque Teno virus infection from
different groups in cardiovascular, tumor and gastroenteritis
patients.
| Group | Patient no. | Gender | Age, years | Groups |
|---|
| Tumor | 2 | F | 65 | 1,4,5 |
|
| 4 | F | 31 | N |
|
| 5 | F | 72 | 1,4,5 |
|
| 6 | F | 38 | 1,5 |
|
| 8 | F | 43 | N |
|
| 9 | M | 66 | 3,4 |
|
| 10 | M | 63 | 1,5 |
|
| 11 | F | 71 | 4 |
|
| 12 | F | 37 | N |
|
| 15 | F | 56 | 1 |
|
| 16 | M | 85 | 3 |
|
| 17 | F | 80 | N |
|
| 18 | M | 63 | 3 |
|
| 19 | M | 62 | 4 |
|
| 20 | F | 71 | 4,5 |
|
| 21 | M | 75 | 4,5 |
|
| 23 | M | 67 | N |
|
| 25 | F | 68 | N |
|
| 29 | M | 65 | 1,4 |
|
| 30 | F | 31 | 4,5 |
|
| 31 | M | 62 | N |
|
| 34 | M | 66 | N |
|
| 36 | F | 66 | 1,4 |
|
| 39 | F | 62 | 4 |
|
| 43 | M | 28 | 5 |
|
| 45 | M | 77 | 4 |
|
| 47 | F | 37 | 4 |
|
| 48 | M | 65 | 3,4 |
|
| 49 | F | 55 | N |
|
| 51 | M | 58 | N |
|
| 58 | F | 59 | N |
|
| 60 | M | 65 | N |
|
| 63 | F | 55 | N |
|
| 64 | M | 50 | N |
|
| 65 | F | 65 | N |
|
| 66 | F | 56 | 1,3 |
| Cardiovascular
disease | 1 | M | 49 | N |
|
| 3 | F | 54 | 4 |
|
| 7 | F | 56 | 3,4 |
|
| 13 | M | 53 | 3,5 |
|
| 14 | F | 54 | 4 |
|
| 22 | M | 69 | N |
|
| 24 | M | 60 | 1 |
|
| 26 | M | 47 | N |
|
| 27 | M | 21 | N |
|
| 33 | F | 64 | N |
|
| 38 | M | 76 | 1,3,4 |
|
| 40 | M | 77 | 1 |
|
| 44 | F | 29 | N |
|
| 46 | F | 79 | 4 |
|
| 50 | M | 44 | N |
|
| 52 | M | 61 | 1,5 |
|
| 53 | M | 69 | N |
|
| 54 | M | 61 | N |
|
| 55 | M | 66 | N |
|
| 56 | M | 42 | N |
|
| 57 | M | 61 | 1 |
|
| 59 | M | 61 | N |
|
| 61 | F | 75 | 3 |
|
| 62 | F | 29 | N |
|
Gastroenteritis | 8 | M | 64 | 4 |
|
| 37 | M | 64 | 4 |
|
| 41 | M | 43 | 5 |
|
| 42 | M | 55 | 5 |
Sequence analysis of TTVs
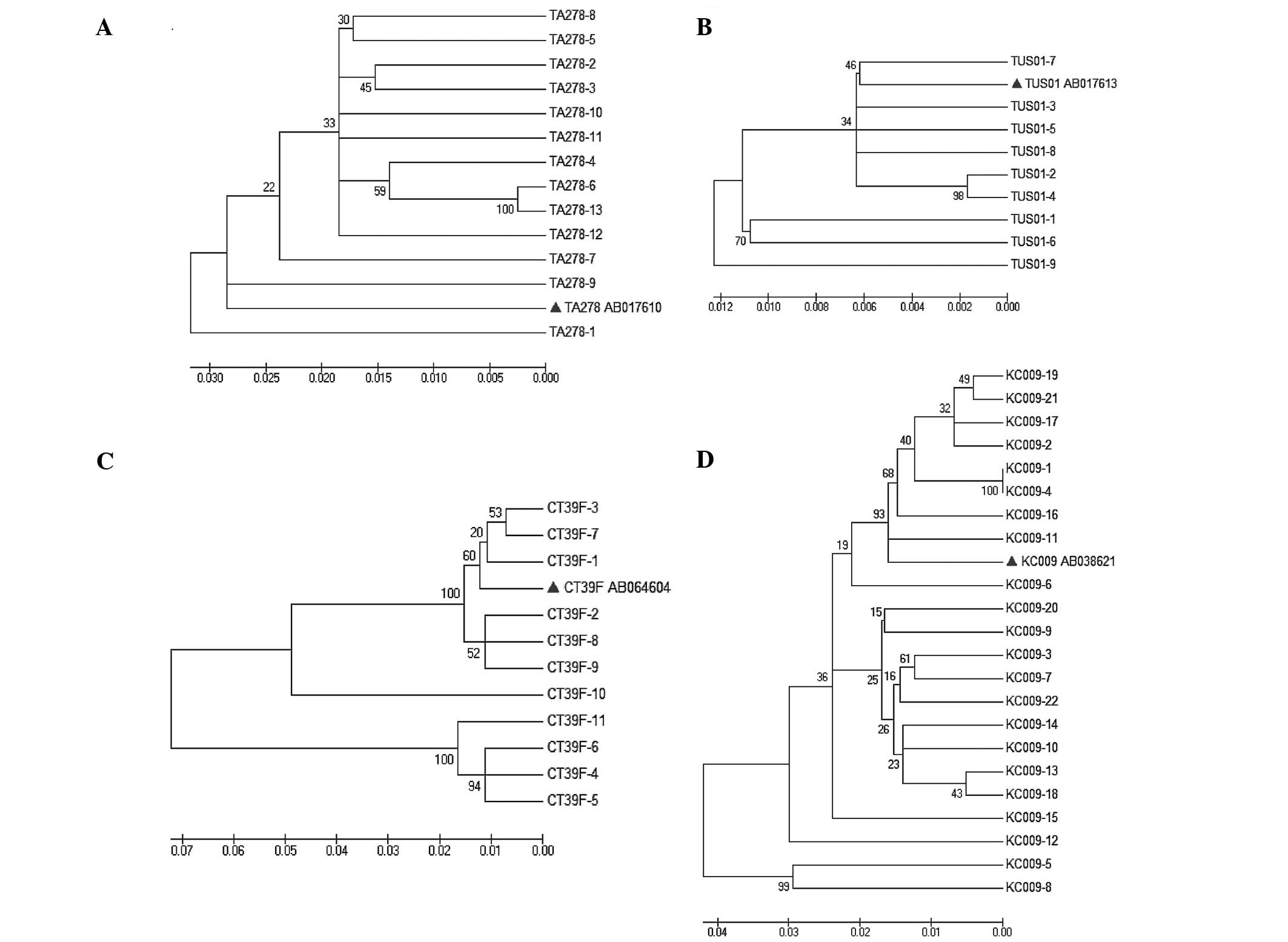
The phylogenetic tree was constructed (Fig. 1) based on the sequences of the isolates
and referenced representative strains (GenBank number and source of
regions are shown). In tree A, all the 13 isolates were found to
share 92–95% nucleotide homology with the genogroup 1 strain TA278
(AB017610). In tree B, the genogroup 3 strain TUS01 (AB017613)
shared 97–99% nucleotide homology with the isolates. The tree C was
grouped into 2 clusters, and the 4 isolates (CT39F-4, 5, 6 and 11)
were found to share 86–88% nucleotide homology with the referenced
genogroup 5 strain CT39F (AB064604), the other isolates were
clustered with the strain CT39F and shared 90–98% nucleotide
homology with it. In tree D, all the isolates shared 92% nucleotide
homology at least with the genogroup 4 strain KC009 (AB038621)
besides one isolate, which shared 85–89% nucleotide homology with
KC009 strain and other isolates.
Discussion
Human TTV is globally distributed, and TTV strains
have been divided into 5 distinct phylogenetic groups (groups 1–5).
PCR is the usual detection method for TTV DNA, and the choice of
primers used may significantly influence the level of detection.
High rates of infection (60–100%) have been identified among
healthy populations worldwide by primers T801 and T935. These were
designed in the 3′ end of the conserved untranslated region (UTR)
(12) that is able to amplify the
genomes of a number of TTV genotypes. However, the phylogenetic
classifications of TTV isolates based on the 2 most prominently
studied regions of the genome (N22 and UTR PCR regions) are
unreliable (26). Therefore,
genogroup-specific or genotype-specific primers were designed to
detect the prevalence of TTV. Genogroups 1 and 3 are the most
widespread, followed by 4 and 5, while genogroup 2 viruses are less
common (20).
Thus far, there are few studies regarding TTV
detection with different genogroup-specific primers. Devalle and
Niel (27) used the oligonucleotide
primers T1S (sense) and T1A (antisense) designed at the 3′ and 5′
ends of the conserved UTR for the first cycle to detect and
differentiate TTV isolates belonging to each of the 5 genomic
groups (groups 1–5). In the second cycle of PCR, T2S (sense),
immediately downstream of T1S, was designed in the UTR. The 5 PCR
assays differed by their internal antisense primers, T2G1A, T2G2A,
T2G3A, T2G4A and T2G5A, which were designed to be specific for TTV
genomic groups 1–5, respectively. The results showed that TTV DNA
from ≥1 genomic group was detected in 11 (46%) blood donors, 13
(54%) hepatitis B virus (HBV) carriers and 24 (100%) human
immunodeficiency virus-1 (HIV-1)-infected patients. The genomic
group 5 TTV was the most prevalent (46%, 33/72), followed by group
3 (43%, 31/72), group 1 (35%, 25/72), group 2 (18%, 13/72) and
group 4 (17%, 12/72), and the prevalence of TTV in HIV-1 patients
was higher compared to HBV carriers and blood donors. Through
aligned submitted full-length TTV nucleotide sequences from
GenBank, Biagini et al (28)
designed the specific primer sets (TTG1S1/R1, TTG1S2/R2- TTG5S1/R1,
TTG5S1/R1) for each of the representative phylogenetic groups to
detect TTV from plasma samples. The above results indicated that
the overall prevalence value for TTV DNA totaled 48%, and TTV
belonging to group 1 was the most frequently detected (34%),
followed by group 3 (24%, TUPB prototype) and group 5 (12%, JT33F
prototype). By contrast, viruses belonging to group 2 (2%, KAV
prototype) and group 4 (2%, JT41F prototype) were only detected
occasionally.
In the present study, a total of 378 patient serum
specimens were collected from different departments (171
cardiovascular disease patients, 192 tumor patients and 15
gastroenteritis patients) and 64 specimens (17%) were positive for
TTV, as detected by the human ELISA detection kit. The prevalence
of TTV was 14.0% (24/171), 18.8% (36/192) and 26.7% (4/15) in
cardiovascular, tumor and gastroenteritis patients, respectively.
The patients aged <30 years have a higher prevalence in
cardiovascular (50.0%, 3/6) and tumor patients (33.3%, 1/3), and
TTV in males, 20.2% (36/178), was more common compared to female
patients, 14.0% (28/200).
Although the ELISA kit can detect TTV Ag, the
genogroups are not clear, and in order to know the most prominent
genogroup, 64 serum specimens were analyzed by nest-PCR with
group-specific primers sets. The results showed that the positive
samples were 48 among the 64 samples (56.3%), but not 100%, which
confirmed that certain other genogroups may exist, or all the
genotypes in one genogroup cannot be detected by the primers.
However, the genogroups were still analyzed in the
positive samples. In the 24 patients with cardiovascular disease,
groups 1 and 4 were most prevalent with the infection of 20.8%
(5/24), followed by group 3 at 16.7% (4/24) and group 5 at 8.3%
(2/24). In the 36 patients with tumors, group 4 was most prevalent
with the infection of 41.2% (15/36), followed by group 1 at 22.2%
(8/36), group 5 at 19.4% (7/36) and 13.9% (5/36). However, in the
patients with gastroenteritis, only groups 4 and 5 were detected
with the prevalence of 50.0% (2/4). The discrepancy between groups
may be due to the study of different populations or by
methodological differences in the protocols used. No samples
belonging to group 2 were detected from all the 64 patients,
indicating that group 2 had a low prevalence, which is consistent
with the previous study.
The phylogenetic tree was constructed (Fig. 1) based on the sequences of the isolates
and referenced representative strains. In tree A, all the 13
isolates were found to share 92–95% nucleotide homology with the
genogroup 1 strain TA278 (AB017610). In tree B, the genogroup 3
strain TUS01 (AB017613) shared 97–99% nucleotide homology with the
isolates. The tree C was grouped into 2 clusters, and the 4
isolates (CT39F-4, 5, 6 and 11) were found to share 86–88%
nucleotide homology with the referenced genogroup 5 strain CT39F
(AB064604). The other isolates were clustered with the strain CT39F
and shared 90–98% nucleotide homology with it. In tree D, all the
isolates shared 92% nucleotide homology at least with the genogroup
4 strain KC009 (AB038621) besides one isolate, which shared 85–89%
nucleotide homology with KC009 strain and other isolates. These
results confirm that genetic variability rather than geographical
variance exists among TTVs in infected humans.
In the present study, different Chinese human TTV
isolates were detected and investigated. These findings provide
novel insights and foundations for further studies to characterize
the territorial presence and prevalence of TTV within China.
References
|
1
|
Nishizawa T, Okamoto H, Konishi K,
Yoshizawa H, Miyakawa Y and Mayumi M: A novel DNA virus (TTV)
associated with elevated transaminase levels in posttransfusion
hepatitis of unknown etiology. Biochem Biophys Res Commun.
241:92–97. 1997. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Okamoto H, Nishizawa T, Takahashi M,
Tawara A, Peng Y, Kishimoto J and Wang Y: Genomic and evolutionary
characterization of TT virus (TTV) in tupaias and comparison with
species-specific TTVs in humans and non-human primates. J Gen
Virol. 82:2041–2050. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Jelcic I, Hotz-Wagenblatt A, Hunziker A,
Zur Hausen H and de Villiers EM: Isolation of multiple TT virus
genotypes from spleen biopsy tissue from a Hodgkin's disease
patient: Genome reorganization and diversity in the hypervariable
region. J Virol. 78:7498–7507. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Ninomiya M, Takahashi M, Hoshino Y,
Ichiyama K, Simmonds P and Okamoto H: Analysis of the entire
genomes of torque teno midi virus variants in chimpanzees:
Infrequent cross-species infection between humans and chimpanzees.
J Gen Virol. 90:347–358. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Leary TP, Erker JC, Chalmers ML, Desai SM
and Mushahwar IK: Improved detection systems for TT virus reveal
high prevalence in humans, non-human primates and farm animals. J
Gen Virol. 80:2115–2120. 1999. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Brassard J, Gagné MJ, Lamoureux L, Inglis
GD, Leblanc D and Houde A: Molecular detection of bovine and
porcine Torque teno virus in plasma and feces. Vet Microbiol.
126:271–276. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Okamoto H and Mayumi M: TT virus:
Virological and genomic characteristics and disease associations. J
Gastroenterol. 36:519–529. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Martínez L, Kekarainen T, Sibila M,
Ruiz-Fons F, Vidal D, Gortázar C and Segalés J: Torque teno virus
(TTV) is highly prevalent in the European wild boar (Sus scrofa).
Vet Microbiol. 118:223–229. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Al Moslih, Perkins H and Hu YW: Genetic
relationship of Torque Teno virus (TTV) between humans and camels
in United Arab Emirates (UAE). J Med Virol. 79:188–191. 2007.
View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Ng TF, Manire C, Borrowman K, Langer T,
Ehrhart L and Breitbart M: Discovery of a novel single-stranded DNA
virus from a sea turtle fibropapilloma by using viral metagenomics.
J Virol. 83:2500–2509. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Ng TF, Suedmeyer WK, Wheeler E, Gulland F
and Breitbart M: Novel anellovirus discovered from a mortality
event of captive California sea lions. J Gen Virol. 90:1256–1261.
2009. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Bendinelli M, Pistello M, Maggi F, Fornai
C, Freer G and Vatteroni ML: Molecular properties, biology and
clinical implications of TT virus, a recently identified widespread
infectious agent of humans. Clin Microbiol Rev. 14:98–113. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Ishikawa T, Hamano Y and Okamoto H:
Frequent detection of TT virus in throat swabs of pediatric
patients. Infection. 27:2981999.PubMed/NCBI
|
|
14
|
Sugiyama K, Goto K, Ando T, Mizutani F,
Terabe K, Kawabe Y and Wada Y: Route of TT virus infection in
children. J Med Virol. 59:204–207. 1999. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Peng YH, Nishizawa T, Takahashi M,
Ishikawa T, Yoshikawa A and Okamoto H: Analysis of the entire
genomes of thirteen TT virus variants classifiable into the fourth
and fifth genetic groups, isolated from viremic infants. Arch
Virol. 147:21–41. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Luo K, He H, Liu Z, Liu D, Xiao H, Jiang
X, Liang W and Zhang L: Novel variants related to TT virus
distributed widely in China. J Med Virol. 67:118–126. 2002.
View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Hallett RL, Clewley JP, Bobet F, McKiernan
PJ and Teo CG: Characterization of a highly divergent TT virus
genome. J Gen Virol. 81:2273–2279. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Hijikata M, Takahashi K and Mishiro S:
Complete circular DNA genome of a TT virus variant (isolate name
SANBAN) and 44 partial ORF2 sequences implicating a great degree of
diversity beyond genotypes. Virology. 260:17–22. 1999. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Okamoto H, Akahane Y, Ukita M, Fukuda M,
Tsuda F, Miyakawa Y and Mayumi M: Fecal excretion of a nonenveloped
DNA virus (TTV) associated with posttransfusion non-A-G hepatitis.
J Med Virol. 56:128–132. 1998. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Maggi F, Andreoli E, Lanini L, Fornai C,
Vatteroni M, Pistello M, Presciuttini S and Bendinelli M:
Relationships between total plasma load of torquetenovirus (TTV)
and TTV genogroups carried. J Clin Microbiol. 43:4807–4810. 2005.
View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Matsubara H, Michitaka K, Horiike N,
Kihana T, Yano M, Mori T and Onji M: Existence of TT virus DNA and
TTV-like mini virus DNA in infant cord blood: Mother-to-neonatal
transmission. Hepatol Res. 21:280–287. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Tanaka Y, Primi D, Wang RY, Umemura T, Yeo
AE, Mizokami M, Alter HJ and Shih JW: Genomic and molecular
evolutionary analysis of a newly identified infectious agent (SEN
virus) and its relationship to the TT virus family. J Infect Dis.
183:359–367. 2001. View
Article : Google Scholar : PubMed/NCBI
|
|
23
|
Okamoto H, Nishizawa T, Ukita M, Takahashi
M, Fukuda M, Iizuka H, Miyakawa Y and Mayumi M: The entire
nucleotide sequence of a TT virus isolate from the United States
(TUS01): Comparison with reported isolates and phylogenetic
analysis. Virology. 259:437–448. 1999. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Muljono DH, Nishizawa T, Tsuda F,
Takahashi M and Okamoto H: Molecular epidemiology of TT virus (TTV)
and characterization of two novel TTV genotypes in Indonesia. Arch
Virol. 146:1249–1266. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Biagini P: Classification of TTV and
related viruses (anelloviruses). Curr Top Microbiol Immunol.
331:21–33. 2009.PubMed/NCBI
|
|
26
|
Niel C, Saback FL and Lampe E: Coinfection
with multiple TT virus strains belonging to different genotypes is
a common event in healthy Brazilian adults. J Clin Microbiol.
38:1926–1930. 2000.PubMed/NCBI
|
|
27
|
Devalle S and Niel C: Distribution of TT
virus genomic groups 1–5 in Brazilian blood donors, HBV carriers
and HIV-1-infected patients. J Med Virol. 72:166–173. 2004.
View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Biagini P, Gallian P, Cantaloube JF,
Attoui H, de Micco P and de Lamballerie X: Distribution and genetic
analysis of TTV and TTMV major phylogenetic groups in French blood
donors. J Med Virol. 78:298–304. 2006. View Article : Google Scholar : PubMed/NCBI
|