Introduction
Colorectal cancer (CRC) is one of the leading causes
of cancer-related death worldwide. An assessment of prognosis based
on features of the resected tumor would permit treating physicians
to qualify the benefit of adjuvant chemotherapy to individual
patients. Currently, anatomic and pathologic staging is still the
most accurate predictor of patient outcome. It would be valuable to
supplement standard clinical and pathologic staging using molecular
markers to more precisely define the subset of patients at highest
or lowest risk of relapse following CRC surgery. This would
facilitate better selection of patients who would benefit most from
adjuvant therapy. One of the most promising molecular markers is
the presence of tumor microsatellite instability (1). We previously reported that expression
of Reg IV and h-prune are prognostic makers for CRC (2,3);
however, these markers cannot completely identify which patients
are at low or high risk for disease recurrence. Therefore,
identification of better prognostic markers for patients with CRC
is important.
We previously performed Serial Analysis of Gene
Expression (SAGE) on four primary gastric cancer samples (4) and identified several gastric
cancer-specific genes (5). Of
these genes, olfactomedin 4 (OLFM4, also known as
GW112 or hGC-1) is a candidate gene for cancer-specific expression,
at least in patients with gastric cancer. OLFM4 was
originally cloned from human hematopoietic myeloid cells (6). Although OLFM4 is predominantly
expressed in bone marrow, the small intestine, colon and prostate
(6), levels of expression are much
lower in normal tissues than in gastric cancer tissues (5). Enhanced olfactomedin 4 expression has
been reported in gastric cancer by Northern blot analysis (7) and by immunostaining (8). Our previous immunohistochemical
analysis revealed that olfactomedin 4 is expressed in 56% of
gastric cancer tissues (9). In
addition, olfactomedin 4 is a secreted protein, and we showed that
serum olfactomedin 4 represents a novel biomarker for gastric
cancer (9). In CRC patients,
preoperative serum levels of olfactomedin 4 were increased in a
small number of samples, and the sensitivities of serum
olfactomedin 4 at stage I–III were lower than those of CEA
(9).
In addition to gastric cancer, OLFM4 mRNA and
olfactomedin 4 protein overexpression have been reported in CRC
(10,11). Olfactomedin 4 inhibits apoptosis
and may have significant roles in the development of cancer
(7). It has been proposed that
olfactomedin 4 can serve as a useful marker for stem cells in the
human small intestine and colon (12). In contrast to these observations,
immunohistochemical analysis has demonstrated that olfactomedin 4
down-regulation is found in late stage CRC cases and in CRC
patients with shorter survival (13). The morphology and actin
distribution of the HT-29 CRC cell line was altered by forced
expression of olfactomedin 4. Forced expression of olfactomedin 4
did not change cell proliferation, but decreased cell adhesion and
migration (13). Our previous
immunohistochemical analysis in gastric cancer revealed that
patients with olfactomedin 4-positive gastric cancer had a better
survival rate than patients with olfactomedin 4-negative gastric
cancer. These results suggest that olfactomedin 4 can inhibit tumor
progression. Thus, the clinical significance of olfactomedin 4
expression in human cancers is controversial and still unclear.
Although immunohistochemical analysis of
olfactomedin 4 has been performed in CRC (13), this study was performed using
tissue microarray. Therefore, detailed expression and distribution
of olfactomedin 4 in CRC has not yet been investigated. In the
present study, we examined the expression and distribution of
olfactomedin 4 in CRC by immunohistochemistry and the relationship
between olfactomedin 4 staining and clinicopathologic
characteristics.
Materials and methods
Tissue samples
In a retrospective study design, 176 primary tumors
were collected from patients diagnosed with CRC who underwent
surgery at Hiroshima University Hospital (Hiroshima, Japan). All
patients underwent curative resection. Only patients without
preoperative radiotherapy or chemotherapy were enrolled in the
study. The patients were comprised of 105 men and 71 women. The
mean age was 63 years (range, 29–89 years). Postoperative follow-up
was scheduled every 1, 2 or 3 months during the first 2 years after
surgery and every 6 months thereafter, unless more frequent
follow-up was deemed necessary. Chest X-rays, chest computed
tomography scans and serum chemistries were performed at every
follow-up visit. Recurrence was evaluated from records at Hiroshima
University Hospital. For immunohistochemical analysis, we used
archival formalin-fixed, paraffin-embedded tissues. Histologic
classification was based on the World Health Organization system.
Tumor staging was performed according to the TNM stage grouping
system (14). Since written
informed consent was not obtained, for strict privacy protection,
identifying information for all samples was removed before
analysis; this procedure is in accordance with the Ethical
Guidelines for Human Genome/Gene Research enacted by the Japanese
Government.
Immunohistochemistry
From each patient, one or two representative tumor
blocks, including the tumor center, invading front and
tumor-associated non-neoplastic mucosa, were examined by
immunohistochemistry. In cases of large, late-stage tumors, two
different sections were examined to include representative areas of
the tumor center as well as of the lateral and deep tumor invasive
front. Olfactomedin 4 was detected immunohistochemically with a
monoclonal antibody raised in our laboratory (9). The specificity of the
anti-olfactomedin 4 antibody has been characterized in detail
(9). A Dako Envision+ Mouse
Peroxidase Detection System (Dako Cytomation, Carpinteria, CA, USA)
was used for immunohistochemical analysis as described previously
(9). In brief, antigen retrieval
was carried out by microwave heating in citrate buffer (pH 6.0) for
30 min. After peroxidase activity was blocked with 3%
H2O2-methanol for 10 min, sections were
incubated with normal goat serum (Dako Cytomation) for 20 min to
block nonspecific antibody binding sites. Sections were incubated
with primary antibody against olfactomedin 4 (1:50) for 1 h at room
temperature, followed by incubations with Envision+ anti-mouse
peroxidase for 1 h. Staining was completed with a 10-min incubation
with the substratechromogen solution. Sections were counterstained
with 0.1% hematoxylin. Negative controls were created by omission
of the primary antibody.
Statistical methods
Correlations between clinicopathologic parameters
and olfactomedin 4 expression were analyzed by the Chi-square test.
Kaplan-Meier survival curves were constructed for olfactomedin
4-positive and olfactomedin 4-negative patients. Survival rates
were compared between olfactomedin 4-positive and olfactomedin
4-negative groups. Differences between survival curves were tested
for statistical significance by the log-rank test (15). The Cox proportional hazards
multivariate model was used to examine the association of clinical
and pathologic factors and the expression of olfactomedin 4 with
survival. A P-value of <0.05 was considered statistically
significant.
Results
Expression and distribution of
olfactomedin 4 in CRC and peritumoral mucosa
We performed immunohistochemical analysis of
olfactomedin 4 in 176 human CRC samples. In CRC tissue,
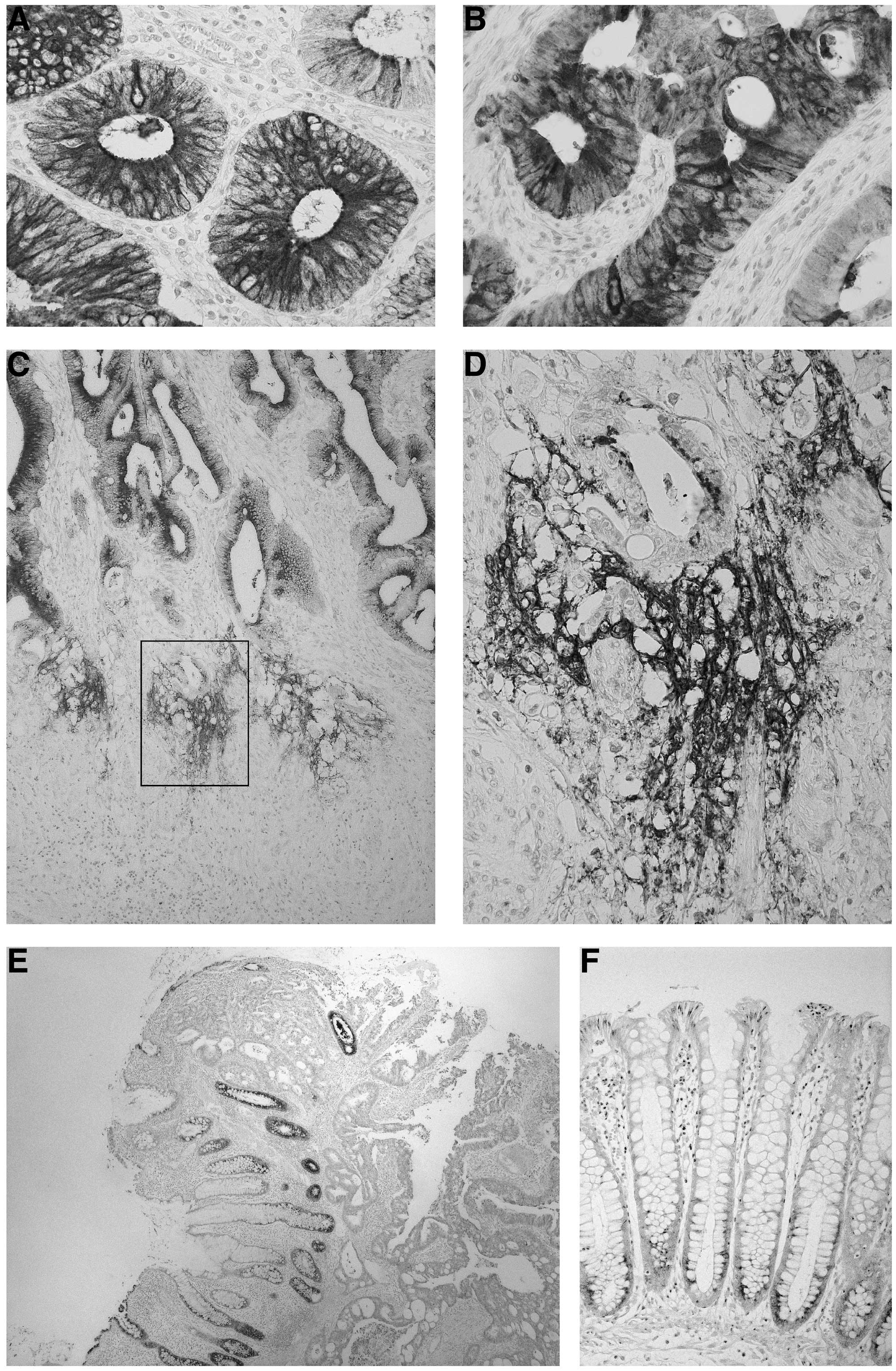
olfactomedin 4 staining was frequently observed in
well-differentiated (Fig. 1A) and
moderately differentiated adenocarcinoma (Fig. 1B). In general, staining for
olfactomedin 4 was detected in the cytoplasm of tumor cells. The
percentage of olfactomedin 4-stained tumor cells ranged from 0 to
80%. It has been reported that a loss/reduction in olfactomedin 4
expression at the front of the invasion is observed in CRC
(13); however, the tendency for
loss of olfactomedin 4 expression at the invasive front was not
observed. In our previous immunohistochemical analysis of gastric
cancer (9), in addition to
cytoplasmic staining, extracellular staining of olfactomedin 4 was
observed. In CRC tissues, extracellular staining of olfactomedin 4
was also observed. Extracellular staining of olfactomedin 4 was
focal, and in general, extracellular staining of olfactomedin 4 was
observed at the invasive front (Fig.
1C). The immunoreactivity for olfactomedin 4 was irregular and
fibrous around tumor cells scattered in the stroma (Fig. 1D).
We then focused on the peritumoral mucosa of CRC.
Notably, strong and extensive olfactomedin 4 staining was detected,
and all peritumoral mucosa samples in the 176 CRC cases were
positive for olfactomedin 4 regardless of the olfactomedin 4
staining in tumor cells. Olfactomedin 4 staining decreased
gradually, moving away from the CRC tissue. In the mucosa closest
to the tumor tissue, almost all epithelial cells showed
olfactomedin 4 staining (Fig. 1E).
In contrast, in the mucosa distant from the tumor tissue, few
epithelial cells showed olfactomedin 4 staining (Fig. 1F). Olfactomedin 4 was expressed in
the basal crypt epithelium in the colon.
Relationship between olfactomedin 4
staining and clinicopathologic characteristics
The relationship of olfactomedin 4 staining with
clinicopathologic characteristics was investigated (Table I). The level of olfactomedin 4
immunoreactivity was first evaluated in tumor cells. When >10%
of tumor cells were stained, the immunostaining was considered
positive for olfactomedin 4. In total, 59 (34%) of the 176 CRC
cases were positive for olfactomedin 4. Olfactomedin 4-positive CRC
cases showed earlier T classification (P=0.0180), N classification
(P=0.0149) and stage (P=0.0144, all by the Chi-square test) than
olfactomedin 4-negative CRC cases (Table I). Olfactomedin 4 staining was not
correlated with age, gender, tumor location, M classification, or
histologic classification. We also examined the relation between
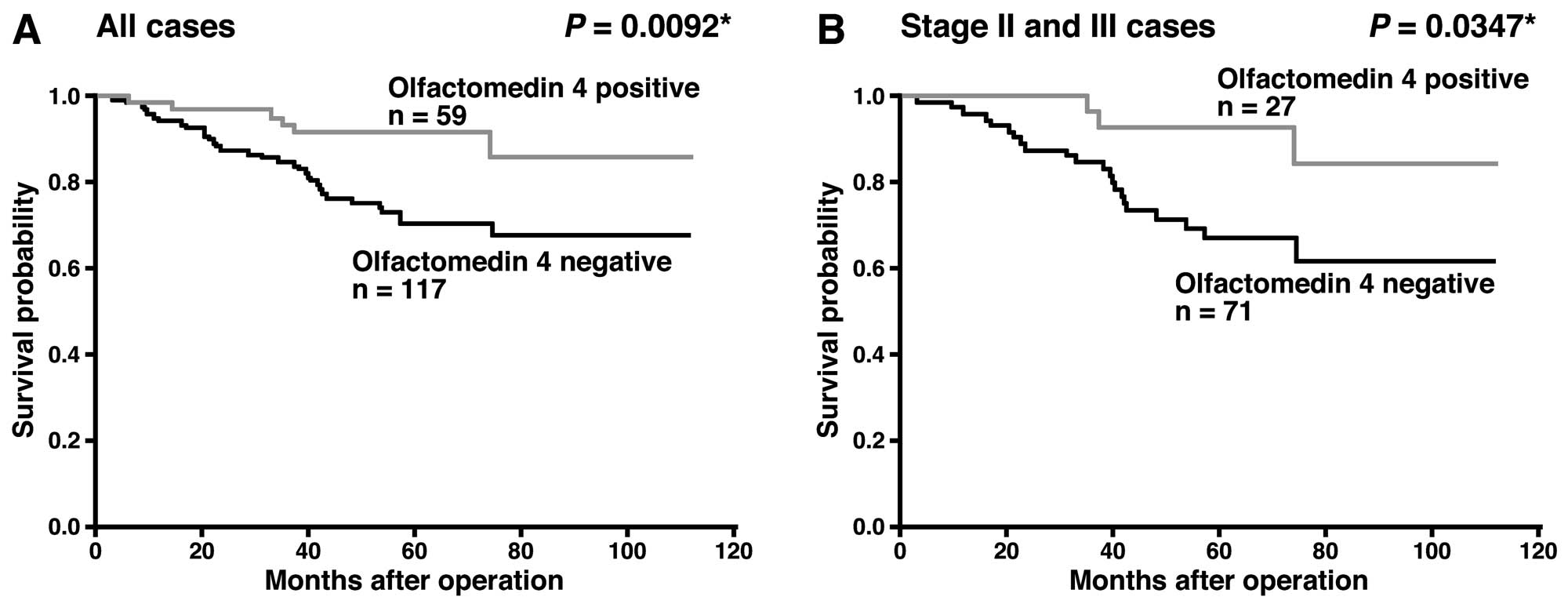
survival and olfactomedin 4 staining in CRC. In the 176 CRC
patients, those with olfactomedin 4-positive CRC had a better
survival rate than patients with olfactomedin 4-negative CRC
(P=0.0092, log-rank test) (Fig.
2A). It is well known that patients with CRC at stage I have a
favorable rate of survival, whereas patients with CRC at stage IV
show a poor rate of survival. However, it is difficult to predict
the survival of patients with stage II or stage III CRC. Therefore,
we analyzed the prognostic value of olfactomedin 4 in patients with
stage II and III CRC. In stage II and III CRC patients (n=98),
those with olfactomedin 4-positive CRC had a better survival rate
than patients with olfactomedin 4-negative CRC (P=0.0347, log-rank
test) (Fig. 2B). We then used Cox
proportional hazards multivariate model to examine the association
of clinicopathologic factors and expression of olfactomedin 4 with
survival. Multivariate analysis indicated that T classification, M
classification and olfactomedin 4 expression were independent
predictors of survival in patients with CRC (Table II).
 | Table ICorrelation of olfactomedin 4
expression with clinicopathologic characteristics of 176 CRC
cases. |
Table I
Correlation of olfactomedin 4
expression with clinicopathologic characteristics of 176 CRC
cases.
| Olfactomedin 4
expression
| |
|---|
| Positive | Negative | P-valuea |
|---|
| Age | | | 0.9551 |
| ≤65 | 31 (33%) | 62 | |
| >65 | 28 (34%) | 55 | |
| Gender | | | 0.7943 |
| Male | 36 (34%) | 69 | |
| Female | 23 (32%) | 48 | |
| Tumor location | | | 0.8075 |
|
Right/transverse | 12 (35%) | 22 | |
|
Left/sigmoid/rectum | 47 (33%) | 95 | |
| T classification | | | 0.0180 |
| T1 | 14 (47%) | 16 | |
| T2 | 15 (39%) | 23 | |
| T3 | 24 (31%) | 54 | |
| T4 | 6 (20%) | 24 | |
| N classification | | | 0.0149 |
| N0 | 43 (41%) | 63 | |
| N1 | 16 (23%) | 54 | |
| M classification | | | 0.6085 |
| M0 | 53 (34%) | 102 | |
| M1 | 6 (29%) | 15 | |
| Stage | | | 0.0144 |
| I | 26 (46%) | 31 | |
| II | 16 (36%) | 29 | |
| III | 11 (21%) | 42 | |
| IV | 6 (29%) | 15 | |
| Histological
classification | | | 0.1973 |
|
Well/moderately | 58 (35%) | 110 | |
|
Poorly/mucinous | 1 (13%) | 7 | |
 | Table IIMultivariate analysis of factors that
influence survival of patients with CRC. |
Table II
Multivariate analysis of factors that
influence survival of patients with CRC.
| Factor | Hazard ratio | (95% CI) | Chi-square test | P-valuea |
|---|
| Age | | | 1.586 | 0.2078 |
| ≤65 | 1 | (Reference) | | |
| >65 | 1.511 | (0.795–2.871) | | |
| Gender | | | 0.708 | 0.4001 |
| Male | 1 | (Reference) | | |
| Female | 0.748 | (0.381–1.470) | | |
| Tumor location | | | 0.368 | 0.5441 |
|
Right/transverse | 1 | (Reference) | | |
|
Left/sigmoid/rectum | 0.745 | (0.287–1.931) | | |
| T
classification | | | 5.995 | 0.0143 |
| T1/2 | 1 | (Reference) | | |
| T3/4 | 4.803 | (1.367–16.869) | | |
| N
classification | | | 3.488 | 0.0618 |
| N0 | 1 | (Reference) | | |
| N1 | 7.327 | (0.906–59.248) | | |
| M
classification | | | 11.161 | 0.0008 |
| M0 | 1 | (Reference) | | |
| M1 | 3.631 | (1.704–7.737) | | |
| Stage | | | 1.032 | 0.3097 |
| I/II | 1 | (Reference) | | |
| III/IV | 3.254 | (0.334–31.713) | | |
| Histologic
classification | | | 0.002 | 0.9632 |
| Well/moderately
differentiated | 1 | (Reference) | | |
| Poorly
differentiated /mucinous | 0.936 | (0.194–4.773) | | |
| Olfactomedin 4
expression | | | 4.486 | 0.0342 |
| Positive | 1 | (Reference) | | |
| Negative | 2.725 | (1.078–6.890) | | |
The level of olfactomedin 4 immunoreactivity was
also evaluated in the tumor-associated stroma. Since extracellular
staining of olfactomedin 4 at the invasive front was frequently
observed, stromal olfactomedin 4 staining was considered positive
when extracellular staining of olfactomedin 4 was stained at the
invasive front. In total, 29 (16%) of the 176 CRC cases were
positive for stromal olfactomedin 4. Stromal olfactomedin 4
staining was not correlated with age, gender, tumor location, T
classification, N classification, M classification, stage, or
histologic classification (data not shown). In the 176 CRC
patients, survival rate was not statistically different between
patients with stromal olfactomedin 4-positive CRC and those with
stromal olfactomedin 4-negative CRC (data not shown).
Discussion
Previously, we performed SAGE on four primary
gastric cancers (4) and identified
several gastric cancer-specific genes (5). Of these genes, olfactomedin 4 is a
candidate gene for cancer-specific expression. In the present
study, we examined the expression and distribution of olfactomedin
4 in CRC by immunohistochemistry and the relationship between
olfactomedin 4 staining and clinicopathologic characteristics.
Although few epithelial cells in colonic mucosa distant from the
CRC tissue showed olfactomedin 4 staining, strong and extensive
olfactomedin 4 staining was found in 34% cases of CRC, and
olfactomedin 4-positive CRC cases showed earlier T classification,
N classification and stage than olfactomedin 4-negative CRC cases.
These results are consistent with results reported previously that
olfactomedin 4 expression is up-regulated in early stage CRC and
down-regulated in advanced stage CRC (13). In our previous study in gastric
cancer, olfactomedin 4-positive cases were found frequently in
early stage cases (9). Taken
together, expression of olfactomedin 4 is an early event, and
loss/reduction of olfactomedin 4 expression is a late event in
gastrointestinal malignancies.
It is generally accepted that apoptosis suppresses
oncogenic transformation. The ability of tumor cell populations to
expand in number is determined, not only by the rate of cell
proliferation, but also by the rate of cell attrition. Apoptosis
represents a major source of this attrition (16). Thus, resistance to apoptosis is a
hallmark of most and perhaps all types of cancer. It has been
reported that olfactomedin 4 interacts with GRIM-19 to attenuate
retinoic acid and interferon β-mediated cellular apoptosis, and
transient expression of olfactomedin 4 promoted tumor growth in
C57/BL/6 mice (7). Therefore,
expression of olfactomedin 4 may contribute to carcinogenesis by
resistance to apoptosis at least in early stage CRC. In contrast,
forced expression of olfactomedin 4 in an HT-29 cell line decreased
cell adhesion and migration (13).
Therefore, it is possible that loss/reduction of olfactomedin 4
expression induces tumor cell invasion in late stage CRC cases.
In the present study, univariate and multivariate
analyses revealed that negative expression of olfactomedin 4 is a
prognostic indicator. Furthermore, negative expression of
olfactomedin 4 correlated with a short survival rate in stage II
and III CRC cases. Patients diagnosed with stage II or III CRC have
variable prognoses, and they are the group that would benefit most
from discovery of a prognostic factor that can identify individuals
for whom adjuvant treatment would be most advantageous. To clarify
whether olfactomedin 4 immunostaining is useful for identification
of patients most likely to benefit from adjuvant treatment,
association between olfactomedin 4 staining and response to
adjuvant therapies should be investigated.
In addition to cytoplasmic olfactomedin 4 staining,
extracellular staining was also observed. Extracellular staining of
olfactomedin 4 at the invasive front was frequently observed.
Observation of the invasive front is important in the analysis of
tumor cells, since it reflects the invasive potential of tumor
cells. It has been reported that the expression of matrilysin in
the invasive front is a promising biomarker predicting nodal
metastasis of CRC (17).
Overexpression of heparanase at the invasive front has been
reported in gastric cancer and high expression of heparanase was a
strong predictor of poor survival (18). These results indicate that the
proteolytic degradation of the extracellular matrix by these
molecules is one of the most important mechanisms in tumor
progression, and the proteolytic degradation occurs at the invasive
front. Although there was no correlation between stromal expression
of olfactomedin 4 and clinicopathologic characteristics, stromal
expression of olfactomedin 4 at the invasive front may partly
contribute to the malignant behavior of CRC, such as local
invasiveness.
Notably, extensive olfactomedin 4 staining was
observed in the peritumoral mucosa of CRC, and olfactomedin 4
staining decreased gradually, moving away from the tumor tissue. It
is well known that the peritumoral mucosa of CRC is often
hyperplastic, and various growth factors, such as transforming
growth factor-α and basic fibroblast growth factor, are increased
in the peritumoral mucosa (19).
Since expression of olfactomedin 4 in the crypt epithelium of
inflamed colonic mucosa has been reported (20), olfactomedin 4 expression may be
induced by growth factors and may function as an antiapoptotic
factor in the peritumoral mucosa of CRC.
In summary, we showed that olfactomedin 4 is a
valuable marker for long survival in patients with CRC. However,
the significance of extracellular staining of olfactomedin 4 at the
invasive front and extensive olfactomedin 4 staining in the
peritumoral mucosa of CRC remains unclear. Since olfactomedin 4 is
a secreted protein, identification of a cell surface receptor for
olfactomedin 4 will further improve our understanding of the basic
biology of olfactomedin 4.
Acknowledgements
We thank Mr Shinichi Norimura for the
excellent technical assistance and advice. We thank the Analysis
Center of Life Science, Hiroshima University for the use of their
facilities. This work was supported, in part, by Grants-in-Aid for
Cancer Research from the Ministry of Education, Culture, Science,
Sports and Technology of Japan, in part, by a Grant-in-Aid for the
Third Comprehensive 10-Year Strategy for Cancer Control and for
Cancer Research from the Ministry of Health, Labour and Welfare of
Japan and in part by a grant (07-23911) from the Princess Takamatsu
Cancer Research Fund.
References
|
1
|
Popat S, Hubner R and Houlston RS:
Systematic review of microsatellite instability and colorectal
cancer prognosis. J Clin Oncol. 23:609–618. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Oue N, Kuniyasu H, Noguchi T, et al: Serum
concentration of Reg IV in patients with colorectal cancer:
overexpression and high serum levels of Reg IV are associated with
liver metastasis. Oncology. 72:371–380. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Kobayashi T, Hino S, Oue N, Asahara T,
Zollo M, Yasui W and Kikuchi A: Glycogen synthase kinase 3 and
h-prune regulate cell migration by modulating focal adhesions. Mol
Cell Biol. 26:898–911. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Oue N, Hamai Y, Mitani Y, et al: Gene
expression profile of gastric carcinoma: identification of genes
and tags potentially involved in invasion, metastasis and
carcinogenesis by serial analysis of gene expression. Cancer Res.
64:2397–2405. 2004. View Article : Google Scholar
|
|
5
|
Aung PP, Oue N, Mitani Y, et al:
Systematic search for gastric cancer-specific genes based on SAGE
data: melanoma inhibitory activity and matrix metalloproteinase-10
are novel prognostic factors in patients with gastric cancer.
Oncogene. 25:2546–2557. 2006. View Article : Google Scholar
|
|
6
|
Zhang J, Liu WL, Tang DC, et al:
Identification and characterization of a novel member of
olfactomedin-related protein family, hGC-1, expressed during
myeloid lineage development. Gene. 283:83–93. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Zhang X, Huang Q, Yang Z, Li Y and Li CY:
GW112, a novel antiapoptotic protein that promotes tumor growth.
Cancer Res. 64:2474–2481. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Liu W, Zhu J, Cao L and Rodgers GP:
Expression of hGC-1 is correlated with differentiation of gastric
carcinoma. Histopathology. 51:157–165. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Oue N, Sentani K, Noguchi T, et al: Serum
olfactomedin 4 (GW112, hGC-1) in combination with Reg IV is a
highly sensitive biomarker for gastric cancer patients. Int J
Cancer. 125:2383–2392. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Koshida S, Kobayashi D, Moriai R, Tsuji N
and Watanabe N: Specific overexpression of OLFM4 (GW112/HGC-1) mRNA
in colon, breast and lung cancer tissues detected using
quantitative analysis. Cancer Sci. 98:315–320. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Conrotto P, Roesli C, Rybak J, et al:
Identification of new accessible tumor antigens in human colon
cancer by ex vivo protein biotinylation and comparative mass
spectrometry analysis. Int J Cancer. 123:2856–2864. 2008.
View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Van der Flier LG, Haegebarth A, Stange DE,
van de Wetering M and Clevers H: OLFM4 is a robust marker for stem
cells in human intestine and marks a subset of colorectal cancer
cells. Gastroenterology. 137:15–17. 2009.PubMed/NCBI
|
|
13
|
Liu W, Liu Y, Zhu J, Wright E, Ding I and
Rodgers GP: Reduced hGC-1 protein expression is associated with
malignant progression of colon carcinoma. Clin Cancer Res.
14:1041–1049. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Sobin LH and Wittekind CH: TNM
Classification of Malignant Tumors. 6th edition. Wiley-Liss; New
York: pp. 65–68. 2002
|
|
15
|
Mantel N: Evaluation of survival data and
two new rank order statistics arising in its consideration. Cancer
Chemother Rep. 50:163–170. 1966.PubMed/NCBI
|
|
16
|
Hanahan D and Weinberg RA: The hallmarks
of cancer. Cell. 100:57–70. 2000. View Article : Google Scholar
|
|
17
|
Kurokawa S, Arimura Y, Yamamoto H, et al:
Tumour matrilysin expression predicts metastatic potential of stage
I (pT1) colon and rectal cancers. Gut. 54:1751–1758. 2005.
View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Takaoka M, Naomoto Y, Ohkawa T, et al:
Heparanase expression correlates with invasion and poor prognosis
in gastric cancers. Lab Invest. 83:613–622. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Kuniyasu H, Yasui W, Shinohara H, et al:
Induction of angiogenesis by hyperplastic colonic mucosa adjacent
to colon cancer. Am J Pathol. 157:1523–1535. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Shinozaki S, Nakamura T, Iimura M, et al:
Upregulation of Reg 1 alpha and GW112 in the epithelium of inflamed
colonic mucosa. Gut. 48:623–629. 2001. View Article : Google Scholar : PubMed/NCBI
|