Introduction
Erythropoietin (EPO) is the core regulator of red
blood cell production (1). The
binding of EPO to functional erythropoietin (EPO) receptor (EPOR)
homodimer triggers several downstream signaling pathways, such as
the Jak2/signal transducer and activator of transcription 5
(STAT5), phosphatidylinositol 3-kinase (PI3K)/protein kinase B
(Akt), Ras/mitogen-activated protein kinase (MAPK) and protein
kinase C pathways (2), resulting
in the proliferation and differentiation of erythroid progenitors.
Apart from erythropoietic cells, EPOR is expressed in many
non-hematopoietic tissues, as well as in cancer (3–5).
The expression of EPO and functional EPOR has also been confirmed
in breast cancer tissue and cell lines (6,7),
the most common type of cancer among females.
Recombinant human EPO (rHuEPO) is used to correct
anemia in chronic kidney disease (CKD) patients (8), chemotherapy-induced anemia (9), as well as other types of anemia. A
number of pre-clinical and clinical trials have verified the use of
rHuEPO as a tissue protective agent in the brain, heart and kidneys
(10–12). The protective effect of EPO has
been proposed to be mediated through the EPOR hetero-receptor,
presumably formed with the common β subunit of the IL-3 receptor
[colony stimulating factor 2 receptor, β, low-affinity (CSF2RB)]
(13). The significance of
receptor tyrosine kinase ephrin type-B receptor 4 (EPHB4) in cancer
development has also been indicated (14), having the potential to be another
EPOR receptor partner (15).
EPHB4 has been shown to be expressed in breast cancer (16). The effect of rHuEPO supportive
care in cancer patients with EPOR-positive cancer is not yet well
understood. Several clinical trials were terminated early due to
rHuEPO-negative effects on recurrence-free survival (RFS) and
overall survival or tumor progression (5,17,18). Decreased patient survival may
result from the rHuEPO effects on increased thrombotic events, the
growth and survival of cancer cells or the attenuated sensitivity
of cancer cells to various types of treatment (5). Nevertheless, the presence of EPO and
EPOR in various cancer tissues and cell lines raises the issue of
whether the use of rHuEPO as supportive care in cancer patients is
appropriate (19).
Breast cancer cells express a variety of growth
factor receptors determining the molecular classification of the
disease (20). The most commonly
expressed are the steroid hormone receptors, estrogen receptor
(ESR), progesterone receptor (PGR) and androgen receptor (AR).
Apart from the classical steroid hormone receptors, that are
members of the nuclear receptor family (ESR, PGR and AR) and upon
which the molecular classification is based, membrane-bound steroid
receptors have been identified (mESR, mPGR and mAR).
Membrane-initiated steroid signaling (MISS) has been implicated in
intracellular signaling (21,22) either in a non-transcriptional
fashion by modifying existing proteins (e.g., phosphorylation) or
by the modulation of gene expression and the production of proteins
(23,24). A cross-talk between membrane and
nuclear steroid receptors has also been indicated which leads to
the amplification of subsequent transcription originating from the
nuclear receptors (25). The
identity of cytoplasmic/membrane steroid receptors has been an
issue of debate. Classical nuclear receptors, ER-α (ESR1) and the
PGRs (PGRA and PGRB), have been observed at the plasma membrane of
breast cancer cells. The palmitoylation of the classical receptors
has been shown to be the signal for the membrane localization and
all membrane-initiated signaling (26). In contrast to classical receptors,
the membrane associated ESR is an atypical G protein-coupled
receptor (GPCR), selectively activating discrete G protein
α- and βγ-subunits that result in rapid signaling
(27). Extra-nuclear signaling
has been shown to modulate the proliferation, survival and invasion
of breast cancer cells (28). The
G protein-coupled estrogen receptor [GPER; also known as G
protein-coupled receptor 30 (GPR30)] has been implicated in
mediating estrogen action at the cell membrane; however, studies
have suggested that GPER may not be a membrane ESR. A role for this
orphan GPCR may involve collaboration with mESR, showing dependence
on both proteins (29).
Studies have indicated the correlation between EPOR
and estrogen and progesterone receptors in breast cancer cells and
patients. The co-expression of EPO, EPOR and mAR protein in breast
cancer patients has been shown to be negatively associated with
disease-free and overall survival (30). In addition, patients with
ESR+/PGR+ tumors and low EPOR expression have
significantly increased RFS following tamoxifen (TAM) treatment
when compared to patients with high EPOR-expressing tumors. On the
other hand, the significant improvement of RFS was detected in
non-treated patients with ESR+ tumors with high EPOR
levels (31). Volgger et
al (32) also confirmed the
positive association between EPOR expression, ESR/PGR status and
decreased local cancer recurrence. These observations suggest the
existence of the active interplay between EPO signaling and the
steroid receptors which may/can furthermore interact with other
cytokine/growth factors, such as epidermal growth factor receptor
(EGFR) and insulin-like growth factor 1 (IGF-I) receptor (33–36). The activation of these receptors
eventually culminates in the activation of MAPK and PI3K signaling
pathways which are also indicative for the receptor tyrosine kinase
EPHB4 (37). The complexity of
the intracellular cross-talk therefore makes it difficult to
decipher the role of the particular receptor and the significance
of a particular correlation.
Therefore, we aimed to assess the applicability of
breast cancer cell lines in the study of the EPO signaling pathway,
particularly as regards the correlation between the expression of
EPOR and that of other cellular receptors. Profound differences in
the expression of estrogen and progesterone receptors and estrogen
metabolizing enzymes in breast cancer cell lines (38) indicate that choosing the proper
cell model is of great importance. The breast cancer cell lines,
MCF-7, MDA-MB- 361, T-47D, MDA-MB-231, Hs578Bst, SKBR3, MCF-10A and
Hs578T were selected for detailed expression analysis of
EPOR, ESR1-2, PGR, GPER, EPHB4,
CSF2RB and EPO. The cell lines were also analyzed for
the responsiveness to rHuEPO induction on the level of cell
proliferation and the activation of EPO signaling pathways. The
phosphorylation of extracellular signal-regulated kinase
(ERK)/MAPK, Akt/PI3K and STAT5/Jak/STAT proteins was also
assessed.
Materials and methods
Cell cultures
The breast cancer cell lines (Table I) were maintained in cell culture
at 37°C in a humidified 5% (v/v) CO2 atmosphere. The
cell lines were obtained from the American Type Culture Collection
(ATCC; Manassas, VA, USA) and were cultured according to the ATCC
recommendations. The receptor status for a specific cell line and
the tumor type are shown in Table
I. Cells were grown in basic growth medium, supplemented with
10% FBS (38). UT7/Epo cells were
used as the positive control of EPO signaling in western blot
analysis. Cells were kindly provided by C. Lacout (Institute of
Cancerology Gustave Roussy, Villejuif, France) and were cultured in
MEM Alpha medium (Sigma, St. Louis, MO, USA), supplemented with 10%
FBS and 2 U/ml rHuEPO (NeoRecormon; Roche, Mannheim, Germany).
 | Table IDetails of the cohort of the selected
breast cancer lines as defined by American Type Culture
Collection. |
Table I
Details of the cohort of the selected
breast cancer lines as defined by American Type Culture
Collection.
| Cell line | Receptor
status | Tissue source | Tumor type |
|---|
| MCF-7 | ESR+,
PGR+ | PE | IDC |
| MDA-MB-361 | ESR+,
PGR− | B | AC |
| T-47D | ESR+,
PGR+ | PE | IDC |
| MDA-MB-231 | ESR+,
PGR− | PE | AC |
| Hs578Bst | ESR−,
PGR− | Adjacent breast
tissue | |
| SKBR3 | ESR−,
PGR− | PE | AC |
| MCF-10A | ESR−,
PGR− | | F |
| Hs578T | ESR−,
PGR− | P.Br | IDC |
Gene expression analysis
RNA isolation
Total RNA was extracted using TRI reagent (Sigma)
and treated with the DNase I (Roche) according to the
manufacturer’s instructions. The quality of the RNA samples was
determined using an Agilent bioanalyzer assuring that all RIN
values were >9.8. Total RNA (1 μg) was transcribed to
cDNA using SuperScript III reverse transcriptase (Invitrogen, USA)
according to the manufacturer’s instructions.
Selection of normalization gene
candidates
According to our previous study (38) peptidylprolyl isomerase A
(PPIA) was selected as the normalization gene from the full
cohort of 16 reference genes that were tested in these cell
lines.
Quantitative real-time PCR (qPCR)
The expression of 9 genes of interest (Table II) and the selected normalization
gene was analyzed using TaqMan® PCR assays. The
expression levels were determined with the exon-spanning hydrolysis
probes (FAM or VIC, dye labeled) that are commercially available as
Assays-on-Demand (Applied Biosystems, Bedford, MA, USA) with
optimized primer and probe concentrations (Table II). Gene expression assays
covering all splice variants of the specific gene were selected;
for GPER 2 assays were required in order to cover all the existing
splice variants. qPCR was performed on a 384-well platform using
the LightCycler® 480 Real-Time PCR System (Roche) and
TaqMan® Universal PCR Master Mix. Universal
thermocycling parameters were applied, as recommended by the
manufacturer (Applied Biosystems). The amplification of specific
PCR products was performed in triplicate in a total reaction
mixture of 5 μl containing 0.25 μl of cDNA template.
Minimum Information for Publication of Quantitative Real-Time PCR
Experiments (MIQE) guidelines were followed in the performance and
interpretation of the qPCR reactions (39).
 | Table IIDetails of genes of interest and
reference genes. |
Table II
Details of genes of interest and
reference genes.
| Gene symbol | Assay ID | Gene name |
|---|
| Genes of
interest |
| ESR1 | Hs00174860_m1 | Estrogen receptor 1
(ESR-α) |
| ESR2 | Hs00230957_m1 | Estrogen receptor 2
(ESR-β) |
| PGR | Hs00172183_m1 | Progesterone
receptor |
| EPOR | Hs00959427_m1 | Erythropoietin
receptor |
| EPO | Hs01071097_m1 | Erythropoietin |
| GPER1 | Hs00173506_m1 | G protein-coupled
estrogen receptor 1 |
| GPER2 | Hs01116133_m1 | G protein-coupled
estrogen receptor 2 |
| EPHB4 | Hs00174752_m1 | Ephrin type-B
receptor 4 |
| CSF2RB | Hs00166144_m1 | Colony stimulating
factor 2 receptor, β, low-affinity (granulocyte-macrophage) |
| Reference gene |
| PPIA | Hs99999904_m1 | Peptidylprolyl
isomerase A (cyclophilin A) |
Western blot analysis
The expression of ERK, Akt and STAT5 proteins and
their phosphorylated forms was determined by western blot analysis
in the cell lysates following rHuEPO treatment. Cells were seeded
on 12-well plates at the concentration of 1×104
cells/well and left in culture until they reached a confluence of
70%. Prior to treatment (24 h) the cells were switched to
serum-free medium and then treated with 5 or 25 U/ml rHuEPO for 5
and 10 min. The culture medium was then aspirated and samples were
fast-frozen in liquid nitrogen.
Cell samples were lysed for 10 min on ice in lysis
buffer as described by Kutuk et al (40) and soluble proteins were recovered
in the supernatant following a 10-min centrifugation (12,000 rpm).
Samples of UT7/Epo cells treated with 2 U/ml rHuEPO were used as
the positive controls. Equal amounts of protein (50 μg) from
each sample were loaded per well. After SDS electrophoresis,
proteins were transferred onto polyvinylidene difluoride (PVDF)
membranes (Immobilon-P; Millipore, Billerica, MA, USA). Membranes
were blocked in a blocking solution (5% BSA in 1 mM PBS, 0.2%
Tween-20) for 1 h and incubated in one of the following antibodies
and dilutions: anti-ERK (1:1,000, no. 9102), anti-Akt (1:600, no.
9272), anti-STAT5 (1:600, no. 9363), anti-P-ERK (1:1,000, no.
9101), anti-P-Akt (1:600, no. 9271) and anti-P-STAT5 (1:600, no.
9351). All antibodies were purchased from Cell Signaling Technology
(Danvers, MA, USA) and were raised against synthetic peptides in
rabbits. As a secondary antibody, peroxidase-conjugated anti-rabbit
IgG (1:5,000, A0545; Sigma) was used and visualized with
chemiluminescence reagent (Pierce ECL Western Blotting Substrate;
Thermo Scientific, Rockford, IL, USA) with a CCD camera (Fujifilm,
Tokyo, Japan). Membranes were densitometrically analyzed using
ImageJ software (National Institutes of Health, Bethesda, MD, USA)
(41) and ratios between
phosphorylated proteins to their non-phosphorylated forms were
calculated and compared between samples. Experiments were repeated
3 times.
Proliferation assays
The proliferation of the MCF-7, MDA-MB-231, SKBR-3
and Hs578T cells was determined using MTT reagent (Sigma). Cells
were seeded on a 96-well plate in 5-plicates at the concentration
of 750 cells/well and left to adhere in the medium. Basic growth
medium supplemented with 10 or 1% of FBS was used. Following a day
in culture, the cells were exposed to 5 U/ml rHuEPO; the control
cells were grown in medium without rHuEPO. Cells were maintained in
culture up to 7 days and cell proliferation was measured every day.
Fold change was calculated by normalizing the proliferation results
with the proliferation measured at day 1. Experiments were repeated
6 times.
Statistical analysis
For correlation analysis, Pearson’s correlation
coefficient was defined with SPSS 19 software (SPSS Inc., Chicago,
IL) using two-tailed analysis. Only correlations with a
significance level of P<0.05 were considered statistically
significant. Between-group linkage was determined using
hierarchical clustering analysis on an interval scale, considering
Euclidean distance. The effect of rHuEPO treatment on cell
proliferation was assessed by two-way analysis of variance (ANOVA).
P<0.05 was considered to indicate a statistically significant
difference.
Results
Expression of EPO, EPOR and its
potential receptor partners
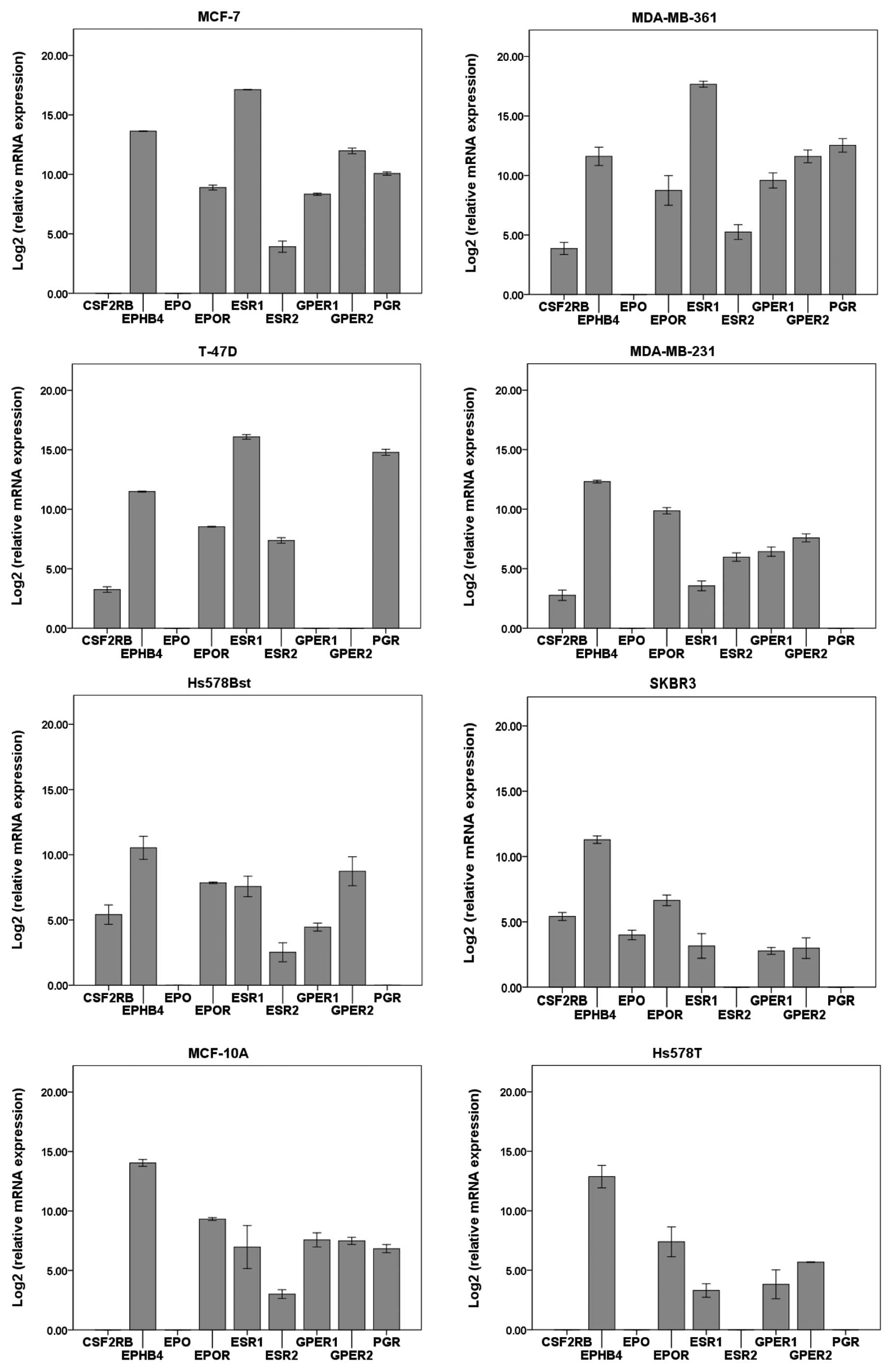
EPO expression was confirmed in the SKBR3
cell line (Fig. 1) while all the
other cell lines were negative for the expression of this gene.
EPOR and EPHB4 were expressed in all of the examined
breast cancer cell lines with EPHB4 expression levels being
substantially higher compared to those of EPOR. On the other
hand, CSF2RB was only weakly expressed in some of the
examined cell lines (Fig. 1).
Expression of genes for estrogen and
progesterone receptors
The expression of genes for ESR1, ESR2
and PGR is comparable with previously published data
(38), confirming the hormone
dependency of the MCF-7, MDA-MB-361 and T-4TD cell lines. In
addition, the MCF-7 and MDA-MB-361 cell lines showed a high
expression of GPER, while T-47D cells were negative for the
expression of this gene. Low levels of both GPER isoforms
were confirmed in the Hs578Bst, SKBR3, MCF-10A and Hs578T cell
lines (Fig. 1).
Correlation analysis
A variety of correlations in the expression of the
analyzed genes were confirmed for the cohort of the analyzed cell
lines (Table III).
 | Table IIIPearson’s correlation coefficients
(rs) calculated for the expression of cellular receptors. |
Table III
Pearson’s correlation coefficients
(rs) calculated for the expression of cellular receptors.
| Receptor | EPOR | EPHB4 | CSF2RB | ESR1 | ESR2 | PGR | GPER1 | GPER2 |
|---|
| EPOR | | | | | | | | |
| r | 1 | 0.385c | −0.708b | 0.275 | 0.276 | −0.404 | 0.549b | 0.426a |
| P-value | | 0.064 | 0.003 | 0.194 | 0.268 | 0.193 | 0.006 | 0.038 |
| N | 24 | 24 | 15 | 24 | 18 | 12 | 24 | 24 |
| EPHB4 | | | | | | | | |
| r | | 1 | −0.541a | 0.014 | −0.144 | −0.871b | 0.468a | 0.285 |
| P-value | | | 0.037 | 0.949 | 0.569 | 0.000 | 0.021 | 0.177 |
| N | | 24 | 15 | 24 | 18 | 12 | 24 | 24 |
| CSF2RB | | | | | | | | |
| r | | | 1 | −0.277 | −0.742b | −0.465 | −0.195 | −0.039 |
| P-value | | | | 0.317 | 0.006 | 0.352 | 0.486 | 0.891 |
| N | | | 15 | 15 | 12 | 6 | 15 | 15 |
| ESR1 | | | | | | | | |
| r | | | | 1 | 0.301 | 0.754b | 0.361 | 0.443a |
| P-value | | | | | 0.224 | 0.005 | 0.083 | 0.030 |
| N | | | | 24 | 18 | 12 | 24 | 24 |
| ESR2 | | | | | | | | |
| r | | | | | 1 | 0.945b | −0.356 | −0.499a |
| P-value | | | | | | 0.000 | 0.148 | 0.035 |
| N | | | | | 18 | 12 | 18 | 18 |
| PGR | | | | | | | | |
| r | | | | | | 1 | −0.539c | −0.368 |
| P-value | | | | | | | 0.071 | 0.239 |
| N | | | | | | 12 | 12 | 12 |
| GPER1 | | | | | | | | |
| r | | | | | | | 1 | 0.884b |
| P-value | | | | | | | | 0.000 |
| N | | | | | | | 24 | 24 |
| GPER2 | | | | | | | | |
| r | | | | | | | | 1 |
| P-value | | | | | | | | |
| N | | | | | | | | 24 |
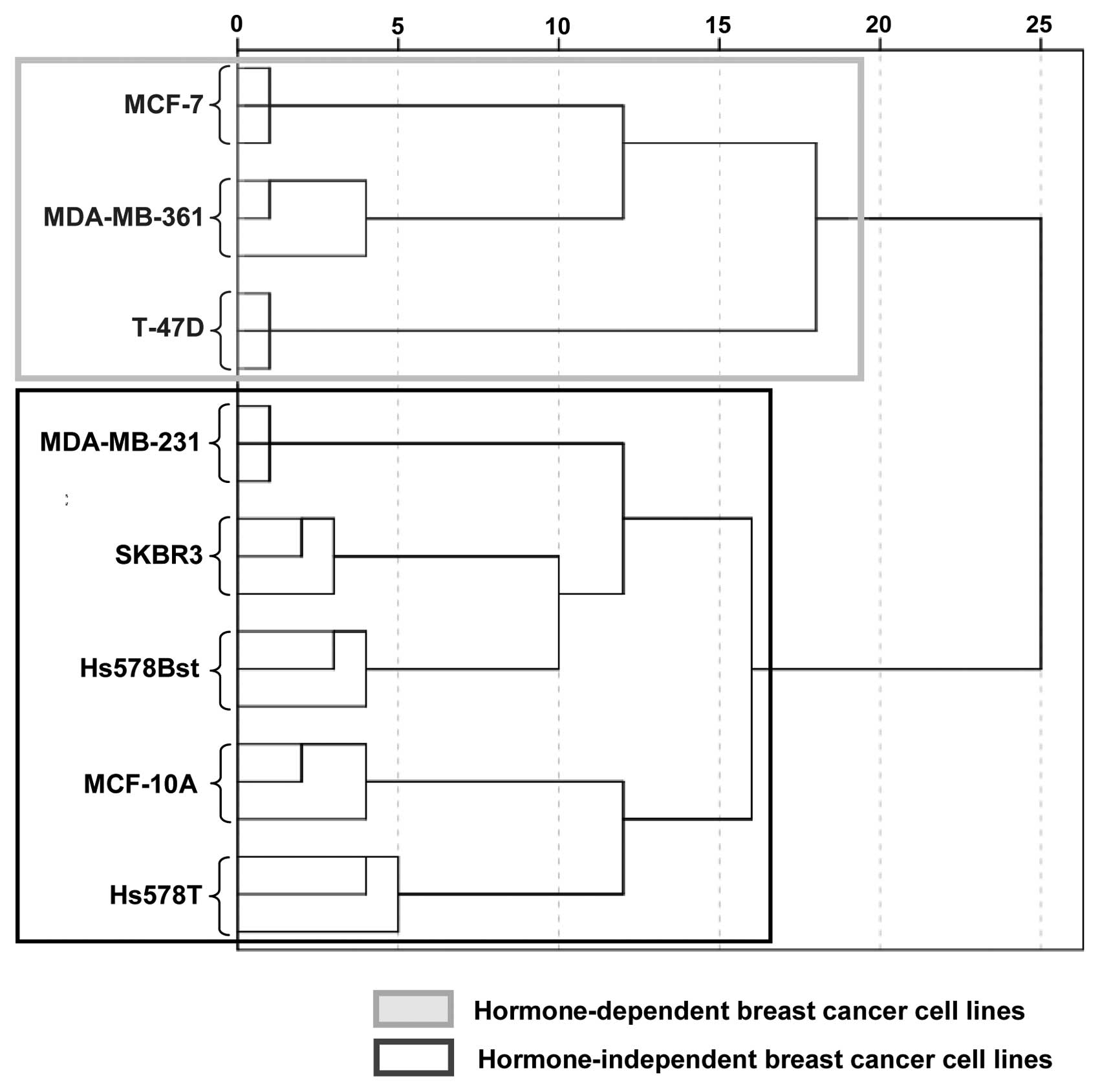
Hierarchical clustering
Based on the gene expression data, the cell lines
were clustered in 5 subgroups (Fig.
2). Principally, the cell lines were stratified in 2 main
clusters: a cluster of hormone-dependent breast cancer cell lines
expressing ESR and PGR (Fig. 2, grey box) and a cluster of
hormone-independent breast cancer cell lines (Fig. 2, white box).
Cell responsiveness to rHuEPO
induction
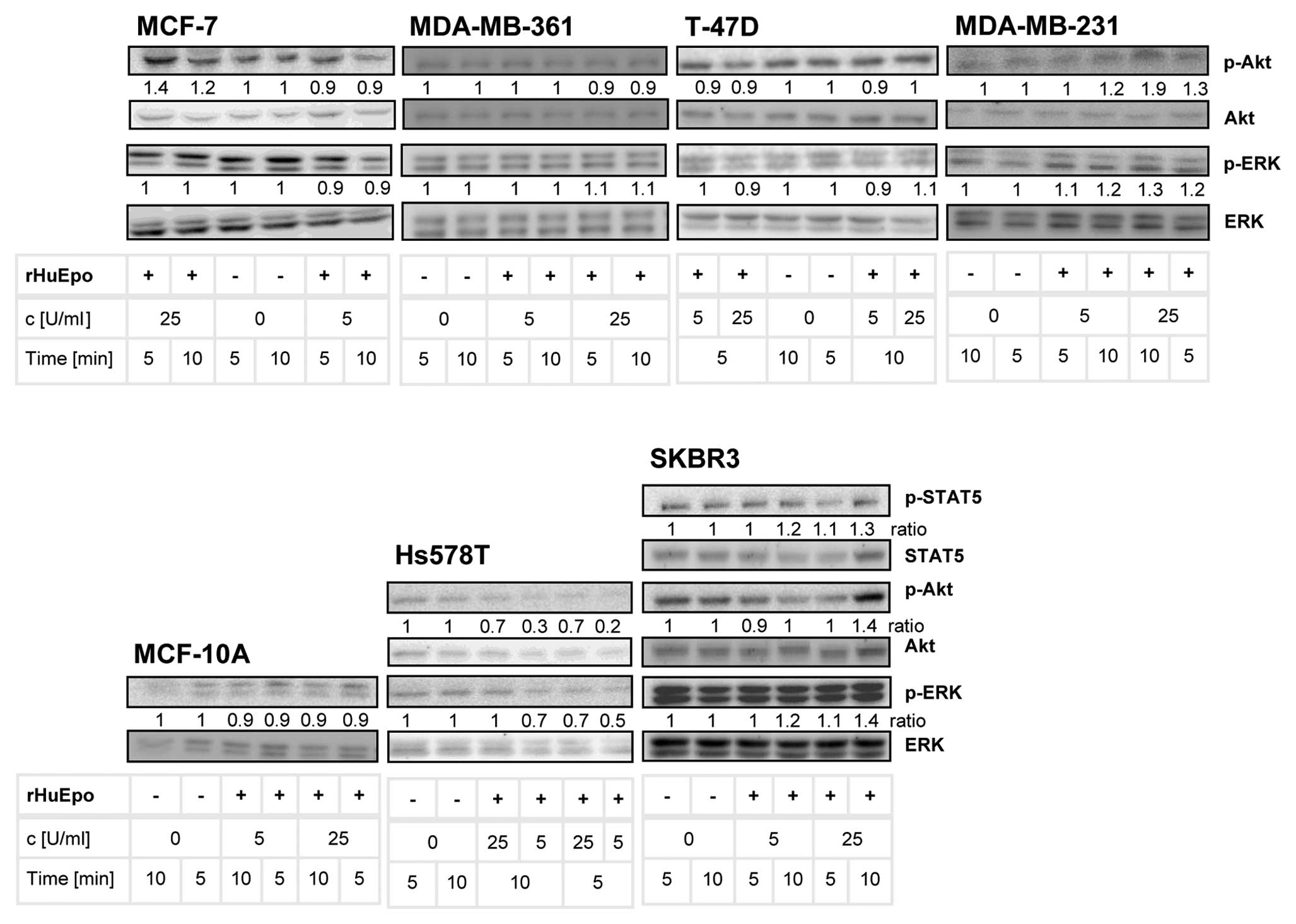
EPO involvement in the activation of
the MAPK, PI3K and STAT5 signaling pathways
EPO has been shown to promote the activation of the
MAPK, PI3K and Jak/STAT signaling pathways (2,42).
Therefore, we evaluated whether rHuEPO treatment promotes the
phosphorylation of ERK (MAPK), Akt (PI3K) and STAT5 (Jak/STAT)
proteins in our breast cancer cell lines. The MDA-MB-231, SKBR3 and
Hs578T cells were the only ones showing some responsiveness to
rHuEPO treatment (Fig. 3), which
was more evident 10 min after rHuEPO induction. A slight increase
in phosphorylated ERK and Akt was detected in the rHuEPO-treated
SKBR3 and MDA-MB-231 cells compared to the controls. An increased
Akt phosphorylation in SKBR3 cells was evident only when using 25
U/ml rHuEPO, otherwise it was independent of the concentration used
(Fig. 3). STAT5 phosphorylation
seems to be cell-specific since it was confirmed only in the SKBR3
cells (Fig. 3); its
phosphorylation was slightly increased 10 min after rHuEPO
induction. In the MCF-10A cells, Akt phosphorylation could not be
confirmed at defined experimental conditions. The Hs578T cells
responded to rHuEPO with a decreased level of phosphorylated ERK
and Akt at both indicated time-points; the decrease was more
pronounced with 5 U/ml rHuEPO.
Cell proliferation
Based on the clustering results, the MCF-7,
MDA-MB-231, SKBR3 and Hs578T cell lines were chosen and analyzed
for their responsiveness to rHuEPO induction in terms of cell
proliferation. We could not confirm any rHuEPO effect on MCF-7 and
MDA-MB-231 cell proliferation under our experimental conditions
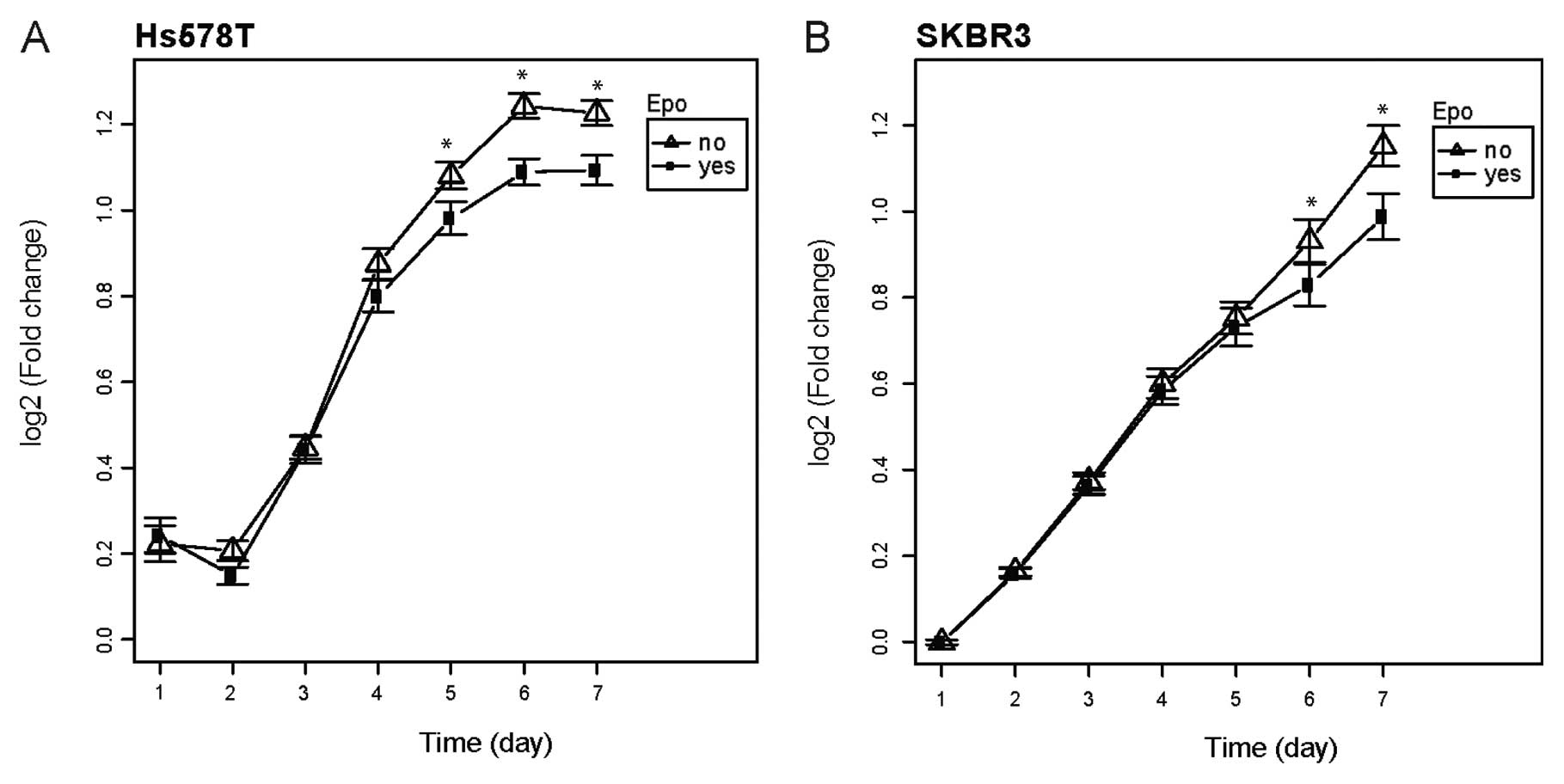
(data not shown). However, the rHuEPO treated Hs578T (p<0.001)
and SKBR3 cells (p=0.014) showed a significant decrease in cell
proliferation when cultivated in the medium supplemented with 1%
FBS (Fig. 4). rHuEPO did not
influence Hs578T and SKBR3 cell proliferation when the cells were
grown in the medium supplemented with 10% FBS (data not shown).
Discussion
Breast cancer is the most common type of cancer
afflicting females in the Western world and as such represents an
important health issue. Estrogens and progesterone have been shown
to be implicated in the development of the disease; therefore, the
determination of the ESR/PGR status is used as a strong predictive
factor. In addition, the EPOR/ESR/PGR status has been suggested to
be of prognostic value for the therapeutic response and RFS
(31). The latest insights into
the pathology of breast cancer has revealed the interaction between
nuclear and membrane ESR, connecting these receptors with the EGFR
and GPCR receptor families (25).
Breast cancer cell lines (Table
I) were therefore analyzed in detail for the gene expression of
EPO, EPOR, potential EPOR receptor partners
(EPHB4 and CSF2RB), several intracellular steroid
receptors (ESR1-2 and PGR) and GPER and the
correlation between the expression of these genes was
determined.
The expression of EPOR was confirmed in all
cell lines in accordance with previous data (6), as was the expression of EPHB4
(16). Furthermore, a correlation
between EPHB4 and EPOR was indicated, proposing its
potential to be involved in the EPOR signaling pathway (Table III). On the other hand,
CSF2RB was weakly expressed only in some of the examined
cell lines and negatively correlated with the expression of
EPOR (Table III), thus
questioning its involvement in the formation of the EPOR-CSF2RB
heteroreceptor in the analyzed cell lines. The expression of
EPO was confirmed only in the SKBR3 cells, suggesting that
in the other selected cell lines endogenous EPO does not act as an
autocrine activator of EPOR and downstream signaling pathways.
The expression levels of genes for ESR
(ESR1 and ESR2) and PGR are in accordance with
the ATCC data (Table I), except
for some of the analyzed cell lines (MDA-MB-361, Hs578Bst, MCF-10A
and Hs578T). The ATCC data is based on the immunohistochemical
protein detection while our data represent mRNA expression levels
that do not necessarily coincide with the functional protein
expression. The ESR receptor genes showed a strong
correlation with the expression of PGR and to a lesser
extent also with that of GPER. A positive correlation
between ESR1 and GPER2 genes in breast cancer cell
lines indicates that these proteins may interact at the plasma
membrane, an observation that has previously been shown in
endometrial and ovarian cancer cells and keratinocytes. In these
cells, the cross-talk between membrane-bound ESR1 and GPER proteins
mediates rapid EGFR-dependent kinase signaling by 17β-estradiol to
c-FOS and cyclin D1 (CCND1) upregulation and
proliferation (43–45). The correlation between GPER
splice variants was confirmed as expected.
Larsson et al (31) reported the positive correlation
between EPOR and ESR/PGR expression in breast cancer patients. We
could not confirm this correlation in our cohort of analyzed cell
lines. On the other hand, we were able to confirm the positive
correlation between EPOR and GPER which is in
agreement with previously published data, connecting GPER to
EPO signaling (30,36). Larsson et al (31) also reported that high EPOR
expression in ESR+/PGR+ breast cancer
patients negatively affects their responses to TAM treatment. TAM
has previously been reported to convert from antagonist to agonist
in breast cancer, depending on the phosphorylation of ESR (46). The enhanced cross-talk between
membrane ESR and EGFR family member receptors, most notably human
epidermal growth factor receptor 2 (HER2), may be associated with
the development of TAM resistance in breast cancer, possibly by
stimulating nuclear ESR phosphorylation (31,47,48). Liang et al (49) reported that EPOR signaling can
circumvent the trastuzumab inhibition of HER2. The inhibition of
ESR by TAM may also be hindered by the crosstalk between HER2, GPER
and EPOR (48) proteins. This
hypothesis needs to be further validated.
Connecting cell response to rHuEPO with
the growth factor expression profile
The MCF-7, MDA-MB-231, SKBR3 and Hs578T cell lines
were selected as representatives for a particular molecular subtype
(Fig. 2) and analyzed for their
responsiveness to rHuEPO treatment. The hormone-dependent MCF-7
(ESR+/PGR+) cells showed no significant
difference in cell proliferation when exposed to rHuEPO compared to
the untreated cells. This result is in agreement with the
observation that rHuEPO cannot activate cell signaling in these
cells (Fig. 3). Similarly, no
change in cell proliferation was detected with the more invasive
MDA-MB-231 (ESR+/PGR−) cells, though the
increase in the phosphorylation of ERK and Akt was confirmed.
MDA-MB-231 cells are PGR-negative and as such represent a more
aggressive breast cancer phenotype which is more independent of
growth factor withdrawal (50).
Furthermore, this cell line expresses a mutated form of tumor
protein 53 (p53) which has previously been implicated in the
regulation of cell survival following exposure to EPO and cisplatin
(51) and serum deprivation
(52). On the other hand, Hs578T
(ESR−/PGR−) and SKBR3
(ESR−/PGR−) cells showed decreased cell
proliferation when treated with rHuEPO in a medium containing a
decreased FBS concentration. Different FBS concentrations were used
on account of a possible mitogenic synergism between rHuEPO and
serum-contained growth factors or cytokines. The decrease in
proliferation was more evident in the Hs578T cells which also
showed decreased levels of ERK and Akt phosphorylation following
rHuEPO induction. The decrease in SKBR3 cell proliferation was
surprising, seeing that the 3 EPO-indicative signaling pathways
were activated upon rHuEPO treatment. The mechanism of action in
the SKBR3 cell line has to be further analyzed.
In conclusion, the molecular classification of
breast cancer is based upon the expression of ESR, PGR and HER2;
however, our results indicate that intracellular receptors may
cross-talk in order to activate signaling pathways or evade their
inhibition. Our current analysis on breast cancer cell lines
confirmed the proposed correlation between GPER and other cellular
receptors, as well as EPOR, characterizing them as a suitable model
for detailed mechanistic analysis. Further analysis of ESR, PGR,
EPOR, EPHB4 and GPER interaction on the protein level needs to be
carried out. The results from the present study suggest that the
expression of EPOR, membrane receptor GPER and EPHB4 may be
considered as an additional classification factor before choosing a
suitable treatment protocol.
Acknowledgements
Authors thank Dr Peter Juvan for
assistance with statistical analysis. Dr Toni Petan and Dr Igor
Križaj are acknowledged for providing the cell lines. This study
was supported by the J3-0124 grant to N.D., J3-4135 grant to T.L.R.
and Young Researcher grants to N.T. and N.H., all from the
Slovenian Research Agency (ARRS).
References
|
1
|
Sytkowski JA: Erythropoietin: Blood, Brain
and Beyond. Wiley-VCH; Boston, MA: 2004, View Article : Google Scholar
|
|
2
|
Debeljak N and Sytkowski AJ: EpoR.
UCSD-Nature Molecule Pages. 2007, View Article : Google Scholar
|
|
3
|
Jelkmann W and Wagner K: Beneficial and
ominous aspects of the pleiotropic action of erythropoietin. Ann
Hematol. 83:673–686. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Debeljak N and Sytkowski AJ:
Erythropoietin: new approaches to improved molecular designs and
therapeutic alternatives. Curr Pharm Des. 14:1302–1310. 2008.
View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Szenajch J, Wcislo G, Jeong JY, Szczylik C
and Feldman L: The role of erythropoietin and its receptor in
growth, survival and therapeutic response of human tumor cells:
from clinic to bench - a critical review. Biochim Biophys Acta.
1806:82–95. 2010.PubMed/NCBI
|
|
6
|
Acs G, Acs P, Beckwith SM, et al:
Erythropoietin and erythropoietin receptor expression in human
cancer. Cancer Res. 61:3561–3565. 2001.PubMed/NCBI
|
|
7
|
Arcasoy MO, Amin K, Karayal AF, et al:
Functional significance of erythropoietin receptor expression in
breast cancer. Lab Invest. 82:911–918. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Eschbach JW, Egrie JC, Downing MR, Browne
JK and Adamson JW: Correction of the anemia of end-stage renal
disease with recombinant human erythropoietin. Results of a
combined phase I and II clinical trial. N Engl J Med. 316:73–78.
1987. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Schrijvers D, De Samblanx H and Roila F:
ESMO Guidelines Working Group: Erythropoiesis-stimulating agents in
the treatment of anaemia in cancer patients:
Erythropoiesis-stimulating agents in the treatment of anaemia in
cancer patients: ESMO Clinical Practice Guidelines for use. Ann
Oncol. 21(Suppl 5): v244–v247. 2010. View Article : Google Scholar
|
|
10
|
Latini R, Brines M and Fiordaliso F: Do
non-hemopoietic effects of erythropoietin play a beneficial role in
heart failure? Heart Fail Rev. 13:415–423. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Moore EM, Bellomo R and Nichol AD:
Erythropoietin as a novel brain and kidney protective agent.
Anaesth Intensive Care. 39:356–372. 2011.PubMed/NCBI
|
|
12
|
Sytkowski AJ: The neurobiology of
erythropoietin. Cell Mol Neurobiol. 31:931–937. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Brines M, Grasso G, Fiordaliso F, et al:
Erythropoietin mediates tissue protection through an erythropoietin
and common beta-subunit heteroreceptor. Proc Natl Acad Sci USA.
101:14907–14912. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Noren NK and Pasquale EB: Paradoxes of the
EphB4 receptor in cancer. Cancer Res. 67:3994–3997. 2007.
View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Jackson DB, Stein M, Voss H, Brock S,
Danes CG and Sood A: Tissue protective erythropoietin receptor
(nepor) and methods to use. Patent WO 2009/068677. Filed November
28, 2008; issued April 18, 2012.
|
|
16
|
Kumar SR, Singh J, Xia G, et al: Receptor
tyrosine kinase EphB4 is a survival factor in breast cancer. Am J
Pathol. 169:279–293. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Henke M, Laszig R, Rube C, et al:
Erythropoietin to treat head and neck cancer patients with anaemia
undergoing radiotherapy: randomised, double-blind,
placebo-controlled trial. Lancet. 362:1255–1260. 2003. View Article : Google Scholar
|
|
18
|
Leyland-Jones B, Semiglazov V, Pawlicki M,
et al: Maintaining normal hemoglobin levels with epoetin alfa in
mainly nonanemic patients with metastatic breast cancer receiving
first-line chemo-therapy: a survival study. J Clin Oncol.
23:5960–5972. 2005. View Article : Google Scholar
|
|
19
|
Sytkowski AJ: Does erythropoietin have a
dark side? Epo signaling and cancer cells. Sci STKE.
2007.pe382007.PubMed/NCBI
|
|
20
|
Schnitt SJ: Classification and prognosis
of invasive breast cancer: from morphology to molecular taxonomy.
Mod Pathol. 23(Suppl 2): S60–S64. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Thomas P, Pang Y, Filardo EJ and Dong J:
Identity of an estrogen membrane receptor coupled to a G protein in
human breast cancer cells. Endocrinology. 146:624–632. 2005.
View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Thomas P: Characteristics of membrane
progestin receptor alpha (mPRalpha) and progesterone membrane
receptor component 1 (PGMRC1) and their roles in mediating rapid
progestin actions. Front Neuroendocrinol. 29:292–312. 2008.
View Article : Google Scholar
|
|
23
|
Levin ER: Integration of the extranuclear
and nuclear actions of estrogen. Mol Endocrinol. 19:1951–1959.
2005. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Bjornstrom L and Sjoberg M: Mechanisms of
estrogen receptor signaling: convergence of genomic and nongenomic
actions on target genes. Mol Endocrinol. 19:833–842. 2005.
View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Vasudevan N and Pfaff DW:
Membrane-initiated actions of estrogens in neuroendocrinology:
emerging principles. Endocr Rev. 28:1–19. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Pedram A, Razandi M, Sainson RC, Kim JK,
Hughes CC and Levin ER: A conserved mechanism for steroid receptor
translocation to the plasma membrane. J Biol Chem. 282:22278–22288.
2007. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Kumar P, Wu Q, Chambliss KL, et al: Direct
interactions with G α i and G βγ mediate nongenomic signaling by
estrogen receptor α. Mol Endocrinol. 21:1370–1380. 2007.
|
|
28
|
Hammes SR and Levin ER: Minireview: Recent
advances in extra-nuclear steroid receptor actions. Endocrinology.
152:4489–4495. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Levin ER: G protein-coupled receptor 30:
estrogen receptor or collaborator? Endocrinology. 150:1563–1565.
2009. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Pelekanou V, Kampa M, Kafousi M, et al:
Erythropoietin and its receptor in breast cancer: correlation with
steroid receptors and outcome. Cancer Epidemiol Biomarkers Prev.
16:2016–2023. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Larsson AM, Jirstrom K, Fredlund E, et al:
Erythropoietin receptor expression and correlation to tamoxifen
response and prognosis in breast cancer. Clin Cancer Res.
15:5552–5559. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Volgger B, Kurz K, Zoschg K, et al:
Importance of erythropoetin receptor expression in tumour tissue
for the clinical course of breast cancer. Anticancer Res.
30:3721–3726. 2010.PubMed/NCBI
|
|
33
|
Filardo EJ, Quinn JA and Sabo E:
Association of the membrane estrogen receptor, GPR30, with breast
tumor metastasis and transactivation of the epidermal growth factor
receptor. Steroids. 73:870–873. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Kampa M, Pelekanou V and Castanas E:
Membrane-initiated steroid action in breast and prostate cancer.
Steroids. 73:953–960. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Pelekanou V, Notas G, Sanidas E, Tsapis A,
Castanas E and Kampa M: Testosterone membrane-initiated action in
breast cancer cells: Interaction with the androgen signaling
pathway and EPOR. Mol Oncol. 4:135–149. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Notas G, Kampa M, Pelekanou V and Castanas
E: Interplay of estrogen receptors and GPR30 for the regulation of
early membrane initiated transcriptional effects: a pharmacological
approach. Steroids. 77:943–950. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Xiao Z, Carrasco R, Kinneer K, et al:
EphB4 promotes or suppresses Ras/MEK/ERK pathway in a
context-dependent manner: implications for EphB4 as a cancer
target. Cancer Biol Ther. 13:630–637. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Hevir N, Trošt N, Debeljak N and Rižner
TL: Expression of estrogen and progesterone receptors and estrogen
metabolizing enzymes in different breast cancer cell lines. Chem
Biol Interact. 191:206–216. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Bustin SA, Benes V, Garson JA, et al: The
MIQE guidelines: minimum information for publication of
quantitative real-time PCR experiments. Clin Chem. 55:611–622.
2009. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Kutuk O, Arisan ED, Tezil T, Shoshan MC
and Basaga H: Cisplatin overcomes Bcl-2-mediated resistance to
apoptosis via preferential engagement of Bak: critical role of
Noxa-mediated lipid peroxidation. Carcinogenesis. 30:1517–1527.
2009. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Abramoff MD, Magelhaes PJ and Ram SJ:
Image Processing with ImageJ. Biophotonics International. 11:36–42.
2004.
|
|
42
|
Jelkmann W, Bohlius J, Hallek M and
Sytkowski AJ: The erythropoietin receptor in normal and cancer
tissues. Crit Rev Oncol Hematol. 67:39–61. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Vivacqua A, Bonofiglio D, Recchia AG, et
al: The G protein-coupled receptor GPR30 mediates the proliferative
effects induced by 17beta-estradiol and hydroxytamoxifen in
endometrial cancer cells. Mol Endocrinol. 20:631–646. 2006.
View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Albanito L, Madeo A, Lappano R, et al: G
protein-coupled receptor 30 (GPR30) mediates gene expression
changes and growth response to 17beta-estradiol and selective GPR30
ligand G-1 in ovarian cancer cells. Cancer Res. 67:1859–1866. 2007.
View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Kanda N and Watanabe S: 17beta-estradiol
stimulates the growth of human keratinocytes by inducing cyclin D2
expression. J Invest Dermatol. 123:319–328. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Michalides R, Griekspoor A, Balkenende A,
et al: Tamoxifen resistance by a conformational arrest of the
estrogen receptor alpha after PKA activation in breast cancer.
Cancer Cell. 5:597–605. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Shou J, Massarweh S, Osborne CK, et al:
Mechanisms of tamoxifen resistance: increased estrogen
receptor-HER2/neu cross-talk in ER/HER2-positive breast cancer. J
Natl Cancer Inst. 96:926–935. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Ignatov A, Ignatov T, Roessner A, Costa SD
and Kalinski T: Role of GPR30 in the mechanisms of tamoxifen
resistance in breast cancer MCF-7 cells. Breast Cancer Res Treat.
123:87–96. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
49
|
Liang K, Esteva FJ, Albarracin C, et al:
Recombinant human erythropoietin antagonizes trastuzumab treatment
of breast cancer cells via Jak2-mediated Src activation and PTEN
inactivation. Cancer Cell. 18:423–435. 2010. View Article : Google Scholar
|
|
50
|
Cui X, Schiff R, Arpino G, Osborne CK and
Lee AV: Biology of progesterone receptor loss in breast cancer and
its implications for endocrine therapy. J Clin Oncol. 23:7721–7735.
2005. View Article : Google Scholar : PubMed/NCBI
|
|
51
|
Trošt N, Juvan P, Serša G and Debeljak N:
Contrasting effect of erythropoietin on breast cancer cell response
to cisplatin induced cytotoxicity. Radiol Oncol. 46:213–225.
2012.PubMed/NCBI
|
|
52
|
Hui L, Zheng Y, Yan Y, Bargonetti J and
Foster DA: Mutant p53 in MDA-MB-231 breast cancer cells is
stabilized by elevated phospholipase D activity and contributes to
survival signals generated by phospholipase D. Oncogene.
25:7305–7310. 2006. View Article : Google Scholar : PubMed/NCBI
|