|
1
|
Siegel R, Desantis C and Jemal A:
Colorectal cancer statistics, 2014. CA Cancer J Clin. 64:104–117.
2014. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Center MM, Jemal A, Smith RA and Ward E:
Worldwide variations in colorectal cancer. CA Cancer J Clin.
59:366–378. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Grivell LA: Mitochondrial DNA. Small,
beautiful and essential. Nature. 341:569–571. 1989. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Lightowlers RN, Chinnery PF, Turnbull DM
and Howell N: Mammalian mitochondrial genetics: Heredity,
heteroplasmy and disease. Trends Genet. 13:450–455. 1997.
View Article : Google Scholar
|
|
5
|
Zhang G, Qu Y, Dang S, Yang Q, Shi B and
Hou P: Variable copy number of mitochondrial DNA (mtDNA) predicts
worse prognosis in advanced gastric cancer patients. Diagn Pathol.
8:1732013. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Cheau-Feng Lin F, Jeng YC, Huang TY, Chi
CS, Chou MC and Chin-Shaw Tsai S: Mitochondrial DNA copy number is
associated with diagnosis and prognosis of head and neck cancer.
Biomarkers. 19:269–274. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
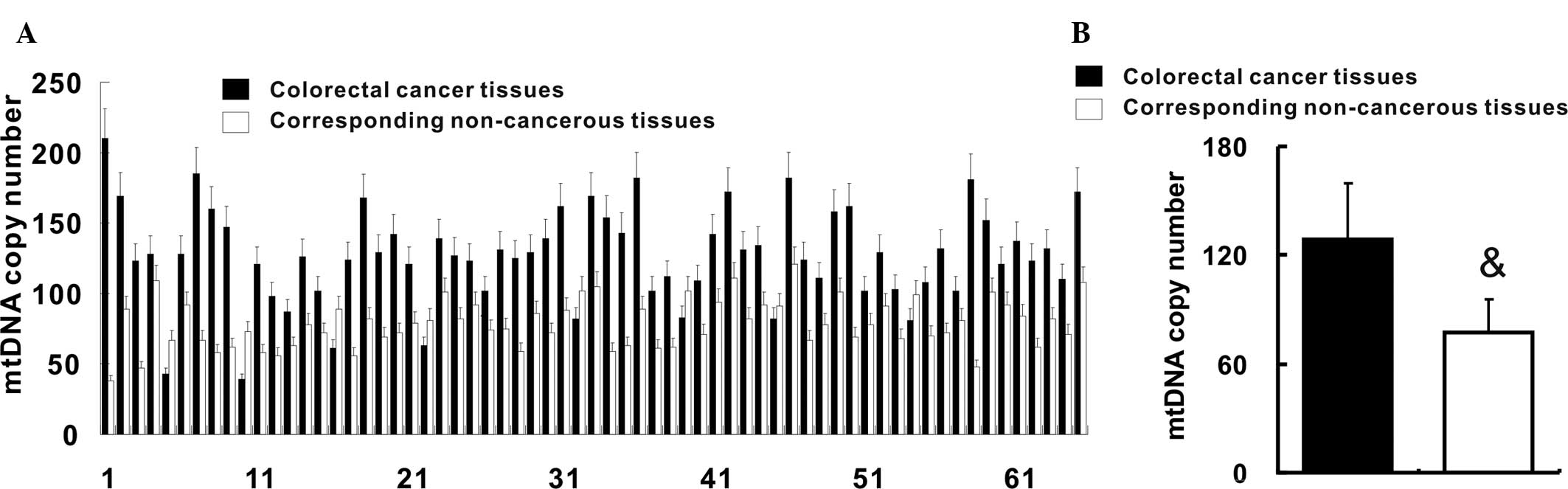
Feng S, Xiong L, Ji Z, Cheng W and Yang H:
Correlation between increased copy number of mitochondrial DNA and
clinicopatho-logical stage in colorectal cancer. Oncol Lett.
2:899–903. 2011.
|
|
8
|
Boland ML, Chourasia AH and Macleod KF:
Mitochondrial dysfunction in cancer. Front Oncol. 3:2922013.
View Article : Google Scholar : PubMed/NCBI
|
|
9
|
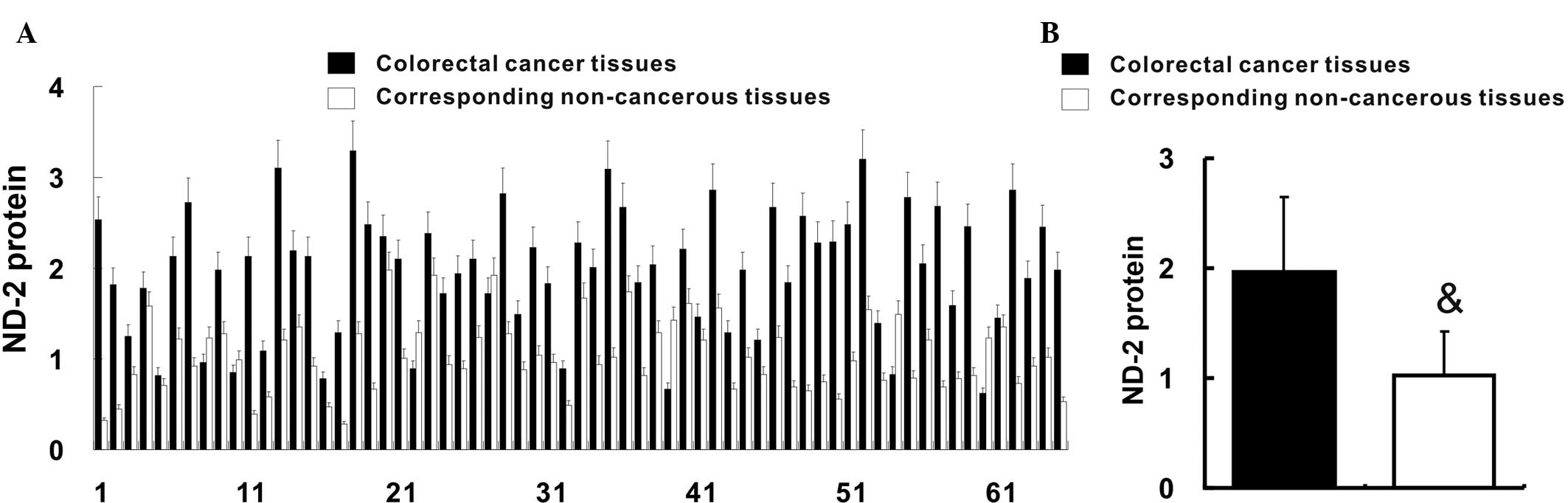
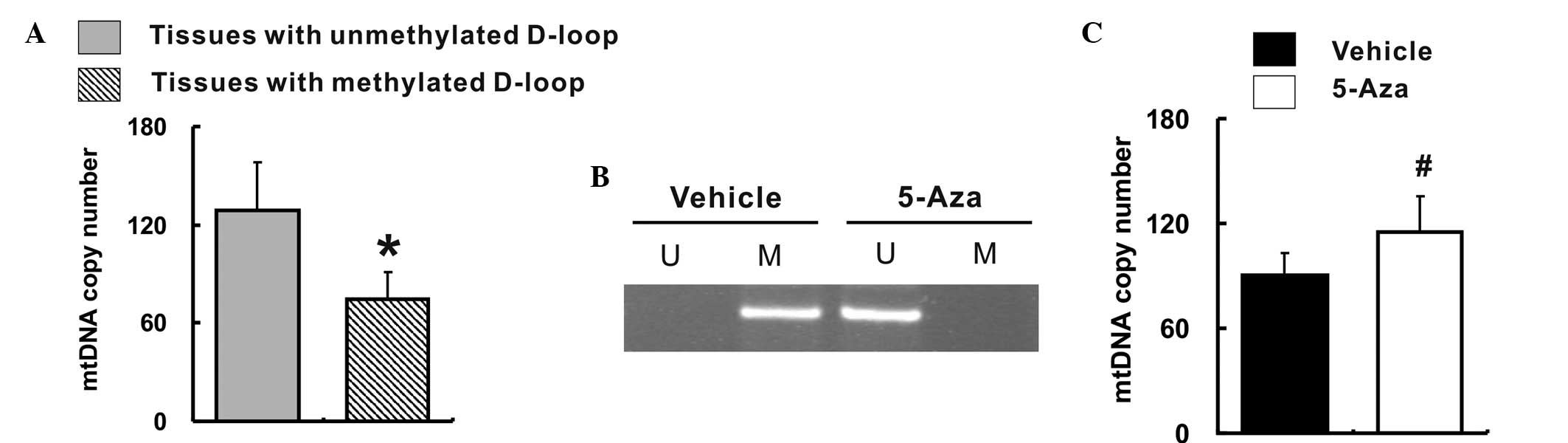
Feng S, Xiong L, Ji Z, Cheng W and Yang H:
Correlation between increased ND2 expression and demethylated
displacement loop of mtDNA in colorectal cancer. Mol Med Rep.
6:125–130. 2012.PubMed/NCBI
|
|
10
|
Bellizzi D, D'Aquila P, Scafone T,
Giordano M, Riso V, Riccio A and Passarino G: The control region of
mitochondrial DNA shows an unusual CpG and non-CpG methylation
pattern. DNA Res. 20:537–547. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Roos-Araujo D, Stuart S, Lea RA, Haupt LM
and Griffiths LR: Epigenetics and migraine; complex mitochondrial
interactions contributing to disease susceptibility. Gene. 543:1–7.
2014. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Esteller M: CpG island hypermethylation
and tumor suppressor genes: A booming present, a brighter future.
Oncogene. 21:5427–5440. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Mates IN, Jinga V, Csiki IE, Mates D, Dinu
D, Constantin A and Jinga M: Single nucleotide polymorphisms in
colorectal cancer: associations with tumor site and TNM stage. J
Gastrointestin Liver Dis. 21:45–52. 2012.PubMed/NCBI
|
|
14
|
Matsubara N: Epigenetic regulation and
colorectal cancer. Dis Colon Rectum. 55:96–104. 2012. View Article : Google Scholar
|
|
15
|
Zhang R, Zhang F, Wang C, Wang S, Shiao YH
and Guo Z: Identification of sequence polymorphism in the D-Loop
region of mitochondrial DNA as a risk factor for hepatocellular
carcinoma with distinct etiology. J Exp Clin Cancer Res.
29:1302010. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Ylikallio E, Tyynismaa H, Tsutsui H, Ide T
and Suomalainen A: High mitochondrial DNA copy number has
detrimental effects in mice. Hum Mol Genet. 19:2695–2705. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Penta JS, Johnson FM, Wachsman JT and
Copeland WC: Mitochondrial DNA in human malignancy. Mutat Res.
488:119–133. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Zhang J, Guo Z, Bai Y, Cui L, Zhang S and
Xu J: Identification of sequence polymorphisms in the displacement
loop region of mitochondrial DNA as a risk factor for renal cell
carcinoma. Biomed Rep. 1:563–566. 2013.
|
|
19
|
Tommasi S, Favia P, Weigl S, Bianco A,
Pilato B, Russo L, Paradiso A and Petruzzella V: Mitochondrial DNA
variants and risk of familial breast cancer: An exploratory study.
Int J Oncol. 44:1691–1698. 2014.PubMed/NCBI
|
|
20
|
Maniura-Weber K, Goffart S, Garstka HL,
Montoya J and Wiesner RJ: Transient overexpression of mitochondrial
transcription factor A (TFAM) is sufficient to stimulate
mitochondrial DNA transcription, but not sufficient to increase
mtDNA copy number in cultured cells. Nucleic Acids Res.
32:6015–6027. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Kurita T, Izumi H, Kagami S, Kawagoe T,
Toki N, Matsuura Y, Hachisuga T and Kohno K: Mitochondrial
transcription factor A regulates BCL2L1 gene expression and is a
prognostic factor in serous ovarian cancer. Cancer Sci.
103:239–444. 2012. View Article : Google Scholar
|
|
22
|
LeBleu VS, O'Connell JT, Gonzalez Herrera
KN, Wikman H, Pantel K, Haigis MC, de Carvalho FM, Damascena A,
Domingos Chinen LT, Rocha RM, et al: PGC-1α mediates mitochondrial
biogenesis and oxidative phosphorylation in cancer cells to promote
metastasis. Nat Cell Biol. 16:992–1003. 2014. View Article : Google Scholar
|
|
23
|
Lim SH, Wu L, Kiew LV, Chung LY, Burgess K
and Lee HB: Rosamines targeting the cancer oxidative
phosphorylation pathway. PloS one. 9:e829342014. View Article : Google Scholar : PubMed/NCBI
|