Introduction
Epigenetics is the study of inherited genetic
changes that occur without altering the DNA sequence. DNA
methylation is a mechanism of epigenetic change, with the most
widely studied epigenetic alteration in human tumor cells being
histone modification and chromatin remodeling (1). Human tumor cells exhibit aberrant DNA
methylation patterns, including the hypermethylation of CpG islands
in tumor suppressor genes (TSGs) and a global loss of DNA
methylation in the genome (2).
These changes are associated with the inactivation of TSGs and the
activation of oncogenes or tumor promoter genes (TPGs), and may
promote tumor progression (3).
Abnormal expression of DNA methyltransferase (DNMT) may be
important in the aberrant DNA methylation that occurs in tumors
(4). Previous studies have
identified elevated DNMT expression in tumors when compared with
control tissue (5–8) and the overexpression of DNMT may
contribute to tumor progression through the
hypermethylation-mediated inactivation of TSGs in CpG islands
(9). In the current study,
expression of DNMTs (DNMT1, DNMT2, DNMT3A, DNMT3B and DNMT3L) and
the DNA methylation pattern of the genome in gastric signet ring
cell carcinoma (SRC) was investigated. Results of this study are
likely to aid future epigenetic studies investigating SRC.
Materials and methods
Ethics statement
All experimental procedures were approved by the
Ethics Committee of the First Affiliated Hospital of Chongqing
Medical University (Chongqing, China). All patients provided
written informed consent.
Clinical specimens
Twenty-eight pairs of human gastric SRC and matched
non-cancerous tissue specimens (gastric mucosa tissue located >5
cm from cancerous areas) were obtained from the Department of
General Surgery of the First Affiliated Hospital of Chongqing
Medical University. Characteristics of these tissues are presented
in Table I. Tissues were fixed in
formalin and embedded in paraffin for immunohistochemistry testing,
and their diagnosis was confirmed by pathological analysis. For the
methylated DNA immunoprecipitation microarray (MeDIP-chip) assay,
one pair of gastric tissues was selected for pathological
examination to verify that the selected cancerous and matched
non-cancerous tissue specimens consisted of >95% cancer and
mucosal cells, respectively. In addition, five pairs of gastric
tissues were selected for MeDIP quantitative real-time PCR
(MeDIP-qPCR) assay to validate MeDIP-chip observations.
 | Table IPositive DNMT expression and
differential clinical characteristics of gastric signet ring cell
carcinoma, n (%). |
Table I
Positive DNMT expression and
differential clinical characteristics of gastric signet ring cell
carcinoma, n (%).
| Parameter | n | DNMT1 expression
(%) | DNMT2 expression
(%) | DNMT3A expression
(%) | DNMT3B expression
(%) | DNMT3L expression
(%) |
|---|
| Gender |
| Male | 12 | 8 (66.7) | 9 (75.0) | 3 (25.0) | 9 (75.0) | 7 (58.3) |
| Female | 16 | 9 (56.3) | 14 (87.5) | 7 (43.8) | 12 (75.0) | 10 (62.5) |
| P-value | | 0.71 | 0.62 | 0.43 | 1.00 | 1.00 |
| Age, years |
| ≥50 | 13 | 7 (53.8) | 10 (76.9) | 5 (38.5) | 10 (76.9) | 7 (53.8) |
| <50 | 15 | 10 (66.7) | 13 (86.7) | 5 (33.3) | 11 (73.3) | 10 (66.7) |
| P-value | | 0.70 | 0.64 | 1.00 | 1.00 | 0.70 |
| Location |
| Upper and
middle | 13 | 6 (46.2) | 11 (84.6) | 5 (38.5) | 11 (84.6) | 7 (53.8) |
| lower | 15 | 11 (73.3) | 12 (80.0) | 5 (33.3) | 10 (66.7) | 10 (66.7) |
| P-value | | 0.25 | 1.00 | 1.00 | 0.40 | 0.70 |
| Tumor size, cm |
| ≥2 | 16 | 12 (75.0) | 13 (81.3) | 7 (43.8) | 13 (81.3) | 11 (68.8) |
| <2 | 12 | 5 (41.7) | 10 (83.3) | 3 (25.0) | 8 (66.7) | 6 (50.0) |
| P-value | | 0.12 | 1.00 | 0.43 | 0.42 | 0.44 |
| Depth of
invasion |
| Mucosa and
muscular | 5 | 1 (20.0) | 4 (80.0) | 2 (40.0) | 2 (40.0) | 2 (40.0) |
| Subserosa and
serosa | 23 | 16 (69.6) | 19 (82.6) | 8 (34.8) | 19 (82.6) | 15 (65.2) |
| P-value | | 0.06 | 1.00 | 1.00 | 0.08 | 0.35 |
| Lymphnode
metastasis |
| Yes | 17 | 14 (82.4) | 13 (76.5) | 5 (29.4) | 14 (82.4) | 11 (64.7) |
| No | 11 | 3 (27.3) | 10 (90.9) | 5 (45.5) | 7 (63.6) | 6 (54.5) |
| P-value | | 0.01* | 0.62 | 0.44 | 0.38 | 0.70 |
| TNM stage |
| I–II | 12 | 4 (33.3) | 9 (75.0) | 4 (33.3) | 8 (66.7) | 5 (41.7) |
| III–IV | 16 | 13 (81.3) | 14 (87.5) | 6 (37.5) | 13 (81.3) | 12 (75.0) |
| P-value | | 0.02* | 0.62 | 1.00 | 0.42 | 0.12 |
| H. pylori
infection |
| Positive | 24 | 16 (66.7) | 21 (87.5) | 9 (37.5) | 19 (79.2) | 16 (66.7) |
| Negative | 4 | 1 (25.0) | 2 (50.0) | 1 (25.0) | 2 (50.0) | 1 (25.0) |
| P-value | | 0.27 | 0.14 | 1.00 | 0.25 | 0.27 |
Immunohistochemistry
The streptavidin-peroxidase (SP) method was adopted
and performed (7). Primary
antibodies against DNMT1 (sc-20701), DNMT2 (sc-20702), DNMT3A
(sc-20703), DNMT3B (sc-20704) and DNMT3L (sc-20705) were purchased
from Santa Cruz Biotechnology, Inc. (Santa Cruz, CA, USA). The SP
and DAB kits were obtained from Beijing Zhongshan Golden Bridge
Biotechnology Co., Ltd. (Beijing, China).
Evaluation of staining
DNMT expression was assessed by scoring the staining
intensity and stained proportion of the cell nucleus. Staining
intensity was recorded as negative = 0, light = 1, moderate = 2 or
strong = 3. The staining proportion was recorded as 1 (≤25%), 2
(≤50%), 3 (≤75%) or 4 (>75%). The two values were multiplied for
each slide to produce a terminal score. If the score was higher in
cancer cells than in matched control cells, this pair of tissues
was marked with a ‘+’ (corresponding to cancer cells that expressed
elevated DNMT levels). The opposite condition was marked with a ‘−’
(corresponding to cancer cells that expressed reduced DNMT levels).
If the scores were equal, the pair was marked with a ‘0’
(corresponding to similar DNMT expression in cancer and control).
Terminal scores of 0–3 were defined as negative expression; 4–12
were defined as positive expression.
MeDIP-chip assay
For the MeDIP-chip assay, the NimbleGen Human DNA
Methylation 385K Promoter Plus CpG Island array was used (Roche
Diagnostics GmbH, Mannheim Germany). This single array design,
includes 28,226 CpG islands and all RefSeq gene promoter regions
(between −800 and +200 bp of the transcription start sites), coated
entirely with ~385,000 probes. Briefly, genomic DNA extraction and
fragmentation, immunoprecipitation [using Biomag™ magnetic beads
(Bangs Laboratories, Inc., Fisher, IN, USA)coupled to a mouse
monoclonal antibody against 5-methylcytidine], whole genome
amplification, DNA labeling and array hybridization, raw data
scanning (with Axon GenePix 4000B microarray scanner; Axon 132
Instruments, Foster City, CA, USA), quality assessment of raw data,
data normalization, data mapping to genomic features (transcripts
and CpG islands) and summarizing for the selected gastric tissue,
were performed by KangChen Bio-tech Inc. (Shanghai, China). Genes
exhibiting differential DNA methylation in CpG islands between SRC
and control tissue from the genome (based on the results supplied
by KangChen Bio-tech) were separated and investigated for
tumor-associated genes.
MeDIP-qPCR
Genomic DNA was extracted from 5 pairs of gastric
tissue using a DNeasy Blood and Tissue kit (Qiagen, Hilden,
Germany) and sonicated to random fragments of 200–1,000 bp with a
Bioruptor sonicator (Diagenode, Denville, NJ, USA). Sonicated DNA
fragments were divided in two, one part was used as input
(normalized control) and the other was prepared for MeDIP. MeDIP
was performed using Biomag magnetic beads coupled to a mouse
monoclonal antibody against 5-methylcytidine. The
immunoprecipitated DNA was eluted and purified by phenol chloroform
extraction and ethanol precipitation for the subsequent qRT-PCR.
qRT-PCR was conducted on an ABI Prism 7900 system (Applied
Biosystems, Foster City, CA, USA) using PCR master mix (Qiagen) and
specific primers were denatured at 95°C for 10 min followed by 40
cycles of 95°C for 10 sec and 60°C for 60 sec. Following this, the
comparative ΔΔCt method was performed. Each MeDIP DNA Ct value was
normalized against the input DNA value for the same qRT-PCR assay
(ΔCt) to account for chromatin sample preparation differences. The
input percentage for each MeDIP fraction was calculated using the
following formula: input percentage = 2(Ctinput −
CtMeDIP) x Fd × 100. Fd represents the input dilution
factor (1/5). The input percentage values represent the DNA
methylation levels of validated genes in this assay.
Statistical analysis
Standard statistical analysis was performed using
the SPSS version 17.0 (SPSS, Inc., Chicago, IL, USA). The Wilcoxon
signed-rank test (for immunohistochemistry analysis), χ2
test, Fisher’s exact test (for association analysis between DNMT
expression and clinical parameters of SRC) and paired t-test (for
MeDIP-qPCR) were used in this study. P<0.05 was considered to
indicate a statistically significant difference.
Results
Expression of DNMT in gastric SRC
tissue
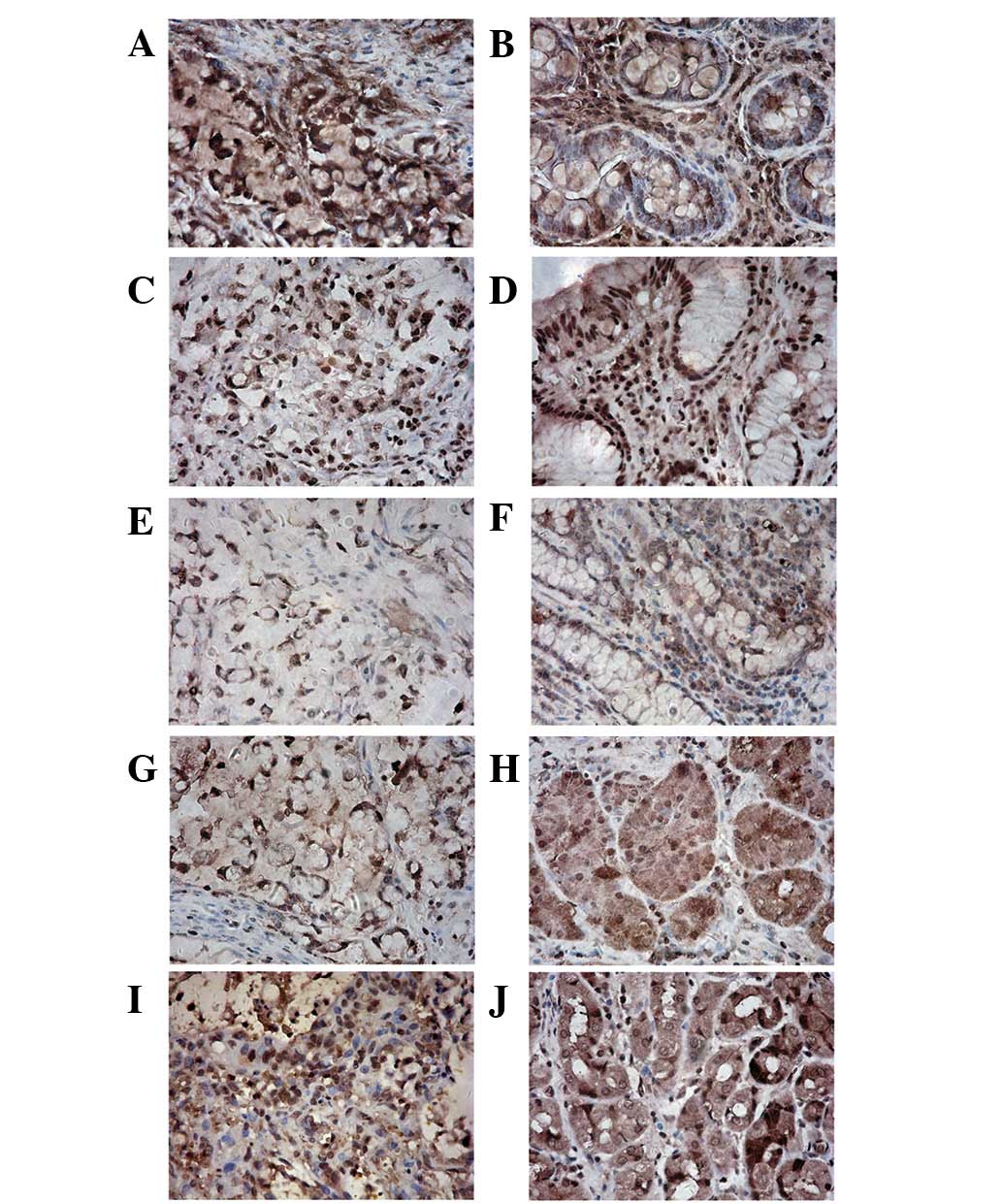
Gastric SRC and matched mucosal tissue expressed
DNMT proteins (Fig. 1). Notably,
DNMT protein distribution was observed primarily in the nucleus of
cancerous tissue and in the nucleus and cytoplasm of control
tissue. DNMT expression in non-cancerous tissue was primarily
focused on the proliferating zone of gastric mucosa. DNMTs perform
DNA methylation in the nucleus; thus, nuclear staining was
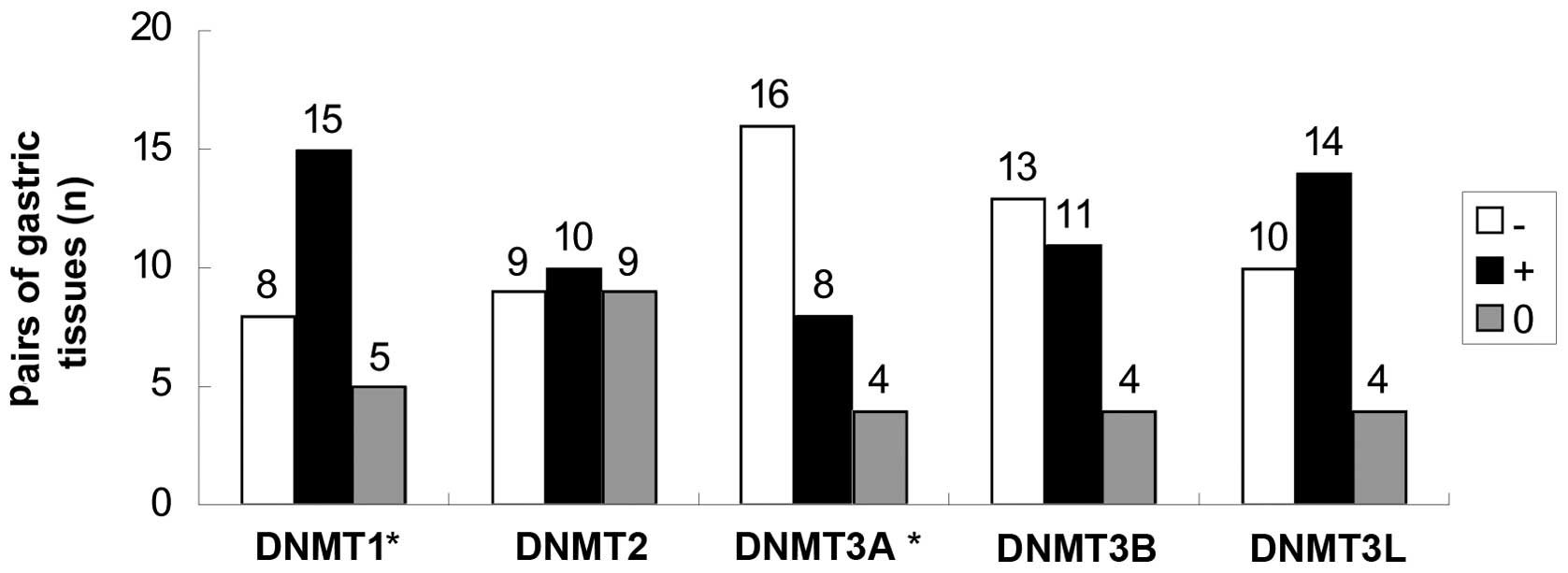
investigated. DNMT1 expression was elevated and DNMT3A expression
was decreased in gastric SRC when compared with matched control
tissue (Fig. 2). However,
expression of other DNMTs did not differ significantly between
cancerous and non-cancerous tissue.
Correlation between DNMT expression and
clinical characteristics of gastric SRC
According to the TNM classification of malignant
tumors, expression of DNMT1 was associated with lymph node
metastasis in gastric SRC (Table
I). SRC samples with features of lymph node metastasis and
attributes of late TNM classification were found to express DNMT1
protein at a higher level than control samples. There was no
association noted in other DNMT expression and demographic
variables of the carcinoma.
Comparison of genomic DNA methylation
between gastric SRC and matched control tissue
DNMT proteins in the selected pair of gastric
tissues were scored by immunohistochemistry assay as follows: DNMT1
‘+’; DNMT2 ‘0’; DNMT3A ‘−’; DNMT3B ‘+’; and DNMT3L ‘−’. In
carcinoma and matched mucosal tissue, characteristics of interest
included local hypermethylation and global hypomethylation of
genomic DNA. Specific CpG islands and gene promoters were
hypermethylated only in carcinoma tissue, whereas other CpG islands
and gene promoters were hypermethylated only in matched mucosa
tissue (Table II), indicating
that these DNA sequences were hypomethylated only in carcinoma
tissue. Following a comprehensive analysis involving separation of
genes exhibiting differential DNA methylation in promoters between
the carcinoma and control from the genome, and searching for
tumor-related genes in these separating genes, gastric SRC was
observed to contain hypermethylated and hypomethylated TSGs,
oncogenes and tumor-promoter genes (TPGs; Table III) (10–47).
 | Table IIOverview of MeDIP-chip data. |
Table II
Overview of MeDIP-chip data.
| Classification | CpG islands
(%) | Gene promoters
(%) |
|---|
| Hypermethylation
only in cancer | 2,832 (10.03) | 1,541 (8.55) |
| Hypermethylation
only in control | 2,273 (8.05) | 913 (5.06) |
| Hypermethylation in
cancer and control | 1,943 (6.88) | 736 (4.08) |
| Hypomethylation in
cancer and control | 21,178 (75.03) | 14,838 (82.31) |
| Hypermethylation in
cancer | 4,775 (16.92) | 2,277 (12.63) |
| Hypermethylation in
control | 4,216 (14.94) | 1,649 (9.15) |
| Total | 28,226 | 18,028 |
 | Table IIIDifferential DNA methylation of
tumor-related genes between SRC and control. |
Table III
Differential DNA methylation of
tumor-related genes between SRC and control.
| Classification | TSG (ref.) | Oncogene and TPG
(ref.) |
|---|
| Hypermethylation
only in cancer | BCL2L11 (10) | APCDD1 (24) |
| BRMS-1 (11) | BCL11A (25) |
| CARS (12) | JUN (26) |
| CDKN1C (13) | LYN (27) |
| CDKN2A (14) | MYB (28) |
| DLC-1 (15) | MYCL1 (29) |
| ING-1 (16) | REL (30) |
| OVCA2 (17) | SRC (31) |
| RASD1 (18) | WNT4 (32) |
| RB-1 (19) | WNT9A (32) |
| SYK (20) | WNT10A (32) |
| | WNT11 (33) |
| | KRAS (34) |
| | VEGFA (35) |
| | RAB6A (36) |
| | RAB8A (37) |
| | RAB27A (38) |
| | RAB32 (39) |
| Hypomethylation
only in cancer | APAF1 (21) | RAB33A (36) |
| CAV2 (22) | ABL2 (40) |
| RASSF1 (23) | FGF18 (41) |
| | EGFL7 (42) |
| | FYN (43) |
| | MYCNOS (44) |
| | RAB3A (45) |
| | TRAF2 (46) |
| | WNT3A (47) |
Validation of the abnormally
hypomethylated genes in gastric SRC by MeDIP-qPCR
ABL2, FGF18, TRAF2, EGFL7 and RAB33A were selected
as validation genes. Their primer sequences are presented in
Table IV. Using MeDIP-qPCR, the
input percentage values of these genes were observed to be
significantly lower in SRC than in matched mucosal tissue (Table V), indicating that the DNA was
abnormally hypomethylated in gastric SRC compared with matched
control tissue. This result was in agreement with the MeDIP-chip
observations. The input percentage values of these genes the in
negative control (non-immune serum) were <0.01.
 | Table IVPrimer sequences of validation
genes. |
Table IV
Primer sequences of validation
genes.
| Gene name | Primer
sequences | Length (bp) |
|---|
| ABL2 |
F:5′ATTTGACAGGTGGAGGTGGGAT3′ | |
|
R:5′CGCTGCTTGAGGTCTTTCGTC3′ | 162 |
| FGF18 |
F:5′GGCTGGGAAACTCCACGAT3′ | |
|
R:5′CCACATTCGCTACTCGCACT3′ | 135 |
| TRAF2 |
F:5′GGAGAATCGCTTGAACCCG3′ | |
|
R:5′GTGTGCTAATCTACTGGGTTGTGC3′ | 138 |
| EGFL7 |
F:5′CTGGTTTCTGGCTGTTTTGG3′ | |
|
R:5′ATGCTCCGTCCTGGGTAATC3′ | 214 |
| RAB33A |
F:5′ACCAGACAAGACTGAAGCCACC3′ | |
|
R:5′CGACAACCGCTAGAGCTATGC3′ | 154 |
 | Table VInput percentage value of validation
genes in signet ring cell carcinoma and control tissues,
MeDIP-qPCR. |
Table V
Input percentage value of validation
genes in signet ring cell carcinoma and control tissues,
MeDIP-qPCR.
| Gene name | Carcinoma
tissue | Control tissue |
|---|
| ABL2 |
0.0114±0.00408a | 1.1698±0.22944 |
| FGF18 |
0.0115±0.00125a | 1.3419±0.15275 |
| TRAF2 |
0.8152±0.20569a | 3.5592±0.40797 |
| EGFL7 |
0.3557±0.06140a | 1.9956±0.31949 |
| RAB33A |
0.0465±0.00735a | 2.1858±0.26880 |
Discussion
According to the World Health Organization, there
are four predominant histological types of gastric adenocarcinoma,
papillary, tubular, mucinous and SRC. SRC is characterized by the
histological appearance of signet ring cells, a large vacuole full
of mucin in the cytoplasm displacing the nucleus to the periphery.
This adenocarcinoma originates from the undifferentiated stem cells
at the gastric gland neck in gastric lamina propria and accounts
for 3.14–29% of gastric cancer (48). SRC is a poorly differentiated
adenocarcinoma with rapid progression and poor prognosis. To date,
the etiology of SRC is unclear and therapy is mainly dependent on
surgical procedures (SRC is non-responsive to chemotherapy).
Epigenetic alterations, including promoter
hypermethylation, lead to chromatin remodeling and the silencing of
tumor-related genes, and are crucial in tumor progression (49). DNA methylation is catalyzed mainly
by DNMTs, including DNMT1, DNMT2, DNMT3A, DNMT3B and DNMT3L
(50–53). Previous studies have indicated that
overexpression of DNMT may contribute to tumor progression through
hypermethylation-mediated TSG inactivation in CpG islands. We
hypothesized that overexpression of DNMT also may be detected in
SRC.
In the current study, it was observed that gastric
SRC and matched mucosal tissue expressed DNMT proteins. DNMT
expression in non-cancerous tissue was primarily focused on the
proliferating zones of gastric mucosa. An abnormal overexpression
of DNMT1 was observed when nuclear staining was taken into
consideration in SRC tissue compared with matched mucosal tissue.
By contrast, DNMT3A expression in SRC was not significant compared
with that in matched controls. For the remaining DNMTs, no
expression difference between SRC and control tissue was noted. The
observations indicate that overexpression of DNMT in SRC tissue was
specific to DNMT1. Following association analysis between the
demographic variables and DNMT expression of SRC, positive
expression of DNMT1 was associated with lymph node metastasis and
late TNM stages of SRC, indicating a potential role of DNMT1
proteins in promoting SRC progression. It was hypothesized that
DNMT1 may function in a similar manner to an oncogene in SRC.
Considering no association was noted between DNMT3A and clinical
characteristics of SRC, the hypothesis that SRC expresses a lower
level of DNMT3A compared with control requires further
investigation. DNMT proteins were distributed in the nucleus and
cytoplasm, particularly in the proliferative zones of normal
gastric mucosa. It is not yet clear if there are unknown substances
in the cytoplasm that may cross-react with DNMT antibodies or if
DNMT proteins function in the cytoplasm. Further investigation is
required to confirm this.
The MeDIP-chip assay revealed local hypermethylation
and global hypomethylation of genomic DNA in SRC and matched
mucosal tissue. The number of hypermethylated CpG islands and gene
promoters in SRC were increased compared with those in control
mucosa (4,775 vs. 4,216 and 2,277 vs. 1,649, respectively; Table II). This observation indicated
that DNA methylation of the genome increased in SRC compared with
matched controls. Notably, in gastric SRC, hypermethylated and
hypomethylated TSGs, oncogenes and TPGs were observed. The
subsequent MeDIP-qPCR assay validated specific MeDIP-chip results.
Tumor-related genes, ABL2, FGF18, TRAF2, EGFL7 and RAB33A, were
abnormally hypomethylated in SRC tissue compared with matched
controls. This observation is an addition to the traditional DNA
methylation theory, which focuses on the hypermethylation of TSG in
tumors and indicates that the aberrant DNA methylation pattern of
the SRC genome is complex. Similar observations were also
demonstrated in liver and pancreatic cancer (54,55).
Specific tumors exhibit abnormal hypomethylation of TPGs and
oncogenes (56,57) and overexpression of these genes due
to hypomethylation, is potentially another epigenetic mechanism for
uncontrollable cancer cell proliferation (55).
In conclusion, gastric SRCs express elevated DNMT1
protein and reduced DNMT3A protein compared with matched gastric
mucosa. A difference between genomic DNA methylation between SRC
and control (gastric mucosa) samples exists, however, it appears to
be complex since it is not limited to hypermethylation of TSGs.
To date, there has been encouraging progression in
the understanding of the role of DNA methylation in tumors.
However, clinical applications based on DNA methylation theory for
diagnosis and treatments of tumors remain scarce. Further
investigation is required to investigate the role of DNMT
inhibitors, including 5-aza-2′-deoxycytidine, in inhibiting cancer
cell proliferation.
References
|
1
|
Esteller M: Cancer Epigenetics for the
21st Century: What’s Next? Genes Cancer. 2:604–606. 2011.
|
|
2
|
Daniel FI, Cherubini K, Yurgel LS, de
Figueiredo MA and Salum FG: The role of epigenetic transcription
repression and DNA methyltransferases in cancer. Cancer.
117:677–687. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Bender CM, Pao MM and Jones PA: Inhibition
of DNA methylation by 5-aza-2′-deoxycytidine suppresses the growth
of human tumor cell lines. Cancer Res. 58:95–101. 1998.
|
|
4
|
Fang JY, Cheng ZH, Chen YX, Lu R, Yang L,
Zhu HY and Lu LG: Expression of Dnmt1, demethylase, MeCP2 and
methylation of tumor-related genes in human gastric cancer. World J
Gastroenterol. 10:3394–3398. 2004.PubMed/NCBI
|
|
5
|
Choi MS, Shim YH, Hwa JY, Lee SK, Ro JY,
Kim JS and Yu E: Expression of DNA methyltransferases in multistep
hepatocarcinogenesis. Hum Pathol. 34:11–17. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Ding WJ, Fang JY, Chen XY and Peng YS: The
expression and clinical significance of DNA methyltransferase
proteins in human gastric cancer. Dig Dis Sci. 53:2083–2089. 2008.
View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Peng DF, Kanai Y, Sawada M, Ushijima S,
Hiraoka N, Kosuge T and Hirohashi S: Increased DNA
methyltransferase 1 (DNMT1) protein expression in precancerous
conditions and ductal carcinomas of the pancreas. Cancer Sci.
96:403–408. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Arai E, Kanai Y, Ushijima S, Fujimoto H,
Mukai K and Hirohashi S: Regional DNA hypermethylation and DNA
methyltransferase (DNMT) 1 protein overexpression in both renal
tumors and corresponding nontumorous renal tissues. Int J Cancer.
119:288–296. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Vertino PM, Yen RW, Gao J and Baylin SB:
De novo methylation of CpG island sequences in human fibroblasts
overexpressing DNA (cytosine-5-)-methyltransferase. Mol Cell Biol.
16:4555–4565. 1996.PubMed/NCBI
|
|
10
|
Klotz DM, Nelson SA, Kroboth K, Newton IP,
Radulescu S, Ridgway RA, Sansom OJ, Appleton PL and Näthke IS: The
microtubule poison vinorelbine kills cells independently of mitotic
arrest and targets cells lacking the APC tumour suppressor more
effectively. J Cell Sci. 125:887–895. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Ohta S, Lai EW, Pang AL, Brouwers FM, Chan
WY, Eisenhofer G, de Krijger R, Ksinantova L, Breza J, Blazicek P,
Kvetnansky R, Wesley RA and Pacak K: Downregulation of metastasis
suppressor genes in malignant pheochromocytoma. Int J Cancer.
114:139–143. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Gilham DE, Debets R, Pule M, Hawkins RE
and Abken H: CAR-T cells and solid tumors: tuning T cells to
challenge an inveterate foe. Trends Mol Med. 18:377–384. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Riccio A and Cubellis MV: Gain of function
in CDKN1C. Nat Genet. 44:737–738. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Harinck F, Kluijt I, van der Stoep N,
Oldenburg RA, Wagner A, Aalfs CM, Sijmons RH, Poley JW, Kuipers EJ,
Fockens P, van Os TA and Bruno MJ: Indication for CDKN2A-mutation
analysis in familial pancreatic cancer families without melanomas.
J Med Genet. 49:362–365. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Guan CN, Zhang PW, Lou HQ, Liao XH and
Chen BY: DLC-1 expression levels in breast cancer assessed by
qRT-PCR are negatively associated with malignancy. Asian Pac J
Cancer Prev. 13:1231–1233. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Liu J, Lin Y, Yang H, Deng Q, Chen G and
He J: The expression of p33(ING1), p53 and autophagy-related gene
Beclin1 in patients with non-small cell lung cancer. Tumour Biol.
32:1113–1121. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Schultz DC, Vanderveer L, Berman DB,
Hamilton TC, Wong AJ and Godwin AK: Identification of two candidate
tumor suppressor genes on chromosome 17p13.3. Cancer Res.
56:1997–2002. 1996.PubMed/NCBI
|
|
18
|
Vaidyanathan G, Cismowski MJ, Wang G,
Vincent TS, Brown KD and Lanier SM: The Ras-related protein
AGS1/RASD1 suppresses cell growth. Oncogene. 23:5858–5863. 2004.
View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Berman SD, Calo E, Landman AS, Danielian
PS, Miller ES, West JC, Fonhoue BD, Caron A, Bronson R, Bouxsein
ML, Mukherjee S and Lees JA: Metastatic osteosarcoma induced by
inactivation of Rb and p53 in the osteoblast lineage. Proc Natl
Acad Sci USA. 105:11851–11856. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Kanwal S, Kayani MA and Faryal R:
Identification of novel SNPs in SYK gene of breast cancer patients:
computational analysis of SNPs in the 5′UTR. Mol Biol Rep.
39:8345–8351. 2012.PubMed/NCBI
|
|
21
|
Behjati R, Kawai K, Inadome Y, Kano J,
Akaza H and Noguchi M: APAF-1 is related to an undifferentiated
state in the testicular germ cell tumor pathway. Cancer Sci.
102:267–274. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Chêne L, Giroud C, Desgrandchamps F,
Boccon-Gibod L, Cussenot O, Berthon P and Latil A: Extensive
analysis of the 7q31 region in human prostate tumors supports TES
as the best candidate tumor suppressor gene. Int J Cancer.
111:798–804. 2004.PubMed/NCBI
|
|
23
|
da Costa Prando E, Cavalli LR and Rainho
CA: Evidence of epigenetic regulation of the tumor suppressor gene
cluster flanking RASSF1 in breast cancer cell lines. Epigenetics.
6:1413–1424. 2011.PubMed/NCBI
|
|
24
|
Takahashi M, Fujita M, Furukawa Y,
Hamamoto R, Shimokawa T, Miwa N, Ogawa M and Nakamura Y: Isolation
of a novel human gene, APCDD1, as a direct target of the
beta-Catenin/T-cell factor 4 complex with probable involvement in
colorectal carcinogenesis. Cancer Res. 62:5651–5656.
2002.PubMed/NCBI
|
|
25
|
Agueli C, Cammarata G, Salemi D, Dagnino
L, Nicoletti R, La Rosa M, Messana F, Marfia A, Bica MG, Coniglio
ML, Pagano M, Fabbiano F and Santoro A: 14q32/miRNA clusters loss
of heterozygosity in acute lymphoblastic leukemia is associated
with up-regulation of BCL11a. Am J Hematol. 85:575–578. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Zhang X, Liu H, Li B, Huang P, Shao J and
He Z: Tumor suppressor BLU inhibits proliferation of nasopharyngeal
carcinoma cells by regulation of cell cycle, c-Jun N-terminal
kinase and the cyclin D1 promoter. BMC Cancer. 12:2672012.
View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Wheeler SE, Morariu EM, Bednash JS, Otte
CG, Seethala RR, Chiosea SI and Grandis JR: Lyn kinase mediates
cell motility and tumor growth in EGFRvIII-expressing head and neck
cancer. Clin Cancer Res. 18:2850–2860. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Persson M, andrén Y, Moskaluk CA, Frierson
HF Jr, Cooke SL, Futreal PA, Kling T, Nelander S, Nordkvist A,
Persson F and Stenman G: Clinically significant copy number
alterations and complex rearrangements of MYB and NFIB in head and
neck adenoid cystic carcinoma. Genes Chromosomes Cancer.
51:805–817. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Xiong F, Wu C, Chang J, Yu D, Xu B, Yuan
P, Zhai K, Xu J, Tan W and Lin D: Genetic variation in an
miRNA-1827 binding site in MYCL1 alters susceptibility to
small-cell lung cancer. Cancer Res. 71:5175–5181. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Gilmore TD and Gerondakis S: The c-Rel
transcription factor in development and disease. Genes Cancer.
2:695–711. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Zhang L, Teng Y, Zhang Y, Liu J, Xu L, Qu
J, Hou K, Yang X, Liu Y and Qu X: C-Src-mediated RANKL-induced
breast cancer cell migration by activation of the ERK and Akt
pathway. Oncol Lett. 3:395–400. 2012.PubMed/NCBI
|
|
32
|
Memarian A, Hojjat-Farsangi M,
Asgarian-Omran H, Younesi V, Jeddi-Tehrani M, Sharifian RA,
Khoshnoodi J, Razavi SM, Rabbani H and Shokri F: Variation in WNT
genes expression in different subtypes of chronic lymphocytic
leukemia. Leuk Lymphoma. 50:2061–2070. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Dwyer MA, Joseph JD, Wade HE, Eaton ML,
Kunder RS, Kazmin D, Chang CY and McDonnell DP: WNT11 expression is
induced by estrogen-related receptor alpha and beta-catenin and
acts in an autocrine manner to increase cancer cell migration.
Cancer Res. 70:9298–9308. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Tanic M, Yanowsky K, Rodriguez-Antona C,
Andrés R, Márquez-Rodas I, Osorio A, Benitez J and Martinez-Delgado
B: Deregulated miRNAs in hereditary breast cancer revealed a role
for miR-30c in regulating KRAS oncogene. PLoS One. 7:e388472012.
View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Qiu JF, Zhang ZQ, Wang Y and You J:
Lentivirus-mediated RNAi knockdown of VEGFA in RKO colorectal
cancer cells decreases tumor formation and growth in vitro and in
vivo. Int J Clin Exp Pathol. 5:290–298. 2012.PubMed/NCBI
|
|
36
|
Seyhan AA, Varadarajan U, Choe S, Liu W
and Ryan TE: A genome-wide RNAi screen identifies novel targets of
neratinib resistance leading to identification of potential drug
resistant genetic markers. Mol Biosyst. 8:1553–1570. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Shen DW and Gottesman MM: RAB8 enhances
TMEM205-mediated cisplatin resistance. Pharm Res. 29:643–650. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Wang JS, Wang FB, Zhang QG, Shen ZZ and
Shao ZM: Enhanced expression of Rab27A gene by breast cancer cells
promoting invasiveness and the metastasis potential by secretion of
insulin-like growth factor-II. Mol Cancer Res. 6:372–382. 2008.
View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Shibata D, Mori Y, Cai K, Zhang L, Yin J,
Elahi A, Hamelin R, Wong YF, Lo WK, Chung TK, Sato F, Karpeh MS Jr
and Meltzer SJ: RAB32 hypermethylation and microsatellite
instability in gastric and endometrial adenocarcinomas. Int J
Cancer. 119:801–806. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Cong XL, Li B, Yang RC, Feng SZ, Chen SJ
and Han ZC: Enhanced growth suppression of Philadephia1 leukemia
cells by targeting bcr3/abl2 and VEGF through antisense strategy.
Leukemia. 19:1517–1524. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Sonvilla G, Allerstorfer S, Stättner S,
Karner J, Klimpfinger M, Fischer H, Grasl-Kraupp B, Holzmann K,
Berger W, Wrba F, Marian B and Grusch M: FGF18 in colorectal tumour
cells: autocrine and paracrine effects. Carcinogenesis. 29:15–24.
2008. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Delfortrie S, Pinte S, Mattot V, Samson C,
Villain G, Caetano B, Lauridant-Philippin G, Baranzelli MC,
Bonneterre J, Trottein F, Faveeuw C and Soncin F: Egfl7 promotes
tumor escape from immunity by repressing endothelial cell
activation. Cancer Res. 71:7176–7186. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Schenone S, Brullo C, Musumeci F, Biava M,
Falchi F and Botta M: Fyn kinase in brain diseases and cancer: the
search for inhibitors. Curr Med Chem. 18:2921–2942. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Jacobs JF, van Bokhoven H, van Leeuwen FN,
Hulsbergen-van de Kaa CA, de Vries IJ, Adema GJ, Hoogerbrugge PM
and de Brouwer AP: Regulation of MYCN expression in human
neuroblastoma cells. BMC Cancer. 9:2392009. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Lankat-Buttgereit B, Fehmann HC, Hering
BJ, Bretzel RG and Göke B: Expression of the ras-related rab3a gene
in human insulinomas and normal human pancreatic islets. Pancreas.
9:434–438. 1994. View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Jang KW, Lee KH, Kim SH, Jin T, Choi EY,
Jeon HJ, Kim E, Han YS and Chung JH: Ubiquitin ligase CHIP induces
TRAF2 proteasomal degradation and NF-κB inactivation to regulate
breast cancer cell invasion. J Cell Biochem. 112:3612–3620.
2011.PubMed/NCBI
|
|
47
|
Kawaguchi-Ihara N, Murohashi I, Nara N and
Tohda S: Promotion of the self-renewal capacity of human acute
leukemia cells by Wnt3A. Anticancer Res. 28:2701–2704.
2008.PubMed/NCBI
|
|
48
|
Charlton A, Blair V, Shaw D, Parry S,
Guilford P and Martin IG: Hereditary diffuse gastric cancer:
predominance of multiple foci of signet ring cell carcinoma in
distal stomach and transitional zone. Gut. 53:814–820. 2004.
View Article : Google Scholar : PubMed/NCBI
|
|
49
|
Rountree MR, Bachman KE, Herman JG and
Baylin SB: DNA methylation, chromatin inheritance and cancer.
Oncogene. 20:3156–3165. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
50
|
Schaefer M, Hagemann S, Hanna K and Lyko
F: Azacytidine inhibits RNA methylation at DNMT2 target sites in
human cancer cell lines. Cancer Res. 69:8127–8132. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
51
|
Ooi SK, Qiu C, Bernstein E, Li K, Jia D,
Yang Z, Erdjument-Bromage H, Tempst P, Lin SP, Allis CD, Cheng X
and Bestor TH: DNMT3L connects unmethylated lysine 4 of histone H3
to de novo methylation of DNA. Nature. 448:714–717. 2007.
View Article : Google Scholar : PubMed/NCBI
|
|
52
|
Bourc’his D, Xu GL, Lin CS, Bollman B and
Bestor TH: Dnmt3L and the establishment of maternal genomic
imprints. Science. 294:2536–2539. 2001.PubMed/NCBI
|
|
53
|
Jia D, Jurkowska RZ, Zhang X, Jeltsch A
and Cheng X: Structure of Dnmt3a bound to Dnmt3L suggests a model
for de novo DNA methylation. Nature. 449:248–251. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
54
|
Stefanska B, Huang J, Bhattacharyya B,
Suderman M, Hallett M, Han ZG and Szyf M: Definition of the
landscape of promoter DNA hypomethylation in liver cancer. Cancer
Res. 71:5891–5903. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
55
|
Tan AC, Jimeno A, Lin SH, Wheelhouse J,
Chan F, Solomon A, Rajeshkumar NV, Rubio-Viqueira B and Hidalgo M:
Characterizing DNA methylation patterns in pancreatic cancer
genome. Mol Oncol. 3:425–438. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
56
|
Dimberg J, Ström K, Löfgren S, Zar N,
Lindh M and Matussek A: DNA promoter methylation status and protein
expression of interleukin-8 in human colorectal adenocarcinomas.
Int J Colorectal Dis. 27:709–714. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
57
|
Radhakrishnan VM, Jensen TJ, Cui H,
Futscher BW and Martinez JD: Hypomethylation of the 14-3-3σ
promoter leads to increased expression in non-small cell lung
cancer. Genes Chromosomes Cancer. 50:830–836. 2011.
|