Introduction
Cyclin-dependent kinase 2-associated protein 1
(CDK2AP1) is a putative growth suppressor gene originally
identified and isolated from normal keratinocytes (1). The human CDK2AP1 gene is 1.6 kb long
and is located on chromosome 12q24.31 (2). The human and rodent polypeptides
encoded by the CDK2AP1 gene share 97% identity, while the mouse and
hamster sequences are identical (3). Human CDK2AP1 encodes a 115-amino acid
peptide with a molecular mass of 12.4 kDa (pI 9.62), namely
p12CDK2AP1(4). Previous
studies have demonstrated that p12CDK2AP1 negatively
regulates cell growth by sequestering the monomeric
non-phosphorylated form of CDK2 and targeting it for proteolysis to
reduce the active pool of CDK2 in cells (5). The protein also reduces CDK2-mediated
retinoblastoma protein phosphorylation by reversing the activity of
TGF-β (6). A previous study has
demonstrated that miR-21 downregulates p12CDK2AP1 to
stimulate cell proliferation and invasion (7). These observations indicate that
p12CDK2AP1 acts as a growth suppressor through
influencing mitosis, the S phase and the cell division cycle. A
study has demonstrated that p12CDK2AP1 is also important
in promoting Oct4 promoter methylation during murine embryonic stem
cell differentiation and thereby downregulates Oct4 expression
levels (8). In addition,
p12CDK2AP1 has been shown to induce the
epithelial-mesenchymal transition of hamster cheek pouch
carcinoma-1 cells by promoting the expression of Twist2 (9).
Other clinical studies have demonstrated that
p12CDK2AP1 is associated with oral squamous cell
carcinoma, breast cancer, esophageal squamous cell carcinoma,
gastric cancer and colorectal cancer (10–14).
It has also been identified that p12CDK2AP1 is involved
in anti-angiogenesis modulation and the suppression of malignant
biological interactions between prostate cancer and bone cells
(15). The possibility of using
p12CDK2AP1 as a therapeutic target has been suggested
(16).
Proteins undertake numerous physiological functions,
such as the regulation of enzymatic activities, the assembly of
cellular components and signal transduction. The elucidation of
protein-protein interactions (PPIs) and whole interaction networks
is vital to understanding biophysical processes. It is well known
that numerous types of cancer are caused by dysfunctions of certain
protein interactions (17).
Studies have demonstrated that a number of proteins interact with
p12CDK2AP1(18,19). However, due to the diverse
properties of proteins and the very different characteristics of
PPIs (20), the proteins that
interact with p12CDK2AP1 are not yet fully
understood.
In the present study, a yeast two-hybrid assay was
used to screen the proteins possibly interacting with
p12CDK2AP1 and a novel unnamed protein product (UPP) was
identified. The interaction between p12CDK2AP1 and this
UPP was verified by glutathione S-transferase (GST) pull-down and
co-immunoprecipitation (Co-IP) assays. Subsequently, the functions
of the UPP in the PPIs with p12CDK2AP1 were studied
in vitro and in vivo. The results demonstrated that
p12CDK2AP1 regulates the expression of the UPP and may be inhibit
cell proliferation by interaction with it.
Materials and methods
Materials
All chemicals were purchased from Sigma-Aldrich (St.
Louis, MO, USA), unless otherwise mentioned. Dulbecco’s modified
Eagle’s medium (DMEM), penicillin-streptomycin, Dulbecco’s
phosphate-buffered saline (PBS) and fetal bovine serum (FBS) were
purchased from Invitrogen (Carlsbad, CA, USA).
Yeast strain and growth medium
The genotypes of the Saccharomyces cerevisiae
strain MaV203 (Invitrogen) used in this study were as follows:
MATα, leu2-3, 112, trp1-901, his3Δ200, ade2-101, gal4Δ, gal80Δ
SPAL10::URA3, GAL1::lacZ, HIS3UAS GAL1::HIS3@LYS2, can1R and cyh2R.
This strain contains a set of non-reverting autotrophic mutations.
A blend of yeast extract, peptone, dextrose and adenine sulfate
medium was used to grow the untransformed yeast strain. Synthetic
dropout minimal medium (Clontech Laboratories, Inc., Victoria,
Australia) with appropriate nutrients was used to separate the
untransformed yeast strain from the transformed.
Plasmid construction
Human liver CDK2AP1 cDNA (Stratagene, La Jolla, CA,
USA) was ligated between the EcoR I and Xho I sites
of a pBD-GAL4 Cam phagemid vector (Stratagene), downstream of the
GAL4 DNA-binding domain. A liver cDNA library was digested by an
interference-resistant helper phage from the Cam Phagemid Vector
kit (Stratagene) and transformed into the Escherichia coli
(E. coli) XLOLR strain (Stratagene). The human CDK2AP1 cDNA
and the UPP gene were cloned into the pGEX-4T-1 GST fusion
expression vector (GE Healthcare, Piscataway, NJ, USA). As there
was no commercially available antibody to detect the UPP, HA and
FLAG protein tags were used to identify the expression of the UPP
and p12CDK2AP1. HA and FLAG mouse monoclonal antibodies
(Zhongshan Golden-Bridge Biotechnology Co., Ltd., Beijing, China)
were used to test the expression of HA-p12CDK2AP1 and
FLAG-UPP in eukaryotes.
Yeast two-hybrid cDNA library screening
assay
A human cDNA library was screened using the pBD-GAL4
plasmid, which was constructed with human full CDK2AP1 cDNA. The
cDNA library bacterial XL1-Blue MRF was digested by a phage. The
two vectors were then cotransfected into the MaV203 S.
cerevisiae strain. Subsequently, positive clones were verified
by distinguishing the His+ phenotype from the
Leu+ and Try+ phenotypes. The screening
procedure fully complied with the manufacturer’s instructions.
Positive interaction was confirmed by testing the dropout culture
media, and then the positive plasmid was transformed into E.
coli JM109 for sequencing.
Pull-down assay
The pGEX 4T-p12CDK2AP1 and pGEX 4T-UPP
plasmids were transformed into BL21 E. coli to express
GST-p12CDK2AP1 and GST-UPP fusion proteins. Isopropyl
β-D-1-thiogalactopyranoside (200 μl, 1 mM) was added to the flasks
and the cells were induced for 5 h. The cells were then harvested
with PBS. Pelleted cells were lysed by sonication in PBS and
Complete Protease Inhibitor Cocktail (Roche, Mannheim, Germany).
Following centrifugation at 12,000 × g for 5 min, the supernatants
were applied to Glutathione Sepharose 4B (GE Healthcare). Thrombin
Protease (GE Healthcare) was added to the GST-UPP tube for 30 min.
The beads were pelleted, washed three times in PBS, resuspended in
Laemmli sample buffer and analyzed by 15% sodium dodecyl
sulfate-polyacrylamide gel electrophoresis (SDS-PAGE).
Cell culture and transfection
Human 293T cells and HeLa cells were cultured under
37°C, 5% CO2 in DMEM supplemented with 10% FBS,
penicillin (10,000 U/ml) and streptomycin (10,000 μg/ml). When the
cells reached 50–60% confluence, Lipofectamine 2000 reagent (Life
Technologies Corp., Rockville, MD, USA) was used to transfect the
pcDNA 3.1-FLAG-UPP and pcDNA 3.1-HA-p12CDK2AP1 plasmids
according to the manufacturer’s instructions. The medium was
changed after 5 h.
Co-IP and western blot analysis
A Protein G Immunoprecipitation kit was used to
perform the Co-IP assay according to the instructions provided with
the kit. The 293T cells were lysed in IP buffer 48 h after
transfection. The lysate was divided into two spin columns and the
samples were analyzed by SDS-PAGE. Proteins were transferred onto
nitrocellulose membranes (Invitrogen) and then incubated with
primary antibodies. The anti-HA antibody (1:10,000/2 μl) was
applied to the UPP membrane, while the anti-FLAG antibody
(1:10,000/2 μl) was applied to the p12CDK2AP1 membrane.
After washing four times with TBST (20 mM Tris-HCl, pH 7.4; 137 mM
NaCl and 0.1% (v/v) Tween 20), the membranes were incubated with
horseradish peroxidase-conjugated goat anti-mouse secondary
antibodies. The ECL Western Blotting Detection system (Amersham
Biosciences, Piscataway, NJ, USA) was used to detect horseradish
peroxidase on the immunoblots. The dried membranes were exposed to
Carestream Kodak Biomax MR film for 2 min at room temperature.
siRNA and qPCR assays
The siRNA sequences used were designed and purchased
from Genepharma Co. (Shanghai, China), and were as follows:
Forward, 5′-UCUUUUCCAAACUGAAUGCGC-3′ and reverse,
5′-GCAUUCAGUUUGGAAAAGACA-3′ for the UPP gene; and forward,
5′-AGGAGAUCAGACCCACGUATT-3′ and reverse 5′-UACGUGGGUCUGAUCUCCUTT-3′
for the p12CDK2AP1 gene. The siRNA oligo was transfected
into the 293T cell line with Lipofectamine 2000. After 24 h, the
cells were harvested and divided into two centrifuge tubes. Total
RNA from the cultured cells was isolated using TRIzol reagent (Life
Technologies Corp., Carlsbad, CA, USA). The primers used to detect
the expression levels were as follows: Forward, 5′-AACGGAACGGAATGC
CAGATCCTA-3 and reverse, 5′-TTCAGAGCCAAGTGA ACCATGGGA-3′ for
p12CDK2AP1; and forward, 5′-TGGTGC ACATCGTGTAACCTGTCT-3′
and reverse, 5′-AAGCTT TCCTTCCTCAGCCACTCT-3′ for the UPP. Following
cDNA synthesis, qPCR was performed using the Bio-Rad Real-Time PCR
Detection system (Bio-Rad, Hercules, CA, USA) with the human
housekeeping cDNA GAPDH as the control. Radio immunoprecipitation
assay buffer cocktail (Dingsheng Biotechnological Co., Beijing,
China) was added to the tube containing 293T cells. Anti-mouse HA
monoclonal antibody was used to perform western blotting to test
the expression of p12CDK2AP1. The human 293T, A-549,
HeLa, MCF-7, ACC-M, fibroblast and human mesenchymal stem cell
lines were incubated to examine the endogenous expression of the
p12CDK2AP1 and UPP genes. The comparative threshold
cycle (CT) method was used to analyze relative changes in gene
expression.
Flow cytometry and cell viability
assay
The UPP-transfected 293T and HeLa cells were
harvested at 12 and 24 h. Prior to flow cytometric examination, the
cells were combined with propidium iodide (1 mg/ml) and RNase (50
μg/ml). The cell cycle was analyzed by flow cytometry (FACSCalibur;
Becton-Dickinson, San Jose, CA, USA), and ModFit and CellQuest
software was utilized. The plasmid pcDNA 3.1-FLAG-UPP was
transfected into 293T cells.
3-(4,5-Dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (20
μl, 5 mg/ml) was applied to the wells after incubation for 24 h.
Dimethyl sulfoxide (Amersco, Solon, OH, USA) was added to each well
to dissolve the formazan crystals for 10 min. The samples were then
transferred to a plate reader (ELx808; BioTek, Shanghai, China) and
the absorbance at 570 nm was measured.
DNA methylation assay
The bisulfite sequencing PCR (BSP) method was used
to examine the UPP gene promoter methylation in the 293T and HeLa
cell lines. The methylation primers used to perform PCR were as
follows: Forward, 5′-GTAGTG TGAGTGAGGGTTTTGATTT-3′ and reverse,
5′-CTAACA ACAAATTACCCCAACTTTC-3′. Streptozotocin buffer (Tris-HCl
10 mM, pH 8.0; EDTA 4 mM; NaCl 0.4 M), SDS and protease K were
applied to harvested 293T and HeLa cells. DNA extraction followed
the standard procedure. Na2S2O5 (5
mol/l) and hydroquinone mixture were used to prepare HeLa and 293T
cell DNA. Subsequently, PCR was performed with the conditions as
follows: 94°C for 5 min, 94°C for 40 sec, 56°C for 40 sec and 72°C
for 50 sec for 34 cycles, and then 72°C for 10 min. An E.Z.N.A.
Cycle Pure kit (Omega Bio-Tek Inc., Norcross, GA, USA) was used to
purify the PCR products. Subsequently, the PCR products were
ligated into a pMD 18T vector (Takara Bio, Inc., Tokyo, Japan) for
sequencing. BiQ Analyzer software was used to statistically analyze
the results (21).
Human tumor xenografts in vivo
The animal experiment followed the Laboratory Animal
Center Institutional Animal Care and Use committee guidelines of
the Peking University Health Science Center. Nude mice (n=36;
Balb/c nude, female, four weeks old) were divided into three groups
at random. The HeLa cells which were stably transfected with the
UPP gene by an Overexpression Lentivector (SBI, Mountain View, CA,
USA) were screened by flow cytometry. HeLa-UPP cells were
subcutaneously injected at a density of 1×105 cells/ml
into the armpits of the nude mice, with the HeLa cell line as the
negative control and the blank plasmid-infected HeLa cells as the
positive control. Once the tumors reached 3–5 mm, all the tumors
were harvested. The weights and volumes of the tumors were then
calculated at the same time.
Statistical analysis
All cell experiments were repeated three times. The
data were calculated as the mean ± standard deviation and the
statistical analyses were performed using SPSS Software, version
19.0 (SPSS, Inc., Chicago, IL, USA). The differences were
considered to be statistically significant when P<0.05, as
determined by Student’s t-test.
Results
Identification of a novel interaction
protein of p12CDK2AP1 by a yeast two-hybrid screening
assay
Interacting cellular partners of
p12CDK2AP1 were searched for by screening a human cDNA
library using a bait construct with the full CDK2AP1 cDNA cloned
in-frame through using a yeast hybrid system. Following screening,
positive clones were identified by selecting with synthetic dropout
minimal medium. A protein that activated yeast transcription was
detected (Fig. 1A). The activating
domain/library plasmids were confirmed by the cotransformation
procedure, and the nucleotide sequence of the protein was obtained
by cloning and sequencing. Through blasting the sequence in the
NCBI BLAST database, a novel protein named UPP (BC006130) was
identified. The UPP is encoded by the Loc93622 gene, which is
located on 4q16.1, and includes 119 amino acids.
Verification of the interaction between
p12CDK2AP1 and the UPP in vitro
Considering the possibility of a false positive
result in the yeast two-hybrid system, the interaction between
p12CDK2AP1 and the UPP was confirmed by GST-pull down.
The prokaryotic expression plasmids pGEX-4T-p12CDK2AP1
and pGEX-4T-UPP were constructed to express GST fusion proteins,
and then the GST-p12CDK2AP1 and GST-UPP proteins were
purified by GST affinity chromatography (Fig. 1B and C). Subsequently, the GST tag
of GST-UPP was cleaved by thrombin protease. The pull-down result
supports the specific interaction of p12CDK2AP1 with the
UPP (Fig. 1D). To confirm this
interaction in mammalian cells, the interaction of
p12CDK2AP1 and the UPP was examined using a Co-IP assay.
The bound proteins were detected by western blotting using
antibodies which were specific to HA and FLAG (1E and F). These
results demonstrated that p12CDK2AP1 interacts with the
UPP in cells.
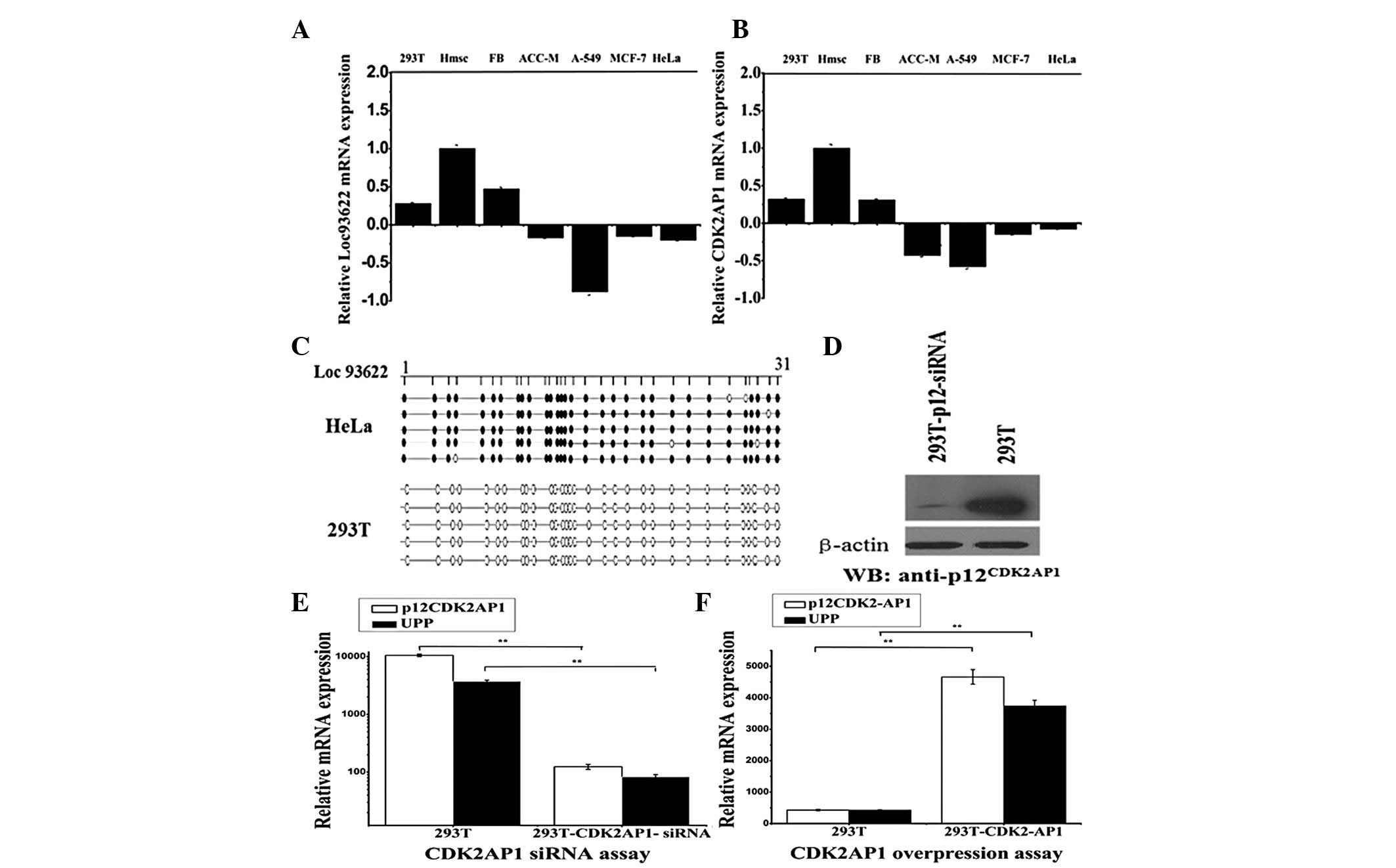
Expression and methylation of the UPP
gene
To analyze the endogenous expression of the UPP
gene, non-neoplastic cell lines (293T, human mesenchymal stem cells
and human fibroblast cells) and human cancer cell lines (ACC-M,
A-549, MCF-7 and HeLa) were tested. The results of the qPCR
demonstrated that the UPP and p12CDK2AP1 were similarly
expressed (Fig. 2A and B), that
is, the UPP expression levels were higher in the non-neoplastic
cell types than in cancer cell lines. Due to the high GC ratio
(63%) of the UPP gene, the UPP gene was searched in the NCBI
database and a CpG island of the gene was identified. The
methylation of the UPP gene was investigated in the 293T and HeLa
cell lines. The DNA of 293T and HeLa cells was extracted and
amplified by BSP methods. The sequencing results demonstrated that
there were 31 methylation sites in the UPP gene. In the DNA of the
HeLa cells, all the methylation sites were methylated, while the
293T DNA was not methylated (Fig.
2C).
CDK2AP1 mediates expression of the UPP
gene
To investigate the association between CDK2AP1 and
the UPP gene, siRNA and overexpression assays were performed.
Following transfection of the p12CDK2AP1 siRNA oligo,
the western blot analysis demonstrated the silencing effect of the
CDK2AP1 gene (Fig. 2D). The
results indicated that the UPP expression levels were decreased
when the CDK2AP1 gene was interfered with (Fig. 2E), while the UPP siRNA did not
downregulate the expression levels of the CDK2AP1 gene (data not
shown). Subsequently, the association between CDK2AP1 and the UPP
was evaluated by overexpressing the two genes in the 293T cell
line. The results revealed that overexpression of the CDK2AP1 gene
significantly increases the expression levels of the UPP gene
(Fig. 2F), while overexpression of
the UPP gene does not affect the expression of CDK2AP1 (data not
shown). Thus, the UPP may function as a downstream effector of
CDK2AP1.
Overexpression of p12CDK2AP1
and UPP modulates the cell cycle and inhibits cell vitality in
vitro
p12CDK2AP1 mediates the cell cycle and
affects cell vitality; thus, whether the UPP gene also shifts the
cell cycle and is associated with cell vitality was investigated.
The UPP and p12CDK2AP1 genes were transfected into 293T
and HeLa cells. After the genes had been transfected for 24 h, the
flow cytometry data indicated that the percentage of cells in the
G2/M phase was decreased significantly from 11.13% to 6.16% in the
293T cells (Fig. 3A) and from
12.15% to 7.65% in the HeLa cells (Fig. 3B). The cell growth curve
demonstrated that overexpression of the UPP also suppresses cell
proliferation (Fig. 3C and D).
After transfection for 5 days, the cell vitality results indicated
that overexpression of the UPP decreased cell survival (Fig. 3E and F).
Stably-transfected UPP inhibits the
growth of human tumor xenografts in vivo
To investigate the mechanisms of action of the UPP
protein in vivo, a HeLa cell line stably transfected with
the UPP gene (Hela-UPP) was constructed (Fig. 4A and B) and transplanted in nude
mice as xenografts. After 62 days, the xenograft tumors were
harvested and fixed (Fig. 4C and
D), and the weight and volume of the tumors were measured. The
results demonstrated that the mean weight of the tumors of the
HeLa-UPP cells (0.38±0.07 g) was lower than that of the control
cells (0.77±0.11 g; Fig. 4E), and
the mean volume of the tumors in the HeLa-UPP cells (109.62±20.40
mm3) was also lower than that of the control group
(228.62±76.58 mm3; Fig.
4F). These results indicated that the UPP effectively inhibits
the growth of human tumor xenografts.
Discussion
Mechanistic studies have demonstrated that
p12CDK2AP1 binds to DNA polymerase α/primase protein,
thereby inhibiting DNA replication (6). In addition, p12CDK2AP1
interacts with the monomeric non-phosphorylated form of the CDK2
protein to inhibit the S phase of the cell cycle (19). Thus, the PPIs of
p12CDK2AP1 have a number of vital roles in physiological
processes. However, the PPIs of p12CDK2AP1 require
further study and the present study sought to identify proteins
which interact with p12CDK2AP1.
Through using the two-hybrid system assay, a novel
protein (UPP, BC006130) was identified to interact with
p12CDK2AP1. The UPP is a 119-amino acid protein and its
molecular mass is ~12.4 kDa. The protein is encoded by the Loc93622
gene, which is located on chromosome 4p16.1. Subsequently, the
interaction was verified using pull-down and Co-IP assays. As the
mechanism of the novel gene and protein has not been studied
previously, the function of the protein was studied. To elucidate
the association between the two genes, silencing and overexpressing
assays were utilized. The results demonstrated that the expression
of the UPP gene was mediated by the p12CDK2AP1 gene.
However, the overexpression and silencing the UPP gene did not
affect the expression of p12CDK2AP1. This indicated that
the p12CDK2AP1 gene may regulate the expression of the
UPP and, as the p12CDK2AP1 gene is a potent
transcription repressor, the UPP gene may be located downstream of
the p12CDK2AP1 gene. Subsequently, the endogenous
expression of the UPP gene in different types of cell was detected.
The expression of the UPP gene was observed to be higher in
non-neoplastic cell lines than in cancer cell lines and, similar to
the expression pattern of the p12CDK2AP1 gene, the
expression of the UPP gene also demonstrated tissue specificity.
This result also suggested that p12CDK2AP1 may mediate
the expression of the UPP gene.
Two studies have demonstrated that the CDK2AP1 gene
is a core subunit of the methyl CpG binding domain protein 2 and 3
NuRD complexes (MBD2/NuRD and MBD3/NuRD) (22,23).
The CDK2AP1 gene is putatively associated with a methylated
promoter as a component of the MBD2/NuRD complex and is associated
with carcinogenesis. Therefore, the CDK2AP1 gene may be involved in
epigenetic mechanisms. In the present study, the BSP method was
used to examine the methylation of the Loc93622 gene in the 293T
and HeLa cell lines. The results demonstrated that the gene is
hypermethylated in HeLa cells, which explained why the expression
levels of the Loc93622 gene were downregulated in the HeLa cells.
The result also verified that the Loc93622 gene was methylated in
certain cell lines in the UCSC Genome Bioinformatics database
(http://genome.ucsc.edu/). While the homologs of
the UPP, MORF4Ap1 and MORF4AP1L1 share sequence identity, they may
have the similar function of interacting with histone acetylases
and acetyltransferases to regulate chromatin dynamics (24). Further studies are required to
address the association between the function of the CDK2AP1 gene
and the Loc93622 gene methylation.
p12CDK2AP1 inhibits cancer cell
proliferation through decreasing the length of the S phase of the
cell cycle and it also changes the proportions of cells in the G1
and S phases. During carcinogenesis, the transcriptional function
of the p12CDK2AP1-encoding gene (CDK2AP1) has been
observed to be lost (25).
Zolochevska et al(15)
studied the CDK2AP1 mechanism in prostate cancer and they
demonstrated that the gene inhibits cancer proliferation and
changes the androgen-responsive pathway. In another study of the
prostate cancer androgen receptor, it was identified that the UPP
gene was upregulated in hormonal therapy-resistant prostate cancer
(26). In the present study,
overexpression of the UPP gene was performed to identify the
functions of the UPP in the cell cycle. The results demonstrated
that the length of the G2/M phase was decreased and that cell
proliferation was suppressed. Moreover, the HeLa cells transfected
with the UPP gene exhibited a lower tumorigenicity than the control
in human tumor xenografts. As an interactive protein of
p12CDK2AP1, the UPP may participate in the inhibition of
cancer cell proliferation with p12CDK2AP1. This study
speculated that p12CDK2AP1 not only controls S phase but
also modulates the G2/M phase of the cell cycle by interacting with
the UPP.
In conclusion, the present study demonstrated that
the UPP is a novel binding protein of p12CDK2AP1. The
interaction was verified by using in vitro and in
vivo assays. Through studying the mechanisms of the interaction
between the UPP and p12CDK2AP1, it was demonstrated that
the expression of the UPP gene was regulated by the
p12CDK2AP1 gene. Furthermore, p12CDK2AP1 may
be involved in the methylation of the UPP gene promoter. The novel
UPP inhibits tumor cell proliferation by regulating the cell cycle.
These findings extend the understanding of the anticancer mechanism
of p12CDK2AP1 in cancer therapy strategies.
Acknowledgements
This study was supported by grants from the National
Natural Science Foundation of China (project no. 81072230) and the
Ministry of Science and Technology of China (project no.
2010CB529403).
References
|
1
|
Todd R, McBride J, Tsuji T, et al: Deleted
in oral cancer-1 (doc-1), a novel oral tumor suppressor gene. FASEB
J. 9:1362–1370. 1995.PubMed/NCBI
|
|
2
|
Daigo Y, Suzuki K, Maruyama O, et al:
Isolation, mapping and mutation analysis of a human cDNA homologous
to the doc-1 gene of the Chinese hamster, a candidate tumor
suppressor for oral cancer. Genes Chromosomes Cancer. 20:204–207.
1997. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Tsuji T, Duh FM, Latif F, et al: Cloning,
mapping, expression, function, and mutation analyses of the human
ortholog of the hamster putative tumor suppressor gene Doc-1. J
Biol Chem. 273:6704–6709. 1998. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Kohno Y, Patel V, Kim Y, et al: Apoptosis,
proliferation and p12(doc-1) profiles in normal, dysplastic and
malignant squamous epithelium of the Syrian hamster cheek pouch
model. Oral Oncol. 38:274–280. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Rosenblatt J, Gu Y and Morgan DO: Human
cyclin-dependent kinase 2 is activated during the S and G2 phases
of the cell cycle and associates with cyclin A. Proc Natl Acad Sci
USA. 89:2824–2828. 1992. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Matsuo K, Shintani S, Tsuji T, et al:
p12(DOC-1), a growth suppressor, associates with DNA polymerase
alpha/primase. FASEB J. 14:1318–1324. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Zheng J, Xue H, Wang T, et al: miR-21
downregulates the tumor suppressor P12 CDK2AP1 and stimulates cell
proliferation and invasion. J Cell Biochem. 112:872–880. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Deshpande AM, Dai YS, Kim Y, et al:
Cdk2ap1 is required for epigenetic silencing of Oct4 during murine
embryonic stem cell differentiation. J Biol Chem. 284:6043–6047.
2009. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Tsuji T, Ibaragi S, Shima K, et al:
Epithelial-mesenchymal transition induced by growth suppressor
p12CDK2-AP1 promotes tumor cell local invasion but suppresses
distant colony growth. Cancer Res. 68:10377–10386. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Cwikla SJ, Tsuji T, McBride J, Wong DT and
Todd R: doc-1-mediated apoptosis in malignant hamster oral
keratinocytes. J Oral Maxillofac Surg. 58:406–414. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Tao K, Fang M, Alroy J and Sahagian GG:
Imagable 4T1 model for the study of late stage breast cancer. BMC
Cancer. 8:2282008. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Hiyoshi Y, Watanabe M, Hirashima K, et al:
p12CDK2-AP1 is associated with tumor progression and a poor
prognosis in esophageal squamous cell carcinoma. Oncol Rep.
22:35–39. 2009.PubMed/NCBI
|
|
13
|
Choi MG, Sohn TS, Park SB, et al:
Decreased expression of p12 is associated with more advanced tumor
invasion in human gastric cancer tissues. Eur Surg Res. 42:223–229.
2009. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Shin J, Yuan Z, Sreeramoju P, et al: A del
T poly T (8) mutation in the 3′ untranslated region (UTR) of the
CDK2-AP1 gene is functionally significant causing decreased mRNA
stability resulting in decreased CDK2-AP1 expression in human
microsatellite unstable (MSI) colorectal cancer (CRC). Surgery.
142:222–227. 2007.PubMed/NCBI
|
|
15
|
Zolochevska O and Figueiredo ML:
Cell-cycle regulators cdk2ap1 and bicalutamide suppress malignant
biological interactions between prostate cancer and bone cells.
Prostate. 71:353–367. 2011. View Article : Google Scholar
|
|
16
|
Figueiredo ML, Kim Y, St John MA and Wong
DT: p12CDK2-AP1 gene therapy strategy inhibits tumor growth in an
in vivo mouse model of head and neck cancer. Clin Cancer Res.
11:3939–3948. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Kar G, Gursoy A and Keskin O: Human cancer
protein-protein interaction network: a structural perspective. PLoS
Comput Biol. 5:e10006012009. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Buajeeb W, Zhang X, Ohyama H, et al:
Interaction of the CDK2-associated protein-1, p12(DOC-1/CDK2AP1),
with its homolog, p14(DOC-1R). Biochem Biophys Res Commun.
315:998–1003. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Shintani S, Ohyama H, Zhang X, et al:
p12(DOC-1) is a novel cyclin-dependent kinase 2-associated protein.
Mol Cell Biol. 20:6300–6307. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Bonetta L: Protein-protein interactions:
Interactome under construction. Nature. 468:851–854. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Bock C, Reither S, Mikeska T, Paulsen M,
Walter J and Lengauer T: BiQ Analyzer: visualization and quality
control for DNA methylation data from bisulfite sequencing.
Bioinformatics. 21:4067–4068. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Le Guezennec X, Vermeulen M, Brinkman AB,
et al: MBD2/NuRD and MBD3/NuRD, two distinct complexes with
different biochemical and functional properties. Mol Cell Biol.
26:843–851. 2006.PubMed/NCBI
|
|
23
|
Spruijt CG, Bartels SJ, Brinkman AB, et
al: CDK2AP1/DOC-1 is a bona fide subunit of the Mi-2/NuRD complex.
Mol Biosyst. 6:1700–1706. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Yochum GS and Ayer DE: Role for the
mortality factors MORF4, MRGX, and MRG15 in transcriptional
repression via associations with Pf1, mSin3A, and Transducin-Like
Enhancer of Split. Mol Cell Biol. 22:7868–7876. 2002. View Article : Google Scholar
|
|
25
|
Peng H, Shintani S, Kim Y and Wong DT:
Loss of p12CDK2-AP1 expression in human oral squamous cell
carcinoma with disrupted transforming growth factor-beta-Smad
signaling pathway. Neoplasia. 8:1028–1036. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Marques RB, Dits NF, Erkens-Schulze S, van
Ijcken WF, van Weerden WM and Jenster G: Modulation of androgen
receptor signaling in hormonal therapy-resistant prostate cancer
cell lines. PLoS One. 6:e231442011. View Article : Google Scholar : PubMed/NCBI
|