Introduction
The ALK fusion gene is a key oncogenic driver
in a subset of patients with non-small cell lung carcinomas
(NSCLCs) (1–5). EML4-ALK, consisting of the N-terminal
portion of EML4 ligated to the intracellular region of the
receptor-type protein tyrosine kinase ALK, has been detected in
~2–7% of NSCLC patients (1–6). EML4-ALK-positive NSCLC is associated
with several characteristics, such as early-onset, a never- or
light-smoking history, adenocarcinoma and mutual exclusiveness with
EGFR or KRAS mutations (7). Recently, crizotinib, a small molecule
inhibitor of ALK, was shown to selectively inhibit the growth of
ALK-positive NSCLC (6–8), indicating that a subclass of NSCLC
patients are likely to benefit clinically from an ALK inhibitor.
Therefore, molecular data regarding oncogenic fusion may have a
significant clinical impact.
Both ROS1 and RET receptor tyrosine kinases have
recently been identified as oncogenic fusions in NSCLC (4,5,9–17).
The expression of these fusion proteins transforms noncancerous
cells (4,14). Among some types of such fusion
forms, SLC34A2-ROS1, EZR-ROS1, CD74-ROS1 and KIF5B-RET are
relatively recurrent (4,5,9–17).
However, since the incidence of ROS1 or RET fusions is less than
that of ALK fusions in NSCLC, only a small number of NSCLC patients
with ROS1 or RET fusions has thus far been identified. In the
present study, to contribute to the elucidation of the
characteristics of ROS1 or RET fusion-positive NSCLC, we examined
114 NSCLCs derived from Japanese patients for the expression of
ROS1 and RET fusion transcripts and pathohistologically and
molecularly characterized those fusions that were detected.
Materials and methods
Primary lung carcinoma
Samples of surgical specimens were obtained from 114
NSCLC patients who underwent surgery for cancer at Mikatahara
Seirei General Hospital and Hamamatsu University Hospital. Informed
consent to use the resected tissues for genetic analysis was
obtained from all the patients and the study was approved by the
Institutional Review Boards (IRBs) of Hamamatsu University School
of Medicine and Mikatahara Seirei General Hospital. The
clinicopathological profiles of the cases are shown in Table I. The staging and histological
classification were based on the World Health Organization
system.
 | Table ISummary of the clinicopathological
profiles of the patients. |
Table I
Summary of the clinicopathological
profiles of the patients.
| Characteristic | n |
|---|
| No. of
patients | 114 |
| Age, years (mean ±
SD) | 68.5±6.0 |
| Gender, n (%) |
| Male | 87 (76.3) |
| Female | 27 (23.7) |
| Smoking, n (%) |
| Current
smokers | 49 (43.0) |
| Ex-smokers | 37 (32.5) |
| Non-smokers | 28 (24.5) |
| Histology, n
(%) |
|
Adenocarcinoma | 69 (60.5) |
| Squamous cell
carcinoma | 39 (34.2) |
| Adenosquamous cell
carcinoma | 4 (3.5) |
| Others | 2 (1.8) |
| Stage, n (%) |
| I | 54 (47.4) |
| II | 33 (28.9) |
| III | 27 (23.7) |
| IV | 0 (0.0) |
Detection of ROS1 and RET gene fusion
transcripts using reverse transcription (RT)-polymerase chain
reaction (PCR)
Total RNA was extracted from the lung tissue samples
using an RNeasy kit (Qiagen, Valencia, CA, USA) and was converted
to first-strand cDNA using a SuperScript First-Strand Synthesis
System for RT-PCR (Invitrogen, Carlsbad, CA, USA) according to the
supplier’s protocol. PCR was performed in 20-μl reaction mixtures
containing HotStarTaq DNA Polymerase (Qiagen). The following PCR
primers were used: 5′-GGGATTGGGATATTGATTTTAC-3′ and 5′-AGCTCA
GCCAACTCTTTGTCT-3′ for the SLC34A2-ROS1 fusion transcript;
5′-ACCGTGGAGAGAGAGAAAGAG-3′ and 5′-AGCTCAGCCAACTCTTTGTCT-3′ for the
EZR-ROS1 fusion transcript; 5′-ATTGGCTCCTGTTTGAAATG-3′ and
5′-TTATAAGCACTGTCACCCCTTC-3′ for the CD74-ROS1 fusion transcript;
and 5′-TCGGCAACTTTAGCGAGTAT-3′ and 5′-TTCTCTTTCAGCATCTTCACG-3′ for
the KIF5B-RET fusion transcript. The PCR products were fractionated
using electrophoresis on an agarose gel and were stained with
ethidium bromide. PCR-amplified products were purified with
ExoSAP-IT (GE Healthcare Bio-Science, Piscataway, NJ, USA) and were
sequenced directly using a BigDye® Terminator Cycle
Sequencing Reaction Kit and the ABI 3130 Genetic Analyzer (both
from Applied Biosystems, Tokyo, Japan).
Immunohistochemical staining
Sections of formalin-fixed, paraffin-embedded tissue
samples were used for immunohistochemical staining performed using
the EnVision (Dako ChemMate) kit (Dako, Kyoto, Japan). The primary
antibodies were as follows: anti-thyroid transcription factor-1
(TTF-1), anti-napsin A, anti-CK14 (all from Novocastra
Laboratories, Newcastle, UK), anti-p63 (Japan Tanner, Osaka,
Japan), anti-CD56 (Novocastra Laboratories), anti-chromogranin A
(Dako), anti-synaptophysin (Novocastra Laboratories) and anti-ROS1
(clone D4D6; Cell Signaling Technology, Beverly, MA, USA).
Hematoxylin and eosin (H&E) staining was also performed.
Mutation search
Genomic DNA was extracted from the lung tissue
samples containing ROS1 fusion transcripts using a DNeasy kit
(Qiagen) and was examined for somatic mutations in the DNA
sequences of mutation cluster regions (hot spots) in the
KRAS, EGFR, BRAF and PIK3CA genes and
in the entire coding sequence of the p53 gene. PCR
amplification was performed in 20-μl reaction mixtures containing
HotStarTaq DNA Polymerase (Qiagen). The following PCR primers were
used: 5′-AAAGGTACTGGTGGAGTATTTG-3′ and 5′-GTCCTGCACCAGTAATATGC-3′
for KRAS (exon 2); 5′-AATCCAGACTGTGTTTCTCC-3′ and
5′-ATATTATATC ATGGCATTAGC-3′ for KRAS (exon 3);
5′-GCAATATCAGCC TTAGGTGCGGTC-3′ and 5′-CATAGAAAGTGAACATTTA
GGATGTG-3′ for EGFR (exon 19); 5′-CTAACGTTCGCCAGC
CATAAGTCC-3′ and 5′-GCTGCGAGCTCACCCAGAAT GTCTGG-3′ for EGFR
(exon 21); 5′-AATTTTTCTTAAG GGGATCTCTTCC-3′ and
5′-GCGAACAGTGAATATTT CCTTTG-3′ for BRAF (exon 11);
5′-ACCTAAACTCTTCAT AATGCTTGCTC-3′ and 5′-CTTCAATGACTTTCTAGTAA
CTCAGCAG-3′ for BRAF (exon 15); 5′-TTAGATTGGTT
CTTTCCTGTCTCTG-3′ and 5′-TCCAATAGGTATGGTAA AAACATGC-3′ for
PIK3CA (exon 9); 5′-GTGACATT TGAGCAAAGACCTG-3′ and
5′-CTGTTCATGGATTGT GCAATTC-3′ for PIK3CA (exon 20);
5′-TTGGAAGTG TCTCATGCTGG-3′ and 5′-AAGAGCAGTCAGAGGACC AGG-3′ for
p53 (exons 2 and 3); 5′-ACCTGGTCCTCT GACTGCTC-3′ and
5′-TTGAAGTCTCATGGAAGCCAG-3′ for p53 (exon 4);
5′-TTGTGCCCTGACTTTCAACTC-3′ and 5′-ACCAGCCCTGTCGTCTCTC-3′ for
p53 (exon 5); 5′-TCAGATAGCGATGGTGAGCAG-3′ and 5′-GGAGGT
CAAATAAGCAGCAGG-3′ for p53 (exon 6); 5′-CTCATC
TTGGGCCTGTGTTATC-3′ and 5′-GAAGAAATCGGTA AGAGGTGGG-3′ for
p53 (exon 7); 5′-CTGCCTCTTGCTT CTCTTTTCC-3′ and
5′-ACTTTCCACTTGATAAGAGGT CCC-3′ for the p53 (exons 8 and 9);
5′-ATACTTACTTCT CCCCCTCCTCTG-3′ and 5′-GGATGAGAATGGAATCCT ATGG-3′
for p53 (exon 10); and 5′-TGATGTCATCTC TCCTCCCTG-3′ and
5′-TTGCAAGCAAGGGTTCAAAG-3′ for p53 (exon 11). Sequencing was
performed as described above.
Detection of ALK gene fusion transcripts
using RT-PCR
PCR was performed in 20-μl reaction mixtures
containing HotStarTaq DNA Polymerase (Qiagen) under the following
conditions: 30 sec at 94°C, 30 sec at 61°C and 70 sec at 72°C for
45 cycles. A total of five different PCR-primer pairs for EML4-ALK
and three PCR-primer pairs for KIF5B-ALK were used for the RT-PCR.
The forward PCR primers were: 5′-CAAGATGGACGGTTTCGCCGGCAGTCTCG-3′
for the sequence at exon 1 of EML4; 5′-ACAAATTCGAGCATC ACCTTCTCC-3′
for the sequence at exon 4 of EML4; 5′-GTG CAGTGTTTAGCATTCTTGGGG-3′
for the sequence at exon 13 of EML4; 5′-CTGTGGGATCATGATCTGAATC
CTG-3′ for the sequence at exon 14 of EML4; 5′-CTTCCT GGCTGTAGG
ATCTCATGAC-3′ for the sequence at exon 19 of EML4;
5′-CACTATTGTAATTTGCTGCTCTCCATC ATC-3′ for the sequence at exon 10
of KIF5B; 5′-AATCTG TCGATGCCCTCAGTGAAG-3′ for the sequence at exon
17 of KIF5B; and 5′-TGATCGCAAACGCTATCAGCAAG-3′ for the sequence at
exon 24 of KIF5B. The reverse PCR primer used was the same, i.e.,
5′-GAGGTCTTGCCAGCAAAG CAGTAG-3′ for the sequence at exon 20 of
ALK.
Results
In the present study, 114 NSCLCs were examined for
SLC34A2-ROS1, EZR-ROS1, CD74-ROS1 and KIF5B-RET fusion transcripts
using RT-PCR and subsequent sequencing analyses. Although the
expression of SLC34A2-ROS1, EZR-ROS1 and KIF5B-RET fusion
transcripts was not detected in any of the carcinomas, CD74-ROS1
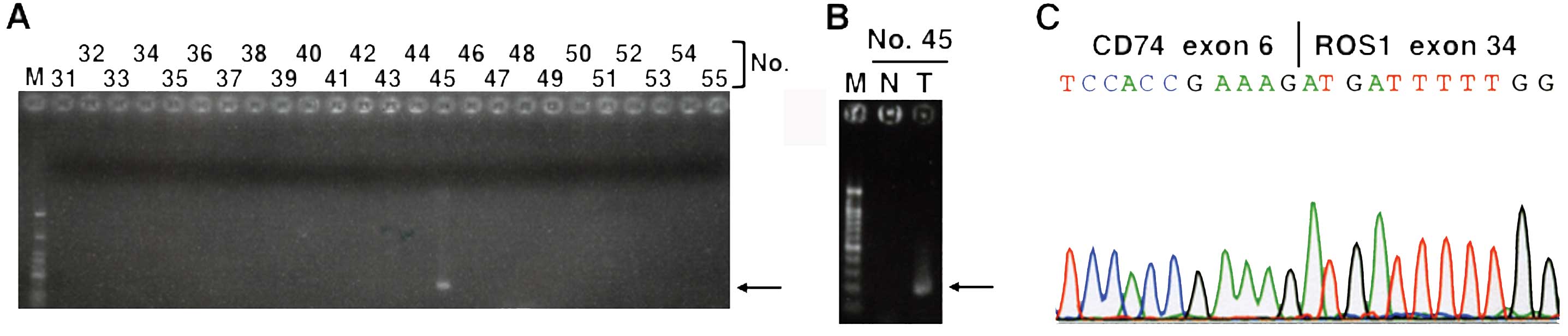
fusion transcripts were detected in one (0.9%) NSCLC (Fig. 1). The CD74-ROS1 fusion transcripts
were expressed in the cancerous tissues but not in the
non-cancerous tissues in the NSCLC case (case no. 45) (Fig. 1B). Sequencing of the RT-PCR product
revealed that the fusion was between exon 6 of CD74 and exon 34 of
ROS1 (Fig. 1C). The fusion was
in-frame and allowed the transmembrane domain and the kinase domain
of ROS1 to be retained. Case no. 45 was a 71-year-old woman who was
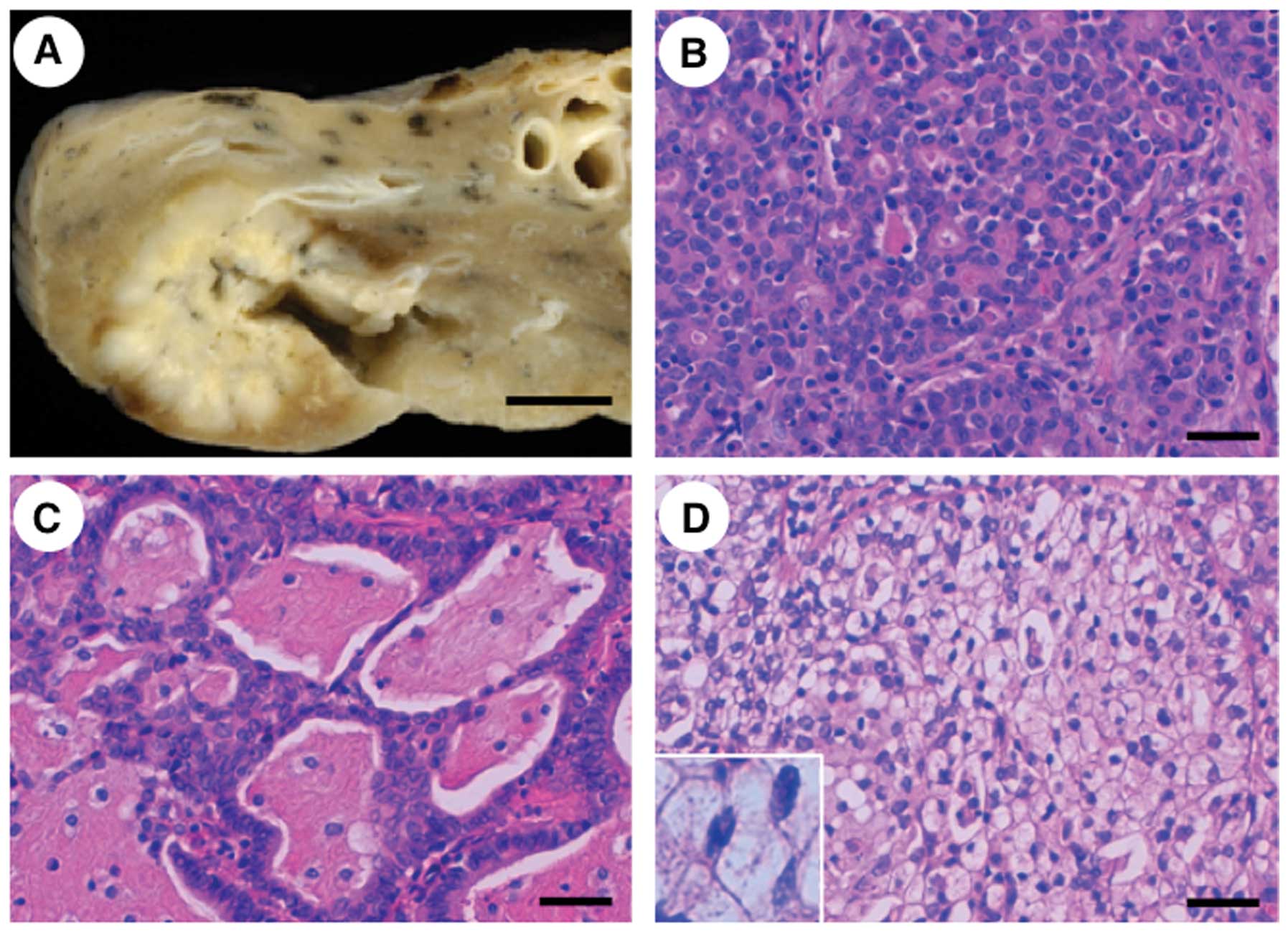
a never-smoker. The NSCLC in this case was well-delineated and 3.0
cm in diameter (Fig. 2A). The
histological classification was adenocarcinoma with a predominantly
acinar pattern (Fig. 2B); a
cribriform structure associated with abundant extracellular mucus
(mucinous cribriform pattern) (Fig.
2C) and a solid growth pattern containing signet-ring cells
(solid signet-ring cell pattern) (Fig.
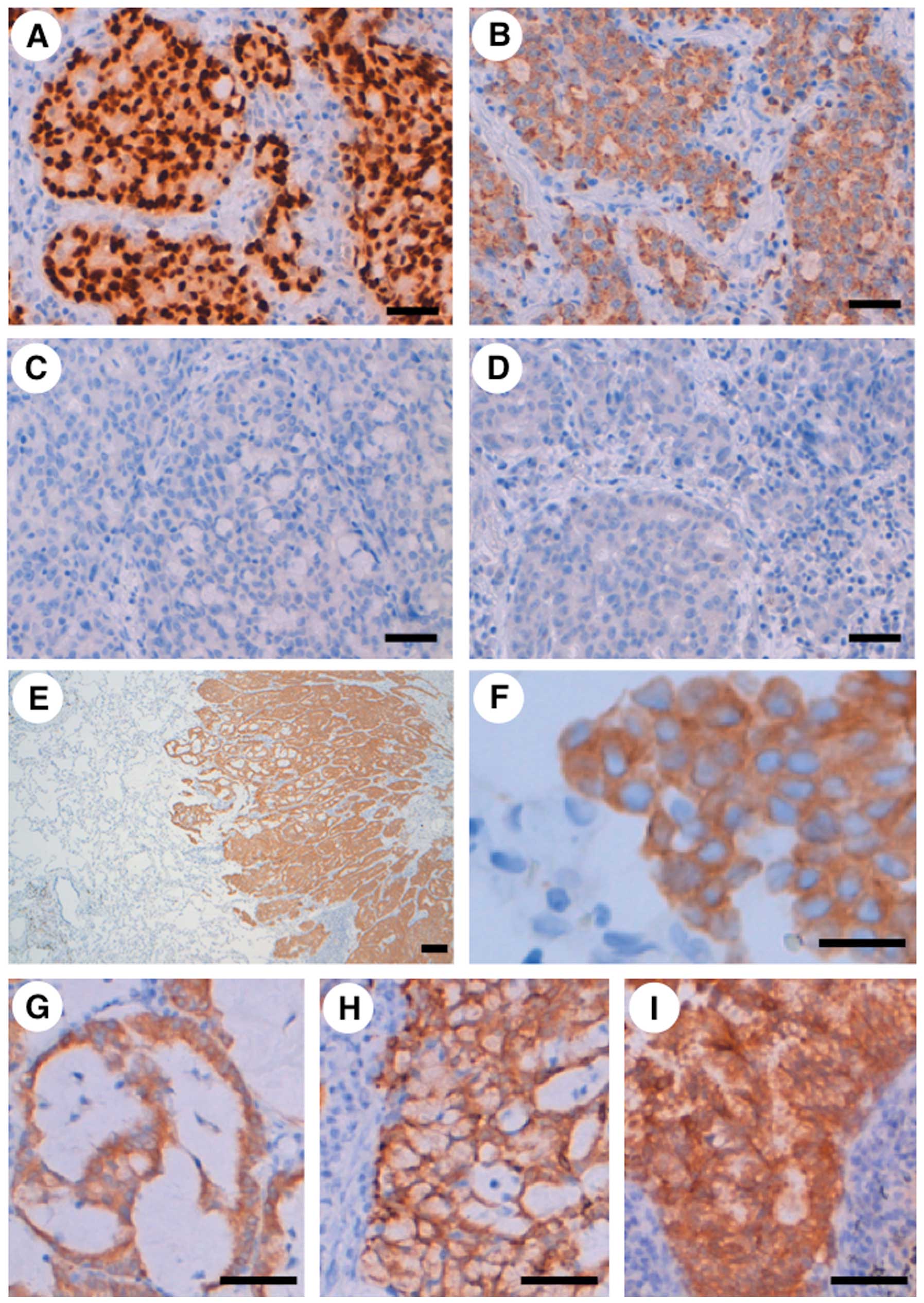
2D) were partly observed. An immunohistochemical study revealed
that the adenocarcinoma was positive for TTF-1 and napsin A, but
negative for CK14, p63, CD56, chromogranin A and synaptophysin
(Fig. 3A–D), indicating that
immunophenotype of the carcinoma was compatible with adenocarcinoma
without neuroendocrine features. Furthermore, the cancer-specific
overexpression of the ROS1 protein was detected using an
immunohistochemical analysis (Fig.
3E–I), in agreement with the cancer-specific expression of the
ROS1 fusion mRNA transcript (Fig.
1B). An increased ROS1 protein signal was detected diffusely in
the adenocarcinoma, including the mucinous cribriform pattern, and
solid signet-ring cell pattern and the ROS1 protein was localized
in the cytoplasm of the carcinoma cells (Fig. 3E–H). ROS1 protein was also highly
expressed in the adenocarcinoma metastases to the lymph node
(Fig. 3I). Additionally, after the
surgical treatment, a brain metastasis of the adenocarcinoma was
observed in the patient. The above findings suggest that CD74-ROS1
fusion transcripts are expressed in a subset of NSCLCs.
We next examined whether the NSCLC case containing
the CD74-ROS1 fusion transcripts also contained previously reported
driver mutations. Mutation cluster regions for KRAS,
EGFR, BRAF and PIK3CA(18–20)
were searched for somatic mutations and the expression of EML4-ALK
and KIF5B-ALK fusion transcripts (1–5) was
examined; however, no somatic mutations in exons 2 and 3 of
KRAS, exons 19 and 21 of EGFR, exons 11 and 15 of
BRAF, or exons 9 and 20 of PIK3CA and no expression
of EML4-ALK or KIF5B-ALK fusion transcripts were detected. We also
performed a mutational analysis of the entire coding region of
p53 of the carcinoma that contained CD74-ROS1 fusion
transcripts. The case was found to be heterozygous for the Arg and
Pro alleles of the p53 p.Arg72Pro genetic polymorphism,
which is associated with a functional difference (21); however, no somatic p53
mutations were detected.
Discussion
In the present study, the expression of CD74-ROS1
fusion transcripts was found in one (0.9%) of the 114 NSCLCs that
were examined, but the expression of SLC34A2-ROS1, EZR-ROS1, or
KIF5B-RET fusion transcripts was not detected in any of the NSCLCs.
The detected fusion occurred between exon 6 of CD74 and exon 34 of
ROS1 and was observed in a non-smoker female. Histologically, the
carcinoma was an adenocarcinoma with a predominant acinar pattern;
a mucinous cribriform pattern and a solid signet-ring cell pattern
were also observed in a portion of the adenocarcinoma. ROS1 protein
was overexpressed in a cancer-specific manner at both the primary
site and the lymph node metastases. No somatic mutations were
detected in the mutation cluster regions of the KRAS,
EGFR, BRAF and PIK3CA genes and the entire
coding region of p53 in the carcinoma, and the expression of
EML4- and KIF5B-ALK fusions was also not detected. These results
suggest that CD74-ROS1 fusion contributes to the carcinogenesis of
this NSCLC case as a driver mutation.
To date, ROS1 fusion transcripts have been detected
in 0.7–1.9% of NSCLC patients (Chinese, Japanese, white and
Caucasian populations) (5,9,10,12,14,15,17).
The analytical methods used in the above-mentioned reports varied
and RT-PCR, fluorescence in situ hybridization and
immunohistochemical analyses have been used to search for NSCLCs
containing ROS1 fusion products (5,9,10,12,14,15,17).
Considering our results (0.9% of Japanese NSCLC patients) and the
results of the above-mentioned previous papers, racial differences
in the frequency of ROS1 fusion-positive NSCLC are thought to be
minimal, although distinct analytical methods were used in the
various reports. Approximately 1.6 million new lung cancers are
diagnosed each year worldwide (22); if NSCLC comprises 90% of these
cancers, 1.45 million new cases of NSCLC are diagnosed every year.
Since the prevalence of ROS1 fusion is ~1% of all NSCLCs, ~14,500
new patients have ROS1 fusion-positive NSCLC. Recently, ROS1
inhibition was shown to lead to the suppression of the
proliferation of cells containing ROS1 fusion in
vitro(12,14); furthermore, crizotinib, a small
molecule inhibitor against ROS1 kinase in addition to ALK kinase,
was shown to exhibit antitumor properties in a patient with NSCLC
containing an ROS1 fusion (12,14).
Thus, patients with ROS1 fusion-positive NSCLC may comprise a novel
subclass that could benefit clinically from ROS1 inhibition.
In our CD74-ROS1 fusion-positive case, no other
oncogenic driver mutations were detected. Oncogenic driver
mutations are responsible for the initiation and progression of
NSCLCs, and they differ from passenger mutations in that they are
also found within the cancer genomes but exist as a by-product of
cancer cell development (10,13,18,19).
In general, oncogenic driver mutations are mutually exclusive
(10). ROS1 fusion-positive cases
reported in two previous studies (4,17) were
negative for alterations in the ALK, RET, EGFR
and KRAS genes. Thus, our results for the driver mutation
search are compatible with the results of these previous reports,
suggesting that ROS1 fusion is mutually exclusive with other
oncogenic driver mutations in NSCLC.
A mucinous cribriform pattern and a solid
signet-ring cell pattern were observed in a portion of the
adenocarcinoma in the present case. Yoshida et al(17) recently reported that one-third of
ROS1 fusion-positive NSCLCs contain a mucinous cribriform pattern
and one-third contain a solid signet-ring cell pattern. Of note,
both patterns are frequently observed in ALK-rearranged NSCLC
(4,23,24).
Thus, both a mucinous cribriform pattern and a solid signet-ring
cell pattern may be common pathohistological characteristics of
ROS1 and ALK fusion-positive NSCLCs. We also showed that the ROS1
protein was highly expressed in both histological patterns in our
case, and an increased ROS1 protein expression in both components
has not previously been demonstrated, suggesting a novel finding of
the present study. These results indicate that ROS1 is involved in
the morphogenesis of the patterns.
EGFR mutations and EML4-ALK fusions are
preferentially associated with NSCLC in non-smokers (4,18,19).
Our CD74-ROS1 fusion-positive case was a never-smoker. In three
previous reports, ROS1 fusion was frequently found in the NSCLCs of
never-smokers (4,12,17);
however, this characteristic was not detected in another report
(14). The accumulation of ROS1
fusion-positive cases with information on the smoking history is
required to better understand the role of ROS1 fusion in
NSCLCs.
In conclusion, our CD74-ROS1 fusion-positive NSCLC
in conjunction with previously detected ROS1 fusion-positive NSCLCs
suggested that ROS1 fusion is involved in the carcinogenesis of a
subset of NSCLCs and may significantly aid in elucidating the
characteristics of ROS1 fusion-positive NSCLC in the future.
Acknowledgements
The authors acknowledge Ms. S. Izumo (Hamamatsu
University School of Medicine) for her technical assistance. The
present study was supported in part by a Grant-in-Aid from the
Ministry of Health, Labour and Welfare (21-1), a Grant-in-Aid from
the Japan Society for the Promotion of Science (22590356), a
Grant-in-Aid from the Ministry of Education, Culture, Sports,
Science and Technology (221S0001) and the Smoking Research
Foundation.
References
|
1
|
Soda M, Choi YL, Enomoto M, Takada S,
Yamashita Y, Ishikawa S, Fujiwara S, Watanabe H, Kurashina K,
Hatanaka H, Bando M, Ohno S, Ishikawa Y, Aburatani H, Niki T,
Sohara Y, Sugiyama Y and Mano H: Identification of the transforming
EML4-ALK fusion gene in non-small-cell lung cancer. Nature.
448:561–566. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Shinmura K, Kageyama S, Tao H, Bunai T,
Suzuki M, Kamo T, Takamochi K, Suzuki K, Tanahashi M, Niwa H, Ogawa
H and Sugimura H: EML4-ALK fusion transcripts, but no NPM-, TPM3-,
CLTC-, ATIC-, or TFG-ALK fusion transcripts, in non-small cell lung
carcinomas. Lung Cancer. 61:163–169. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Shinmura K, Kageyama S, Igarashi H, Kamo
T, Mochizuki T, Suzuki K, Tanahashi M, Niwa H, Ogawa H and Sugimura
H: EML4-ALK fusion transcripts in immunohistochemically
ALK-positive non-small cell lung carcinomas. Exp Ther Med.
1:271–275. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Takeuchi K, Soda M, Togashi Y, Suzuki R,
Sakata S, Hatano S, Asaka R, Hamanaka W, Ninomiya H, Uehara H, Lim
Choi Y, Satoh Y, Okumura S, Nakagawa K, Mano H and Ishikawa Y: RET,
ROS1 and ALK fusions in lung cancer. Nat Med. 18:378–381. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Lipson D, Capelletti M, Yelensky R, Otto
G, Parker A, Jarosz M, Curran JA, Balasubramanian S, Bloom T,
Brennan KW, Donahue A, Downing SR, Frampton GM, Garcia L, Juhn F,
Mitchell KC, White E, White J, Zwirko Z, Peretz T, Nechushtan H,
Soussan-Gutman L, Kim J, Sasaki H, Kim HR, Park SI, Ercan D,
Sheehan CE, Ross JS, Cronin MT, Jänne PA and Stephens PJ:
Identification of new ALK and RET gene fusions from colorectal and
lung cancer biopsies. Nat Med. 18:382–384. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Casaluce F, Sgambato A, Maione P, Rossi A,
Ferrara C, Napolitano A, Palazzolo G, Ciardiello F and Gridelli C:
ALK inhibitors: a new targeted therapy in the treatment of advanced
NSCLC. Target Oncol. 8:55–67. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Shaw AT and Solomon B: Targeting
anaplastic lymphoma kinase in lung cancer. Clin Cancer Res.
17:2081–2086. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
O’Bryant CL, Wenger SD, Kim M and Thompson
LA: Crizotinib: a new treatment option for ALK-positive non-small
cell lung cancer. Ann Pharmacother. 47:189–197. 2013.PubMed/NCBI
|
|
9
|
Rikova K, Guo A, Zeng Q, Possemato A, Yu
J, Haack H, Nardone J, Lee K, Reeves C, Li Y, Hu Y, Tan Z, Stokes
M, Sullivan L, Mitchell J, Wetzel R, Macneill J, Ren JM, Yuan J,
Bakalarski CE, Villen J, Kornhauser JM, Smith B, Li D, Zhou X, Gygi
SP, Gu TL, Polakiewicz RD, Rush J and Comb MJ: Global survey of
phosphotyrosine signaling identifies oncogenic kinases in lung
cancer. Cell. 131:1190–1203. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Li C, Fang R, Sun Y, Han X, Li F, Gao B,
Iafrate AJ, Liu XY, Pao W, Chen H and Ji H: Spectrum of oncogenic
driver mutations in lung adenocarcinomas from East Asian never
smokers. PLoS One. 6:e282042011. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Kohno T, Ichikawa H, Totoki Y, Yasuda K,
Hiramoto M, Nammo T, Sakamoto H, Tsuta K, Furuta K, Shimada Y,
Iwakawa R, Ogiwara H, Oike T, Enari M, Schetter AJ, Okayama H,
Haugen A, Skaug V, Chiku S, Yamanaka I, Arai Y, Watanabe S, Sekine
I, Ogawa S, Harris CC, Tsuda H, Yoshida T, Yokota J and Shibata T:
KIF5B-RET fusions in lung adenocarcinoma. Nat Med. 18:375–377.
2012. View
Article : Google Scholar : PubMed/NCBI
|
|
12
|
Bergethon K, Shaw AT, Ou SH, Katayama R,
Lovly CM, McDonald NT, Massion PP, Siwak-Tapp C, Gonzalez A, Fang
R, Mark EJ, Batten JM, Chen H, Wilner KD, Kwak EL, Clark JW,
Carbone DP, Ji H, Engelman JA, Mino-Kenudson M, Pao W and Iafrate
AJ: ROS1 rearrangements define a unique molecular class of lung
cancers. J Clin Oncol. 30:863–870. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Seo JS, Ju YS, Lee WC, Shin JY, Lee JK,
Bleazard T, Lee J, Jung YJ, Kim JO, Shin JY, Yu SB, Kim J, Lee ER,
Kang CH, Park IK, Rhee H, Lee SH, Kim JI, Kang JH and Kim YT: The
transcriptional landscape and mutational profile of lung
adenocarcinoma. Genome Res. 22:2109–2119. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Davies KD, Le AT, Theodoro MF, Skokan MC,
Aisner DL, Berge EM, Terracciano LM, Cappuzzo F, Incarbone M,
Roncalli M, Alloisio M, Santoro A, Camidge DR, Varella-Garcia M and
Doebele RC: Identifying and targeting ROS1 gene fusions in
non-small cell lung cancer. Clin Cancer Res. 18:4570–4579. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Rimkunas VM, Crosby KE, Li D, Hu Y, Kelly
ME, Gu TL, Mack JS, Silver MR, Zhou X and Haack H: Analysis of
receptor tyrosine kinase ROS1-positive tumors in non-small cell
lung cancer: identification of a FIG-ROS1 fusion. Clin Cancer Res.
18:4449–4457. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Yokota K, Sasaki H, Okuda K, Shimizu S,
Shitara M, Hikosaka Y, Moriyama S, Yano M and Fujii Y:
KIF5B/RET fusion gene in surgically-treated adenocarcinoma
of the lung. Oncol Rep. 28:1187–1192. 2012.
|
|
17
|
Yoshida A, Kohno T, Tsuta K, Wakai S, Arai
Y, Shimada Y, Asamura H, Furuta K, Shibata T and Tsuda H:
ROS1-rearranged lung cancer: a clinicopathologic and molecular
study of 15 surgical sases. Am J Surg Pathol. 37:554–562. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Schmid K, Oehl N, Wrba F, Pirker R, Pirker
C and Filipits M: EGFR/KRAS/BRAF mutations in primary lung
adenocarcinomas and corresponding locoregional lymph node
metastases. Clin Cancer Res. 15:4554–4560. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Pao W and Girard N: New driver mutations
in non-small-cell lung cancer. Lancet Oncol. 12:175–180. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Ulivi P, Romagnoli M, Chiadini E, Casoni
GL, Capelli L, Gurioli C, Zoli W, Saragoni L, Dubini A, Tesei A,
Amadori D and Poletti V: Assessment of EGFR and
K-ras mutations in fixed and fresh specimens from
transesophageal ultrasound-guided fine needle aspiration in
non-small cell lung cancer patients. Int J Oncol. 41:147–152.
2012.
|
|
21
|
Thomas M, Kalita A, Labrecque S, Pim D,
Banks L and Matlashewski G: Two polymorphic variants of wild-type
p53 differ biochemically and biologically. Mol Cell Biol.
19:1092–1100. 1999.PubMed/NCBI
|
|
22
|
Jemal A, Bray F, Center MM, Ferlay J, Ward
E and Forman D: Global cancer statistics. CA Cancer J Clin.
61:69–90. 2011. View Article : Google Scholar
|
|
23
|
Rodig SJ, Mino-Kenudson M, Dacic S, Yeap
BY, Shaw A, Barletta JA, Stubbs H, Law K, Lindeman N, Mark E, Janne
PA, Lynch T, Johnson BE, Iafrate AJ and Chirieac LR: Unique
clinicopathologic features characterize ALK-rearranged lung
adenocarcinoma in the western population. Clin Cancer Res.
15:5216–5223. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Jokoji R, Yamasaki T, Minami S, Komuta K,
Sakamaki Y, Takeuchi K and Tsujimoto M: Combination of
morphological feature analysis and immunohistochemistry is useful
for screening of EML4-ALK-positive lung adenocarcinoma. J Clin
Pathol. 63:1066–1070. 2010. View Article : Google Scholar : PubMed/NCBI
|