Introduction
Prostate cancer (PCa) is one of the most common
malignant tumors in males and the fifth leading cause of
cancer-related mortality worldwide, and the second leading cause in
the male population (1,2). Because the onset of PCa is insidious,
early diagnosis is difficult, and many patients have metastatic
disease at the time of diagnosis (3,4) and
death from PCa occurs largely in patients with aggressive
androgen-insensitive metastatic disease (5). Upon sustained hormone-ablation
therapy, invasive PCa frequently acquires castration-resistant
features and gains metastatic potential, which accounts for high
patient morbidity (6). Even though
initially certain PCa cases are androgen-dependent (ADPC),
eventually, if sufficient time is allowed to elapse, virtually all
prostate cancers become androgen-independent (AIPC). The mechanisms
leading to androgen independence have been extensively studied, yet
the mechanisms are still not fully understood (7,8).
Prostate-specific membrane antigen (PSMA), a
transmembrane-carboxypeptidase (9,10), as
a prostate cancer-specific biomarker and potential target for
diagnosis and treatment, has been correlated with aggressive
disease and has obtained wide attention (1,11,12).
It has been confirmed that PSMA is PCa-specific and is highly
expressed in PCa. Its expression is further increased in poorly
differentiated, advanced, hormone-refractory metastatic PCa,
therefore, it is an ideal marker for prostate cancer diagnosis and
a target for treatment (13,14).
The high expression of PSMA in PCa and its correlation with
pathological classification and prognosis indicate that PSMA may
play a regulatory role in PCa development and metastasis.
Although a high level of PSMA is usually associated
with PCa progression, the role of PSMA in tumor migration and
invasiveness is still uncertain, and contradictory results exist
(15–17). A recent study by Ghosh et
al(17) revealed that PSMA
suppresses PC-3 cell invasiveness. Previous studies by our group
found that PSMA and its spliced variant are highly expressed in PCa
tissues (11), suppress prostate
cancer cell proliferation, migration and invasiveness, and blockage
of PSMA in LNCap cells promotes the invasion and migration of
prostate cancer cells (12). Yet,
the mechanism of PSMA involved in the process of metastasis of PCa
remains unclear. In the present study, in order to screen and
identify metastasis-related genes that are possibly involved in PCa
metastasis regulated by PSMA, we used a tumor metastasis PCR array
to analyze the expression of 84 tumor metastasis-related genes in
LNCap cells (a known PSMA-positive prostate cancer cell line),
si-PSMA LNCap cells and PC-3 cells (known prostate cancer cell line
that does not express PSMA), respectively, and we compared the
common differentially expressed genes to screen the possible
PSMA-related genes. Three of these candidate genes were further
confirmed in prostate cancer cell lines and also in clinical
prostate samples. Our findings may provide clues for exploring the
role of PSMA in PCa metastasis and its molecular mechanism.
Materials and methods
Cell lines, tissue samples and
reagents
Prostate cancer cell lines LNCap, 22RV1, PC-3 and
DU145 were obtained from the ATCC (American Type Culture
Collection, Manassas, VA, USA) and cultured in RPMI-1640 medium
(Gibco-BRL, Invitrogen Corp., Grand Island, NY, USA) with 1 mM
sodium pyruvate, 100 U/ml penicillin G, 100 μg/ml streptomycin
(Sigma Corp. of America, Ronkonkoma, NY, USA), 10% fetal bovine
serum (FBS; Tianjin Hao Yang Biological Manufacture Co., Ltd.,
Beijing, China). Lipofectamine™ 2000 transfection reagent, TRIzol
and the One-Step Fluorescence assay kit were products of Invitrogen
(Carlsbad, CA, USA).
Prostate tissue samples including normal prostate
(n=12), primary prostate cancer (n=47), and benign prostatic
hypertrophy (BPH) (n=26), were obtained at the Department of
Urology, the First Affiliated Hospital of Sun Yat-sen University,
Guangzhou, China. All tissue samples were stored at −80°C until
being further processed. The Gleason score was used to evaluate
tumor malignancy, and the prostate cancer samples were then divided
into two categories according to their Gleason score (Table I). Informed content for the
scientific use of biologic materials was obtained from all patients
in accordance with the requirements of the Medical Ethics Review
Board of Sun Yat-sen University.
 | Table IPathological information regarding the
prostate samples used in real-time PCR identification of the MMP3,
MTSS1 and CDH6 genes. |
Table I
Pathological information regarding the
prostate samples used in real-time PCR identification of the MMP3,
MTSS1 and CDH6 genes.
| Tissue type | Gleason score | No. of tissue samples
analyzed |
|---|
| Normal prostate | | 12 |
| Benign prostatic
hypertrophy (BPH) | | 26 |
| Primary prostate
cancer | 4, 5, 6 | 17 |
| Primary prostate
cancer | 7, 8, 9 | 30 |
Reverse transcription and real-time
PCR
One step TaqMan real-time RT-PCR was used to
evaluate the PSMA silencing effect. Specific probes and primers of
PSMA were designed by Beacon Designer 7.01 Demo, and the β-actin
gene served as the internal control. Primer and probe sequences are
shown in Table II. One step
real-time PCR amplification reaction of PSMA contained 1 μl
SuperScript III RT/Platinum Taq Mix, 1 μl RNaseOut, 25 μl 2X
reaction mixture (Invitrogen), 0.5 μM of the forward and reverse
primers and 0.5 μM of the probe, and 3 μl of RNA and nuclease-free
water to a final volume of 50 μl. Real-time RT-PCR was performed on
an ABI 7500 real-time PCR system (Applied Biosystems, Foster City,
CA, USA) using thermal cycling conditions as follows: 50°C for 15
min, 95°C for 2 min; 95°C for 15 sec, 60°C for 30 sec, 40 cycles.
All PCR reactions were performed in duplicate.
 | Table IIReal-time PCR primer and probe
sequences. |
Table II
Real-time PCR primer and probe
sequences.
| Primer/Taq | Primer and TaqMan
probe sequences |
|---|
| TaqMan real-time
PCR | PSMA-F |
5′-GAAGTCTCAAAGTGCCCTACAATG-3′ |
| PSMA-R |
5′-ACCCATGAGTCCCGGTGAC-3′ |
| PSMA-Taq | 5′-(FAM)
CTGTCTGGTTCCACTGCTCCTCTGA (Eclipse)-3′ |
| Actin-F |
5′-GACTACCTCATGAAGATCCTCACC-3′ |
| Actin-R |
5′-TCTCCTTAATGTCACGCACGATT-3′ |
| Actin-Taq | 5′-(FAM)
CGGCTACAGCTTCACCACCACGGC (Eclipse)-3′ |
| SYBR-Green-based
real-time PCR | PSMA-F |
5′-ACACCGCTGATGTACAGCTTGGT-3′ |
| PSMA-R |
5′-TGCTTATCCTGGGCATGCCACTG-3′ |
| MMP3-F |
5′-GGCAGGCAAGACAGCAAGGCAT-3′ |
| MMP3-R |
5′-GCTGAGCAAACTGCCACGCAC-3′ |
| MTSS1-F |
5′-GCGAGAACCGGACCCCAACG-3′ |
| MTSS1-R |
5′-AGGCCTGGTGGCAGCCGATA-3′ |
| CDH6-F |
5′-TTTTGGGTGGGCCAGCCCTACC-3′ |
| CDH6-R |
5′-CAGAGAGCTCCAGGGCCCTTTTCT-3′ |
| Actin-F |
5′-CAGAGCCTCGCCTTTGCCGATCC-3′ |
| Actin-R |
5′-CCTTGCACATGCCGGAGCCGT-3′ |
Reverse transcription (RT) and real-time PCR based
on SYBR-Green detection was used to identify candidate PSMA-related
genes in PCa cell lines and tissues. Total RNA was extracted by
TRIzol reagent following the manufacturer’s instructions. Briefly,
PCa cell total RNA was extracted directly in a 3.5 cm diameter
culture dish by adding 1 ml TRIzol reagent, and the total RNA of
the PCa clinical samples that were cut into ~0.5 cm pieces, adding
the appropriate volume of pre-cooling TRIzol regeant at v/v 10:1.
RT reaction was carried out in a final volume of 20 μl using the
Invitrogen reverse transcription kit (Invitrogen) according to the
manufacturer’s instructions. First, the reaction mixture containing
500 ng/μl Oligo (dT)18 (1 μl), total RNA (1.5 μg), 10 mM
each deoxynucleoside triphosphate (dNTP) (1 μl) and nuclease-free
water up to a volume of 13 μl was incubated for 5 min at 65°C,
placed on ice for more than 1 min, then 5X first strand buffer (4
μl), 0.1 M DTT (1 μl), RNase inhibitor (1 μl), 200 U/μl SuperScript
III reverse transcriptase (1 μl) were added to a final reaction
volume of 20 μl. The thermal cycling parameters for the RT reaction
were as follows: 60 min at 50°C and 15 min at 70°C. Products of RT
reactions were stored at −20°C until further analysis. The mRNA
expression of candidate genes was determined using the ABI PRISM
7500 detection system (Applied Biosystems) and SYBR Premix Taq™
(Invitrogen). The primer sequences are shown in Table II. The reaction system contained 2X
SYBR Premix Ex Taq™ II (10 μl), 0.4 μM of each forward and reverse
primers, 50X ROX reference dye 0.4 μl and 2 μl of the synthesized
cDNA in a 20 μl reaction volume. Thermal cycling conditions were as
follows: 95°C for 30 sec, followed by 40 cycles of 95°C for 5 sec,
60°C for 30 sec. All PCR reactions were performed in duplicate and
repeated three times. Melting curve analysis was used to control
the amplification specificity. Target gene expression is shown as
the fold increase or decrease relative to the expression of
β-actin. Fold-changes in target gene mRNA expression were
determined as 2−ΔΔCt using the same calculation formula
as determined in the microarray analysis.
PSMA gene silencing by siRNA in LNCap
cells
Three pairs of siRNAs for PSMA (GenBank ID: M99487)
were designed by online software (http://jura.wi.mit.edu/bioc/siRNAext/siRNA_search.cgi?tasto=10925811)
(Table III). Designed siRNAs and
the positive and negative control siRNAs were synthesized and
supplied by Guangzhou RiboBio Co., Ltd. (Guangzhou, China). The day
before transfection, LNCap cells were collected, digested with
0.25% trypsin and counted, and then seeded onto 6-well plates
(Corning Incorporated Costar, Corning, NY, USA) at the initial
density of 2×105 cells/well and cultured with
antibiotic-free medium. The transfection was performed by
Lipofectamine™ 2000 with the concentration of siRNA at 50 nM, and
LNCap cells transfected with the negative control siRNA
(supplemented by RiboBio) served as the control. Total protein and
total RNA were extracted 72 h after transfection. One step
real-time RT-PCR was used to evaluate the PSMA silencing effect at
the mRNA level. Silencing effect of PSMA at the protein level was
identified by western blotting. The monoclonal antibody (YPSMA,
ab19071; Abcam Inc., Cambridge, MA, USA) used in the western
blotting recognized 716–723 amino acid residues of the C terminus
of PSMA. The antibody dilution ratio used in the western blotting
was 1:1,000.
 | Table IIIsiRNA sequences targeting the PSMA
gene. |
Table III
siRNA sequences targeting the PSMA
gene.
| siRNA duplex | Nucleotide
sequences | Position in the
PSMA gene coding sequencea |
|---|
| siRNA-1 | Sense
5′-CUGAGAACAUCAAGAAGUUdTdT-3′ | |
| Antisense
3′-dTdTGACUCUUGUAGUUCUUCAA-5′ | 461–479 |
| siRNA-2 | Sense
5′-GGCAAAUCUCUUUAUGAAAdTdT-3′ | |
| Antisense
3′-dTdTCCGUUUAGAGAAAUACUUU-5′ | 1729–1747 |
| siRNA-3 | Sense
5′-GCGAUCUAGUGUAUGUUAAdTdT-3′ | |
| Antisense
3′-dTdTCGCUAGAUCACAUACAAUU-5′ | 776–794 |
Microarray analysis
Human tumor metastasis PCR array (SABiosciences,
Qiagen, Gaithersburg, MD, USA) was designed to analyze 96 genes by
real-time PCR, including 84 genes known to be involved in tumor
metastasis, 4 housekeeping genes (HK genes) and 8 quality control
genes. Tumor metastasis-related genes selected for this array
encode several classes of protein factors including cell adhesion,
ECM components, cell cycle, cell growth and proliferation,
apoptosis, transcription factors and regulators and other genes
related to tumor metastasis.
Two groups were designed for the tumor metastasis
PCR array analysis: the PSMA(−) group was used to analyze the
differentially expressed genes between the LNCap and PC-3 cells,
and the si-PSMA group was used to analyze the differentially
expressed genes between the si-PSMA LNCap and LNCap cells. The mean
value of the replicates for each group was calculated and expressed
as the cycle threshold (Ct). The amount of gene expression was then
calculated as the difference (ΔCt) between the Ct value obtained
for the target gene and the Ct value obtained for the HK genes. The
value of ΔCt was calculated for each pathway-focused gene in each
group using the following calculation formula: ΔCt (control group)
= average Ct of the target gene − average Ct of the HK genes for
the control group array; ΔCt (experimental group) = average Ct of
the target gene − average Ct of the HK genes for the experimental
group array. The ΔΔCt was calculated for each gene across two PCR
arrays (or groups), ΔΔCt = ΔCt (experimental group) − ΔCt (control
group). The fold-change for each gene from the control group to the
experimental group was calculated as 2−ΔΔCt.
Candidate gene identification in the PCa
cell lines and tissues
Based on the comparison of the common genes between
the PSMA(−) group and the si-PSMA group, and the previous studies
and reports, the MMP3, MTSS1 and CDH6 genes were chosen and further
analyzed by reverse transcription (RT) and SYBR-Green-based
real-time PCR in PCa cell lines including the PSMA-positive cell
lines (LNCap and 22RV1) and the PSMA-negative cell lines (PC-3 and
DU145), and also 85 clinical samples of prostate tissues, including
12 normal prostate, 26 BPH and 47 prostate cancer samples. The
pathological information regarding the prostate tissues is shown in
Table I.
Statistical analysis
The PSMA mRNA expression levels were not distributed
normally, therefore the log-transformed relative expression levels
were used to obtain sufficiently normally distributed values. The
differences in expression of MMP3, MTSS1 and CDH6 genes among
normal prostate, BPH and PCa were evaluated using one-way ANOVA.
Spearman correlation coefficient was used to evaluate the
correlation between Gleason score and the levels of MMP3, MTSS1,
CDH6 and PSMA mRNA expression, and also between PSMA and MMP3,
MTSS1 and CDH6. P-values <0.05 were considered to indicate a
statistically significant result. All statistical analyses were
performed using SPSS11.5 software (SPSS Inc., Chicago, IL,
USA).
Results
Construction of si-PSMA LNCap cells
Stable PSMA silenced LNCap cell lines (si-PSMA LNCap
cells) were successfully obtained by transfection with the siRNA
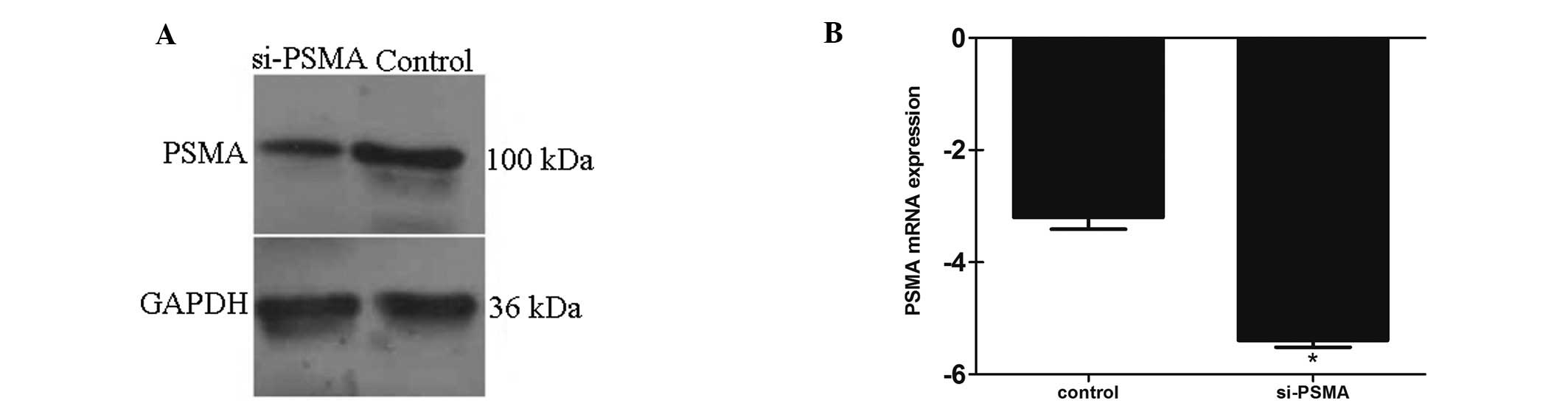
sequences. Seventy-two hours after transfection with the siRNAs,
the PSMA gene expression level was detected by western blotting and
real-time PCR. Real-time PCR showed that the PSMA mRNA expression
level was downregulated >4-fold by siRNA-3, and the western blot
analysis showed that the PSMA protein expression level was reduced
68% by siRNA-3 (Chi-square test, P<0.01 compared to the negative
control; Fig. 1). siRNA-1 and
siRNA-2 exhibited a weaker silencing effect than siRNA-3,
therefore, siRNA-3 was chosen for further study.
Screening of differentially expressed
tumor metastasis-related genes between LNCap and PC-3 cells by
tumor metastasis PCR array
In order to gain better understanding of the
molecular mechanism involved in the metastatic progression of
androgen-independent prostate cancer cells, we performed a
comprehensive metastasis-related gene expression profile analysis
of PSMA-positive LNCap and PSMA-negative PC-3 cells, namely the
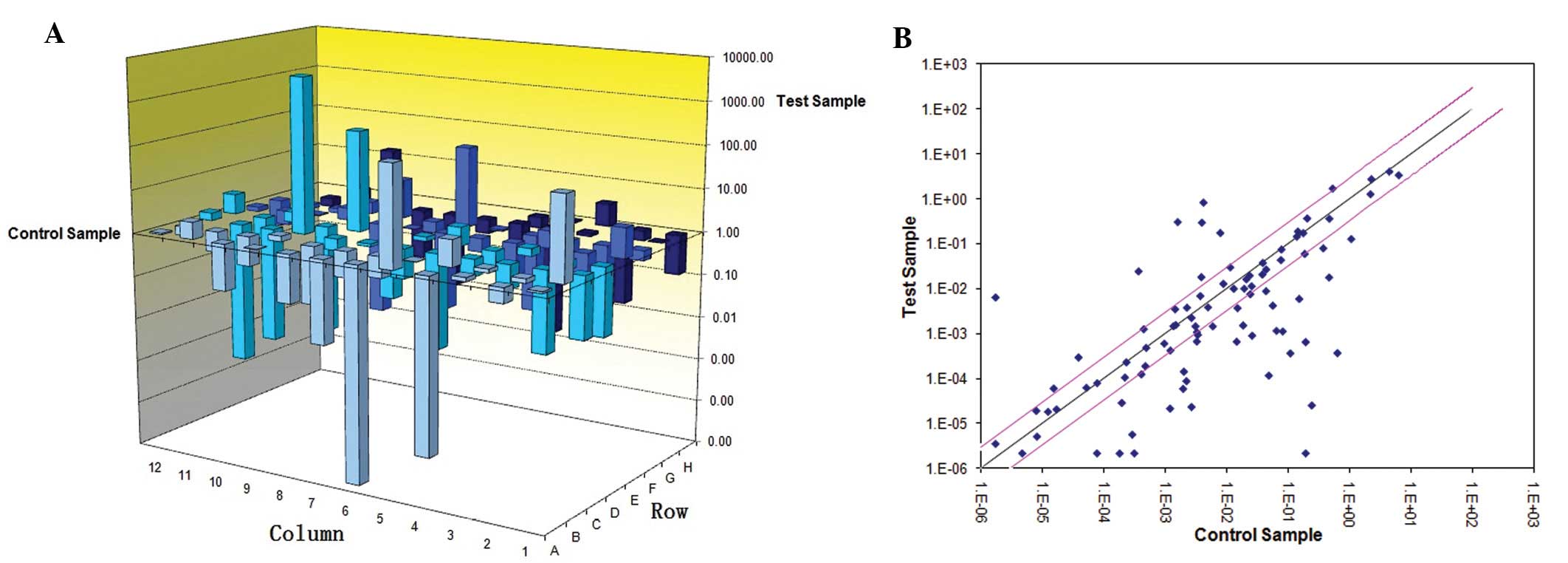
PSMA(−) group, using a tumor metastasis PCR array. Genes whose
expression was upregulated or downregulated >2-fold, were
considered significantly differentially expressed and were selected
for further analysis. Forty-one genes had significantly higher
expression in the PC-3 cells (P<0.05; Fig. 2), among which 23 genes were
expressed >10-fold (Table IV).
The genes with the highest upregulated expression in the PC-3 cells
were CDH11 (87,443.2-fold). There were 15 genes with downregulated
expression in the PC-3 cells when compared to the LNCap cells;
among which the expression levels of 6 genes (TP53, PTEN, CTNNA1,
CDH1, MDM2 and IGF1) were >10-fold (Table IV). Most of the significant genes
identified in the present study have not been previously reported
in prostate cancer.
 | Table IVTumor metastasis-related genes
upregulated or downregulated >10-fold in PC-3 cells compared
with LNCap cells. |
Table IV
Tumor metastasis-related genes
upregulated or downregulated >10-fold in PC-3 cells compared
with LNCap cells.
| Gene | Description | Fold-change:
upregulation or downregulation |
|---|
| CDH11 | Cadherin 11, type
2, OB-cadherin (osteoblast) | 87,443.2 |
| CD44 | CD44 molecule
(Indian blood group) | 94,50.01 |
| IL1B | Interleukin 1,
β | 16,76.25 |
| IL18 | Interleukin 18
(interferon-γ-inducing factor) | 407.47 |
| TGFβ1 | Transforming growth
factor, β1 | 292.18 |
| MET | Met proto-oncogene
(hepatocyte growth factor receptor) | 290.7 |
| HGF | Hepatocyte growth
factor (hepapoietin A; scatter factor) | 142.19 |
| MTSS1 | Metastasis
suppressor 1 | 112.47 |
| CDH6 | Cadherin 6, type 2,
K-cadherin (fetal kidney) | 82.12 |
| FXYD5 | FXYD domain
containing ion transport regulator 5 | 71.28 |
| PLAUR | Plasminogen
activator, urokinase receptor | 54.97 |
| MMP9 | Matrix
metallopeptidase 9 (gelatinase B, 92 kDa gelatinase, 92 kDa type IV
collagenase) | 54.91 |
| TIMP4 | TIMP
metallopeptidase inhibitor 4 | 50.48 |
| ITGA7 | Integrin, α7 | 35.24 |
| SYK | Spleen tyrosine
kinase | 32.42 |
| MCAM | Melanoma cell
adhesion molecule | 28.16 |
| FN1 | Fibronectin 1 | 25.31 |
| NME4 | Non-metastatic
cells 4, protein expressed in | 24.95 |
| MMP13 | Matrix
metallopeptidase 13 (collagenase 3) | 24.72 |
| ETV4 | Ets variant 4 | 21.47 |
| TNFSF10 | Tumor necrosis
factor (ligand) superfamily, member 10 | 13.78 |
| CDKN2A | Cyclin-dependent
kinase inhibitor 2A (melanoma, p16, inhibits CDK4) | 13.06 |
| COL4A2 | Collagen, type IV,
α2 | 11.86 |
| TP53 | Tumor protein
p53 | −22.72 |
| PTEN | Phosphatase and
tensin homolog | −67.89 |
| CTNNA1 | Catenin
(cadherin-associated protein), α1, 102 kDa | −77.89 |
| CDH1 | Cadherin 1, type 1,
E-cadherin (epithelial) | −195.5 |
| MDM2 | MDM2 p53 binding
protein homolog (mouse) | −204.1 |
| IGF1 | Insulin-like growth
factor 1 (somatomedin C) | −3,806 |
Screening of differentially expressed
tumor metastasis-related genes between LNCap and si-PSMA LNCap
cells by the tumor metastasis PCR array
We further analyzed the differential expression of
tumor metastasis-related genes in the si-PSMA LNCap cells in which
PSMA expression was blocked. Tumor metastasis PCR array analysis
revealed that 10 genes were upregulated (namely CDH6, CXCL12, IL18,
IL8RB, IGF1, MMP13, MMP3, MTSS1, TSHR and ITGA7) and 4 genes were
downregulated (CCL7, ITGB3, MDM2 and MMP2) significantly in the
si-PSMA LNCap cells when compared to the PSMA non-silenced LNCap
cells. The differentially expressed genes and their description are
documented in Table V. Consistent
with other reports, the IGF1 gene has been previously reported to
be upregulated in C4-2 cells when compared with LNCap cells
(18). Among the total 14 genes
that were significantly altered in the si-PSMA LNCap cells, the
MDM2 gene showed the maximum downregulated change when PSMA
expression was suppressed (Table
V), consistent with a previous report (19).
 | Table VTumor metastasis-related genes
upregulated or downregulated in the si-PSMA LNCap cells. |
Table V
Tumor metastasis-related genes
upregulated or downregulated in the si-PSMA LNCap cells.
| Gene | Description | Fold-change:
upregulation or downregulation |
|---|
| CDH6 | Cadherin 6, type 2,
K-cadherin (fetal kidney) | 2.88 |
| CXCL12 | Chemokine (C-X-C
motif) ligand 12 (stromal cell-derived factor 1) | 2.51 |
| IGF1 | Insulin-like growth
factor 1 (somatomedin C) | 2.04 |
| IL18 | Interleukin 18
(interferon-γ-inducing factor) | 2.04 |
| IL8RB | Interleukin 8
receptor, β | 2.22 |
| ITGA7 | Integrin, α7 | 2.25 |
| MMP13 | Matrix
metallopeptidase 13 (collagenase 3) | 2.24 |
| MMP3 | Matrix
metallopeptidase 3 (stromelysin 1, progelatinase) | 2.40 |
| MTSS1 | Metastasis
suppressor 1 | 2.05 |
| TSHR | Thyroid stimulating
hormone receptor | 2.82 |
| CCL7 | Chemokine (C-C
motif) ligand 7 | −2.23 |
| ITGB3 | Integrin, β3
(platelet glycoprotein IIIa, antigen CD61) | −2.05 |
| MDM2 | MDM2 p53 binding
protein homolog (mouse) | −80.35 |
| MMP2 | Matrix
metallopeptidase 2 (gelatinase A, 72 kDa gelatinase, 72 kDa type IV
collagenase) | −2.29 |
Screening of tumor metastasis-related
genes possibly related to PSMA by tumor metastasis PCR array
By comparison of the differentially expressed genes
between the PSMA(−) group and the si-PSMA group, we aimed to
identify those common genes that may be regulated by or related to
PSMA. From the tumor metastasis PCR array results, we found that
expression of 6 genes (namely IL18, MTSS1, MMP13, ITGA7, MMP3 and
CDH6) was significantly upregulated in both the si-PSMA LNCap and
the PC-3 cells, although their expression levels were different.
Expression of 2 genes (MDM2 and MMP2) was downregulated to a
similar extent in both the PC-3 and the si-PSMA LNCap cells. The
results indicate that these 8 genes may be related to the role of
PSMA in tumor metastasis.
The genes differently expressed between the PSMA(−)
and si-PSMA groups included 2 genes (CCL7 and ITGB3) that were
upregulated significantly in the si-PSMA LNCap cells but not in the
PC-3 cells. Expression of 13 genes was significantly lower in the
PC-3 cells but not in the si-PSMA LNCap cells. Moreover, expression
of IGF1 and CXCL12 was significantly upregulated in si-PSMA LNCap
cells, but significantly downregulated in the PC-3 cells. The ΔCt
changes in 28 genes were <2 and showed no significant change in
both the PC-3 cells and the si-PSMA LNCap cells.
Verification of tumor metastasis-related
gene expression in prostate cancer cell lines and clinical prostate
samples
Three candidate genes, namely MMP3, CDH6 and MTSS1,
were chosen for further analysis and identification in the PCa cell
lines and prostate tissues by real-time RT-PCR. PCa cell lines used
for verification included the PSMA-positive LNCap and 22RV1 cells,
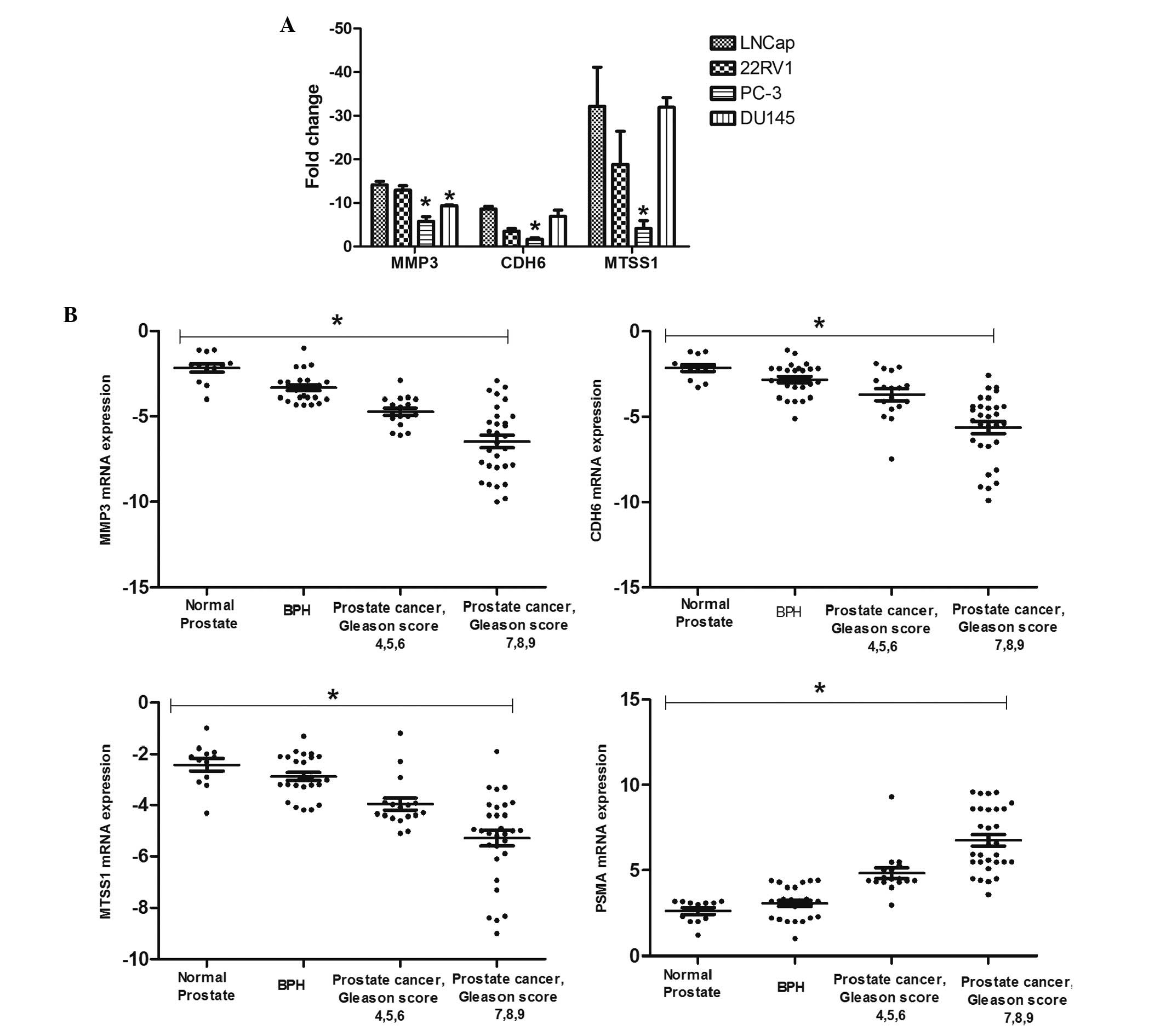
and the PSMA-negative PC-3 and DU145 cells. Consistent with the
results of the tumor metastasis PCR array analysis based on LNCap
and PC-3 cells, the MMP3 mRNA expression level was downregulated in
the PSMA-positive cells including LNCap and 22RV1, and upregulated
in the PSMA-negative cells including PC-3 and DU145 (P<0.05;
Fig. 3A). CDH6 and MTSS1 levels
were significantly higher in the PC-3 cells when compared with the
other 3 cell lines (P<0.05; Fig.
3A). Real-time PCR confirmed the microarray results for MMP3,
CDH6 and MTSS1 in the LNCap and PC-3 cells, and the PSMA-positive
22RV1 cells showed the same differential expression trend as that
in LNCap cells. In contrast, the expression trend of CDH6 and MTSS1
in the PSMA-negative DU145 cells was different from that in the
PC-3 cells.
The results of real-time RT-PCR identification in
clinical samples showed that the relative MMP3, CDH6 and MTSS1
expression level in normal prostate, BPH and PCa tissues decreased
with the development of prostate cancer, and showed a contrary
trend with the expression level of PSMA. Expression levels of MMP3,
MTSS1 and CDH6 were significantly lower in PCa compared with BPH
and normal prostate, respectively (P<0.05). Correlation analysis
demonstrated a negative correlation between the MMP3, MTSS1 and
CDH6 mRNA expression and Gleason score (for MMP3, r=−0.5826,
P<0.0001; for MTSS1, r=−0.4727, P<0.0001; for CDH6,
r=−0.4774, P<0.0001; Fig. 3B).
Negative correlation was also observed between the levels of PSMA
and MMP3, MTSS1 and CDH6, indicating that these genes may be
involved in the regulation of prostate cancer metastasis mediated
by PSMA (for MMP3, r=−0.6811, P<0.05; for CDH6, r=−0.5430,
P<0.05; for MTSS1, r=−0.5540, P<0.05) (Fig. 3B).
Discussion
Prostate cancer metastasis is a complicated and
continuous process, which includes tumor cell shedding, invasion,
migration, adhesion and proliferation, a process that involves many
factors (20,21). Many tumor metastasis-related genes
that can promote the tumor metastasis process have been reported
(20,21), but few tumor metastasis-related
genes in prostate cancer have been identified. Therefore, further
study of metastasis-related genes is needed to understand the
molecular mechanisms of PCa metastasis. In our previous studies, we
found that PSMA negatively regulates the process of metastasis in
PCa (12), and blocking of PSMA
promotes the invasion and migration of LNCap cells (12), yet the underlying mechanism is still
unknown. Therefore, to identify the metastasis-related genes
possibly regulated by PSMA, in the present study, we screened 84
metastasis-related genes by tumor metastasis PCR array analysis.
The differentially expressed genes between LNCap and PSMA-silenced
LNCap cells, and between PSMA-positive LNCap cells and
PSMA-negative PC-3 cells were analyzed, and genes possibly related
to PSMA were screened by comparison of the common differentially
expressed genes in both the si-PSMA group and the PSMA(−)
group.
Tumor metastasis PCR array analysis was chosen for
the screening of PSMA-related tumor metastasis-related genes. This
functional classification gene chip can detect a large number of
genes in a parallel analysis, has the characteristics of
high-throughput, is fast, accurate and sensitive, and shows
superiority in the analysis of disease-related genes and therefore
is widely used in oncobiology studies. The 84 genes selected for
this array encode many classes of protein factors that are related
to tumor metastasis. The results showed that when compared to the
PSMA-positive LNCap cells, there were 41 genes upregulated and 15
genes downregulated in the PSMA-negative PC-3 cells. To further
identify the significant screened genes that were related to PSMA,
we performed gene silencing in PSMA-expressing LNCap cells by
siRNA, to ascertain how the PSMA blockage can affect the
tumor-metastasis gene expression. The results showed that
expression of 14 genes was significantly altered in the si-PSMA
LNCap cells, among which 10 genes were upregulated and 4 genes were
downregulated. By comparison of the differentially expressed genes
between the two groups, we found that 6 genes (IL18, MTSS1, MMP13,
ITGA7, MMP3 and CDH6) were upregulated and 2 genes (MDM2 and MMP2)
were downregulated in both the PSMA-negative PC-3 cells and the
si-PSMA group, suggesting that these genes may be involved in the
regulation of prostate cancer cell migration and invasiveness
mediated by PSMA. Among the 8 screened PSMA-related genes, CDH6,
MDM2 and MMPs were previously reported to be involved in PCa
metastasis (13,22,23),
whereas other genes found in the present study have not yet been
reported.
IL-18 is a cytokine that mediates the immune
response and we deduced that its upregulation is very likely to be
related to tumor immunity. MTSS1 (metastasis tumor suppressor-1) is
a novel putative metastasis-suppressor gene, and its three
alternatively splice variants have been confirmed to be
downregulated in PCa cell lines and clinical samples, and may
contribute to tumor growth and development, as well as metastasis
(24). CDH6 (cadherin-6) is a
calcium-dependent transmembrane glycoprotein, highly expressed in
liver and kidney cancer, and plays an important role in the process
of cancer metastasis and invasion. Yet, to date, it has not been
reported in PCa (25). MMP2 and
MMP3 gene expression levels and enzyme activity were found to be
elevated in brain metastases of breast cancer (26). MMP13 also belongs to the MMP family,
and high expression of MMP13 may be an important target for breast
cancer (27). MDM2 gene expression
is increased in prostate cancer (19), and its function and pathway have
been elucidated (22). ITGA7
belongs to the integrin α chain family (28), and has been shown to interact with
FHL2 and FHL3 (29). It has also
been reported that ITGA7 interacts with high temperature
requirement A2 (HtrA2) to induce cell death in human PCa cell lines
PC-3 and DU145 (30). Based on the
previous reports of these genes, MMP3, MTSS1 and CDH6 were chosen
for further verification in prostate cancer cell lines and clinical
samples.
Real-time PCR identification results showed that,
consistent with the results of gene chip analysis based on LNCap
cells and PC-3 cells, MMP3 was expressed at a higher level in PC-3
and DU145 than in LNCap and 22RV1 cells (P<0.05). Expression
levels of CDH6 and MTSS1 were upregulated in PC-3 cells when
compared to LNCap, 22RV1 and DU145 cells. Differential expression
results of MMP3, MTSS1 and CDH6 shown in tumor metastasis PCR array
analysis were confirmed by real-time PCR in the PSMA positive LNCap
and 22RV1 cells, and PSMA-negative PC-3 cells, while in DU145, the
expression level of CDH6 and MTSS1 was different from that in the
PSMA-negative PC-3 cells. Since DU145 was derived from metastatic
prostate cancer cells, its metastasis gene expression profile may
be different. The underlying mechanism needs further study.
To further identify the expression of these genes in
clinical prostate tissues, we detected their expression level in
normal prostate, BPH and PCa tissues by real-time PCR. We found
that expression levels of MMP3, CDH6 and MTSS1 genes were high in
normal prostate tissues, relatively high in BPH tissues, but
significantly decreased in PCa samples, and moreover, their
expression decreased with the Gleason score and showed high
correlation to tumors. It was noteworthy that the expression of
these genes showed a negative correlation with PSMA expression
level, which suggests that MMP3, CDH6 and MTSS1 may be involved in
the regulatory mechanism of tumor metastasis by PSMA, and may also
have potential significance as indicators of malignant potential of
PCa. Thus, we deduced that PSMA may suppress the migration and
invasiveness of PCa cells through directly or indirectly
interacting with MMP3, CDH6 and MTSS1. These results may provide
important insights into the molecular mechanism of PCa metastasis.
However, further studies are needed to elucidate the possible
pathway by which PSMA regulates PCa metastasis and how it interacts
with these molecules.
Genes showing discordant change in the PSMA(−) and
the si-PSMA groups were noted. CCL7 and ITGB3 genes were
upregulated in the si-PSMA LNCap cells, while the expression in
PC-3 cells was not significantly changed. IGF1 and CXCL12 genes
were highly expressed in the si-PSMA LNCap cells, but at a
significantly lower level in the PC-3 cells. Thirteen genes were
significantly expressed at a lower level in PC-3 cells but not in
the si-PSMA LNCap cells. These genes are reported to play a role in
the process of carcinogenesis and tumor metastasis (31–33).
Although some of the genes screened in this study were previously
reported, e.g., upregulation of CXCL12 is a major mechanism
underlying SLUG-mediated migration and invasion of PCa cells
(34), most of the differentially
expressed genes were reported to be involved in the metastasis of a
variety of tumors types, but not in PCa. The exact role of these
genes in PCa metastasis remains to be explored.
In conclusion, the present study screened
differentially expressed tumor metastasis-related genes in LNCap
and PC-3 cells and in si-PSMA LNCap and LNCap cells by gene chip
technique. Eight genes were screened as PSMA-related by comparison
of the common differentially expressed genes, and 3 were further
identified in PCa cell lines and clinical samples. These findings
provide insight into the regulatory mechanism of the suppression of
PCa metastasis by PSMA and the possible interacting proteins.
Evidence is provided of the pathway implicated in the suppression
of PCa metastasis by PSMA, and provides important clues for further
exploration of the molecular mechanisms of PCa metastasis.
Acknowledgements
The present study was supported by grants from the
Natural Science Foundation of China (nos. 81071760, 30772503 and
30371426), and the Natural Science Foundation of Guangdong
Province, China (no. 021907).
References
|
1
|
Chang SS: Overview of prostate-specific
membrane antigen. Rev Urol. 6(Suppl 10): S13–S18. 2004.PubMed/NCBI
|
|
2
|
Kypta R, Unda M and Carracedo A: Is the
bench getting closer to the bedside in the war on cancer? A quick
look at prostate cancer. Front Endocrinol. 3:532012. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Nelson WG, De Marzo AM and Isaacs WB:
Prostate cancer. N Engl J Med. 349:366–381. 2003. View Article : Google Scholar
|
|
4
|
Paller CJ, Carducci MA and Philips GK:
Management of bone metastases in refractory prostate cancer - role
of denosumab. Clin Interv Aging. 7:363–372. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Jemal A, Tiwari RC, Murray T, Ghafoor A,
Samuels A, Ward E, Feuer EJ and Thun MJ: Cancer statistics. CA
Cancer J Clin. 54:8–29. 2004.
|
|
6
|
Germain A, Richardson R, Moul DE, Mammen
O, Haas G, Forman SD, Rode N, Begley A and Nofzinger EA:
Placebo-controlled comparison of prazosin and cognitive-behavioral
treatments for sleep disturbances in US Military Veterans. J
Psychosom Res. 72:89–96. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Schroder FH: Progress in understanding
androgen-independent prostate cancer (AIPC): a review of potential
endocrine-mediated mechanisms. Eur Urol. 53:1129–1137. 2008.
View Article : Google Scholar
|
|
8
|
Kasper S and Cookson MS: Mechanisms
leading to the development of hormone-resistant prostate cancer.
Urol Clin North Am. 33:201–210. vii2006. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Pinto JT, Suffoletto BP, Berzin TM, Qiao
CH, Lin S, Tong WP, May F, Mukherjee B and Heston WD:
Prostate-specific membrane antigen: a novel folate hydrolase in
human prostatic carcinoma cells. Clin Cancer Res. 2:1445–1451.
1996.PubMed/NCBI
|
|
10
|
Robinson MB, Blakely RD, Couto R and Coyle
JT: Hydrolysis of the brain dipeptide
N-acetyl-L-aspartyl-L-glutamate. Identification and
characterization of a novel N-acetylated alpha-linked acidic
dipeptidase activity from rat brain. J Biol Chem. 262:14498–14506.
1987.PubMed/NCBI
|
|
11
|
Cao KY, Mao XP, Wang DH, Xu L, Yuan GQ,
Dai SQ, Zheng BJ and Qiu SP: High expression of PSM-E correlated
with tumor grade in prostate cancer: a new alternatively spliced
variant of prostate-specific membrane antigen. Prostate.
67:1791–1800. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Cao KY, Xu L, Zhang DM, Zhang XM, Zhang T,
He X, Wang Z, Feng FS, Qiu SP and Shen GX: New alternatively
spliced variant of prostate-specific membrane antigen PSM-E
suppresses the proliferation, migration and invasiveness of
prostate cancer cells. Int J Oncol. 40:1977–1985. 2012.PubMed/NCBI
|
|
13
|
Ghosh A and Heston WD: Tumor target
prostate specific membrane antigen (PSMA) and its regulation in
prostate cancer. J Cell Biochem. 91:528–539. 2004. View Article : Google Scholar
|
|
14
|
Frigerio B, Fracasso G, Luison E,
Cingarlini S, Mortarino M, Coliva A, Seregni E, Bombardieri E,
Zuccolotto G, Rosato A, Colombatti M, Canevari S and Figini M: A
single-chain fragment against prostate specific membrane antigen as
a tool to build theranostic reagents for prostate cancer. Eur J
Cancer. pii: S0959-8049(13)00087-7. View Article : Google Scholar : 2013. View Article : Google Scholar
|
|
15
|
Zhao LY, Mao XP, Chao KY, Guo SJ and Qiu
SP: Prostate-specific membrane antigen can promote in vivo osseous
metastasis of prostate cancer cells in mice. Braz J Med Biol Res.
45:737–745. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Mannweiler S, Amersdorfer P, Trajanoski S,
Terrett JA, King D and Mehes G: Heterogeneity of prostate-specific
membrane antigen (PSMA) expression in prostate carcinoma with
distant metastasis. Pathol Oncol Res. 15:167–172. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Ghosh A, Wang X, Klein E and Heston WD:
Novel role of prostate-specific membrane antigen in suppressing
prostate cancer invasiveness. Cancer Res. 65:727–731.
2005.PubMed/NCBI
|
|
18
|
Xie BX, Zhang H, Wang J, Pang B, Wu RQ,
Qian XL, Yu L, Li SH, Shi QG, Huang CF and Zhou JG: Analysis of
differentially expressed genes in LNCaP prostate cancer progression
model. J Androl. 32:170–182. 2010.PubMed/NCBI
|
|
19
|
Wang G, Firoz EF, Rose A, Blochin E,
Christos P, Pollens D, Mazumdar M, Gerald W, Oddoux C, Lee P and
Osman I: MDM2 expression and regulation in prostate cancer racial
disparity. Int J Clin Exp Pathol. 2:353–360. 2009.PubMed/NCBI
|
|
20
|
Kohn EC: Invasion and metastasis: biology
and clinical potential. Pharmacol Ther. 52:235–244. 1991.
View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Duffy MJ, McGowan PM and Gallagher WM:
Cancer invasion and metastasis: changing views. J Pathol.
214:283–293. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Kong L, Yuan Q, Zhu H, Li Y, Guo Q, Wang
Q, Bi X and Gao X: The suppression of prostate LNCaP cancer cells
growth by Selenium nanoparticles through Akt/Mdm2/AR controlled
apoptosis. Biomaterials. 32:6515–6522. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Roomi MW, Monterrey JC, Kalinovsky T, Rath
M and Niedzwiecki A: Inhibition of invasion and MMPs by a nutrient
mixture in human cancer cell lines: a correlation study. Exp Oncol.
32:243–248. 2011.PubMed/NCBI
|
|
24
|
Loberg RD, Neeley CK, Adam-Day LL, Fridman
Y, St John LN, Nixdorf S, Jackson P, Kalikin LM and Pienta KJ:
Differential expression analysis of MIM (MTSS1) splice variants and
a functional role of MIM in prostate cancer cell biology. Int J
Oncol. 26:1699–1705. 2005.PubMed/NCBI
|
|
25
|
Inoue T, Inoue YU, Asami J, Izumi H,
Nakamura S and Krumlauf R: Analysis of mouse Cdh6 gene regulation
by transgenesis of modified bacterial artificial chromosomes. Dev
Biol. 315:506–520. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Mendes O, Kim HT and Stoica G: Expression
of MMP2, MMP9 and MMP3 in breast cancer brain metastasis in a rat
model. Clin Exp Metastasis. 22:237–246. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Chang HJ, Yang MJ, Yang YH, Hou MF, Hsueh
EJ and Lin SR: MMP13 is potentially a new tumor marker for breast
cancer diagnosis. Oncol Rep. 22:1119–1127. 2009.PubMed/NCBI
|
|
28
|
Wang W, Wu W, Desai T, Ward DC and Kaufman
SJ: Localization of the alpha 7 integrin gene (ITGA7) on human
chromosome 12q13: clustering of integrin and Hox genes implies
parallel evolution of these gene families. Genomics. 26:568–570.
1995. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Samson T, Smyth N, Janetzky S, Wendler O,
Müller JM, Schüle R, von der Mark H, von der Mark K and Wixler V:
The LIM-only proteins FHL2 and FHL3 interact
with α- and β-subunits of the muscle α7β1
integrin receptor. J Biol Chem. 279:28641–28652. 2004.PubMed/NCBI
|
|
30
|
Zhu ZH, Yu YP, Zheng ZL, Song Y, Xiang GS,
Nelson J, Michalopoulos G and Luo JH: Integrin alpha 7 interacts
with high temperature requirement A2 (HtrA2) to induce prostate
cancer cell death. Am J Pathol. 177:1176–1186. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Mazzoccoli G, Sothern RB, Pazienza V,
Piepoli A, Muscarella LA, Giuliani F and Tarquini R: Circadian
aspects of growth hormone-insulin-like growth factor axis function
in patients with lung cancer. Clin Lung Cancer. 13:68–74. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Montagnani Marelli M, Moretti RM, Procacci
P, Motta M and Limonta P: Insulin-like growth factor-I promotes
migration in human androgen-independent prostate cancer cells via
the αvβ3 integrin and PI3-K/Akt signaling.
Int J Oncol. 28:723–730. 2006.PubMed/NCBI
|
|
33
|
Nickerson T, Chang F, Lorimer D, Smeekens
SP, Sawyers CL and Pollak M: In vivo progression of LAPC-9 and
LNCaP prostate cancer models to androgen independence is associated
with increased expression of insulin-like growth factor I (IGF-I)
and IGF-I receptor (IGF-IR). Cancer Res. 61:6276–6280.
2001.PubMed/NCBI
|
|
34
|
Uygur B and Wu WS: SLUG promotes prostate
cancer cell migration and invasion via CXCR4/CXCL12 axis. Mol
Cancer. 10:1392011. View Article : Google Scholar : PubMed/NCBI
|