Introduction
Vascular endothelial growth factor (VEGF) is one of
the key modulators of angiogenesis. The VEGF gene is located
on chromosome 6p21.3 and is organized as eight exons separated by
seven introns (1,2). Alternative exon splicing was
initially shown to result in the generation of four major isoforms
that were 121, 165, 189 and 206 amino acids in length (2). The VEGF promoter, which spans
2.36 kb, contains several transcription factor binding sites,
including Sp1/Sp3, activating protein (AP)-2, Egr-1, signal
transducer and activator of transcription-3 and hypoxia-inducing
factor-1. These transcription factor binding sites are highly
conserved in mice, rats and humans. These studies indicated that
the VEGF promoter is critical in the regulation of VEGF
expression. Several studies showed an association between
VEGF polymorphisms and breast cancer
susceptibility/aggressiveness, as well as levels of VEGF expression
[reviewed in (3)].
Haplotype analysis showed that −1498C was linked
with −634G (4). These two
polymorphisms were associated with breast cancer susceptibility and
aggressiveness (5–9). However, these polymorphisms were not
located in the established transcription factor binding sites.
In vitro models suggested a haplotypic effect of the
polymorphic VEGF promoter on basal and stimulated promoter
activity (10). In the present
study, VEGF mRNA and VEGF protein expression in breast
cancer tissue were determined and were correlated with various
clinicopathological parameters. To verify the functional role of
VEGF polymorphisms at the −634 and −1498 positions,
site-directed mutagenesis was performed to generate different
VEGF genotypes and to exclude other functional polymorphisms
that may be in linkage disequilibrium with the polymorphisms of
interest. The transcriptional activities of these polymorphisms
were determined by a dual-luciferase assay.
Materials and methods
Study population
The study population was recruited from the Division
of Head-Neck and Breast Surgery, Department of Surgery, Faculty of
Medicine, Siriraj Hospital (Bangkok, Thailand) between 2000 and
2003. Patients with newly diagnosed breast cancer, aged ≥18 years
with ability to provide informed consent were included. Patients
with a history of other cancers were excluded. At recruitment,
informed consent was obtained, and each participant was interviewed
to collect detailed information with regard to demographic
characteristics. This study was approved by the Siriraj Ethics
Committee on Research (Bangkok, Thailand).
Genotyping of VEGF polymorphisms
Genomic DNA was obtained from peripheral blood
according to a standard method. Briefly, venous blood samples were
drawn into EDTA-containing tubes. The leukocyte cell pellet
obtained from the buffy coat was resuspended with TE 20-5 solution,
then digested with proteinase K (Promega Corporation, Madison, WI,
USA) at 37°C overnight. DNA was isolated by the addition of phenol
and chloroform-isoamyl alcohol (24:1) and centrifugation. The
VEGF −634G/C polymorphisms were genotyped on allele
refractory mutation system-polymerase chain reaction (PCR) using
primers as follows: Forward, 5′-CATTGATCCGGGTTTTATCCC-3′; reverse
−634G, 5′-CACTCACTTTGTCCCTGTAG-3′; reverse −634C, 5′-CAC
TCACTTTGTCCCTGTAC-3′; forward control, 5′-AGA TGGTCCCTCACCTTCCT-3′;
and reverse control, 5′-GTC TACCCTCCTGAGCTTGC-3′.
VEGF-1498C/T polymorphisms were genotyped by PCR-restriction
fragment length polymorphisms. The forward primer was 5′-TGTGCGTGT
GGGGTTGAGCG-3′ and the reverse primer was 5′-TACGTG
CGGACAGGGCCTGA-3′. The products were digested with the BstUI
restriction enzyme (New England Biolabs, Beverly, MA, USA). −1498C
products were digested and resulted in 155 and 20 bp fragments.
Representative PCR products were sequenced to validate the
assay.
Evaluation of VEGF expression and
microvessel density (MVD) in breast cancer tissue
The levels of VEGF mRNA expression in breast
cancer tissue were determined by qPCR as described previously
(11). VEGF protein expression was
determined by immunohistochemistry as follows: Paraffin-embedded
sections were stained with a monoclonal mouse antibody to human
VEGF clone VG1 (dilution, 1:100; incubation time, 1 h; Diagnostic
BioSystems, Pleasanton CA, USA) and a monoclonal antibody to CD31
(dilution, 1:300; incubation time, 16 h; Dako, Glostrup, Denmark).
The immunohistochemical data was evaluated by two pathologists who
had no knowledge of the patients’ characteristics and/or clinical
outcome. Expression of VEGF was assessed semiquantitatively using
an immunohistochemical score (H score). The score was calculated by
multiplying the percentage of positive carcinoma cells by the
staining intensity (0, negative; 1, weak; 2, moderate; and 3,
strong; as determined subjectively by the two pathologists). The
average of the H scores from two pathologists was used. The median
of the scores was used as the cutoff level to categorize tumors
into low- (below the median H score) and high-expressing tumors
(above the median H score), with regard to VEGF. MVD was expressed
as the average number of microvessels per ×200 field. The three
most intense areas of angiogenesis were identified and microvessels
were counted. A single microvessel was defined as any brown
immunostained endothelial cell that was separated from adjacent
microvessels and other connective tissue elements. Large vessels
with thick muscular walls were not counted, and the presence of a
lumen was not required for scoring as a microvessel. The median of
the MVD was used as the cutoff level to categorize tumors into low-
and high-MVD tumors.
Construction of plasmids
pCR2.1 plasmids were purchased from Invitrogen Life
Technologies (Grand Island, NY, USA). pGL3-Basic and pRL-SV40
plasmids were purchased from Promega Corporation (Madison, WI,
USA). The VEGF promoter was amplified from human genomic DNA
using the following primers: Forward primer, 5′-CAGGACTAGTGC
ACGAATGA-3′ and reverse primer, 5′-CTGTCTGTCTGT CCGTCAG-3′. The PCR
reaction was conducted in a 100-μl reaction containing 5 units of
ProofStart DNA polymerase (Qiagen, Hilden, Germany), 1X PCR buffer
(containing 1.5 mM MgCl2), 0.2 mM dNTPs, 0.4 μM of each
primer, 1X Q-solution and 800 ng genomic DNA. The 3′ A-overhang was
added to the purified PCR product by means of a Qiagen A-addition
kit (Qiagen). This product was immediately ligated into pCR2.1
plasmid using T4 DNA ligase (Life Technologies Corporations, Grand
Island, NY, USA). The ligated plasmid was transformed into
Escherichia coli (E. coli) strain DH5α and
propagated. The constructed plasmids were fully sequenced to
exclude PCR errors. The promoter was excised using HindIII
and XhoI (both from New England Biolabs) prior to ligation
into the pGL3-Basic vector. The promoter-reporter plasmid was
resequenced to confirm correct orientation of the promoter. This
plasmid was used as a template to generate other plasmids
containing different polymorphisms.
Site-directed mutagenesis
Plasmids containing different polymorphisms were
amplified by ProofStart DNA polymerase (Qiagen). The DNA primers
were as follows: −1498Mut-C forward, 5′-GTGGGGTTGAGGGCGTTGGAGCGGGG-3′ and reverse,
5′-CCCCGCTCCAACGCCCTCAACCCCAC-3′; -634Mut-C
forward, 5′-GAGCAGCGAAAGCGACAGGGGC AAAGTG-3′ and
reverse, 5′-CACTTTGCCCCTGTCGCT TTCGCTGCTC-3′. The
underlined base indicated the site of mutagenesis. The amplified
products were digested by the addition of 10 units of DpnI
(Stratagene, Santa Clara, CA, USA) directly to the reaction. The
DpnI-treated DNA was transformed into E. coli. All
plasmids were fully sequenced to exclude PCR errors and to confirm
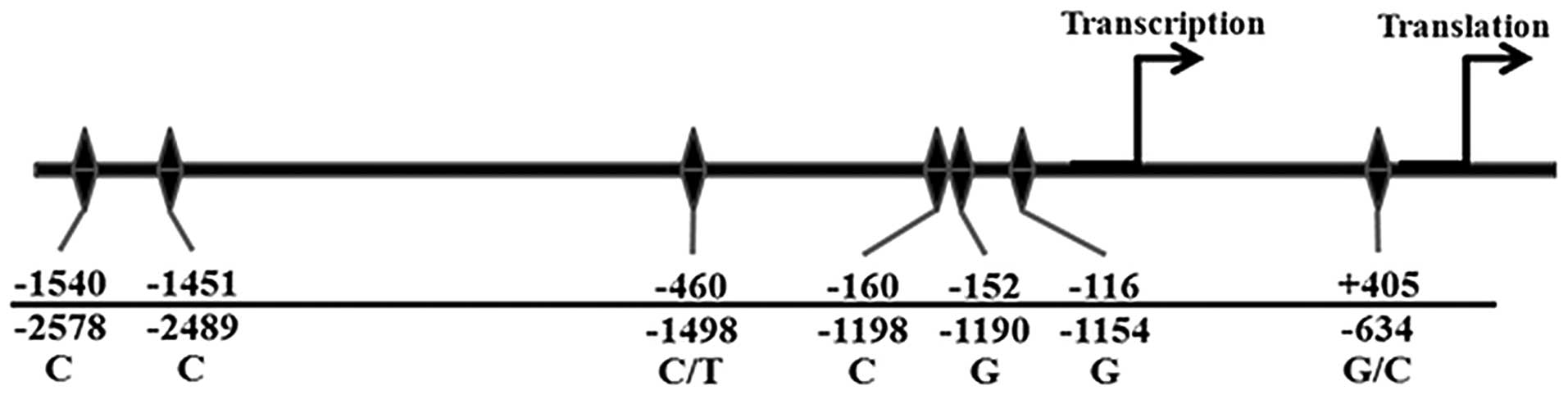
the presence of the polymorphisms. The polymorphisms in other
positions are illustrated in Fig.
1.
Cell culture and DNA transfection
MCF7 breast carcinoma cells (derived from the
American Type Culture Collection, Manassas, VA, USA), were cultured
in Dulbecco’s modified Eagle’s medium with Ham’s Nutrient Mixture
F12 containing 10% fetal calf serum (Life Technologies, Inc.,
Middlesex, UK) at 37°C and in 5% CO2. For transient
transfection, 5×104 cells were placed in 24-well plates
and grown to 60–70% confluence. Transfection was conducted using
Lipofectamine reagent Life Technologies, Inc. (Grand Island, NY,
USA) according to the manufacturer’s instructions. Cells were
co-transfected with VEGF promoter-luciferase plasmid and
Renilla luciferase plasmid (pRL-SV40). After 5 h of
incubation, fetal calf serum was added to a final concentration of
10% with or without 10−7 M phorbol myristate acetate,
and incubated for 16 h prior to evaluation of luciferase
activity.
Dual-luciferase reporter assay
The cells were washed with phosphate-buffered saline
and passive lysis buffer (Promega Corporation) was added to each
well. The dual-luciferase reporter assay was performed according to
the manufacturer’s instructions. The experiments were performed in
sextuplicate and repeated on three independent occasions.
Statistical analysis
The level of mRNA was calculated as the ratio of
tissue sample to corresponding β-actin and then corrected as a
ratio to the MDA-MB231 on the same scan. All luciferase results
were expressed as a ratio to the luciferase activity of the empty
vector and were normalized by Renilla luciferase activity.
Analysis of variance was used to evaluate the difference in mRNA
expression among different genotypes and the difference in
luciferase activity among haplotypes. Scheffe’s post hoc test was
performed to compare the difference between each pair of
haplotypes. P<0.05 was considered to indicate a statistically
significant difference.
Results
Genotyping of VEGF polymorphisms
The distribution of the VEGF genotype among breast
cancer patients is summarized in Table
I. The characteristics of the patients were summarized in our
published data (12). Due to
inadequate specimens and poor tissue quality, certain breast cancer
specimens were not included in the analysis.
 | Table IDetermination of the distribution of
VEGF genotypes among breast cancer patients. |
Table I
Determination of the distribution of
VEGF genotypes among breast cancer patients.
| A, qPCR
(total=124) |
|---|
|
|---|
| VEGF
gentoype | No. of patients
(%) |
|---|
| −1498 |
| TT | 65 (52.42) |
| CT | 53 (42.74) |
| CC | 6 (4.84) |
| −634 |
| GG | 57 (45.97) |
| GC | 55 (44.35) |
| CC | 12 (9.68) |
|
| B,
Immunohistochemistry (total=108) |
|
| VEGF
gentoype | No. of patients
(%) |
|
| −1498 |
| TT | 58 (53.70) |
| CT | 44 (40.74) |
| CC | 6 (5.56) |
| −634 |
| GG | 47 (43.52) |
| GC | 50 (46.30) |
| CC | 11 (10.18) |
VEGF expression in breast cancer
tissue
Patients with the −634CC genotype had significantly
higher VEGF mRNA in breast cancer tissue than those with
−634GG or −634GC genotypes (P<0.001). High VEGF mRNA was
previously shown to be associated with a tumor size >2 cm [odds
ratio (OR), 2.476; 95% confidence interval (CI), 1.047–5.858,
P=0.039], the presence of lymphovascular invasion (OR, 2.406; 95%
CI, 1.142–5.070; P=0.021), and the presence of axillary nodal
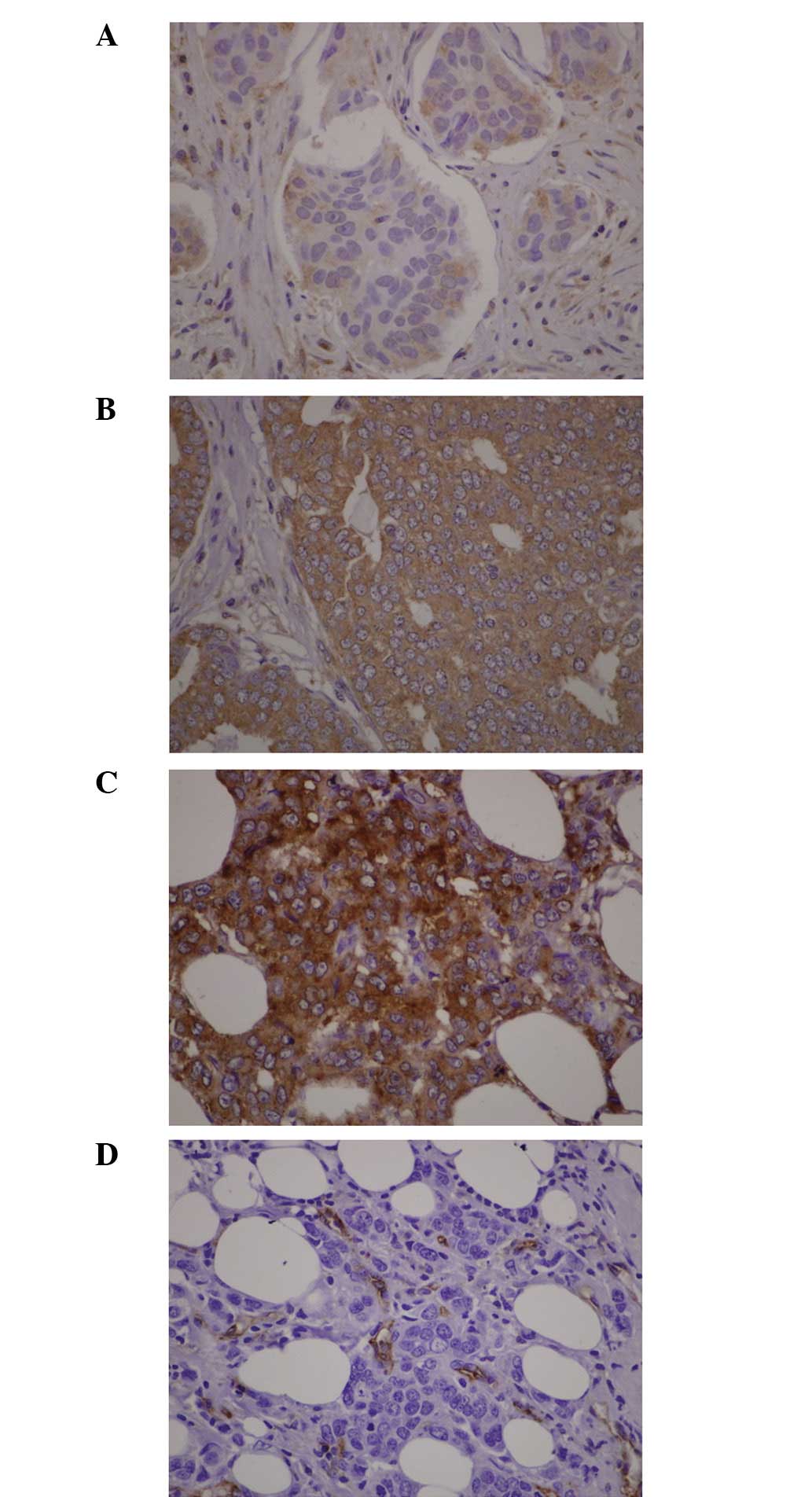
metastasis (OR, 2.288; 95% CI, 1.110–4.713; P=0.025) (12). Fig.
2 shows representative VEGF immunohistochemistry and staining
of endothelial cells. All specimens were VEGF positive. The
percentage of positive carcinoma cells ranged from 10 to 100%. The
intensity was expressed as weak, moderate or strong staining. The H
score ranged from 10 to 300. The median of the H score was 141.25.
At this cutoff level, 54 breast carcinoma specimens were classified
as exhibiting high VEGF expression. No association between
VEGF genotype and H score was observed. High VEGF H-scores
were associated with higher MVD counts (P=0.021) and the
correlation between the H score and MVD count was statistically
significant (Pearson correlation, 0.203; P=0.035). No significant
difference was observed between MVD and any of the four
polymorphisms.
Comparison of transcriptional activity of
different VEGF promoter genotypes
Comparison of the VEGF promoter and empty
vector activity revealed that the VEGF promoter increased
basal luciferase activity of the pGL3-plasmid (Table II). The VEGF promoter
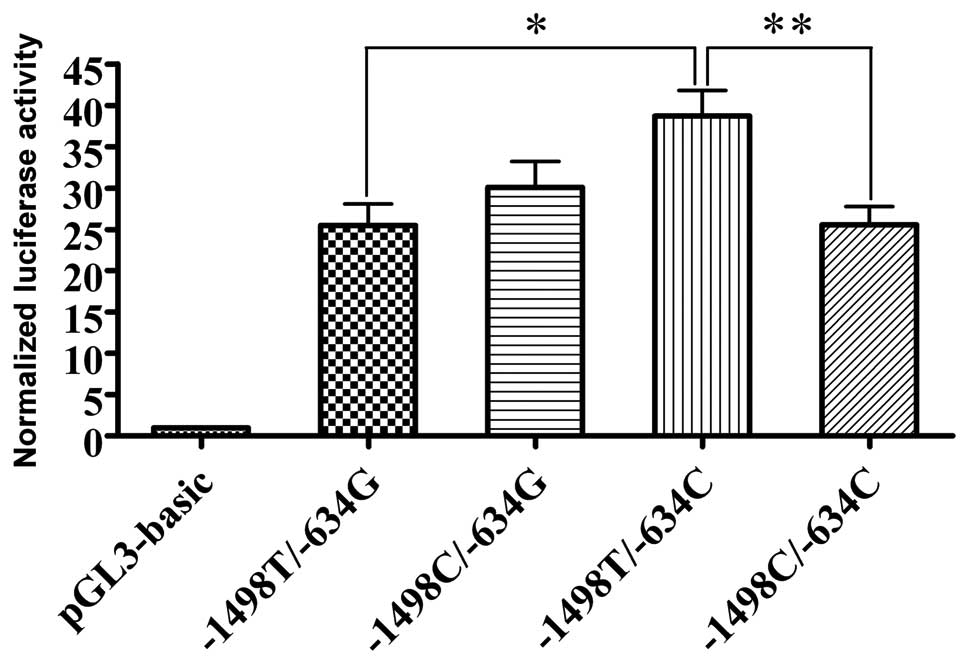
bearing −1498T/−634C had significantly higher promoter activity
when compared with −1498T/−634G and −1498C/−634C (P=0.014 and
0.015, respectively; Fig. 3). To
examine mechanisms that VEGF polymorphisms use to alter
transcription, phorbol ester, which is known to stimulate the AP-1
site was added into the culture medium. Notably, phorbol ester
increased transcriptional activity of the internal control,
Renilla luciferase, more than the VEGF promoter-pGL3
Luc. This resulted in a decrease in normalized luciferase activity
when compared with the basal luciferase activity of each
VEGF promoter genotype.
 | Table IINormalized luciferase activity from
three independent experiments. |
Table II
Normalized luciferase activity from
three independent experiments.
| Normalized luciferase
activity, mean (standard deviation) |
|---|
|
|
|---|
| −460T+405G | −460C+405G | −460T+405C | −460C+405C |
|---|
| I | 15.12 (1.55) | 18.56 (3.43) | 26.05 (3.83) | 18.04 (1.60) |
| II | 25.98 (4.05) | 29.21 (7.18) | 39.41 (8.85) | 22.91 (2.79) |
| III | 37.32 (9.69) | 45.03 (9.78) | 53.14 (5.98) | 37.79 (6.18) |
| Average | 25.48 (10.65) | 30.10 (12.79) | 38.73 (12.78) | 25.57 (9.13) |
| P-valuea | 0.003 | | | |
Discussion
The significant correlation between the −634CC
genotype and high levels of VEGF mRNA expression in breast
cancer tissues was demonstrated and was concordant with in
vitro promoter activity. No correlation was identified between
VEGF protein expression and VEGF genotype or mRNA
expression. The findings of the present study and previous studies
(13–15) showed no correlation between VEGF
protein expression which was determined by immunohistochemistry and
VEGF genotype or mRNA expression. Failure to identify the
association may be due to differences in scoring systems and the
lack of reproducibility of subjective scoring in the determination
of VEGF by immunohistochemistry (16). Although the levels of VEGF protein
expression were not evaluated in the patients that were enrolled,
several clinical trials concerning bevacizumab treatment showed
satisfactory results in terms of objective response rate and
progression-free survival (17–20).
The majority of the patients in the present study had a VEGF
intensity of 2+ and the median proportion of positive cells was
80%. This evidence suggests that almost all of the patients had
relatively high VEGF levels and the expression of VEGF occurred in
a dynamic manner as it varied with time.
In the current study, promoters bearing different
haplotypes were generated by site-directed mutagenesis, which
allowed desired loci to change and be compared, without altering
the functional polymorphisms that may be in linkage disequilibrium
with the loci of interest. Alteration of −634G to C resulted in
increased luciferase activity; thus, −634G/C polymorphisms may have
a direct effect on transcriptional activity. However, no
transcription factor binding motif was identified at this position
(21). The transcription factor
binding motif predicted by the MatInspector Online Tool found no
potential transcription factor bound to this position (22). Identification of the transcription
factor binding site using TFSEARCH version 1.3 (http://mbs.cbrc.jp/research/db/TFSEARCH.html) revealed
that −634G was the potential binding site for myeloid zinc finger
protein 1 (MZF1), which is expressed in hematopoietic progenitor
cells that are committed to myeloid lineage differentiation
(23). Watson et al
(4) reported that alteration from
G to C diminished the potential binding capacity. However, MZF1 may
not have any role in the breast cancer cells used in the current
study. Transcriptional activity assessed in GI-1 human glioma cell
lines and Jurkat human lymphoblastic T-lymphocyte cell lines
revealed that constructions bearing −1154G/−634C haplotypes
exhibited higher luciferase activity than those bearing
−1154G/−634G haplotypes (24).
This indicated a direct effect of the alteration from G to C at-634
position on promoter activity, and polymorphisms at this position
may regulate promoter activity at the post-transcriptional level. G
to C alterations may affect the internal ribosome entry site and
enhance transcription of the large VEGF isoform (395 amino acids)
(25).
Stevens et al (10) constructed different haplotypes by
direct amplification of genetic DNA bearing different haplotypes.
This method could not ensure that the polymorphisms other than
those at a specific position were identical. However, due to the
high linkage disequilibrium of the VEGF promoter, this method had
the advantage that the functional polymorphisms that linked with
the position of interest remained linked as a block. It was
reported that haplotypes containing −1198T/−1190G/−634G had higher
basal VEGF promoter activity than haplotypes containing
−1198C/−1190A/−634G. In the current study, alteration from T to C
at position −1498 significantly decreased the VEGF promoter
activity. The interactions between these two positions contributed
to the difference in promoter activity and susceptibility
to/aggressiveness of breast cancer.
In conclusion, the present study demonstrated the
association between mRNA expression and breast cancer
aggressiveness. VEGF polymorphisms altered the expression by
modification of VEGF promoter activity. These findings
suggested that VEGF polymorphisms influence growth and
invasion of breast cancer cells through increased transcriptional
activity and lead to increased angiogenesis. Genotyping of
VEGF as a potential marker for identification of the
high-risk patients may therefore improve the outcome of breast
cancer treatment.
Acknowledgements
This study was supported by the Research and
Development Fund, Faculty of Medicine Siriraj Hospital Medical
School, Mahidol University (Bangkok, Thailand).
References
|
1
|
Vincenti V, Cassano C, Rocchi M and
Persico G: Assignment of the vascular endothelial growth factor
gene to human chromosome 6p21.3. Circulation. 93:1493–1495. 1996.
View Article : Google Scholar
|
|
2
|
Tischer E, Mitchell R, Hartman T, et al:
The human gene for vascular endothelial growth factor. Multiple
protein forms are encoded through alternative exon splicing. J Biol
Chem. 266:11947–11954. 1991.PubMed/NCBI
|
|
3
|
Sa-Nguanraksa D and O-Charoenrat P: The
role of vascular endothelial growth factor A polymorphisms in
breast cancer. Int J Mol Sci. 13:14845–14864. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Watson CJ, Webb NJ, Bottomley MJ and
Brenchley PE: Identification of polymorphisms within the vascular
endothelial growth factor (VEGF) gene: correlation with variation
in VEGF protein production. Cytokine. 12:1232–1235. 2000.
View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Oliveira C, Lourenço GJ, Silva PM, et al:
Polymorphisms in the 5′- and 3′-untranslated region of the VEGF
gene and sporadic breast cancer risk and clinicopathologic
characteristics. Tumour Biol. 32:295–300. 2011.
|
|
6
|
Jin Q, Hemminki K, Enquist K, et al:
Vascular endothelial growth factor polymorphisms in relation to
breast cancer development and prognosis. Clin Cancer Res.
11:3647–3653. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Balasubramanian SP, Cox A, Cross SS,
Higham SE, Brown NJ and Reed MW: Influence of VEGF-A gene variation
and protein levels in breast cancer susceptibility and severity.
Int J Cancer. 121:1009–1016. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Schneider BP, Wang M, Radovich M, et al:
Association of vascular endothelial growth factor and vascular
endothelial growth factor receptor-2 genetic polymorphisms with
outcome in a trial of paclitaxel compared with paclitaxel plus
bevacizumab in advanced breast cancer: ECOG 2100. J Clin Oncol.
26:4672–4678. 2008. View Article : Google Scholar
|
|
9
|
Lu H, Shu XO, Cui Y, et al: Association of
genetic polymorphisms in the VEGF gene with breast cancer survival.
Cancer Res. 65:5015–5019. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Stevens A, Soden J, Brenchley PE, Ralph S
and Ray DW: Haplotype analysis of the polymorphic human vascular
endothelial growth factor gene promoter. Cancer Res. 63:812–816.
2003.PubMed/NCBI
|
|
11
|
O-charoenrat P, Rhys-Evans P, Modjtahedi H
and Eccles SA: Vascular endothelial growth factor family members
are differentially regulated by c-erbB signaling in head and neck
squamous carcinoma cells. Clin Exp Metastasis. 18:155–161. 2000.
View Article : Google Scholar
|
|
12
|
Sa-Nguanraksa D, Chuangsuwanich T,
Pongpruttipan T, et al: Vascular endothelial growth factor −634G/C
polymorphism is associated with increased breast cancer risk and
aggressiveness. Mol Med Rep. 8:1242–1250. 2013.
|
|
13
|
Kostopoulos I, Arapantoni-Dadioti P, Gogas
H, et al: Evaluation of the prognostic value of HER-2 and VEGF in
breast cancer patients participating in a randomized study with
dose-dense sequential adjuvant chemotherapy. Breast Cancer Res
Treat. 96:251–261. 2006. View Article : Google Scholar
|
|
14
|
Ludovini V, Sidoni A, Pistola L, et al:
Evaluation of the prognostic role of vascular endothelial growth
factor and microvessel density in stages I and II breast cancer
patients. Breast Cancer Res Treat. 81:159–168. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
MacConmara M, O’Hanlon DM, Kiely MJ,
Connolly Y, Jeffers M and Keane FB: An evaluation of the prognostic
significance of vascular endothelial growth factor in node positive
primary breast carcinoma. Int J Oncol. 20:717–721. 2002.PubMed/NCBI
|
|
16
|
Maae E, Nielsen M, Steffensen KD, Jakobsen
EH, Jakobsen A and Sørensen FB: Estimation of immunohistochemical
expression of VEGF in ductal carcinomas of the breast. J Histochem
Cytochem. 59:750–760. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Miller KD, Chap LI, Holmes FA, et al:
Randomized phase III trial of capecitabine compared with
bevacizumab plus capecitabine in patients with previously treated
metastatic breast cancer. J Clin Oncol. 23:792–799. 2005.
View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Miller K, Wang M, Gralow J, et al:
Paclitaxel plus bevacizumab versus paclitaxel alone for metastatic
breast cancer. N Engl J Med. 357:2666–2676. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Robert NJ, Diéras V, Glaspy J, et al:
RIBBON-1: randomized, double-blind, placebo-controlled, phase III
trial of chemotherapy with or without bevacizumab for first-line
treatment of human epidermal growth factor receptor 2-negative,
locally recurrent or metastatic breast cancer. J Clin Oncol.
29:1252–1260. 2011. View Article : Google Scholar
|
|
20
|
O’Shaughnessy JA and Brufsky AM: RiBBON 1
and RiBBON 2: phase III trials of bevacizumab with standard
chemotherapy for metastatic breast cancer. Clin Breast Cancer.
8:370–373. 2008.PubMed/NCBI
|
|
21
|
Pagès G and Pouysségur J: Transcriptional
regulation of the Vascular Endothelial Growth Factor gene - a
concert of activating factors. Cardiovasc Res. 65:564–573.
2005.PubMed/NCBI
|
|
22
|
Awata T, Inoue K, Kurihara S, et al: A
common polymorphism in the 5′-untranslated region of the VEGF gene
is associated with diabetic retinopathy in type 2 diabetes.
Diabetes. 51:1635–1639. 2002.
|
|
23
|
Morris JF, Hromas R and Rauscher FJ III:
Characterization of the DNA-binding properties of the myeloid zinc
finger protein MZF1: two independent DNA-binding domains recognize
two DNA consensus sequences with a common G-rich core. Mol Cell
Biol. 14:1786–1795. 1994.PubMed/NCBI
|
|
24
|
Awata T, Kurihara S, Takata N, et al:
Functional VEGF C-634G polymorphism is associated with development
of diabetic macular edema and correlated with macular retinal
thickness in type 2 diabetes. Biochem Biophys Res Commun.
333:679–685. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Huez I, Bornes S, Bresson D, Créancier L
and Prats H: New vascular endothelial growth factor isoform
generated by internal ribosome entry site-driven CUG translation
initiation. Mol Endocrinol. 15:2197–2210. 2001. View Article : Google Scholar : PubMed/NCBI
|