|
1
|
Nugent M, Edney B, Hammerness PG, Dain BJ,
Maurer LH and Rigas JR: Non-small cell lung cancer at the extremes
of age: Impact on diagnosis and treatment. Ann Thorac Surg.
63:193–197. 1997. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Yang SP, Luh KT, Kuo SH and Lin CC:
Chronological observation of epidemiological characteristics of
lung cancer in Taiwan with etiological consideration-a 30-year
consecutive study. Jpn J Clin Oncol. 14:7–19. 1984.PubMed/NCBI
|
|
3
|
Andriani F, Roz E, Caserini R, Conte D,
Pastorino U, Sozzi G and Roz L: Inactivation of both FHIT and p53
cooperate in deregulating proliferation-related pathways in lung
cancer. J Thorac Oncol. 7:631–642. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Toyooka S, Tsuda T and Gazdar AF: The TP53
gene, tobacco exposure and lung cancer. Hum Mutat. 21:229–239.
2003. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Shigematsu H, Lin L, Takahashi T, Nomura
M, Suzuki M, Wistuba II, Fong KM, Lee H, Toyooka S, Shimizu N, et
al: Clinical and biological features associated with epidermal
growth factor receptor gene mutations in lung cancers. J Natl
Cancer Inst. 97:339–346. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Martin P, Kelly CM and Carney D: Epidermal
growth factor receptor-targeted agents for lung cancer. Cancer
Control. 13:129–140. 2006.PubMed/NCBI
|
|
7
|
Eberhard DA, Johnson BE, Amler LC, Goddard
AD, Heldens SL, Herbst RS, Ince WL, Jänne PA, Januario T, Johnson
DH, et al: Mutations in the epidermal growth factor receptor and in
KRAS are predictive and prognostic indicators in patients with
non-small-cell lung cancer treated with chemotherapy alone and in
combination with erlotinib. J Clin Oncol. 23:5900–5909. 2005.
View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Yamamoto H, Shigematsu H, Nomura M,
Lockwood WW, Sato M, Okumura N, Soh J, Suzuki M, Wistuba II, Fong
KM, et al: PIK3CA mutations and copy number gains in human lung
cancers. Cancer Res. 68:6913–6921. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Wong DW, Leung EL, So KK, Tam IY, Sihoe
AD, Cheng LC, Ho KK, Au JS, Chung LP, Pik Wong M, et al: The
EML4-ALK fusion gene is involved in various histologic types of
lung cancers from nonsmokers with wild-type EGFR and KRAS. Cancer.
115:1723–1733. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Castro-Rivera E, Ran S, Thorpe P and Minna
JD: Semaphorin 3B (SEMA3B) induces apoptosis in lung and breast
cancer, whereas VEGF165 antagonizes this effect. Proc Natl Acad Sci
USA. 101:11432–11437. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Tomizawa Y, Sekido Y, Kondo M, Gao B,
Yokota J, Roche J, Drabkin H, Lerman MI, Gazdar AF, Minna JD, et
al: Inhibition of lung cancer cell growth and induction of
apoptosis after reex-pression of 3p21. 3 candidate tumor suppressor
gene SEMA3B. Proc Natl Acad Sci USA. 98:13954–13959. 2001.
View Article : Google Scholar
|
|
12
|
Brambilla E, Constantin B, Drabkin H and
Roche J: Semaphorin SEMA3F localization in malignant human lung and
cell lines: A suggested role in cell adhesion and cell migration.
Am J Pathol. 156:939–950. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Wachi S, Yoneda K and Wu R:
Interactome-transcriptome analysis reveals the high centrality of
genes differentially expressed in lung cancer tissues.
Bioinformatics. 21:4205–4208. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Lu TP, Tsai MH, Lee JM, Hsu CP, Chen PC,
Lin CW, Shih JY, Yang PC, Hsiao CK, Lai LC, et al: Identification
of a novel biomarker, SEMA5A, for non-small cell lung carcinoma in
nonsmoking women. Cancer Epidemiol Biomarkers Prev. 19:2590–2597.
2010. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
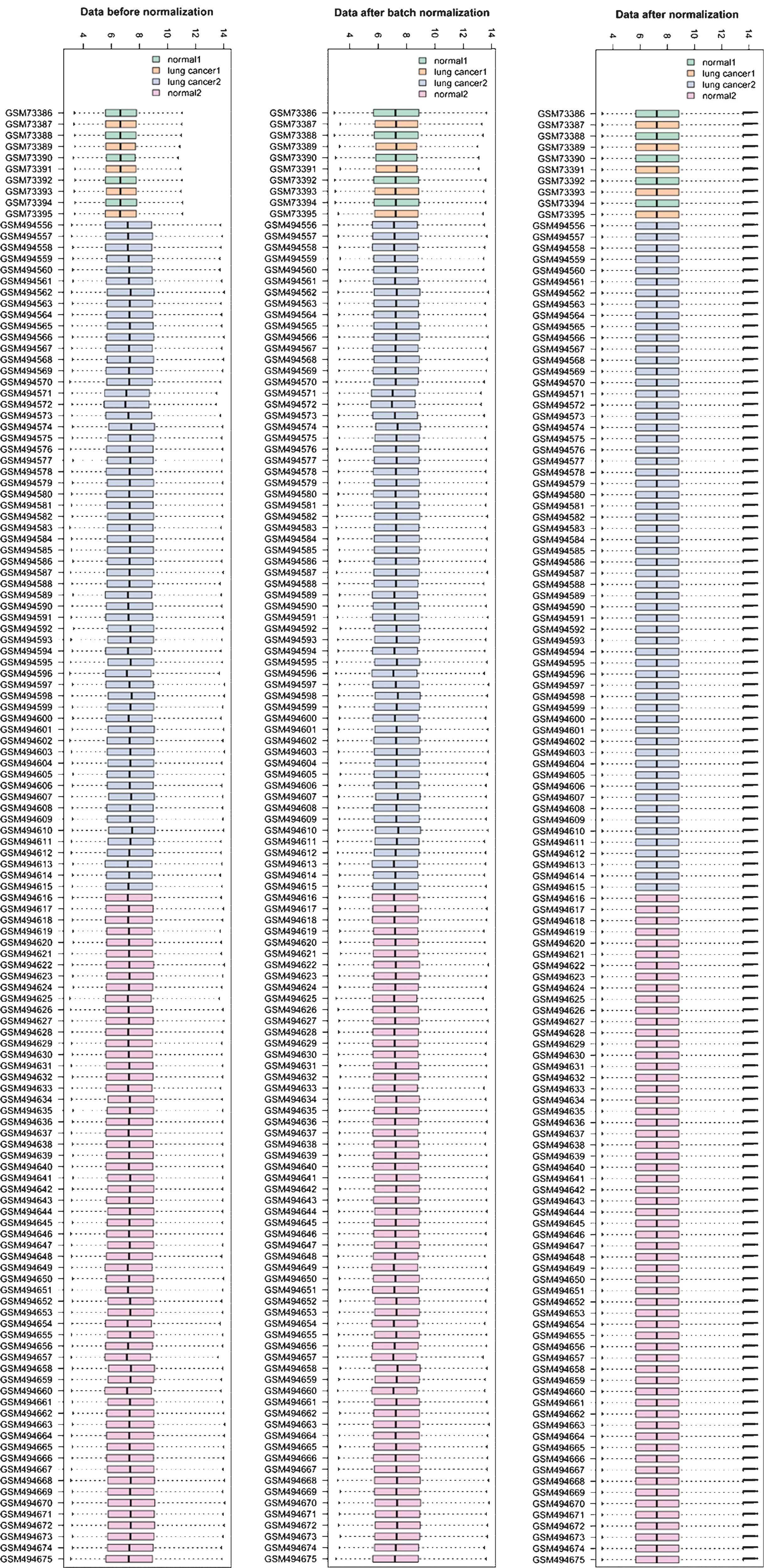
Gautier L, Cope L, Bolstad BM and Irizarry
RA: Affy-analysis of Affymetrix GeneChip data at the probe level.
Bioinformatics. 20:307–315. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Smyth GK: Linear models and empirical
bayes methods for assessing differential expression in microarray
experiments. Stat Appl Genet Mol Biol. 3:2004.
|
|
17
|
Hulsegge I, Kommadath A and Smits MA:
Globaltest and GOEAST: Two different approaches for Gene Ontology
analysis. BMC Proc. 3(Suppl 4): S102009. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Ogata H, Goto S, Sato K, Fujibuchi W, Bono
H and Kanehisa M: KEGG: Kyoto encyclopedia of genes and genomes.
Nucleic Acids Res. 27:29–34. 1999. View Article : Google Scholar
|
|
19
|
Dennis G Jr, Sherman BT, Hosack DA, Yang
J, Gao W, Lane HC and Lempicki RA: DAVID: Database for annotation,
visualization and integrated discovery. Genome Biol. 4:P32003.
View Article : Google Scholar
|
|
20
|
Franceschini A, Szklarczyk D, Frankild S,
Kuhn M, Simonovic M, Roth A, Lin J, Minguez P, Bork P, von Mering
C, et al: STRING v9.1: Protein-protein interaction networks, with
increased coverage and integration. Nucleic Acids Res.
41:D808–D815. 2013. View Article : Google Scholar :
|
|
21
|
Szklarczyk D, Franceschini A, Kuhn M,
Simonovic M, Roth A, Minguez P, Doerks T, Stark M, Muller J, Bork
P, et al: The STRING database in 2011: Functional interaction
networks of proteins, globally integrated and scored. Nucleic Acids
Res. 39:D561–D568. 2011. View Article : Google Scholar :
|
|
22
|
Kohl M, Wiese S and Warscheid B:
Cytoscape: Software for visualization and analysis of biological
networks. Methods Mol Biol. 696:291–303. 2011. View Article : Google Scholar
|
|
23
|
Bader GD and Hogue CW: An automated method
for finding molecular complexes in large protein interaction
networks. BMC Bioinformatics. 4:22003. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Maere S, Heymans K and Kuiper M: BiNGO: A
cytoscape plugin to assess overrepresentation of gene ontology
categories in biological networks. Bioinformatics. 21:3448–3449.
2005. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Wingender E, Dietze P, Karas H and Knüppel
R: TRANSFAC: A database on transcription factors and their DNA
binding sites. Nucleic Acids Res. 24:238–241. 1996. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Shannon P, Markiel A, Ozier O, Baliga NS,
Wang JT, Ramage D, Amin N, Schwikowski B and Ideker T: Cytoscape: A
software environment for integrated models of biomolecular
interaction networks. Genome Res. 13:2498–2504. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Schafer ZT and Brugge JS: IL-6 involvement
in epithelial cancers. J Clin Invest. 117:3660–3663. 2007.
View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Kishimoto T: Interleukin-6: From basic
science to medicine-40 years in immunology. Annu Rev Immunol.
23:1–21. 2005. View Article : Google Scholar
|
|
29
|
Hodge DR, Hurt EM and Farrar WL: The role
of IL-6 and STAT3 in inflammation and cancer. Eur J Cancer.
41:2502–2512. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Chung TD, Yu JJ, Kong TA, Spiotto MT and
Lin JM: Interleukin-6 activates phosphatidylinositol-3 kinase,
which inhibits apoptosis in human prostate cancer cell lines.
Prostate. 42:1–7. 2000. View Article : Google Scholar
|
|
31
|
Takizawa H, Ohtoshi T, Ohta K, Yamashita
N, Hirohata S, Hirai K, Hiramatsu K and Ito K: Growth inhibition of
human lung cancer cell lines by interleukin 6 in vitro: A possible
role in tumor growth via an autocrine mechanism. Cancer Res.
53:4175–4181. 1993.PubMed/NCBI
|
|
32
|
Knüpfer H and Preiss R: Significance of
interleukin-6 (IL-6) in breast cancer (review). Breast Cancer Res
Treat. 102:129–135. 2007. View Article : Google Scholar
|
|
33
|
Giri D, Ozen M and Ittmann M:
Interleukin-6 is an autocrine growth factor in human prostate
cancer. Am J Pathol. 159:2159–2165. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Sica A, Allavena P and Mantovani A: Cancer
related inflammation: The macrophage connection. Cancer Lett.
267:204–215. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Benveniste EN: Role of
macrophages/microglia in multiple sclerosis and experimental
allergic encephalomyelitis. J Mol Med (Berl). 75:165–173. 1997.
View Article : Google Scholar
|
|
36
|
Firestein GS: Evolving concepts of
rheumatoid arthritis. Nature. 423:356–361. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Coussens LM, Fingleton B and Matrisian LM:
Matrix metal-loproteinase inhibitors and cancer: Trials and
tribulations. Science. 295:2387–2392. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Kim JH, Lee JY, Lee KT, Lee JK, Lee KH,
Jang KT, Heo JS, Choi SH and Rhee JC: RGS16 and FosB underexpressed
in pancreatic cancer with lymph node metastasis promote tumor
progression. Tumor Biol. 31:541–548. 2010. View Article : Google Scholar
|
|
39
|
Kataoka F, Tsuda H, Arao T, Nishimura S,
Tanaka H, Nomura H, Chiyoda T, Hirasawa A, Akahane T, Nishio H, et
al: EGRI and FOSB gene expressions in cancer stroma are independent
prognostic indicators for epithelial ovarian cancer receiving
standard therapy. Gene Chromosome Cancer. 51:300–312. 2012.
View Article : Google Scholar
|
|
40
|
Yoon CH, Kim MJ, Lee H, Kim RK, Lim EJ,
Yoo KC, Lee GH, Cui YH, Oh YS, Gye MC, et al: PTTG1 oncogene
promotes tumor malignancy via epithelial to mesenchymal transition
and expansion of cancer stem cell population. J Biol Chem.
287:19516–19527. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Shibata Y, Haruki N, Kuwabara Y, Nishiwaki
T, Kato J, Shinoda N, Sato A, Kimura M, Koyama H, Toyama T, et al:
Expression of PTTG (pituitary tumor transforming gene) in
esophageal cancer. Jpn J Clin Oncol. 32:233–237. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Ghayad SE, Vendrell JA, Bieche I, Spyratos
F, Dumontet C, Treilleux I, Lidereau R and Cohen PA: Identification
of TACC1, NOV and PTTG1 as new candidate genes associated with
endocrine therapy resistance in breast cancer. J Mol Endocrinol.
42:87–103. 2009. View Article : Google Scholar
|
|
43
|
Hamid T, Malik MT and Kakar SS: Ectopic
expression of PTTG1/securin promotes tumorigenesis in human
embryonic kidney cells. Mol Cancer. 4:32005. View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Li H, Yin C, Zhang B, Sun Y, Shi L, Liu N,
Liang S, Lu S, Liu Y, Zhang J, et al: PTTG1 promotes migration and
invasion of human non-small cell lung cancer cells and is modulated
by miR-186. Carcinogenesis. 34:2145–2155. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Panguluri SK, Yeakel C and Kakar SS: PTTG:
An important target gene for ovarian cancer therapy. J Ovarian Res.
1:62008. View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Ma S, Guan XY, Beh PS, Wong KY, Chan YP,
Yuen HF, Vielkind J and Chan KW: The significance of LMO2
expression in the progression of prostate cancer. J Pathol.
211:278–285. 2007. View Article : Google Scholar
|
|
47
|
Nakata K, Ohuchida K, Nagai E, Hayashi A,
Miyasaka Y, Kayashima T, Yu J, Aishima S, Oda Y, Mizumoto K, et al:
LMO2 is a novel predictive marker for a better prognosis in
pancreatic cancer. Neoplasia. 11:712–719. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Yamada Y, Pannell R, Forster A and
Rabbitts TH: The LIM-domain protein Lmo2 is a key regulator of
tumour angiogenesis: A new anti-angiogenesis drug target. Oncogene.
21:1309–1315. 2002. View Article : Google Scholar : PubMed/NCBI
|