|
1
|
Sung H, Ferlay J, Siegel RL, Laversanne M,
Soerjomataram I, Jemal A and Bray F: Global cancer statistics 2020:
GLOBOCAN estimates of incidence and mortality worldwide for 36
cancers in 185 countries. CA Cancer J Clin. 71:209–249. 2021.
View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Singal AG, Lampertico P and Nahon P:
Epidemiology and surveillance for hepatocellular carcinoma: New
trends. J Hepatol. 72:250–261. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Faivre S, Rimassa L and Finn RS: Molecular
therapies for HCC: Looking outside the box. J Hepatol. 72:342–352.
2020. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Sangro B, Sarobe P, Hervás-Stubbs S and
Melero I: Advances in immunotherapy for hepatocellular carcinoma.
Nat Rev Gastroenterol Hepatol. 18:525–543. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
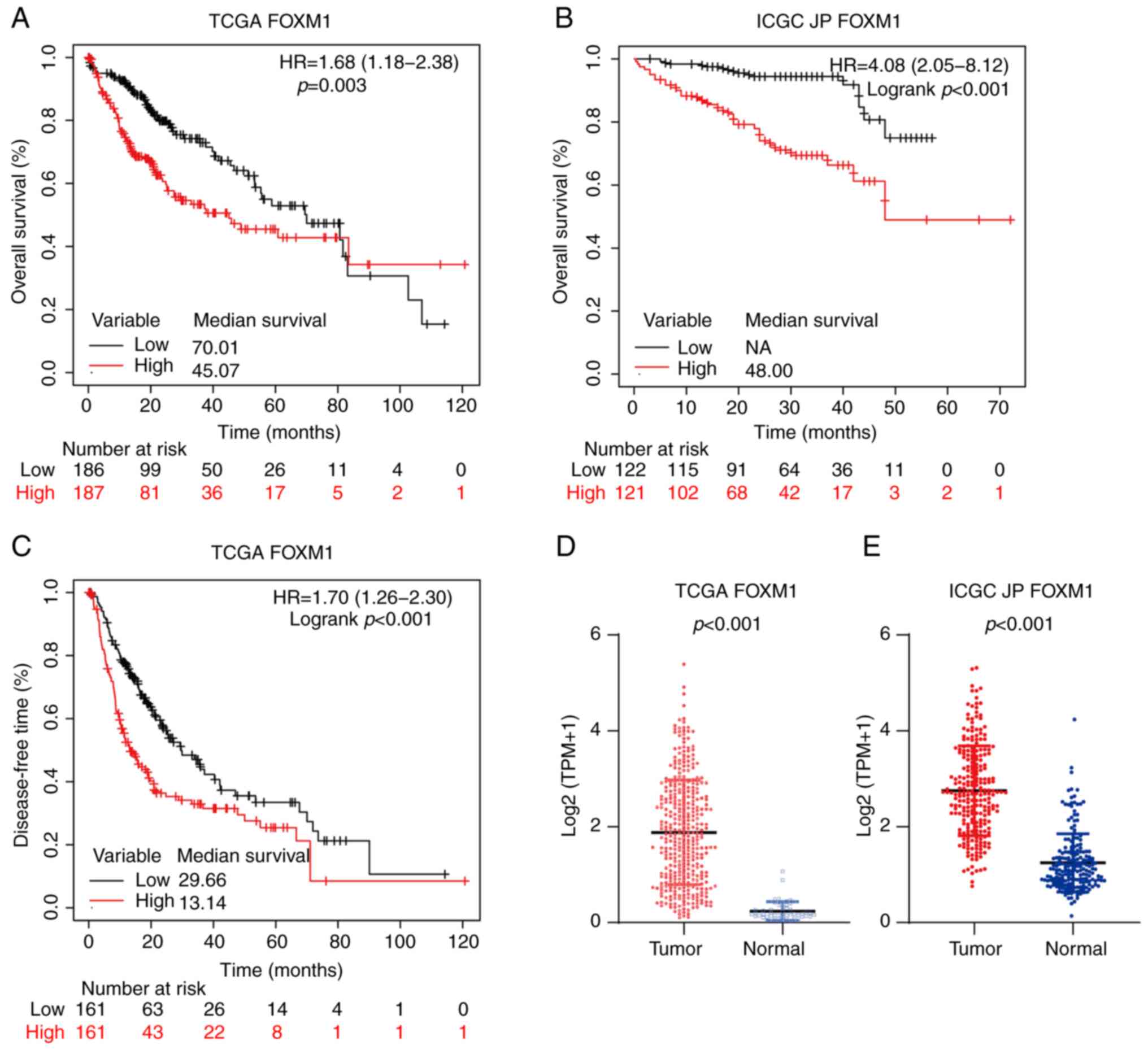
Song BN and Chu IS: A gene expression
signature of FOXM1 predicts the prognosis of hepatocellular
carcinoma. Exp Mol Med. 50:e4182018. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Teng L, Wang K, Liu Y, Ma Y, Chen W and Bi
L: Based on integrated bioinformatics analysis identification of
biomarkers in hepatocellular carcinoma patients from different
regions. Biomed Res Int. 2019:17423412019. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Agarwal R, Narayan J, Bhattacharyya A,
Saraswat M and Tomar AK: Gene expression profiling, pathway
analysis and subtype classification reveal molecular heterogeneity
in hepatocellular carcinoma and suggest subtype specific
therapeutic targets. Cancer Genet. 216–217. 37–51. 2017.PubMed/NCBI
|
|
8
|
Sun W, Su Q, Cao X, Shang B, Chen A, Yin H
and Liu B: High expression of polo-like kinase 1 is associated with
early development of hepatocellular carcinoma. Int J Genomics.
2014:3121302014. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Seyedabadi S, Saidijam M, Najafi R,
Mousavi-Bahar SH, Jafari M, MohammadGanji S and Mahdavinezhad A:
Assessment of CEP55, PLK1 and FOXM1 expression in patients with
bladder cancer in comparison with healthy individuals. Cancer
Invest. 36:407–414. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Dibb M, Han N, Choudhury J, Hayes S,
Valentine H, West C, Sharrocks AD and Ang YS: FOXM1 and polo-like
kinase 1 are co-ordinately overexpressed in patients with gastric
adenocarcinomas. BMC Res Notes. 8:6762015. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Wang R, Song Y, Xu X, Wu Q and Liu C: The
expression of Nek7, FoxM1, and Plk1 in gallbladder cancer and their
relationships to clinicopathologic features and survival. Clin
Transl Oncol. 15:626–632. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Lam EW, Brosens JJ, Gomes AR and Koo CY:
Forkhead box proteins: Tuning forks for transcriptional harmony.
Nat Rev Cancer. 13:482–495. 2013. View
Article : Google Scholar : PubMed/NCBI
|
|
13
|
Wang IC, Chen YJ, Hughes D, Petrovic V,
Major ML, Park HJ, Tan Y, Ackerson T and Costa RH: Forkhead box M1
regulates the transcriptional network of genes essential for
mitotic progression and genes encoding the SCF (Skp2-Cks1)
ubiquitin ligase. Mol Cell Biol. 25:10875–10894. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Laoukili J, Kooistra MR, Brás A, Kauw J,
Kerkhoven RM, Morrison A, Clevers H and Medema RH: FoxM1 is
required for execution of the mitotic programme and chromosome
stability. Nat Cell Biol. 7:126–136. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Chen YJ, Dominguez-Brauer C, Wang Z, Asara
JM, Costa RH, Tyner AL, Lau LF and Raychaudhuri P: A conserved
phosphorylation site within the forkhead domain of FoxM1B is
required for its activation by cyclin-CDK1. J Biol Chem.
284:30695–30707. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Fu Z, Malureanu L, Huang J, Wang W, Li H,
van Deursen JM, Tindall DJ and Chen J: Plk1-dependent
phosphorylation of FoxM1 regulates a transcriptional programme
required for mitotic progression. Nat Cell Biol. 10:1076–1082.
2008. View
Article : Google Scholar : PubMed/NCBI
|
|
17
|
Zhang J, Yuan C, Wu J, Elsayed Z and Fu Z:
Polo-like kinase 1-mediated phosphorylation of forkhead box protein
M1b antagonizes its SUMOylation and facilitates its mitotic
function. J Biol Chem. 290:3708–3719. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Pilarsky C, Wenzig M, Specht T, Saeger HD
and Grützmann R: Identification and validation of commonly
overexpressed genes in solid tumors by comparison of microarray
data. Neoplasia. 6:744–750. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Kim IM, Ackerson T, Ramakrishna S,
Tretiakova M, Wang IC, Kalin TV, Major ML, Gusarova GA, Yoder HM,
Costa RH and Kalinichenko VV: The Forkhead Box m1 transcription
factor stimulates the proliferation of tumor cells during
development of lung cancer. Cancer Res. 66:2153–2161. 2006.
View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Wang Z, Park HJ, Carr JR, Chen YJ, Zheng
Y, Li J, Tyner AL, Costa RH, Bagchi S and Raychaudhuri P: FoxM1 in
tumorigenicity of the neuroblastoma cells and renewal of the neural
progenitors. Cancer Res. 71:4292–4302. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Hu G, Yan Z, Zhang C, Cheng M, Yan Y, Wang
Y, Deng L, Lu Q and Luo S: FOXM1 promotes hepatocellular carcinoma
progression by regulating KIF4A expression. J Exp Clin Cancer Res.
38:1882019. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Chai N, Xie HH, Yin JP, Sa KD, Guo Y, Wang
M, Liu J, Zhang XF, Zhang X, Yin H, et al: FOXM1 promotes
proliferation in human hepatocellular carcinoma cells by
transcriptional activation of CCNB1. Biochem Biophys Res Commun.
500:924–929. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Egawa M, Yoshida Y, Ogura S, Kurahashi T,
Kizu T, Furuta K, Kamada Y, Chatani N, Hamano M, Kiso S, et al:
Increased expression of Forkhead box M1 transcription factor is
associated with clinicopathological features and confers a poor
prognosis in human hepatocellular carcinoma. Hepatol Res.
47:1196–1205. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Sun HC, Li M, Lu JL, Yan DW, Zhou CZ, Fan
JW, Qin XB, Tang HM and Peng ZH: Overexpression of Forkhead box M1
protein associates with aggressive tumor features and poor
prognosis of hepatocellular carcinoma. Oncol Rep. 25:1533–1539.
2011.PubMed/NCBI
|
|
25
|
Combes G, Alharbi I, Braga LG and Elowe S:
Playing polo during mitosis: PLK1 takes the lead. Oncogene.
36:4819–4827. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Colicino EG and Hehnly H: Regulating a key
mitotic regulator, polo-like kinase 1 (PLK1). Cytoskeleton
(Hoboken). 75:481–494. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Barr FA, Silljé HH and Nigg EA: Polo-like
kinases and the orchestration of cell division. Nat Rev Mol Cell
Biol. 5:429–440. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Xie S, Xie B, Lee MY and Dai W: Regulation
of cell cycle checkpoints by polo-like kinases. Oncogene.
24:277–286. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
He Z, Wu J, Dang H, Lin H, Zheng H and
Zhong D: Polo-like kinase 1 contributes to the tumorigenicity of
BEL-7402 hepatoma cells via regulation of Survivin expression.
Cancer Lett. 303:92–98. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Park JS, Sohn HJ, Park GS, Chung YJ and
Kim TG: Induction of antitumor immunity using dendritic cells
electroporated with Polo-like kinase 1 (Plk1) mRNA in murine tumor
models. Cancer Sci. 102:1448–1454. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Jang HR, Shin SB, Kim CH, Won JY, Xu R,
Kim DE and Yim H: PLK1/vimentin signaling facilitates immune escape
by recruiting Smad2/3 to PD-L1 promoter in metastatic lung
adenocarcinoma. Cell Death Differ. 28:2745–2764. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Fristrup N, Ulhøi BP, Birkenkamp-Demtröder
K, Mansilla F, Sanchez-Carbayo M, Segersten U, Malmström PU,
Hartmann A, Palou J, Alvarez-Múgica M, et al: Cathepsin E, maspin,
Plk1, and survivin are promising prognostic protein markers for
progression in non-muscle invasive bladder cancer. Am J Pathol.
180:1824–1834. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Takai N, Hamanaka R, Yoshimatsu J and
Miyakawa I: Polo-like kinases (Plks) and cancer. Oncogene.
24:287–291. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Cheng L, Wang C and Jing J: Polo-like
kinase 1 as a potential therapeutic target for osteosarcoma. Curr
Pharm Des. 21:1347–1350. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
He ZL, Zheng H, Lin H, Miao XY and Zhong
DW: Overexpression of polo-like kinase1 predicts a poor prognosis
in hepatocellular carcinoma patients. World J Gastroenterol.
15:4177–4182. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Tian L, Yao K, Liu K, Han B, Dong H, Zhao
W, Jiang W, Qiu F, Qu L, Wu Z, et al: PLK1/NF-κB feedforward
circuit antagonizes the mono-ADP-ribosyltransferase activity of
PARP10 and facilitates HCC progression. Oncogene. 39:3145–3162.
2020. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Gumireddy K, Reddy MV, Cosenza SC,
Boominathan R, Baker SJ, Papathi N, Jiang J, Holland J and Reddy
EP: ON01910, a non-ATP-competitive small molecule inhibitor of
Plk1, is a potent anticancer agent. Cancer Cell. 7:275–286. 2005.
View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Steegmaier M, Hoffmann M, Baum A, Lénárt
P, Petronczki M, Krssák M, Gürtler U, Garin-Chesa P, Lieb S, Quant
J, et al: BI 2536, a potent and selective inhibitor of polo-like
kinase 1, inhibits tumor growth in vivo. Curr Biol. 17:316–322.
2007. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Rudolph D, Steegmaier M, Hoffmann M,
Grauert M, Baum A, Quant J, Haslinger C, Garin-Chesa P and Adolf
GR: BI 6727, a Polo-like kinase inhibitor with improved
pharmacokinetic profile and broad antitumor activity. Clin Cancer
Res. 15:3094–3102. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Schmittgen TD and Livak KJ: Analyzing
real-time PCR data by the comparative C(T) method. Nat Protoc.
3:1101–1108. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Qiu P and Sheng J: A two-stage procedure
for comparing hazard rate functions. J R Stat Soc Series B (Stat
Methodol). 70:191–208. 2008.PubMed/NCBI
|
|
42
|
Li H, Han D, Hou Y, Chen H and Chen Z:
Statistical inference methods for two crossing survival curves: A
comparison of methods. PLoS One. 10:e01167742015. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Duffy AG, Ulahannan SV, Makorova-Rusher O,
Rahma O, Wedemeyer H, Pratt D, Davis JL, Hughes MS, Heller T,
ElGindi M, et al: Tremelimumab in combination with ablation in
patients with advanced hepatocellular carcinoma. J Hepatol.
66:545–551. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Finn RS, Qin S, Ikeda M, Galle PR, Ducreux
M, Kim TY, Kudo M, Breder V, Merle P, Kaseb AO, et al: Atezolizumab
plus bevacizumab in unresectable hepatocellular carcinoma. N Engl J
Med. 382:1894–1905. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Ren Z, Xu J, Bai Y, Xu A, Cang S, Du C, Li
Q, Lu Y, Chen Y, Guo Y, et al: Sintilimab plus a bevacizumab
biosimilar (IBI305) versus sorafenib in unresectable hepatocellular
carcinoma (ORIENT-32): A randomised, open-label, phase 2–3 study.
Lancet Oncol. 22:977–990. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Cariani E and Missale G: Immune landscape
of hepatocellular carcinoma microenvironment: Implications for
prognosis and therapeutic applications. Liver Int. 39:1608–1621.
2019. View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Oura K, Morishita A, Tani J and Masaki T:
Tumor Immune microenvironment and immunosuppressive therapy in
hepatocellular carcinoma: A review. Int J Mol Sci. 22:58012021.
View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Thorsson V, Gibbs DL, Brown SD, Wolf D,
Bortone DS, Ou Yang TH, Porta-Pardo E, Gao GF, Plaisier CL, Eddy
JA, et al: The immune landscape of cancer. Immunity.
48:812–830.e14. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
49
|
Newman AM, Liu CL, Green MR, Gentles AJ,
Feng W, Xu Y, Hoang CD, Diehn M and Alizadeh AA: Robust enumeration
of cell subsets from tissue expression profiles. Nat Methods.
12:453–457. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
50
|
Kondĕlková K, Vokurková D, Krejsek J,
Borská L, Fiala Z and Ctirad A: Regulatory T cells (TREG) and their
roles in immune system with respect to immunopathological
disorders. Acta Medica (Hradec Kralove). 53:73–77. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
51
|
Hatzioannou A, Boumpas A, Papadopoulou M,
Papafragkos I, Varveri A, Alissafi T and Verginis P: Regulatory T
cells in autoimmunity and cancer: A duplicitous lifestyle. Front
Immunol. 12:7319472021. View Article : Google Scholar : PubMed/NCBI
|
|
52
|
Matthews HK, Bertoli C and de Bruin RAM:
Cell cycle control in cancer. Nat Rev Mol Cell Biol. 23:74–88.
2022. View Article : Google Scholar : PubMed/NCBI
|
|
53
|
Kashyap D, Garg VK, Sandberg EN, Goel N
and Bishayee A: Oncogenic and tumor suppressive components of the
cell cycle in breast cancer progression and prognosis.
Pharmaceutics. 13:5692021. View Article : Google Scholar : PubMed/NCBI
|
|
54
|
Mross K, Dittrich C, Aulitzky WE,
Strumberg D, Schutte J, Schmid RM, Hollerbach S, Merger M, Munzert
G, Fleischer F and Scheulen ME: A randomised phase II trial of the
Polo-like kinase inhibitor BI 2536 in chemo-naïve patients with
unresectable exocrine adenocarcinoma of the pancreas-a study within
the Central European society anticancer drug research (CESAR)
collaborative network. Br J Cancer. 107:280–286. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
55
|
Garcia-Manero G, Fenaux P, Al-Kali A, Baer
MR, Sekeres MA, Roboz GJ, Gaidano G, Scott BL, Greenberg P,
Platzbecker U, et al: Rigosertib versus best supportive care for
patients with high-risk myelodysplastic syndromes after failure of
hypomethylating drugs (ONTIME): A randomised, controlled, phase 3
trial. Lancet Oncol. 17:496–508. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
56
|
Pujade-Lauraine E, Selle F, Weber B,
Ray-Coquard IL, Vergote I, Sufliarsky J, Del Campo JM, Lortholary
A, Lesoin A, Follana P, et al: Volasertib versus chemotherapy in
platinum-resistant or -refractory ovarian cancer: A randomized
phase II groupe des investigateurs nationaux pour l'Etude des
cancers de l'Ovaire study. J Clin Oncol. 34:706–713. 2016.
View Article : Google Scholar : PubMed/NCBI
|