Gastric cancer (GC)
Cancer is a group of genetic diseases characterized
by the uncontrolled proliferation of heterogeneous cell populations
with the ability to invade tissues locally and remotely from the
site of origin (metastasis), responding to stimuli from the
adjacent microenvironment and from the host organism. During
oncogenesis, cancer not only escapes the host regulatory
mechanisms, but also gains the ability to affect local and systemic
homeostasis (1).
Cancer is the second leading cause of death
worldwide; in 2022, 9,743,832 deaths were attributed to this
disease, and it is estimated that the number of prevalent cases in
5 years will be 53,504,187 (2).
Worldwide, GC is estimated to be the fifth most common cancer type
in both sexes, ranking sixth for new cases, with 640,850 cases per
year, and fourth in terms of mortality rate (GloboCan, 2020)
(3).
The etiology of GC is complex, primarily due to
genetic alterations and a set of factors (diet, lifestyle, genetics
and socioeconomic factors). Significantly, 80% of cases are
associated with infection with Helicobacter pylori (4,5).
Of note, <3% of GC cases are attributed to
heredity causes, and these types of GC cancer cases include
Hereditary GC (HGC), proximal polyposis of the stomach, and
hereditary non-polyposis colorectal cancer (5). HGC is the best-known familial GC and
is characterized by the loss of the CDH1 gene. Hereditary Diffuse
GC (HDGC) is rare, with an incidence rate ranging from 0.3–3.1% in
Korea and Japan. However, excluding the small portion of cases of
familial GC syndrome, the risk of GC in those with a family history
is three times higher vs. those with no history, which is higher
than that for other adult solid cancer cases except ovarian cancer
(6). Although family history is a
systematically reported risk factor in GC, the molecular basis for
familial clustering is unclear. Family members have shared exposure
to carcinogens, (such as cigarette smoke and alcohol consumption),
along with similar levels of hygiene, dietary habits (salty, spicy,
and smoked foods), bacterial virulence, including Helicobacter
pylori CagA+ and also genetic susceptibility
(7).
Invasive GC is preceded by a long precancerous
period, which can last for decades and thus provides ample
opportunities to detect and treat precancerous lesions. When these
lesions progress to an advanced stage, periodic endoscopic
follow-ups should be performed to identify the lesions before they
become invasive (8,9). Anatomic demarcations, histological
differences, or both can be used to distinguish benign lesions from
invasive lesions. Most relevant is the distinction between
adenocarcinomas that arise from the cardia (the part of the stomach
closest to the esophagus, cardia-GC) and other parts of the stomach
(non-cardia-GC) (10). Currently,
there are different histological classifications of GC, the most
commonly used are the Lauren (11),
Nakamura et al (12), Ming
(8), Goseki et al (13), and the World Health Organization
(WHO) classification systems (14).
GC can be divided into diffuse and intestinal GC
based on its histological appearance as well as cardia and
non-cardia-GC according to location. The epidemiological and
molecular characteristics of GC differ according to the
histological type and location of the tumor (6).
Intestinal-type GC (IGC) has well-defined structures
or ductal chords surrounded by a zone of desmoplastic stroma. It
forms adhesions or fibrous joint tissue within the tumor, with
mixed inflammatory infiltration (15). IGC is observed more frequently in
older adult patients and follows Correa's precancerous chain, which
includes the following states: Atrophic gastritis, intestinal
metaplasia and dysplasia (16).
Tumor cells often have nuclei that are polymorphic and isochromatic
with a coarse chromatin pattern; mitotic figures are thus easily
detected. Intestinal-type carcinomas must be well or moderately
differentiated, and diffuse-type adenocarcinomas have solitary or
small groups of tumor cells without forming glandular structures
(15).
Furthermore, Diffuse-type GC (DGC) occurs more
frequently in younger patients, and there are no associations with
atrophic gastritis or intestinal metaplasia (16). Clear cytoplasmic vacuoles may
sometimes be observed. These cells that contain mucus push the
nucleus to the periphery of the cell (signet ring cell carcinoma).
The stroma formed is usually extensive, making it difficult to
identify separate tumor cells on standard hematoxylin and eosin
stains; additional keratin staining reveals the true extent of the
tumor (15).
The pathogenicity of IGC has been well characterized
and studied. However, diffuse GC (DGC) remains undefined, is
considered genetically determined, and is less associated with
environmental factors and the inflammatory cascade. Additionally, a
minor proportion of DGC cases (1–3%) are inherently linked and
associated with germline alterations in cell physiology, known as
HDGC (7).
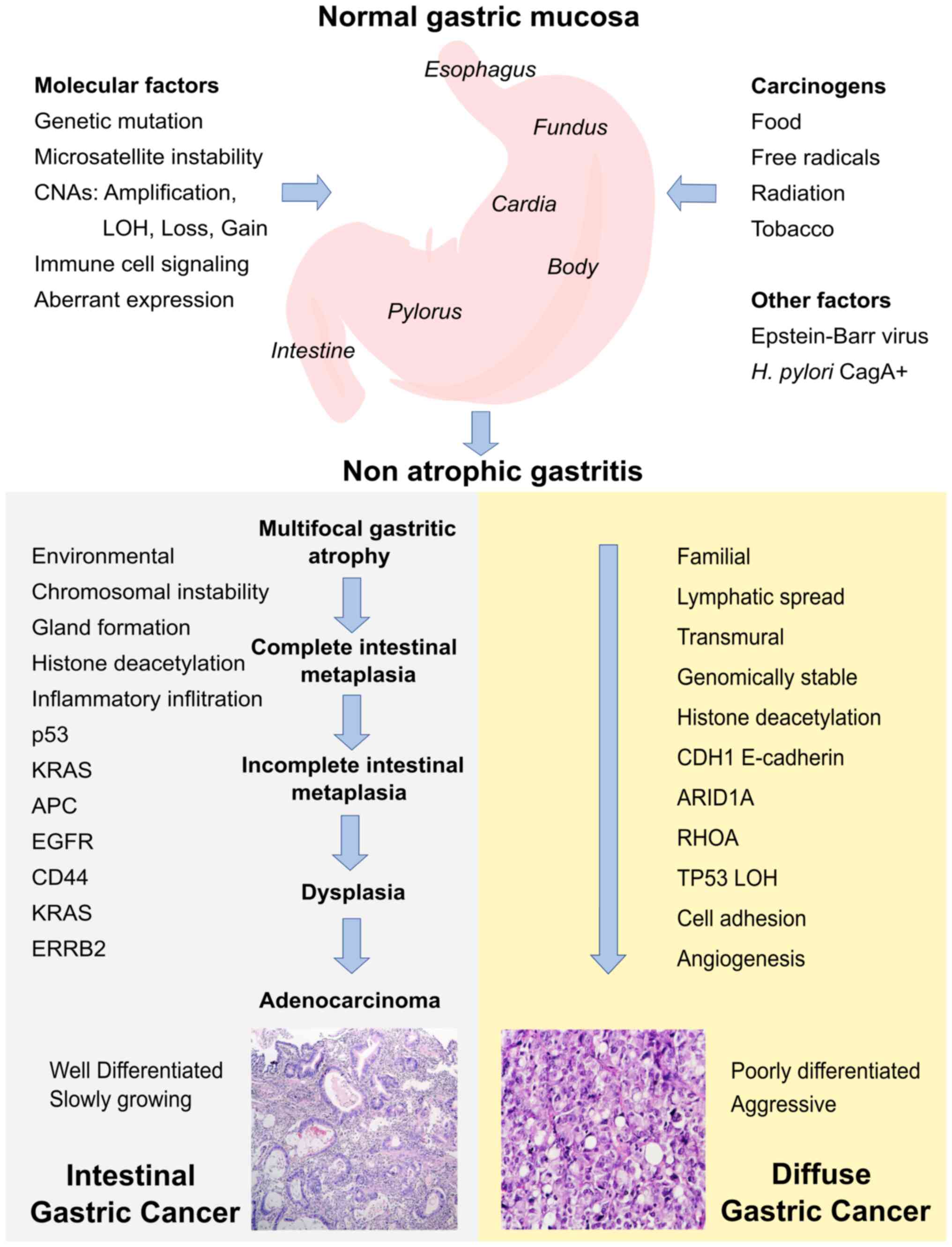
The stages of the precancerous cascade illustrated
in Fig. 1 have been well
characterized from the histopathological point of view. It is
postulated that the progression from one stage to the next is
determined by etiological factors linked to the inflammatory
process and decades of progression (9). The stages of the precancerous cascade
are non-atrophic gastritis, multifocal atrophic gastritis, complete
intestinal metaplasia, incomplete intestinal metaplasia, dysplasia
and adenocarcinoma (9).
Among the molecular pathogenesis, chromosome
instability is involved (aneuploid, translocation, amplification,
deletions and loss of heterozygosity), fusion genes,
microsatellites instability (hypermethylation of promoters of DNA
repair genes) and changes in gene expression profile (4,5,17).
Chromosomes and copy number alterations
(CNA)
In humans, the genome consists of 23 pairs of
chromosomes [22 autosomes (44 chromosomes and one pair of sex
chromosomes (XX or XY), for a total of 46 chromosomes] located in
the nucleus, as well as a small chromosome in each mitochondrion.
Each human chromosome has a short arm (‘p’ for ‘petit’) and a long
arm (‘q’ for ‘queue’), separated by a centromere. The ends of
chromosomes are called telomeres. Each chromosome arm is divided
into regions, or cytogenetic bands, that can be observed using a
microscope and special stains. The cytogenetic bands are labeled
p1, p2, p3, q1, q2, q3, counting from the centromere to the
telomeres. At higher resolutions, sub-bands can be identified
within the bands. The sub-bands are also numbered from the
centromere out toward the telomere. For example, the cytogenetic
map location of the MYC gene is 8q24.21, which indicates it is
located on chromosome 8, q arm, band 24, sub-band 21 and
sub-sub-band 2. The ends of the chromosomes are labeled ptel and
qtel. For example, the notation 8qtel refers to the end of the long
arm of chromosome 8 (18).
CNA represent a genetic variation class involving
cumulative somatic variations. CNA are defined as non-inherited
genetic alterations in somatic cells (19). These unbalanced structural variants
usually contain gains or losses. Their interpretation and the CNA
report continue to be a topic of interest in health and disease and
have an essential role in GC (19,20).
The majority of gastric adenocarcinomas, similar to numerous other
types of solid tumors, exhibit defects in the maintenance of genome
stability, resulting in DNA CNA that may be analyzed using
comparative genomic hybridization (21) and sequencing (22). Based on the aforementioned, it is a
widespread phenomenon among humans, and several studies have
focused on understanding these genomic alterations that are
responsible for cancer. They may be used for its diagnosis and
prognosis (23).
In the present review, CNA research on samples of
patients with GC is discussed. In a previous investigation, it was
noted that the 8q24 cytoband exhibited alterations (24), and thus this cytoband's role in this
neoplasia is reviewed below. The complete sequence of chromosome 8
has 145,138,636 bases, the NCBI RefSeq access key corresponding to
the GRCh38.p14 version of the human genome is NC_000008.11, the
chromosome has 34 cytobands, 2,388 genes, and 4,651 proteins.
Chromosome 8 has 103 genes related to cancer; 22 of these genes are
in cytoband 8q24, and seven of these genes are reported to be
associated with stomach cancer (25,26)
(Table I; 27–49).
 | Table I.Gastric cancer-related genes. |
Table I.
Gastric cancer-related genes.
|
| Gene | ID | Chr | Summary | (Refs.) |
|---|
| 1 | ADGRB1a | 575 | 8q24.3 | Angiogenesis is
controlled by a local balance between stimulators and inhibitors of
new vessel growth and is suppressed under normal physiologic
conditions. Angiogenesis is essential for the growth and metastasis
of solid tumors. | (27) |
| 2 | AGO2 | 27161 | 8q24.3 | This gene encodes a
member of the Argonaute family of proteins which play a role in RNA
interference. The encoded protein is highly basic and contains a
PAZ domain and a PIWI domain. It may interact with Dicer1 and play
a role in short-interfering-RNA-mediated gene silencing. Multiple
transcript variants encoding different isoforms have been found for
this gene. | (28) |
| 3 | CYP11B1 | 1584 | 8q24.3 | This gene encodes a
member of the cytochrome P450 superfamily of enzymes. The
cytochrome P450 proteins are monooxygenases that catalyze numerous
reactions in drug metabolism and synthesize cholesterol, steroids,
and other lipids. | (29,30) |
| 4 | CYP11B2 | 1585 | 8q24.3 | This gene encodes a
member of the cytochrome P450 superfamily of enzymes. The
cytochrome P450 proteins are monooxygenases that catalyze numerous
reactions in drug metabolism and synthesize cholesterol, steroids,
and other lipids. | (31) |
| 5 | HSF1 | 3297 | 8q24.3 | The product of this
gene is a transcription factor rapidly induced after temperature
stress and binds heat shock promoter elements (HSE). This protein
plays a role in the regulation of lifespan. | (32) |
| 6 | RECQL4 | 9401 | 8q24.3 | The protein encoded
by this gene is a DNA helicase that belongs to the RecQ helicase
family. | (33) |
| 7 | SCRIB | 23513 | 8q24.3 | The mammalian
protein is involved in tumor suppression pathways. As a scaffold
protein involved in cell polarization processes, this protein binds
to numerous other proteins. | (34) |
| 8 | PTK2a | 5747 | 8q24.3 | This gene encodes a
cytoplasmic protein tyrosine kinase, which is concentrated in the
focal adhesions between cells growing in the presence of
extracellular matrix constituents. | (35) |
| 9 | PTP4A3a | 11156 | 8q24.3 | This gene encodes a
member of the protein-tyrosine phosphatase family. Protein tyrosine
phosphatases are cell signaling molecules that play regulatory
roles in various cellular processes. | (36) |
| 10 | PSCAa | 8000 | 8q24.3 | This gene encodes a
glycosylphosphatidylinositol-anchored cell membrane glycoprotein.
This gene is up- regulated in numerous prostate cancers and is also
detected in bladder and pancreas cancers. This gene includes a
polymorphism that results in an upstream start codon in some
individuals; this polymorphism is thought to be associated with a
risk for certain gastric and bladder cancers-alternative splicing
results in multiple transcript variants. | (37) |
| 11 | EXT1 | 2131 | 8q24.11 | This gene encodes
an endoplasmic reticulum-resident type II transmembrane
glycosyltransferase involved in the chain elongation step of
heparan sulfate biosynthesis. | (38) |
| 12 | RAD21 | 5885 | 8q24.11 | This protein is a
nuclear phospho-protein, which becomes hyperphosphorylated in the
cell cycle M phase. The highly regulated association of this
protein with mitotic chromatin, specifically at the centromere
region, suggests its role in sister chromatid cohesion in mitotic
cells. | (39) |
| 13 | CCN3 | 4856 | 8q24.12 | The protein encoded
by this gene is a small, secreted cysteine-rich (NOV) protein and a
member of the CCN family of regulatory proteins. CNN family
proteins associate with the extracellular matrix and play an
important role in cardiovascular and skeletal development,
fibrosis, and cancer development. | (40) |
| 14 | TNFRSF11B | 4982 | 8q24.12 | The protein encoded
by this gene is a member of the TNF-receptor superfamily. | (41) |
| 15 | HAS2 | 3037 | 8q24.13 | Hyaluronan or
hyaluronic acid (HA) is a high molecular weight unbranched
polysaccharide synthesized by various organisms, from bacteria to
mammals, and is a constituent of the extracellular matrix. Changes
in the serum concentration of HA are associated with inflammatory
and degenerative arthropathies such as rheumatoid arthritis. | (42) |
| 16 | MTSS1a | 9788 | 8q24.13 | Predicted to be
involved in the cellular response to fluid shear stress, negative
regulation of epithelial cell proliferation, and urogenital system
development. Predicted to act upstream of or within several
processes, including actin filament polymerization; adherents
junction maintenance; and magnesium ion homeostasis, which are in
the actin cytoskeleton. | (43) |
| 17 | RNF139 | 11236 | 8q24.13 | This gene was found
to be interrupted by a t(3:8) translocation in a family with
hereditary renal and non-medullary thyroid cancer. Studies of the
Drosophila counterpart suggested that this protein may interact
with tumor suppressor protein VHL and COPS5/JAB1, a protein
responsible for the degradation of tumor suppressor
CDKN1B/P27KIP. | (44) |
| 18 | CCDC26 | 137196 | 8q24.21 | No data | (45) |
| 19 | MYCa | 4609 | 8q24.21 | This gene is a
proto-oncogene and encodes a nuclear phosphoprotein that plays a
role in cell cycle progression, apoptosis, and cellular
transformation. | (46) |
| 20 | PVT1a | 5820 | 8q24.21 | This gene
represents a long non-coding RNA locus identified as a candidate
oncogene. Increased copy number and overexpression of this gene are
associated with numerous cancers, including breast and ovarian
cancers, acute myeloid leukemia, and Hodgkin lymphoma. | (47) |
| 21 | CCN4 | 8840 | 8q24.22 | This gene encodes a
WNT1 inducible signaling pathway (WISP) protein (WISP1) subfamily
member, which belongs to the connective tissue growth factor (CTGF)
family. It is expressed at a high level in fibroblast cells and
overexpressed in colon tumors. It also attenuates p53-mediated
apoptosis in response to DNA damage by activating the Akt
kinase. | (48) |
| 22 | NDRG1 | 10397 | 8q24.22 | The protein encoded
by this gene is a cytoplasmic protein involved in stress responses,
hormone responses, cell growth, and differentiation. The encoded
protein is necessary for p53-mediated caspase activation and
apoptosis. | (49) |
8q24 cytoband genes and single nucleotide
polymorphisms (SNPs)
Alterations in the 8q24 cytoband have been
associated with multiple conditions, including cancer; several
mechanisms have been proposed, including chromosomal translocations
(50), viral integration (51), identification of nucleotide
variations (51–53), and variations in the number of
copies (24). The 8q24.21 cytoband
is one of the most studied due to its association with various
types of cancer or complex diseases; this locus has very few
protein-coding genes and is rich in long non-coding RNAs (lncRNAs)
(54). The latter play a range of
roles in transcription and translation; however, a number of the
identified variants in these regions have been insufficiently
studied. The study of this locus has presented several difficulties
since most of the lncRNAs in 8q24.21 are not evolutionarily
conserved, and there are no mouse orthologs (55).
Currently, repositories such as ENSEMBL (56) and UCSC (57) allow the consultation of different
cytobands of the human genome in their different versions.
Additionally, BioMart is a tool that provides an interface for
accessing database collections, which allows annotating or
obtaining genomics, genetics and proteomics records, amongst other
tools. With the BioMart data mining tool (58), an analysis of 8q24 (GRCh38.p13) was
performed; it has a length of 138.9 megabase pairs (Mbp) and seven
segments, of which q24.3 is the longest (6.2 Mbp) and the one with
the most significant number of coding genes, pseudogenes and small
RNAs, and q24.11 is the smallest (1.6 Mbp) (Table II).
 | Table II.Description of the 8q24 cytoband.
Information obtained from BioMart. |
Table II.
Description of the 8q24 cytoband.
Information obtained from BioMart.
|
| Cytobands |
|---|
|
|
|
|---|
| Features | q24.11 | q24.12 | q24.13 | q24.21 | q24.22 | q24.23 | q24.3 |
|---|
| Start Chr | 116700000 | 118300000 | 121500000 | 126300000 | 130400000 | 135400000 | 138900000 |
| End Chr | 118300000 | 121500000 | 126300000 | 130400000 | 135400000 | 138900000 | 145138636 |
| Length (Mbp) | 1.6 | 3.2 | 4.8 | 4.1 | 5.0 | 3.5 | 6.2 |
| Protein coding | 7 | 13 | 26 | 5 | 16 | 2 | 103 |
| rRNA
pseudogene | 0 | 1 | 0 | 0 | 0 | 0 | 1 |
| lncRNA | 4 | 11 | 37 | 29 | 26 | 8 | 68 |
| miRNA | 1 | 1 | 6 | 7 | 3 | 0 | 16 |
| snoRNA | 1 | 1 | 2 | 2 | 2 | 0 | 2 |
| snRNA | 0 | 1 | 5 | 6 | 0 | 2 | 1 |
| miscRNA | 2 | 2 | 2 | 3 | 4 | 0 | 4 |
| Processed
pseudogene | 1 | 7 | 15 | 6 | 13 | 2 | 10 |
| Unprocessed
pseudogene | 1 | 0 | 0 | 0 | 1 | 1 | 2 |
| Transcribe
processed pseudogene | 0 | 0 | 1 | 1 | 0 | 0 | 0 |
| Polymorphic
pseudogene | 0 | 0 | 0 | 0 | 0 | 0 | 1 |
| Transcribe unitary
pseudogene | 0 | 0 | 0 | 0 | 0 | 0 | 2 |
| Transcribe
unprocessed pseudogene | 0 | 0 | 0 | 0 | 0 | 0 | 2 |
| Unitary
pseudogene | 0 | 0 | 0 | 0 | 0 | 0 | 1 |
The 8q24 cytoband has 509 genes: 172 coding genes
and 337 non-coding genes; among the non-coding ones, lncRNAs,
microRNAs, miscellaneous RNAs, small nucleolar RNAs, small nuclear
RNAs, and processed, unprocessed, polymorphic pseudogenes were
identified (Table SI).
The most studied genes of this cytoband are those
belonging to the MYC family (8q24.21: c-MYC, l-MYC, and n-MYC). The
c-MYC proto-oncogene is affected in almost 20% of the different
types of cancer, and it is suspected that it may be related to the
functioning of other genes (59).
It is a coding gene that participates in cell division and
multiplication, maturation and apoptosis. To date, when searching
for information on ‘8q24 cytoband AND cancer’ in PubMed (https://www.ncbi.nlm.nih.gov/pmc/), 248 articles
were found (April 2023). When exploring the combination ‘8q24
cytoband AND GC’, 73 articles were found; however, when performing
an advanced search (selecting the ‘Title and Abstract’ options) in
PubMed with the combination ‘8q24 AND cancer’ a total of 1,553
articles were found, whereas when using ‘8q24 AND GC’ as the search
term, only 50 articles were returned (Table SII). Based on these analyses,
Fig. 2 was constructed, which
serves as an axis for the following parts of the present
review.
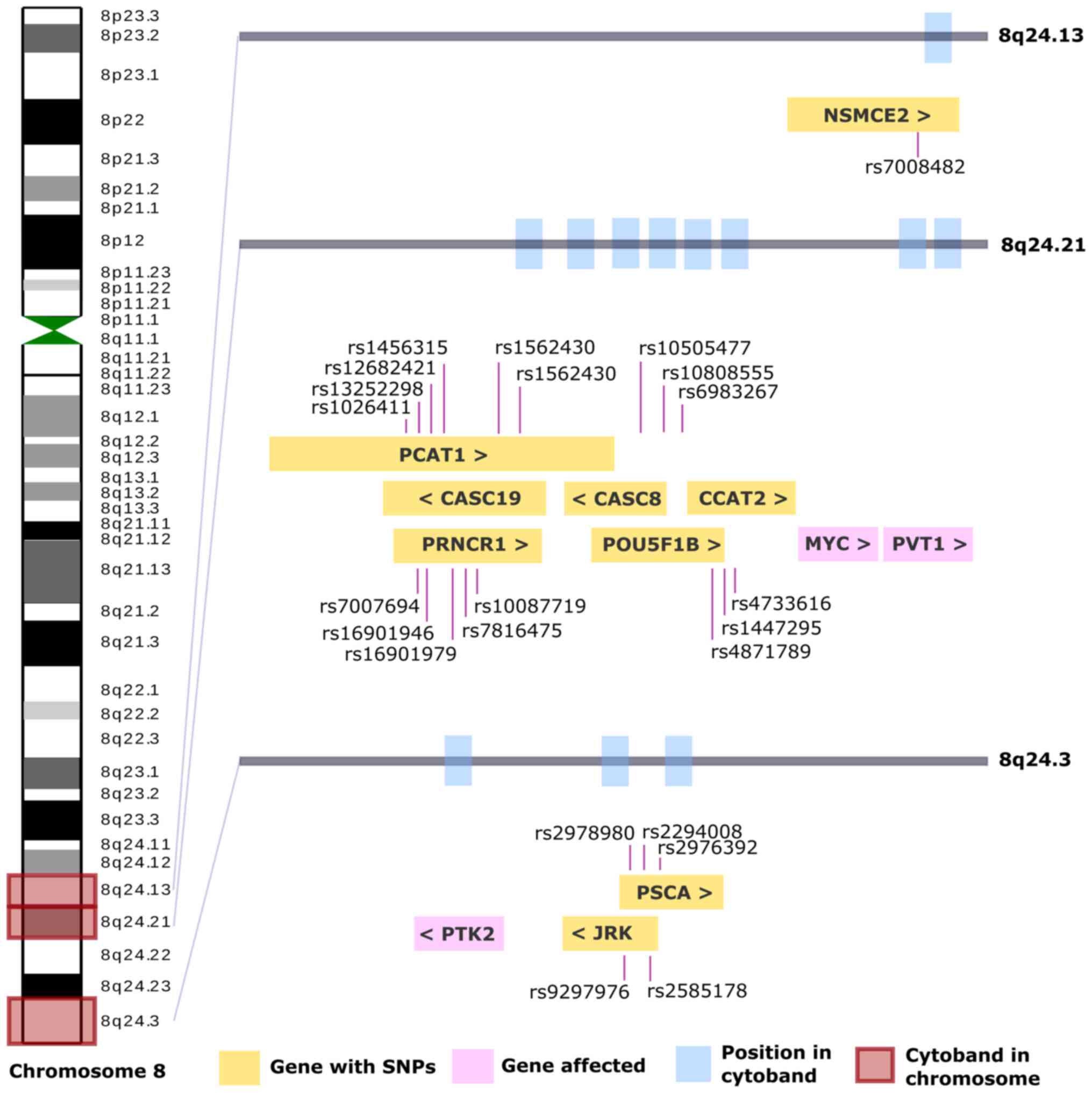 | Figure 2.The complete chromosome 8 is shown on
the left side, the centromeres are shown in green, and of the 40
cytobands, three are highlighted in red as they are most commonly
affected in GC. Each affected cytoband has a gray bar with blue
rectangles, indicating the affected genes. In the yellow
rectangles, the genes and their SNP identified in GC are observed,
and in pink, those that present an alteration, such as CNV. GC,
gastric cancer; SNP, single nucleotide polymorphism; CNV, copy
number variation; NSMCE2, NSE2 (MMS21) homolog, SMC5-SMC6 complex
SUMO ligase; PCAT1, prostate cancer associated transcript 1;
CASC19, cancer susceptibility 19; CASC8, cancer susceptibility 8;
CCAT2, colon cancer associated transcript 2; PRNCR1, prostate
cancer associated non-coding RNA 1; POU5F1B, POU class 5 homeobox
1B; PVT1, Pvt1 oncogene; PSCA, prostate stem cell antigen; PTK2,
protein tyrosine kinase 2; JRK, jrk helix-turn-helix protein; >,
sense; <, antisense. |
A total of three genes (pink rectangles) are
demonstrated in Fig. 2: MYC, PVT1
and PTK2, which have been reported to be altered in GC (Table I), and nine genes (yellow
rectangles) in which SNP alterations in GC have been described; the
cytobands in which they were located were 8q24.13, 8q24.21 and
8q24.3. The information on all these cytobands can be found in
Table SIII.
A total of seven genes related to GC are in cytoband
8q24 (ADGRB1, MTSS1, MYC, PSCA, PTK2, PTP4A3, and PVT1; Table I). The present bibliographic
analysis determined the most frequently cited 12 genes (NSMCE2,
PCAT1, CASC19, CASC8, CCAT2, PRNCR1, POU5F1B, PSCA, JRK, MYC, PVT1
and PTK2) (Table SIII). The genes
that were present in both search strategies were MYC, PVT1, PTK2
and PSCA, and of these, the PSCA gene was the most cited in GC
articles.
The PSCA gene has four SNPs: i) rs2978980, which is
located at a functional enhancer in the 8q24.3 GC-susceptibility
locus (60,61); ii) rs2294008, the T allele of which
is a risk allele for diffuse-type GC (62); iii) rs2976392, which is associated
with alterations in apoptosis/proliferation (63); iv) rs9297976, which can potentially
be recommended as a criterion for identifying high-risk groups for
the development of GC (64).
SNPs and the risk of cancer
A SNP is a genomic variant at a single base position
in DNA. Numerous studies have focused on identifying the mechanisms
of specific SNPs in a genome on health, disease, drug responses,
and other traits in SNPs can occur in promoters, exons, introns,
untranslated regions (UTRs), amongst other regions; the molecular
mechanisms that may be affected are described in Table III.
 | Table III.Molecular mechanisms of region-based
SNP on cancer susceptibility. |
Table III.
Molecular mechanisms of region-based
SNP on cancer susceptibility.
| SNP regions | Possible molecular
mechanism | Unclear issues |
|---|
| Promoter | Genetic regulation:
promoter activity (TATA box, transcription-factor binding
ability) | The interaction
between genetic and epigenetic elements |
|
| Epigenetic
regulation: DNA methylation, histone modification | Effect of SNPs on
DNA methylation status |
| Exons | Non-synonymous
cSNPs: coding protein structure and function | Detail mechanism at
the biochemical and cellular level |
|
| Synonymous cSNPs:
secondary structure conformation translation dynamics | Mechanism of the
kinetics of translation |
| Introns | cis-regulatory
elements mRNA splicing genomic imprinting long non-coding RNAs
chromatin looping | Detail functions of
cis-regulatory elements and splicing |
| UTRs | 5′-UTRs: protein
translation and transcription activity | How SNPs in the
5′-UTR affect the efficiency of translation |
|
| 3′-UTRs: Regulate
mRNA degradation and translation | How does the 3′-UTR
affect miRNA binding sites |
| Non-definite
regions | Long-range
cis-regulation tRNA and rRNA | The ways
polymorphisms affect long-range cis-regulation, tRNA and rRNA |
These alterations, as shown, affect the control of
transcription and/or translation of the genes. These sequence
changes can be insignificant or very relevant, affecting key
molecules or ‘checkpoints’ of cellular homeostasis and leading to a
predisposition to specific diseases such as various types of
cancer, particularly in GC.
In the present review, 12 genes present in cytoband
8q24 related to GC (NSMCE2, PCAT1, CASC19, CASC8, CCAT2, PRNCR1,
POU5F1B, PSCA, JRK, MYC, PVT1 and PTK2) are discussed. The PSCA
gene was cited more frequently than others; it has four known SNPs
associated with GC (rs2978980, rs2294008, rs2976392 and rs9297976).
Thus, these SNPs should be further studied in different populations
to determine their risk value in patients with GC.
Conclusions
Alterations in cytoband 8q24 can occur at the
structural level (deletions, insertions and translocations, among
other types), at the functional level (genes and proteins), and at
the SNP level, which can translate to a risk of suffering from a
disease depending on the base that is mutated, that is, if it
occurs in an oncogene or tumor suppressor gene and the function of
the affected gene. In GC, mutations in the PSCA gene and the
presence of SNPs are consistent and are thus deserved of further
study.
Supplementary Material
Supporting Data
Supporting Data
Supporting Data
Acknowledgements
The authors would like to thank Ms. Alejandra García
Bejarano and Ms. Araceli Peralta Aguilar (Medical Research Unit in
Oncological Diseases, Oncology Hospital, Century XXI National
Medical Center, Mexican Social Security Institute) for support in
bibliographic organization, and Dr Penélope Aguilera [Cerebral
Vascular Pathology Laboratory, National Institute of Neurology and
Neurosurgery (INNN)] for the English editing and critical revision
of the manuscript.
Funding
Funding: No funding was received.
Availability of data and materials
All data generated or analyzed during this study are
included in this published article.
Authors' contributions
VLS and HAVS, participated in the analysis of
results, preparation, writing, and discussion of the manuscript.
MERT was responsible for the design of the present study. VLS, HAVS
and MERT supervised, critically reviewed, edited and wrote the
manuscript. Data authentication is not applicable. All authors read
and approved the final version of the manuscript.
Ethics approval and consent to
participate
Not applicable.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Slominski RM, Raman C, Chen JY and
Slominski AT: How cancer hijacks the body's homeostasis through the
neuroendocrine system. Trends Neurosci. 46:263–275. 2023.
View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Global Cancer Observatory: 900 World Fact
Sheet. https://gco.iarc.who.int/media/globocan/factsheets/populations/900-world-fact-sheet.pdfFebruary
22–2024
|
|
3
|
Li J: Digestive cancer incidence and
mortality among young adults worldwide in 2020: A population-based
study. World J Gastrointest Oncol. 14:278–294. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
McLean MH and El-Omar EM: Genetics of
gastric cancer. Nat Rev Gastroenterol Hepatol. 11:664–674. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Nagini S: Carcinoma of the stomach: A
review of epidemiology, pathogenesis, molecular genetics, and
chemoprevention. World J Gastrointest Oncol. 4:156–169. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Choi YJ and Kim N: Gastric cancer and
family history. Korean J Intern Med. 31:1042–1053. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Ansari S, Gantuya B, Tuan VP and Yamaoka
Y: Diffuse astric cancer: A summary of analogous contributing
factors for its molecular pathogenicity. Int J Mol Sci.
19:24242018. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Ming SC: Gastric carcinoma. A
pathobiological classification. Cancer. 39:2475–2485. 1977.
View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Correa P: Human gastric carcinogenesis: A
multistep and multifactorial process-first american cancer society
award lecture on cancer epidemiology and Prevention. Cancer Res.
52:6735–6740. 1992.PubMed/NCBI
|
|
10
|
Karimi P, Islami F, Anandasabapathy S,
Freedman ND and Kamangar F: Gastric cancer: Descriptive
epidemiology, risk factors, screening, and prevention. Cancer
Epidemiol Biomarkers Prev. 23:700–713. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Lauren P: The two histological main types
of gastric carcinoma: Diffuse and so-called intestinal-type
carcinoma. An attempt at a histo-clinical classification. Acta
Pathol Microbiol Scand. 64:31–49. 1965. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Nakamura K, Sugano H and Takagi K:
Carcinoma of the stomach in incipient phase: Its histogenesis and
histological appearances. Gan. 59:251–258. 1968.PubMed/NCBI
|
|
13
|
Goseki N, Takizawa T and Koike M:
Differences in the mode of the extension of gastric cancer
classified by histological type: New histological classification of
gastric carcinoma. Gut. 33:606–612. 1992. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Nagtegaal ID, Odze RD, Klimstra D, Paradis
V, Rugge M, Schirmacher P, Washington KM, Carneiro F and Cree IA;
WHO Classification of Tumours Editorial Board, : The 2019 WHO
classification of tumours of the digestive system. Histopathology.
76:182–188. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Hartgrink HH, Jansen EPM, van Grieken NC
and van de Velde CJ: Gastric cancer. Lancet. 374:477–490. 2009.
View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Jang BG and Kim WH: Molecular pathology of
gastric carcinoma. Pathobiology. 78:302–310. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Suárez-Arriaga MC, Ribas-Aparicio RM and
Ruiz-Tachiquín ME: MicroRNAs in hereditary diffuse gastric cancer.
Biomed Rep. 5:151–154. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
An International System for Human
Cytogenetic Nomenclature, . Recommendations of the International
Standing Committee on Human Cytogenetic Nomenclature, Memphis,
Tennessee, USA. Karger Medical and Scientific Publishers; 1995
|
|
19
|
Wallander K, Eisfeldt J, Lindblad M,
Nilsson D, Billiau K, Foroughi H, Nordenskjöld M, Liedén A and Tham
E: Cell-free tumour DNA analysis detects copy number alterations in
gastro-oesophageal cancer patients. PLoS One. 16:e02454882021.
View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Han B, Ren D, Mao B, Song X, Yang W, Zhang
H and Gao F: Tumor copy number alteration (CNA) burden as a
prognostic factor for overall survival in Chinese gastric cancers.
J Clin Oncol. 37 (Suppl 15):e155552019.(In Chinese). View Article : Google Scholar
|
|
21
|
Milne AN and Offerhaus GJ: Early-onset
gastric cancer: Learning lessons from the young. World J
Gastrointest Oncol. 2:59–64. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Gordeeva V, Sharova E, Babalyan K,
Sultanov R, Govorun VM and Arapidi G: Benchmarking germline CNV
calling tools from exome sequencing data. Sci Rep. 11:144162021.
View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Iafrate AJ, Feuk L, Rivera MN, Listewnik
ML, Donahoe PK, Qi Y, Scherer SW and Lee C: Detection of
large-scale variation in the human genome. Nat Genet. 36:949–951.
2004. View
Article : Google Scholar : PubMed/NCBI
|
|
24
|
Larios-Serrato V, Martínez-Ezquerro JD,
Valdez-Salazar HA, Torres J, Camorlinga-Ponce M, Piña-Sánchez P and
Ruiz-Tachiquín ME: Copy number alterations and
epithelial-mesenchymal transition genes in diffuse and intestinal
gastric cancers in Mexican patients. Mol Med Rep. 25:1912022.
View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Cancer Genetic Web, . Genes Linked with
Cancer in Chromosome 8. https://www.cancer-genetics.org/clinkc08.htmApril
30–2023
|
|
26
|
Logsdon GA, Vollger MR, Hsieh P, Mao Y,
Liskovykh MA, Koren S, Nurk S, Mercuri L, Dishuck PC, Rhie A, et
al: The structure, function and evolution of a complete human
chromosome 8. Nature. 593:101–107. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
ADGRB1 adhesion G protein-coupled receptor
B1 [Homo sapiens (human)]-Gene-NCBI, . https://www.ncbi.nlm.nih.gov/gene?Db=gene&Cmd=DetailsSearch&Term=575April
30–2023
|
|
28
|
AGO2 argonaute RISC catalytic component 2
[Homo sapiens (human)]-Gene-NCBI, . https://www.ncbi.nlm.nih.gov/gene/27161April
30–2023
|
|
29
|
CYP11B1 cytochrome P450 family 11
subfamily B member 1 [Homo sapiens (human)]-Gene-NCBI, . https://www.ncbi.nlm.nih.gov/gene?Db=gene&Cmd=DetailsSearch&Term=1584April
30–2023
|
|
30
|
Slominski RM, Tuckey RC, Manna PR, Jetten
AM, Postlethwaite A, Raman C and Slominski AT: Extra-adrenal
glucocorticoid biosynthesis: Implications for autoimmune and
inflammatory disorders. Genes Immun. 21:150–168. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
CYP11B2 cytochrome P450 family 11
subfamily B member 2 [Homo sapiens (human)]-Gene-NCBI, . https://www.ncbi.nlm.nih.gov/gene?Db=gene&Cmd=DetailsSearch&Term=1585April
30–2023
|
|
32
|
HSF1 heat shock transcription factor 1
[Homo sapiens (human)]-Gene-NCBI, . https://www.ncbi.nlm.nih.gov/gene?Db=gene&Cmd=DetailsSearch&Term=3297April
30–2023
|
|
33
|
RECQL4 RecQ like helicase 4 [Homo sapiens
(human)]-Gene-NCBI, . https://www.ncbi.nlm.nih.gov/gene?Db=gene&Cmd=DetailsSearch&Term=5965April
30–2023
|
|
34
|
SCRIB scribble planar cell polarity
protein [Homo sapiens (human)]-Gene-NCBI, . https://www.ncbi.nlm.nih.gov/gene?Db=gene&Cmd=DetailsSearch&Term=23513April
30–2023
|
|
35
|
PTK2 protein tyrosine kinase 2 [Homo
sapiens (human)]-Gene-NCBI, . https://www.ncbi.nlm.nih.gov/gene?Db=gene&Cmd=DetailsSearch&Term=5747April
30–2023
|
|
36
|
PTP4A3 protein tyrosine phosphatase 4A3
[Homo sapiens (human)]-Gene-NCBI, . https://www.ncbi.nlm.nih.gov/gene?Db=gene&Cmd=DetailsSearch&Term=11156April
30–2023
|
|
37
|
PSCA prostate stem cell antigen [Homo
sapiens (human)]-Gene-NCBI, . https://www.ncbi.nlm.nih.gov/gene?Db=gene&Cmd=DetailsSearch&Term=8000April
30–2023
|
|
38
|
EXT1 exostosin glycosyltransferase 1 [Homo
sapiens (human)]-Gene-NCBI, . https://www.ncbi.nlm.nih.gov/gene?Db=gene&Cmd=DetailsSearch&Term=2131April
30–2023
|
|
39
|
RAD21 RAD21 cohesin complex component
[Homo sapiens (human)]-Gene-NCBI, . https://www.ncbi.nlm.nih.gov/gene?Db=gene&Cmd=DetailsSearch&Term=5885April
30–2023
|
|
40
|
CCN3 cellular communication network factor
3 [Homo sapiens (human)]-Gene-NCBI, . https://www.ncbi.nlm.nih.gov/gene?Db=gene&Cmd=DetailsSearch&Term=4856April
30–2023
|
|
41
|
TNFRSF11B TNF receptor superfamily member
11b [Homo sapiens (human)]-Gene-NCBI, . https://www.ncbi.nlm.nih.gov/gene?Db=gene&Cmd=DetailsSearch&Term=4982April
30–2023
|
|
42
|
HAS2 hyaluronan synthase 2 [Homo sapiens
(human)]-Gene-NCBI, . https://www.ncbi.nlm.nih.gov/gene?Db=gene&Cmd=DetailsSearch&Term=3037April
30–2023
|
|
43
|
MTSS1 MTSS I-BAR domain containing 1 [Homo
sapiens (human)]-Gene-NCBI, . https://www.ncbi.nlm.nih.gov/gene?Db=gene&Cmd=DetailsSearch&Term=9788April
30–2023
|
|
44
|
RNF139 ring finger protein 139 [Homo
sapiens (human)]-Gene-NCBI, . https://www.ncbi.nlm.nih.gov/gene?Db=gene&Cmd=DetailsSearch&Term=11236April
30–2023April 30–2023
|
|
45
|
CCDC26 CCDC26 long non-coding RNA [Homo
sapiens (human)]-Gene-NCBI, . https://www.ncbi.nlm.nih.gov/gene?Db=gene&Cmd=DetailsSearch&Term=137196April
30–2023
|
|
46
|
MYC MYC proto-oncogene, bHLH transcription
factor [Homo sapiens (human)]-Gene-NCBI, . https://www.ncbi.nlm.nih.gov/gene?Db=gene&Cmd=DetailsSearch&Term=4609April
30–2023
|
|
47
|
PVT1 Pvt1 oncogene [Homo sapiens
(human)]-Gene-NCBI, . https://www.ncbi.nlm.nih.gov/gene?Db=gene&Cmd=DetailsSearch&Term=5820April
30–2023
|
|
48
|
CCN4 cellular communication network factor
4 [Homo sapiens (human)]-Gene-NCBI, . https://www.ncbi.nlm.nih.gov/gene?Db=gene&Cmd=DetailsSearch&Term=8840April
30–2023
|
|
49
|
NDRG1 N-myc downstream regulated 1 [Homo
sapiens (human)]-Gene-NCBI, . https://www.ncbi.nlm.nih.gov/gene?Db=gene&Cmd=DetailsSearch&Term=10397April
30–2023
|
|
50
|
Zech L, Haglund U, Nilsson K and Klein G:
Characteristic chromosomal abnormalities in biopsies and
lymphoid-cell lines from patients with Burkitt and non-Burkitt
lymphomas. Int J Cancer. 17:47–56. 1976. View Article : Google Scholar : PubMed/NCBI
|
|
51
|
zur Hausen A, Brink AA, Craanen ME,
Middeldorp JM, Meijer CJ and van den Brule AJ: Unique transcription
pattern of Epstein-Barr virus (EBV) in EBV-carrying gastric
adenocarcinomas: Expression of the transforming BARF1 gene. Cancer
Res. 60:2745–2748. 2000.PubMed/NCBI
|
|
52
|
Gómez-Carpintero García A, Vidal Esteban
A, Bermejo Gómez A and Púa Torrejón RC: Phenotype and variations
associated with the deletion of the 1q44 cytoband and the
pathogenic duplication in the 9q32q34.3 cytobands. BMJ Case Rep.
13:e2319382020. View Article : Google Scholar : PubMed/NCBI
|
|
53
|
Brisbin AG, Asmann YW, Song H, Tsai YY,
Aakre JA, Yang P, Jenkins RB, Pharoah P, Schumacher F, Conti DV, et
al: Meta-analysis of 8q24 for seven cancers reveals a locus between
NOV and ENPP2 associated with cancer development. BMC Med Genet.
12:1562011. View Article : Google Scholar : PubMed/NCBI
|
|
54
|
Wilson C and Kanhere A: 8q24.21 locus: A
paradigm to link non-coding RNAs, genome polymorphisms and cancer.
Int J Mol Sci. 22:10942021. View Article : Google Scholar : PubMed/NCBI
|
|
55
|
Sun Y and Ma L: New insights into long
non-coding RNA MALATI in cancer and metastasis. Cancers (Basel).
11:2162019. View Article : Google Scholar : PubMed/NCBI
|
|
56
|
Cunningham F, Allen JE, Allen J,
Alvarez-Jarreta J, Amode MR, Armean IM, Austine-Orimoloye O, Azov
AG, Barnes I, Bennett R, et al: Ensembl 2022. Nucleic Acids Res.
50((D1)): D988–D995. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
57
|
Kent WJ, Sugnet CW, Furey TS, Roskin KM,
Pringle TH, Zahler AM and Haussler D: The human genome browser at
UCSC. Genome Res. 12:996–1006. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
58
|
Smedley D, Haider S, Ballester B, Holland
R, London D, Thorisson G and Kasprzyk A: BioMart-biological queries
made easy. BMC Genomics. 10:222009. View Article : Google Scholar : PubMed/NCBI
|
|
59
|
Prochownik EV: Regulation of normal and
neoplastic proliferation and metabolism by the extended Myc
network. Cells. 11:39742022. View Article : Google Scholar : PubMed/NCBI
|
|
60
|
Zheng Y, Lei T, Jin G, Guo H, Zhang N,
Chai J, Xie M, Xu Y, Wang T, Liu J, et al: LncPSCA in the 8q24.3
risk locus drives gastric cancer through destabilizing DDX5. EMBO
Rep. 22:e527072021. View Article : Google Scholar : PubMed/NCBI
|
|
61
|
Yang W, Zhang T, Song X, Dong G, Xu L and
Jiang F: SNP-target genes interaction perturbing the cancer risk in
the post-GWAS. Cancers (Basel). 14:56362022. View Article : Google Scholar : PubMed/NCBI
|
|
62
|
Iwasaki RL, Ishiya K, Kanzawa-Kiriyama H,
Kawai Y, Gojobori J and Satta Y: Evolutionary history of the risk
of SNPs for diffuse-type gastric cancer in the Japanese population.
Genes (Basel). 11:7752020. View Article : Google Scholar : PubMed/NCBI
|
|
63
|
Duan F, Song C, Shi J, Wang P, Ye H, Dai
L, Zhang J and Wang K: Identification and epidemiological
evaluation of gastric cancer risk factors: Based on a field
synopsis and meta-analysis in Chinese population. Aging (Albany
NY). 13:21451–21469. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
64
|
Turdikulova S, Dalimova D, Abdurakhimov A,
Adilov B, Navruzov S, Yusupbekov A, Djuraev M, Abdujapparov S,
Egamberdiev D and Mukhamedov R: Association of rs2294008 and
rs9297976 polymorphisms in PSCA gene with gastric cancer
Ssusceptibility in uzbekistan. Cent Asian J Glob Health.
5:2272016.PubMed/NCBI
|
|
65
|
Deng N, Zhou H, Fan H and Yuan Y: Single
nucleotide polymorphisms and cancer susceptibility. Oncotarget.
8:110635–110649. 2017. View Article : Google Scholar : PubMed/NCBI
|