|
1
|
Aryee MJ, Liu W, Engelmann JC, Nuhn P,
Gurel M, Haffner MC, Esopi D, Irizarry RA, Getzenberg RH, Nelson
WG, Luo J, Xu J, Isaacs WB, Bova GS and Yegnasubramanian S: DNA
methylation alterations exhibit intra-individual stability and
inter-individual heterogeneity in prostate cancer metastases. Sci
Transl Med. View Article : Google Scholar : 2013.
|
|
2
|
Jones PA and Baylin SB: The fundamental
role of epigenetic events in cancer. Nat Rev Genet. 3:85–93.
2002.
|
|
3
|
Jones PA and Baylin SB: The epigenomics of
cancer. Cell. 128:683–692. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Rishi V, Bhattacharya P, Chatterjee R,
Rozenberg J, Zhao J, Glass K, Fitzgerald P and Vinson C: CpG
methylation of half-CRE sequences creates C/EBPα binding sites that
activate some tissue-specific genes. Proc Natl Acad Sci.
107:20311–20316. 2010.PubMed/NCBI
|
|
5
|
Ziller MJ, Gu H, Müller F, Donaghey J,
Tsai LT, Kohlbacher O, De Jager PL, Rosen ED, Bennett DA, Bernstein
BE, Gnirke A and Meissner A: Charting a dynamic DNA methylation
landscape of the human genome. Nature. 500:477–481. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Cheng X and Blumenthal RM: Mammalian DNA
methyltransferases: a structural perspective. Structure. 3:341–350.
2008. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Sigalotti L, Fratta E, Coral S, Cortini E,
Covre A, Nicolay HJ, Anzalone L, Pezzani L, Di Giacomo A, Fonsatti
E, Colizzi F, Altomonte M, Calabrò L and Maio M: Epigenetic drugs
as pleiotropic agents in cancer treatment: biomolecular aspects and
clinical applications. J Cell Physiol. 212:330–344. 2007.
View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Asada K, Asada R, Yoshiji H, Fukui H,
Floyd RA and Kotake Y: DNA cytosine methylation profile in various
cancer-related genes is altered in cultured rat hepatocyte cell
lines as compared with primary hepatocytes. Oncol Rep.
15:1241–1248. 2006.
|
|
9
|
Feinberg AP and Vogelstein B:
Hypomethylation distinguishes genes of some human cancers from
their normal counterparts. Nature. 301:89–92. 1983. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Lee B and Muller MT: SUMOylation enhances
DNA methyltransferase 1 activity. Biochem J. 421:449–461. 2009.
View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Ling Y, Sankpal UT, Robertson AK, McNally
JG, Karpova T and Robertson KD: Modification of de novo DNA
methyltransferase 3a (Dnmt3a) by SUMO-1 modulates its interaction
with histone deacetylases (HDACs) and its capacity to repress
transcription. Nucleic Acids Res. 32:598–610. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Kanai Y and Hiroshida S: Alterations of
DNA methylation associated with abnormalities of DNA
methyltransferases in human cancers during transition from a
precancerous to a malignant state. Carcinogenesis. 28:2434–2442.
2007. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Oh BK, Kim H, Park HJ, Shim YH, Choi J,
Park C and Park YN: DNA methyltransferase expression and DNA
methylation in human hepatocellular carcinoma and their
clinicopathological correlation. Int J Mol Med. 20:65–73.
2007.PubMed/NCBI
|
|
14
|
Choi MS, Shim YH, Hwa JY, Lee SK, Ro JY,
Kim JS and Yu E: Expression of DNA methyltransferases in multistep
hepatocarcinogenesis. Hum Pathol. 34:11–18. 2013. View Article : Google Scholar
|
|
15
|
Yoshimasa S, Yae K, Tohru N, Michiie S,
Hidetsugu S, Hiromasa I and Setsuo H: Increased protein expression
of DNA methyltransferase (DNMT) 1 is significantly correlated with
the malignant potential and poor prognosis of human hepatocellular
carcinomas. Int J Cancer. 105:527–532. 2003. View Article : Google Scholar
|
|
16
|
Raddatz G, Gao Q, Bender S, Jaenisch R and
Lyko F: Dnmt3a protects active chromosome domains against
cancer-associated hypomethylation. PLoS Genet. 8:e10031462012.
View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Yan XJ, Xu J, Gu ZH, Pan CM, Lu G, Shen Y,
Shi JY, Zhu YM, Tang L, Zhang XW, Liang WX, Mi JQ, Song HD, Li KQ,
Chen Z and Chen SJ: Exome sequencing identifies somatic mutations
of DNA methyltransferase gene DNMT3A in acute monocytic leukemia.
Nat Genet. 43:309–315. 2011. View
Article : Google Scholar : PubMed/NCBI
|
|
18
|
Hlady RA, Novakova S, Opavska J,
Klinkebiel D, Peters LS, Bies J, Hannah J, Iqbal J, Anderson KM,
Siebler HM, Smith LM, Greiner TC, Bastola D, Joshi S, Lockridge O,
Simpson MA, Felsher DW, Wagner KU, Chan WC, Christman JK and
Opavsky R: Loss of Dnmt3b function upregulates the tumor modifier
Ment and accelerates mouse lymphomagenesis. J Clin Invest.
122:163–177. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Trinh BN, Long TI, Nickel AE, Shibata D
and Laird PW: DNA methyltransferase deficiency modifies cancer
susceptibility in mice lacking DNA mismatch repair. Mol Cell Biol.
22:2906–2917. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Pérez-Carreón JI, López-García C,
Fattel-Fazenda S, Arce-Popoca E, Alemán-Lazarini L,
Hernández-García S, Le Berrey V, Sokoly S, Francoisy JM and
Villa-Treviño S: Gene expression profile related to the progresión
of preneoplastic nodules toward hepatocellular carcinoma in rats.
Neoplasia. 8:373–383. 2006.
|
|
21
|
Rotstein J, Macdonald PD, Rabes HM and
Farber E: Cell cycle kinetics of rat hepatocytes in early putative
preneoplastic lesions in hepatocarcinogenesis. Cancer Res.
44:2913–2917. 1984.PubMed/NCBI
|
|
22
|
Sánchez-Pérez Y, Carrasco-Legleu C,
García-Cuellar C, Pérez-Carreón JI, Hernández-García S,
Salcido-Neyoy M, Alemán-Lazarini L and Villa-Treviño S: Oxidative
stress in carcinogenesis. Correlation between lipid peroxidation
and induction of preneoplastic lesions in rat hepatocarcinogenesis.
Cancer Lett. 217:25–32. 2005.PubMed/NCBI
|
|
23
|
Solt D and Farber E: New principle for the
analysis of chemical carcinogenesis. Nature. 263:701–703. 1976.
View Article : Google Scholar
|
|
24
|
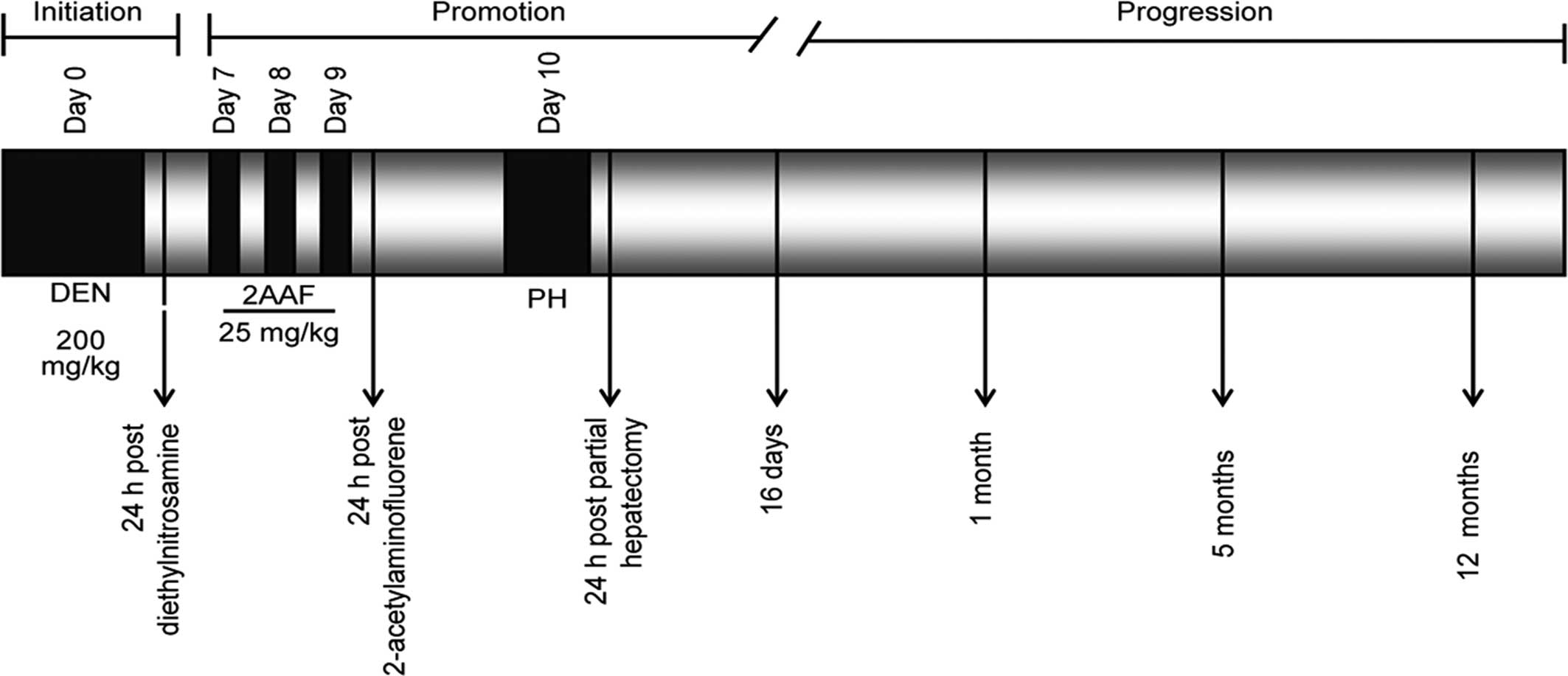
Solt DB, Medine A and Farber E: Rapid
emergence of carcinogen-induced hyperplastic lesions in a new model
for the sequential analysis of liver carcinogenesis. Am J Pathol.
88:595–618. 1977.PubMed/NCBI
|
|
25
|
Solt DB, Cayama E, Tsuda H, Enomoto K, Lee
G and Farber E: Promotion of liver cancer development by brief
exposure to dietary 2-acetylaminofluorene plus partial hepatectomy
or carbon tetrachloride. Cancer Res. 43:188–191. 1983.PubMed/NCBI
|
|
26
|
Pascale RM, Simile MM, De Miglio MR,
Muroni MR, Calvisi DF, Asara G, Casabona D, Frau M, Seddaiu MA and
Feo F: Cell cycle deregulation in liver lesions of rats with and
without genetic predisposition to hepatocarcinogenesis. Hepatology.
35:1341–1350. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
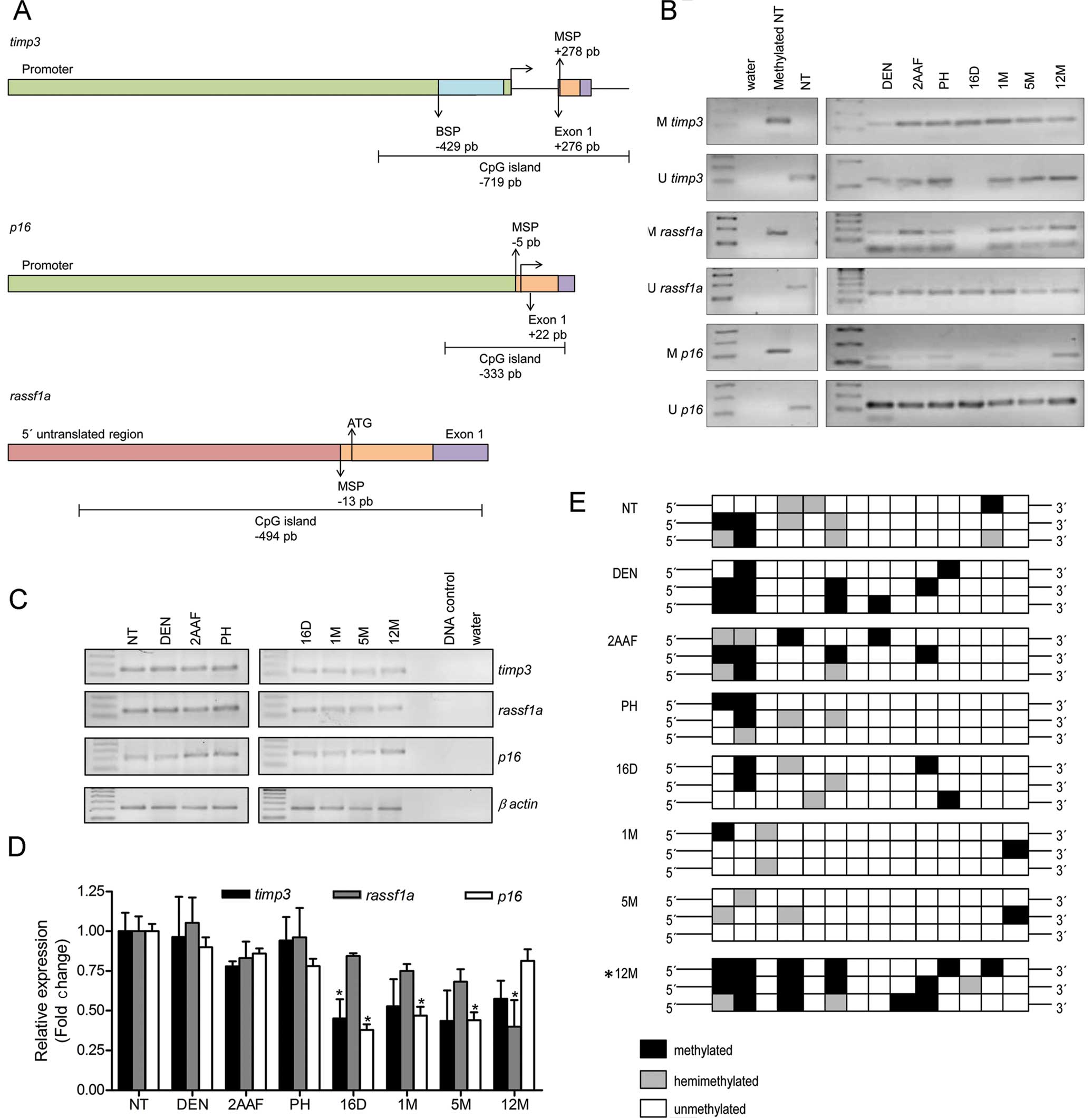
Wen BL, Ao L, Zhou ZY, Cui ZH, Zhou YH,
Yuan XY, Xiang YL, Cao J and Liu JY: CpG island hypermethylation of
multiple tumor suppressor genes associated with loss of their
protein expression during rat lung carcinogenesis induced by
3-methylcholanthrene and diethylnitrosamine. Biochem Biophys Res
Commun. 402:507–514. 2010. View Article : Google Scholar
|
|
28
|
Agathanggelou A, Cooper WN and Latif F:
Role of the Ras-association domain family 1 tumor suppressor gene
in human cancers. Cancer Res. 65:3497–3508. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Du YP, Peng JS, Sun A, Tang ZH, Ling WH
and Zhu HL: Assessment of the effect of betaine on p16 and
c-myc DNA methylation and mRNA expression in a chemical
induced rat liver cancer model. BMC Cancer. View Article : Google Scholar : 2009.
|
|
30
|
Li J, Knobloch TJ, Poi MJ, Zhang Z, Davis
AT, Muscarella P and Weghorst CM: Genetic alterations of
RDINK4/ARF enhancer in human cancer cells. Mol Carcinog.
53:211–218. 2014.
|
|
31
|
Gutierrez-Reyes G, del Carmen Garcia de
Leon M, Varela-Fascinetto G, Valencia P, Pérez Tamayo R, Rosado CG,
Labonne BF, Rochilin NM, Garcia RM, Valadez JA, Latour GT, Corona
DL, Diaz GR, Zlotnik A and Kershenobich D: Cellular senescence in
livers from children with end stage liver disease. PLoS One.
5:e102312010. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Enomoto K and Farber E: Kinetics of
phenotypic maturation of remodeling of hyperplastic nodules during
liver carcinogenesis. Cancer Res. 42:2330–2335. 1982.PubMed/NCBI
|
|
33
|
Ziech D, Franco R, Pappa A and
Panayiotidis MI: Reactive oxygen species (ROS) - induced genetic
and epigenetic alterations in human carcinogenesis. Mutat Res.
711:167–173. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Torres L, Avila MA, Carretero MV, Latasa
MU, Caballería J, López-Rodas G, Boukaba A, Lu SC, Franco L and
Mato JM: Liver-specific methionine adenosyltransferase MAT1A gene
expression is associated with a specific pattern of promoter
methylation and histone acetylation: implications for MAT1A
silencing during transformation. FASEB J. 14:95–102. 2004.
|
|
35
|
Yang H, Huang ZZ, Zeng Z, Chen C, Selby RR
and Lu SC: Role of promoter methylation in increased methionine
adenosyltransferase 2A expression in human liver cancer. Am J
Physiol Gastrointest Liver Physiol. 280:184–190. 2001.PubMed/NCBI
|
|
36
|
Takayama K, Shimoda N, Takanaga S, Hozumi
S and Kikuchi Y: Expression patterns of dnmt3aa, dnmt3ab and dnmt4
during development and fin regeneration in zebrafish. Gene Expr
Patterns. 14:105–110. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Takasugi M, Hayakawa K, Arai D and Shiota
K: Age- and sex-dependent DNA hypomethylation controlled by growth
hormone in mouse liver. Mech Ageing Dev. 134:331–337. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Berman BP, Weisenberger DJ, Aman JF,
Hinoue T, Ramjan Z, Liu Y, Noushmehr H, Lange CPE, van Dijk MC,
Tollenaar RAEM, van den Berg D and Laird PW: Regions of focal DNA
hypermethylation and long-range hypomethylation in colorectal
cancer coincide with nuclear lamina-associated domains. Nat Genet.
44:40–46. 2012. View
Article : Google Scholar : PubMed/NCBI
|
|
39
|
Hervouet E, Lalier L, Debien D, Cheray M,
Geairon A, Rogniaux H, Loussouarn D, Martin SA, Vallette FM and
Cartron PF: Disruption of Dnmt1/PCNA/UHRF1 interactions promotes
tumorigenesis from human and mice glial cells. PLoS One.
5:e113332010. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Mortusewicz O, Schermelleh L, Walter J,
Cardoso MC and Leonhardt H: Recruitment of DNA methyltransferase I
to DNA repair sites. Proc Natl Acad Sci. 102:8905–8909. 2005.
View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Gazin C, Wajapeyee N, Gobeil S, Virbasius
CM and Green MR: An elaborate pathway required for Ras-mediated
epigenetic silencing. Nature. 449:1073–1077. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Liu X, Gao Q, Li P, Zhao Q, Zhang J, Li J,
Koseki H and Wong J: UHRF1 targets DNMT1 for DNA methylation
through cooperative binding of hemi methylated DNA and methylated
H3K9. Nat Commun. View Article : Google Scholar : 2013.PubMed/NCBI
|
|
43
|
Shamma A, Suzuki M, Hayashi N, Kobayashi
M, Sasaki N, Nihiuchi T, Doki Y, Okamoto T, Kohno S, Muranaka H,
Kitajima S, Yamamoto K and Takahashi C: ATM mediates pRB function
to control DNMT1 protein stability and DNA methylation. Mol Cell
Biol. 33:3113–3124. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Cartron PF, Blanquart C, Hervouet E,
Gregoire M and Vallette FM: HDAC1-mSin3a-NCOR1, Dnmt3b-HDAC1-Egr1
and Dnmt1-PCNA-UHRF1-G9a regulate the NY-ESO1 gene expression. Mol
Oncol. 7:452–463. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Xionga Y, Dowdya SC, Xueb A, Shujuana J,
Eberhardtc NL, Podratza KC and Jianga SW: Opposite alterations of
DNA methyltransferase gene expression in endometrioid and serous
endometrial cancers. Gynecol Oncol. 96:601–609. 2005. View Article : Google Scholar : PubMed/NCBI
|