Introduction
Over 90% of lip and oral cavity cancers are squamous
cell carcinomas, with 300,000 new cases annually worldwide. Based
on the classification of the cancer cell type, it is associated
with squamous cell carcinoma of the head and neck, esophagus, lung,
cervix, uterus, skin and mucosa at other sites; almost 2 million
new cases of these types of cancer are reported each year (1). Locoregional treatment is frequently used
for squamous cell carcinoma and the standard treatment currently
consists of surgery plus adjuvant chemoradiation. Despite the
recent introduction of molecular-targeted therapy, treatment
options depend on the disease stage (2). Direct cause of fatality from oral
squamous cell carcinoma (OSCC) is uncontrollable locoregional
recurrence at the surgically treated site (such as resection of
primary cancer and neck dissection) or distant metastasis.
Locoregional and hematogenous dissemination in the early stage of
cancer also remains difficult to control with standard treatment
(3). To improve the control rate,
local plus systemic treatment strategies against micro-metastasis
are required from the outset of treatment.
Our previous studies conducted comprehensive
integrin gene expression analysis to identify prognostic biomarkers
that predict the clinical behavior of OSCC and identified that the
two gene expression ratios, ITGA3/CD9 and
ITGB4/JUP, reflect malignant behavior associated with
lymph node metastasis (LNM) and distant metastasis, respectively
(3–5).
In the present study, the diagnostic value of cyclins,
cyclin-dependent kinases (CDKs) and CDK inhibitors (CDKIs) gene
expression was investigated since these molecules play central
roles in cell cycle control, have been extensively studied, mediate
carcinogenesis and can be used as prognostic factors of cancer
(6,7).
These genes became of particular interest recently as possible
molecular targets of cancer therapy (8). Therefore, the association of the
expression of 11 cyclin-related genes in OSCC tissues with clinical
events were investigated to examine their value as biomarkers.
Materials and methods
Patients and specimens
Tumor samples were collected at biopsy from 77
patients with OSCC (49 men, 28 women; mean age, 66 years; range,
22–86 years) who underwent surgical treatment between 1999 and 2012
at the Dental Department of Niigata University Medical and Dental
Hospital (Niigata, Japan); the Special Dental Care and Oral Surgery
Department of Shinshu University Hospital (Nagano, Japan); and the
Division of Oral Surgery of Nagaoka Red Cross Hospital (Nagaoka,
Japan) (Table I). The treatment
modalities included local resection, composite resection (resection
of primary oral cancer, a portion of the oral floor and mandible,
and reconstruction with tissue transplantation and neck dissection)
and composite resection with radiation therapy with or without
intravenous adjuvant chemotherapy.
 | Table I.Demographic and clinicopathological
characteristics of 77 patients with oral squamous cell
carcinoma. |
Table I.
Demographic and clinicopathological
characteristics of 77 patients with oral squamous cell
carcinoma.
| Clinicopathological
factors | Values |
|---|
| Total, n | 77 |
| Observation period,
days [range (mean)] | 144–2,762
(1,510) |
| Age, years [range
(mean)] | 22–86 (66) |
| Gender, n (%) |
|
|
Male | 49 (63.6) |
|
Female | 28 (36.4) |
| Site, n (%) |
|
|
Tongue | 38 (49.4) |
|
Gingiva | 27 (35.1) |
| Oral
floor | 9
(11.7) |
| Buccal
mucosa | 3 (3.9) |
| Tumor
sizea, mm [range
(mean)] | 8–52 (27.1) |
| ≤20 mm,
n (%) | 25 (32.5) |
| 21–30
mm, n (%) | 26 (33.8) |
| 31–40
mm, n (%) | 18 (23.4) |
| >40
mm, n (%) | 8
(10.4) |
| Tumor
statusb, n (%) |
|
| T1 | 24 (31.2) |
| T2 | 35 (45.5) |
| T3 | 3 (3.9) |
| T4 | 15 (19.5) |
| Lymph node
metastasisc, n (%) |
|
| N0 | 37 (48.1) |
| N1 | 10 (13.0) |
| N2 | 30 (39.0) |
| N3 | 0 (0.0) |
| Histological grade
(YK4)d, n (%) |
|
|
2–3 | 31 (40.3) |
|
4c-d | 46 (59.7) |
| Margin
statuse, n (%) |
|
|
Negative | 72 (93.5) |
|
Positive | 5 (6.5) |
| Primary site
recurrence, n (%) |
|
|
Negative | 71 (92.2) |
|
Positive | 6 (7.8) |
| Distant metastasis,
n (%) |
|
|
Negative | 72 (93.5) |
|
Positive | 5 (6.5) |
| Disease-specific
deathf, n (%) |
|
|
Alive | 63 (81.8) |
|
Succumbed | 14 (18.2) |
The study was performed in accordance with the
guidelines of the Declaration of Helsinki and the protocol was
approved by the Research Ethics Committee of Niigata University
Medical and Dental Hospital, the Research Ethics Committee of
Shinshu University Hospital and the Ethics Committee of Nagaoka Red
Cross Hospital. Patients provided written informed consent to
participate in the study.
Total RNA extraction from carcinoma
tissue
Cancer tissue specimens were preserved by immersion
in RNAlater solution (Invitrogen Life Technologies, Tokyo, Japan).
Extraction of total RNA was performed using the RNeasy Lipid Tissue
Mini kit (Qiagen, Tokyo, Japan) following homogenization by
TissueLyser LT (Qiagen) in QIAzol Lysis reagent according to the
manufacturer's instructions. Synthesis of first-strand cDNA was
performed by reverse transcription using total RNA (0.2–1 µg) as a
template (SuperScript III®; Invitrogen Life Technologies).
Gene expression analysis by reverse
transcription-quantitative polymerase chain reaction (RT-qPCR)
RT-qPCR was performed on a Thermal Cycler Dice® Real
Time System II (Takara Bio Inc., Shiga, Japan) using cDNA
synthesized from the patients' cancer specimens and TaqMan probes
for cyclins, CDKs, CDKIs, integrins and integrin-related genes
(TaqMan Gene Expression Assays; Invitrogen Life Technologies)
according to the following protocol: 600 sec at 95°C, followed by
extension for 15 sec at 95°C and 60 sec at 60°C. Relative standard
curves representing several 10-fold cDNA dilutions
(1:10:100:1,000:10,000:100,000) from an OSCC tissue sample were
used for the linear regression analysis of other samples. The
manufacturer's TaqMan probe assay IDs are as follows: CCNA1:
Hs00171105_m1; CCND1: Hs00277039_m1; CCND2:
Hs00153380_m1; CCNE1: Hs00233356_m1; CDK1:
Hs00938777_m1; CDK2: Hs00608082_m1; CDK4:
Hs00364847_m1; CDKN2A: Hs00923894_m1; CDKN1A:
Hs00355782_m1; CDKN1B: Hs01597588_m1; CDKN1C:
Hs00175938_m1; ITGA3: Hs00233722_m1; ITGB4:
Hs01103172_g1; CD9: Hs01124027_m1; JUP:
Hs00158408_m1; and GAPDH: Hs02758991_g1.
Statistical analysis
Expression ratios of all the possible combinations
of the 12 genes (11 cyclin-related genes and GAPDH; 66
combinations in total) and expression ratios of two combinations of
the integrin genes (ITGA3/CD9 and
ITGB4/JUP) were calculated for each of the 77 OSCC
cases. Each gene expression ratio was categorized by the cut-off
point estimated from the receiver operating characteristic curve
for each clinical event. The response variables used were LNM
determined by histopathological examination of the surgical
specimen, primary site recurrence (PSR) following surgery, distant
metastasis following successful surgery for the primary cancer and
disease-specific death (DSD) from uncontrollable OSCC. The
clinicopathological parameters were age, gender, tumor size, tumor
category (Union for International Cancer Control
Tumor-Node-Metastasis Classification), histopathological mode of
invasion (9) and positive surgical
margins on histological examination. Distant metastasis following
locoregional failure was regarded as locoregional recurrence.
The influence of the gene expression ratios and
clinicopathological parameters on LNM, PSR, distant metastasis and
DSD was examined by univariate analyses (Mann-Whitney U test
followed by log-rank test) to optimize the combination of variables
for subsequent multivariate analysis. Analyses using a Cox
proportional hazards model and Kaplan-Meier (K–M) curves were
performed for the events of LNM, PSR, distant metastasis and DSD
(or disease-specific survival in K–M curves) using SPSS 21.0 (IBM
Japan Ltd., Tokyo, Japan). Time to each event was calculated from
the date of first visit to the date of neck dissection, diagnosis
of recurrence and date of DSD or final observation for surviving
patients. P<0.05 was considered to indicate a statistically
significant difference.
Results
Univariate analysis
The gene expression ratios and clinicopathological
parameters that showed significance for each clinical event on the
Mann-Whitney U test were further tested by the log-rank test. This
revealed that the four clinical events were influenced by three
CDK-related gene expression ratios, as well as the two integrin
gene expression ratios, tumor size, surgical margin status and
invasive histology (Table II). For
multivariate analysis, these gene expression ratios were used in
double or triple combinations to group ‘double-positive’ or
‘triple-positive’ cases.
 | Table II.Final log-rank test results for the
risks of lymph node metastasis, primary site recurrence, distant
metastasis and disease-specific death in oral squamous cell
carcinoma. |
Table II.
Final log-rank test results for the
risks of lymph node metastasis, primary site recurrence, distant
metastasis and disease-specific death in oral squamous cell
carcinoma.
| Log-rank
(Mantel-Cox) | Lymph node
metastasis | Primary site
recurrence | Distant
metastasis | Disease-specific
deathd |
|---|
| Gene expression
ratios |
|
|
|
|
|
CDK2/CDKN1A | – | 0.019 | 0.017 | 0.002 |
|
CDK1/CDKN1B | 0.008 | 0.047 | – | 0.041 |
|
CCNE1/CDK2 | 0.002 | – | 0.012 | 0.005 |
| Integrin gene
expression ratios |
|
|
|
|
|
ITGB4/JUP | – | – | 0.008 | – |
|
ITGA3/CD9 | 0.005 | – | – | 0.011 |
| Clinicopathological
parameters |
|
|
|
|
| Size,
>20 mma | 0.047 | 0.057 | – | – |
|
Positive marginb | 0.002 | <0.001 | – | 0.001 |
|
Invasive
histologyc | 0.005 | 0.032 | 0.051 | 0.001 |
LNM
In the Cox proportional hazards model of LNM
(Table IIIA), the single parameters
(high-ITGA3/CD9 and high-CDK1/CDKN1B)
were independent significant variables in addition to tumor size
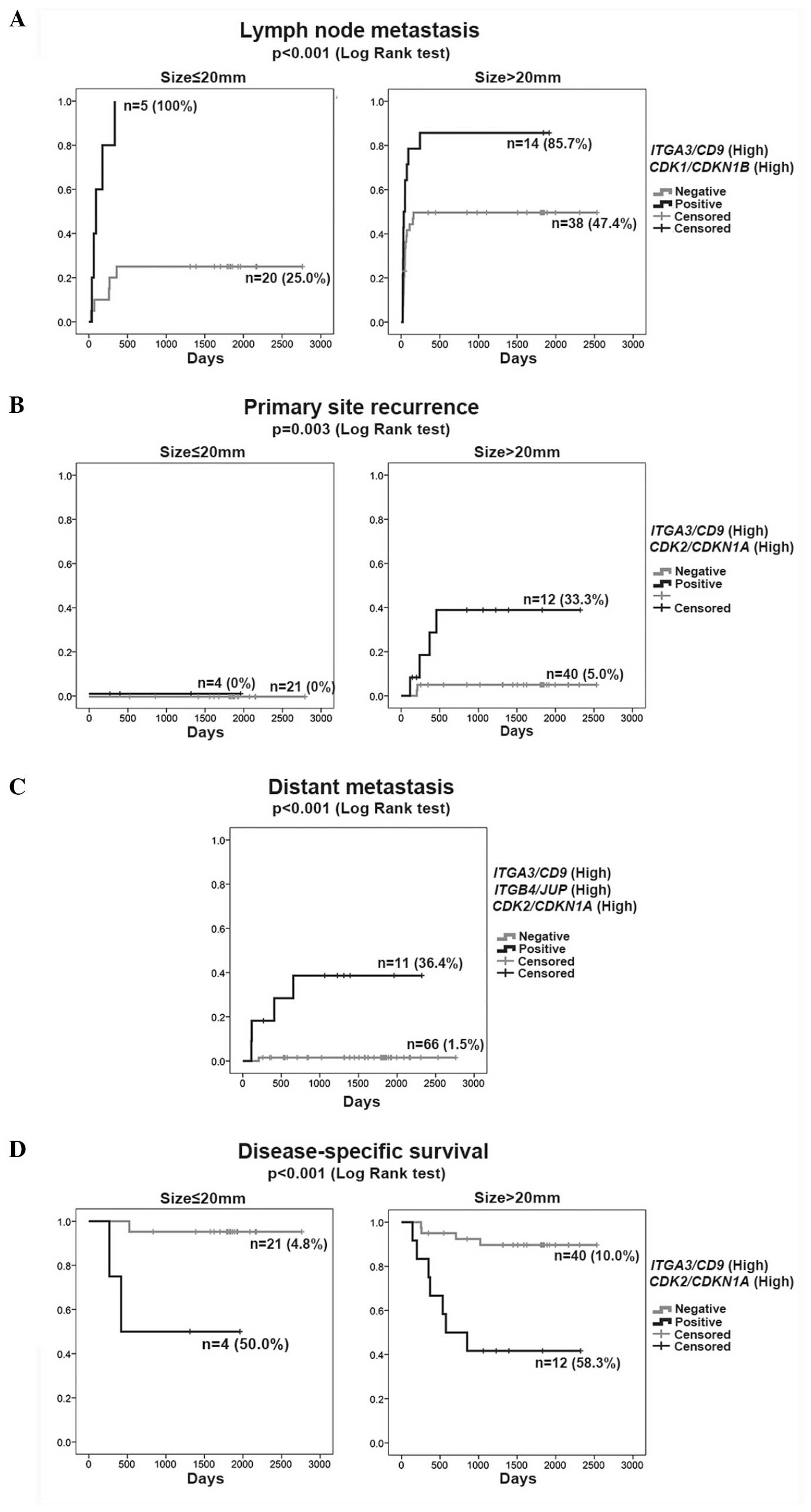
(size, >20 mm). The rate of LNM was represented by the K–M curve
(one minus cumulative survival) between groups of double-positive
cases (high-ITGA3/CD9 and
high-CDK1/CDKN1B) and the remaining (negative) cases
according to size (≤20 or >20 mm) (Fig. 1A). The double-positive cases
(high-ITGA3/CD9 and high-CDK1/CDKN1B)
consistently exhibited a high rate of LNM (>85%) irrespective of
tumor size, but the negative cases revealed an increasing rate of
LNM with larger tumor size (>20 mm).
 | Table III.Cox proportional hazard models for
the risks of lymph node metastasis, primary site recurrence,
distant metastasis and fatality in oral squamous cell
carcinoma. |
Table III.
Cox proportional hazard models for
the risks of lymph node metastasis, primary site recurrence,
distant metastasis and fatality in oral squamous cell
carcinoma.
| A, Lymph node
metastasis |
|
|
|
|
|
|
|
|---|
|
|---|
|
|
|
|
|
|
| 95% CI for OR |
|---|
|
|
|
|
|
|
|
|
|---|
| Variable | B | SE | Wald | P-value | OR | Lower limit | Upper limit |
|---|
|
High-ITGA3/CD9 | 1.057 | 0.341 |
9.627 | 0.002 |
2.878 | 1.476 |
5.613 |
|
High-CDK1/CDKN1B | 0.853 | 0.332 |
6.609 | 0.010 |
2.346 | 1.225 |
4.494 |
| Size >20
mma | 0.623 | 0.367 |
2.875 | 0.090 |
1.864 | 0.908 |
3.829 |
|
| B, Primary site
recurrence |
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 95% CI for OR |
|
|
|
|
|
|
|
|
| Variable | B | SE | Wald | P-value | OR | Lower limit | Upper limit |
|
|
High-ITGA3/CD9 and
High-CDK2/CDKN1A | 2.352 | 0.883 |
7.089 | 0.008 | 10.505 | 1.860 |
59.328 |
| Positive
marginb | 2.291 | 0.903 |
6.444 | 0.011 |
9.886 | 1.686 |
57.980 |
|
| C, Distant
metastasis |
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 95% CI for OR |
|
|
|
|
|
|
|
|
| Variable | B | SE | Wald | P-value | OR | Lower limit | Upper limit |
|
|
High-ITGB4/JUP,
High-ITGA3/CD9 and High-CDK2/CDKN1A | 3.350 | 1.119 |
8.960 | 0.003 | 28.496 | 3.179 | 255.464 |
|
| D, Disease-specific
deathc |
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 95% CI for OR |
|
|
|
|
|
|
|
|
| Variable | B | SE | Wald | P-value | OR | Lower limit | Upper limit |
|
|
High-ITGA3/CD9 and
High-CDK2/CDKN1A | 2.330 | 0.569 | 16.750 | <0.001 | 10.274 | 3.367 |
31.350 |
| Positive
margin | 1.919 | 0.686 |
7.835 | 0.005 |
6.817 | 1.778 |
26.139 |
PSR
The Cox proportional hazards model identified
double-positive (high-ITGA3/CD9 and
high-CDK2/CDKN1A) and positive margin to be
independent significant variables for PSR (Table IIIB). Although positive margin is a
clinical event following surgery, it was included in the analysis
due to its considerable influence on the risk for PSR. The rate of
PSR was represented by the K–M curves (one minus cumulative
survival) between groups of double-positive cases
(high-ITGA3/CD9 and high-CDK2/CDKN1A)
and the remaining negative cases according to size, ≤20 or >20
mm (Fig. 1B). The double-positive
group (high-ITGA3/CD9 and
high-CDK2/CDKN1A) >20 mm in size clearly showed a
higher rate of PSR independent of the primary therapeutic
intervention (positive margin), indicating diagnostic value of the
high-ITGA3/CD9 and high-CDK2/CDKN1A as
a biomarker at the outset of treatment planning.
Distant metastasis
The Cox proportional hazards model identified that
triple-positive (high-ITGB4/JUP,
high-ITGA3/CD9 and high-CDK2/CDKN1A)
was the only significant variable for distant metastasis (Table IIIC). Tumor size, histopathological
parameters and LNM did not exhibit a significant influence in
multivariate analysis in the presence of the triple-positive
(high-ITGB4/JUP, high-ITGA3/CD9 and
high-CDK2/CDKN1A) influence. The rate of distant
metastasis was presented by the K–M curves (one minus cumulative
survival) between the groups of triple-positive cases
(high-ITGB4/JUP, high-ITGA3/CD9 and
high-CDK2/CDKN1A) and the remaining negative cases
(Fig. 1C). Among all 77 cases, 4 of 5
cases that developed distant metastasis were extracted from 11
cases based on their triple-positive status
(high-ITGB4/JUP, high-ITGA3/CD9 and
high-CDK2/CDKN1A). A false-negative case had a tumor
size of 38 mm, suggesting the higher diagnostic reliability of the
triple-positive status (high-ITGB4/JUP,
high-ITGA3/CD9 and high-CDK2/CDKN1A) in
the early detection of distant metastasis.
DSD
The Cox proportional hazards model found
double-positive (high-ITGA3/CD9 and
high-CDK2/CDKN1A) and positive margin to be
independent significant variables (Table IIID). The cumulative survival was
represented by the K–M curves between the groups of double-positive
cases (high-ITGA3/CD9 and
high-CDK2/CDKN1A) and the remaining negative cases
according to size (≤20 or >20 mm) (Fig. 1D). The risk of DSD was significantly
higher in double-positive cases (high-ITGA3/CD9 and
high-CDK2/CDKN1A) with a small influence from tumor
size. Positive margin was associated with fatality in
double-positive cases (high-ITGA3/CD9 and
high-CDK2/CDKN1A) (data not shown).
Discussion
In our previous study, we demonstrated that there
are two OSCC metastatic traits: Simple LNM and uncontrollable
locoregional spread and distant metastasis via the circulation,
leading to fatality. We proposed that a biomarker system using
integrin gene expression ratios of high-ITGA3/CD9 for
locoregional spread and high-ITGB4/JUP for
hematogenous dissemination is effective for distinguishing these
two types of risks (3). With the aim
of improving diagnostic reliability, RT-qPCR was performed to
determine the expression levels of 11 genes of cyclin-related
proteins (cyclins, CDKs and CDKIs) for calculating the gene
expression ratios of 66 gene combinations and assessed their
diagnostic value in combination with the previously reported
integrin gene expression biomarkers. Four gene expression ratios
(ITGA3/CD9, ITGB4/JUP,
CDK1/CDKN1B and CDK2/CDKN1A) are
predictive of major clinical events with a high degree of
reliability in OSCC patients.
Two gene expression ratio patterns,
high-ITGA3/CD9 and high-CDK1/CDKN1B,
reflected the LNM risk with a high degree of reliability. While LNM
occurred in ~50% of OSCC cases, it may not respond to therapy in
certain cases. Therefore, predicting the risk of LNM with
biomarkers would be helpful in selecting optimal treatment. The
finding that <90% of patients with high-ITGA3/CD9
and high-CDK1/CDKN1B eventually had LNM irrespective
of the primary tumor size (Fig. 1A)
suggests that LNM can be detected before it becomes clinically
evident.
As PSR is strongly influenced by clinical factors
(such as site of the lesion and the extent of involvement within
the oral cavity) and factors associated with treatment (such as
positive margin), it is hard to conduct an accurate risk assessment
prior to starting therapy. In the present study, the analysis using
the Cox proportional hazard model showed that concurrent
high-ITGA3/CD9 and high-CDK2/CDKN1A and
positive surgical margins are significant predictors of PSR
(Table IIIB). This suggests that
biological propensities are important underlying risk factors for
PSR. By contrast, clear surgical margins reflect the relative
success of surgery and therefore strongly affect PSR, but it cannot
serve as a preoperative parameter for PSR risk assessment. As
demonstrated by the K–M curve for PSR, the combination of
high-ITGA3/CD9 and high-CDK2/CDKN1A
without consideration of another clinical factor estimates the risk
of PSR well (Fig. 1B), and the ratios
will therefore provide surgeons with useful information for
determining the extent of surgical resection.
Distant metastasis was identified in ~6.5% of
patients (Table I), which is lower
than the rate (~10%) observed in clinical settings. The low distant
metastasis rate may be due to our definition of distant metastasis,
in which distant metastasis following PSR was classified as PSR.
Achieving a highly reliable estimate of the risk of distant
metastasis based on tumor progression and histological findings is
difficult and numerous patients may be receiving unnecessary
postoperative chemotherapy due to this. The analysis using the Cox
proportional hazard model showed that none of the
clinicopathological parameters were significant risk factors for
distant metastasis, while the combination of
high-ITGB4/JUP, high-ITGA3/CD9 and
high-CDK2/CDKN1A was the only significant risk factor
for distant metastasis. This suggests that there is a biological
propensity for distant metastasis and it is difficult to predict
distant metastasis based on clinical parameters and histological
findings. Reliable analysis of distant metastasis risks will enable
appropriate selection of chemotherapy, molecular-targeted therapy
and/or immunotherapy for patients with highly malignant cancer in
the early stage when maximum therapeutic effect is expected. In
addition, it may eliminate unnecessary adjuvant chemoradiation in
the majority of patients.
The combination of positve margins,
high-ITGA3/CD9 and high-CDK2/CDKN1A
were shown to contribute to the risk of DSD. The influence of tumor
size on prognosis was relatively small. By contrast, as
demonstrated by analyses using the Cox proportional hazard model
and the K–M curve (Table IIID and
Fig. 1D), a malignant propensity
indicated by the status of high-ITGA3/CD9 and
high-CDK2/CDKN1A plus successful surgical resection
directly affected the prognosis.
The present study demonstrated that CDK gene
expression ratios, CDK1/CDKN1B and
CDK2/CDKN1A, may be used as parameters for assessing
the risk of LNM, PSR, distant metastasis and DSD reliably,
particularly in combination with integrin-related gene expression
ratios, ITGA3/CD9 and ITGB4/JUP, whose
value has been confirmed previously (3). The cell cycle is regulated by a
mechanism involving the coordinated actions of CDK, cyclin and CDKI
(8,10). Dysregulated function of these
molecules is the most frequently observed anomaly in malignant
tumors (7,8,11). CDK1 is
the key factor in the G2/M phase transition and can be
considered as a cell proliferation index. Upregulation of
CDK1 is observed in OSCC and its overexpression is
associated with malignant behavior. CDK1 is therefore expected to
be an indicator of poor prognosis (6,12). CDK2
plays various roles in DNA synthesis, modulation of G2
progression and in G1/S transition (13). CDK2 overexpression was shown to
be associated with malignant biological behavior and a poor
prognosis in patients with head and neck carcinoma (14). CDKN1B (p27) and CDKN1A (p21) are CDKIs
belonging to the Cip/Kip family that inhibit kinase activity of the
cyclin E/A-CDK2 complex, thereby blocking the G1/S
transition and inducing apoptosis (15,16).
Downregulation and increased degradation of these CDKIs were
observed in cancerous tissues and their association with malignant
behavior of squamous cell carcinoma was also demonstrated (17–19).
Specific associations of CDK-CDKI gene expression ratios
with major clinical events (LNM, PSR, distant metastasis and DSD)
suggest that cell cycle regulation mechanisms involving these
molecules have effects on invasion, dissemination and tumor
growth.
The present study demonstrated the feasibility of
using four combinations of CDK and integrin gene expression status
as a simple and relatively inexpensive assessment of the risk of
locoregional spread or hematogenous dissemination. Highly reliable
risk assessment prior to treatment enables selection of optimal
therapy and use of novel therapeutic strategies, thereby improving
control rates. Further studies using a large group of subjects and
prospective clinical studies, as well as improved understanding of
biological mechanisms, are essential for showing the clinical value
of the biomarker system and application in the clinical
setting.
Acknowledgements
The present study was supported by the Japan
Society for the Promotion of Science (grant no. 25463108). The
manuscript was translated and edited by ThinkSCIENCE K.K., Tokyo,
Japan.
Glossary
Abbreviations
Abbreviations:
|
OSCC
|
oral squamous cell carcinoma
|
|
LNM
|
lymph node metastasis
|
|
PSR
|
primary site recurrence
|
|
DSD
|
disease-specific death
|
|
K–M curve
|
Kaplan-Meier curve
|
|
CDK
|
cyclin-dependent kinase
|
|
CDKI
|
cyclin-dependent kinase inhibitor
|
|
RT-qPCR
|
reverse transcription-quantitative
polymerase chain reaction
|
References
|
1
|
International Agency for Research on
Cancer, . GLOBOCAN 2012: Estimated cancer incidence, mortality and
prevalence worldwide in 2012. http://globocan.iarc.fr/Pages/fact_sheets_population.aspxDecember
30–2014
|
|
2
|
NCCN, . Clinical practice guidelines in
oncology head and neck cancers. Version 2. http://www.nccn.org/professionals/physician_gls/f_guidelines.aspDecember
30–2014
|
|
3
|
Nagata M, Noman AA, Suzuki K, Kurita H,
Ohnishi M, Ohyama T, Kitamura N, Kobayashi T, Uematsu K, Takahashi
K, et al: ITGA3 and ITGB4 expression biomarkers estimate the risks
of locoregional and hematogenous dissemination of oral squamous
cell carcinoma. BMC Cancer. 13:4102013. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Nagata M, Fujita H, Ida H, Hoshina H,
Inoue T, Seki Y, Ohnishi M, Ohyama T, Shingaki S, Kaji M, et al:
Identification of potential biomarkers of lymph node metastasis in
oral squamous cell carcinoma by cDNA microarray analysis. Int J
Cancer. 106:683–689. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Kurokawa A, Nagata M, Kitamura N, Noman
AA, Ohnishi M, Ohyama T, Kobayashi T, Shingaki S and Takagi R:
Oral, Maxillofacial Pathology and Surgery Group: Diagnostic value
of integrin alpha3, beta4 and beta5 gene expression levels for the
clinical outcome of tongue squamous cell carcinoma. Cancer.
112:1272–1281. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Chen X, Zhang FH, Chen QE, Wang YY, Wang
YL, He JC and Zhou J: The clinical significance of CDK1 expression
in oral squamous cell carcinoma. Med Oral Patol Oral Cir Bucal.
20:e7–e12. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Bonelli P, Tuccillo FM, Borrelli A,
Schiattarella A and Buonaguro FM: CDK/CCN and CDKI alterations for
cancer prognosis and therapeutic predictivity. BioMed Res Int.
2014:3610202014. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Malumbres M and Barbacid M: Cell cycle
kinases in cancer. Curr Opin Genet Dev. 17:60–65. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Yamamoto E, Kohama G, Sunakawa H, Iwai M
and Hiratsuka H: Mode of invasion, bleomycin sensitivity, and
clinical course in squamous cell carcinoma of the oral cavity.
Cancer. 51:2175–2180. 1983. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Malumbres M: Physiological relevance of
cell cycle kinases. Physiol Rev. 91:973–1007. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Shintani S, Mihara M, Nakahara Y, Kiyota
A, Ueyama Y, Matsumura T and Wong DT: Expression of cell cycle
control proteins in normal epithelium, premalignant and malignant
lesions of oral cavity. Oral Oncol. 38:235–243. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Chang JT, Wang HM, Chang KW, Chen WH, Wen
MC, Hsu YM, Yung BY, Chen IH, Liao CT, Hsieh LL, et al:
Identification of differentially expressed genes in oral squamous
cell carcinoma (OSCC): Overexpression of NPM, CDK1 and NDRG1 and
underexpression of CHES1. Int J Cancer. 114:942–949. 2005.
View Article : Google Scholar : PubMed/NCBI
|
|
13
|
De Boer L, Oakes V, Beamish H, Giles N,
Stevens F, Somodevilla-Torres M, Desouza C and Gabrielli B: Cyclin
A/cdk2 coordinates centrosomal and nuclear mitotic events.
Oncogene. 27:4261–4268. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Dong Y, Sui L, Tai Y, Sugimoto K and
Tokuda M: The overexpression of cyclin-dependent kinase (CDK) 2 in
laryngeal squamous cell carcinomas. Anticancer Res. 21:103–108.
2001.PubMed/NCBI
|
|
15
|
Pérez-Sayáns M, Suárez-Peñaranda JM,
Gayoso-Diz P, Barros-Angueira F, Gándara-Rey JM and García-García
A: The role of p21Waf1/CIP1 as a Cip/Kip type cell-cycle regulator
in oral squamous cell carcinoma (Review). Med Oral Patol Oral Cir
Bucal. 18:e219–e225. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Kudo Y, Kitajima S, Ogawa I, Miyauchi M
and Takata T: Down-regulation of Cdk inhibitor p27 in oral squamous
cell carcinoma. Oral Oncol. 41:105–116. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Bandoh N, Hayashi T, Takahara M, Kishibe
K, Ogino T, Katayama A, Imada M, Nonaka S and Harabuchi Y: Loss of
p21 expression is associated with p53 mutations and increased cell
proliferation and p27 expression is associated with apoptosis in
maxillary sinus squamous cell carcinoma. Acta Otolaryngol.
125:779–785. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Ohashi K, Nemoto T, Eishi Y, Matsuno A,
Nakamura K and Hirokawa K: Expression of the cyclin dependent
kinase inhibitor p21WAF1/CIP1 in oesophageal squamous cell
carcinomas. Virchows Arch. 430:389–395. 1997. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Gao L, Gu W, Zheng J, Ren W, Chang S, Wang
X, Li S, Song T, Huang C and Zhi K: Clinicopathological and
prognostic significance of p27 expression in oral squamous cell
carcinoma: A meta-analysis. Int J Biol Markers. 28:e329–e335. 2013.
View Article : Google Scholar : PubMed/NCBI
|