Introduction
According to the World Health Organisation (WHO),
cancer is the 2nd leading cause of death worldwide causing 9
million deaths in 2016, 0.4 million of which were attributable to
alcohol consumption (4.2% of all cancer deaths) (1). The cancer burden is rising with an
incidence of 18.1 million in 2018 and a predicted increase to an
incidence of 29.5 million in 2040(2). Japan must also tackle this increasing
cancer burden with a reported 361,400 deaths attributable to cancer
in 2014 alone (3). The highest
incidence rates amongst men for lung cancer are seen in Eastern
Asia, including Japan with rates above 40 per 100,000(4). Incidence rates for known
alcohol-related cancers such as stomach, liver and colorectal are
markedly elevated in Eastern Asia (4). Researching the effect of genetic
polymorphisms in alcohol-related cancer risk may help to tackle
this global burden, as acknowledged by the WHO. Thus, research
moves forward with the anticipation that SNP investigation may
create genetic screening strategies identifying individuals at risk
of cancer to provide appropriate lifestyle and clinical advice
(5).
SNP rs1229984 is a missense variant located on exon
3 chr4:99318162 (GRCh38.p12) in the ADH1B gene on chromosome
4q23. It involves a single substitution of nucleotide cytosine (C)
to thymine (T), resulting in an amino acid change from arginine to
histidine in the β subunit of the ADH enzyme, hence this SNP
is also referred to as Arg48His mutation. Those with SNP rs1229984
can be said to have mutant ‘His allele’, with the wild-type
classified as ‘Arg allele’. Individuals with SNP rs1229984 (His
allele) metabolise ethanol to acetaldehyde 70- to 80-fold faster
than individuals without due to increased enzymatic function
(6). Acetaldehyde is a carcinogen
that can promote cancer development through multiple mechanisms,
thus any mutation that increases levels of acetaldehyde, such as
rs1229984, may confer increased cancer risk (7). Previous evidence explores the diverse
role of rs1229984 in alcohol metabolism, alcohol drinking
behaviours and in cancer risk (8).
With regard to the effect of rs1229984 on alcohol drinking
behaviours, evidence suggests that SNP presence confers a strongly
protective effect against alcohol dependence (9-11).
However, conclusions about rs1229984 and associations with cancer
are conflicting with some reporting SNP presence conferring
increased risk, decreased risk or no association with overall
cancer. This suggests a scientific need to clarify the role of
rs1229984 in cancer risk.
When looking at global allelic distributions of
rs1229984, research suggests that Arg allele is more prevalent than
His allele. Contrastingly, in the Japanese population mutant His
allele is more common conferring SNP rs1229984 presence. Overall,
SNP rs1229984 is most prevalent in the Eastern hemisphere and rarer
in Western populations. Thus, studies specifically in the Japanese
sub-population may be important in establishing the effect of
rs1229984 on cancer risk due to higher SNP prevalence here than in
the global population (12).
In the present study, rs1229984 status was
classified as genotypes Arg/Arg, Arg/His or His/His representing
wild-type homozygous; heterozygous or mutant homozygous individuals
respectively.
The primary aim of this study was to clarify any
association between SNP rs1229984 and overall cancer in a Japanese
population. Our secondary aim was to identify any associations
between rs1229984 and specific cancer phenotypes.
Patients and methods
Study participants
Experiments were conducted through genotyping DNA
samples from consecutive autopsy cases registered in the internet
database of Japanese single-nucleotide polymorphisms for geriatric
research (JG-SNP database). The JG-SNP database is a collection of
pathological data and samples from consecutive autopsy cases
employed in Tokyo Metropolitan Geriatric Medical Centre (Tokyo,
Japan) since 1995(13). Autopsies
were performed on 40% of all patients who died at the Tokyo
Metropolitan Geriatric Hospital between 1995-2004(14). Genomic DNA was extracted from the
kidney renal cortex of consecutive autopsy patients by
phenol-chloroform methods. Genomic DNA was isolated from kidney
cortex tissue by proteinase K digestion followed by
phenol-chloroform extraction. All pathological assessments at
autopsy and genotyping experiments were performed in a double-blind
manner for both pathologists and clinicians. Autopsy studies
included medical information, such as the presence of undiagnosed
latent cancers, thus providing a unique database for research on
genetic polymorphisms. At the time of sampling in 2004
approximately 1,800 participants were present in the database for
use in this study. A total of 1,359 samples remained for use in
this study after accounting for loss of samples, poor DNA quality
(checked via gel electrophoresis) and contamination of samples.
Totally, 1,359 consecutive autopsy cases were analysed.
Cancer phenotype data of participants was also
extracted from the JG-SNP database. Patients were defined as
‘cases’ or ‘controls’ depending on cancer status. Participants were
defined as ‘cases’ if 1 or more cancer sites were recorded or
‘controls’ if the recorded value was 0 where no cancer was
identified. Cases and controls were unmatched.
The proportion of different diseases in our study
participants did not greatly differ from those in the census data
of Ministry of Health, Labour and Welfare (Tokyo, Japan) (15), allowing the geriatric autopsy samples
to be validly used in genotyping analysis.
Informed consent for the use of autopsy samples and
patient clinical data for this study was obtained from the family
of study participants at the time of autopsy. Clinical data such as
drinking and smoking habit were extracted from patient medical
records with informed consent. The study protocol was approved by
The Ethics Committees of Tokyo Geriatric Hospital and Tokyo Medical
and Dental University and authorized by TMDU Research Ethics
Committee (approval no. 2016-011-02).
Sample preparation and genotyping
All autopsy cases were eligible if genomic DNA
samples were available of adequate quality. DNA quality was checked
via gel electrophoresis and random sampling before genotyping.
Since the concentration of the DNA can affect the genotyping
results, we quantified the concentration using Thermo Scientific
NanoDrop One™ for all the samples used in our study. In a pilot
study, RT-PCR was performed on diluted stock DNA samples to assess
different concentrations of DNA. This showed concentrations of
2.5-5 ng/µl yielded best genotyping results. All 1,359 samples were
mapped to 384 plates and diluted samples were transferred using a
multichannel pipette-man. This experiment used a dry genotyping
protocol according to the manufacturer's guidelines (Thermo Fisher
Scientific, Inc.) thus plates were dried overnight in a
dehumidifier (16). The reaction
mixture was made by adding TaqMan Drug Metabolism Genotyping Assay
(ID C—2688467_20) to the 384 plates. This genotyping assay
contained two TaqMan MGB probes and forward and reverse primers.
Each probe had fluorescent reporter dye VIC or FAM attached to the
5'end and a non-fluorescent quencher and MGB molecule bound to the
3'end. The probe with VIC dye bound to the complimentary region on
the target DNA strand for Arg allele/nucleotide C (wild-type). The
probe with FAM dye bound to the complimentary region on the DNA
strand for His allele/nucleotide T (SNP presence). This
substitution polymorphism can be denoted as context sequence
(VIC/FAM): GCCACTAACCACGTGGTCATCTGTG[C/T]GAC
AGATTCCTACAGCCACCATCTA. Further information regarding sequences of
primers and probes was requested from manufacturers, who stated
that these are commercial secrets and only the assay ID is
available (assay ID: C-2688467_20; Thermo Fisher Scientific, Inc.).
A total of 2.5 µl of TaqMan Universal PCR Master Mix, 0.25 µl of
20X SNP genotyping assay and 2.25 µl of DNase-free water was added
to the dried 384 plates, creating total 5 µl reaction mixture. The
samples were mixed using a microplate mixer and spun down in a
centrifuge. Centrifuging was performed at room temperature at 1,500
rpm/min for 5 min. The reaction mixture is photosensitive, so light
exposure was avoided. The plates were transferred to LightCycler
480 instrument (Roche Diagnostics, Penzberg, Germany) to run
qualitative RT-PCR. PCR settings were specified in the
manufacturer's guide for a total of 50 cycles as follows: i) Pre
incubation: 90 degrees for 10 min; ii) denaturation: 92 degrees for
15 sec and iii) annealing and extension: 60 degrees for 1 min.
Qualitative results were generated by the
thermocycler through detection of fluorescent light emitted by both
reporter dyes VIC and FAM, where the wavelength was ~551 and ~517
nm, respectively. This allowed samples to be genotyped where green
light emission (~551 nm) indicated the sample was of genotype CC,
where blue was genotype TT (~517 nm) and where red was heterozygous
CT (17). All 1,359 samples obtained
were genotyped leading to a 100% genotyping success rate. All
primers and probes were purchased from Thermo Fisher Scientific,
Inc.
Statistical analysis
Statistical analysis was performed blinded post
genotyping using SPSS version 19 (IBM Corp). Two statistical tests
were used to compare baseline characteristics between cases and
controls. Statistical significance in categorical variables was
evaluated using the Chi-square test and continuous variables were
evaluated using analysis of variance (ANOVA). The Bonferroni
correction used to adjust for multiple testing was not used in
statistical analysis due to its conservative nature (18). This correction creates a more
stringent criterion for ‘statistical significance’ by adjusting
probability values (P), aiming to reduce risk of type I error when
performing multiple statistical analyses. This is done by adjusting
the conventional P<0.05 for population size. However, use can
eliminate important significant findings and lead to an increase in
type II errors. Therefore, in this study statistical significance
was defined as results with P<0.05 in accordance with
conventional standards. Hardy-Weinberg Equilibrium (HWE) was
calculated to determine if genotype frequencies observed in the
study population differed from expected population frequencies.
Odds ratios (OR) and 95% confidence intervals (CI) were calculated
using multinomial regression analysis to determine any association
between genotype status and odds of cancer, in both overall and
specific cancers. Confounding factors such as sex, smoking status,
drinking status, and age at death were adjusted for in regression
analysis, generating crude and adjusted ORs.
Results
Patient demographics
General demographics of the study population
including age at death, sex, smoking status and drinking status are
described in Table I. The study
population comprised of 816 (60%) cases and 543 (40%) controls.
There were 732 (54%) males and 627 (46%) females with a mean age at
death of 80.1 (±8.87) (Table I).
 | Table IGeneral demographics of study
participants. |
Table I
General demographics of study
participants.
| Characteristics | Total number (%) | Number of cases
(%) | Number of controls
(%) | P-valuea |
|---|
| Number | 1,359 | 816(60) | 543(40) | |
| Ageb | 80.1 (±8.87) | 79.7 (±8.46) | 80.6 (±9.62) | 0.11 |
| Sex | | | | |
|
Male | 732(54) | 462(37) | 270(63) | |
|
Female | 627(46) | 354(44) | 273(56) | 0.01a |
| Smoking status | | | | |
|
Smoker | 650(48) | 408(50) | 242(45) | |
|
Non-smoker | 624(46) | 369(45) | 255(47) | 0.18 |
|
Unknown | 85(6) | 39(5) | 46 | |
| Alcohol
consumption | | | | |
|
Drinker | 452(33) | 296(36) | 156(29) | |
|
Non-drinker | 823(61) | 480(59) | 343(63) | 0.01a |
|
Unknown | 84(6) | 40(5) | 44(8) | |
‘Smokers’ were defined as patients who have ever
smoked, and ‘non-smokers’ as those who have never smoked. The
number of smokers was 650 (48%) and number of non-smokers was 624
(46%). There was no significant difference in smoking status
between cases and controls as P=0.1837 (Table I).
‘Drinkers’ were defined as those who have ever
consumed alcohol and ‘non-drinkers’ as those who have never
consumed alcohol. The number of drinkers was 452 (33%) and number
of non-drinkers was 823 (61%). Results showed drinking status was
significantly different between cases and controls as P=0.0122
(Table I).
Overall, sex (P=0.0125) and alcohol consumption
(P=0.0122) were the only variables significantly different between
cases and controls as shown in bold (Table I). All other variables showed no
significant difference between cases and controls as P>0.05
(Table I).
Genotyping results and allelic
counts
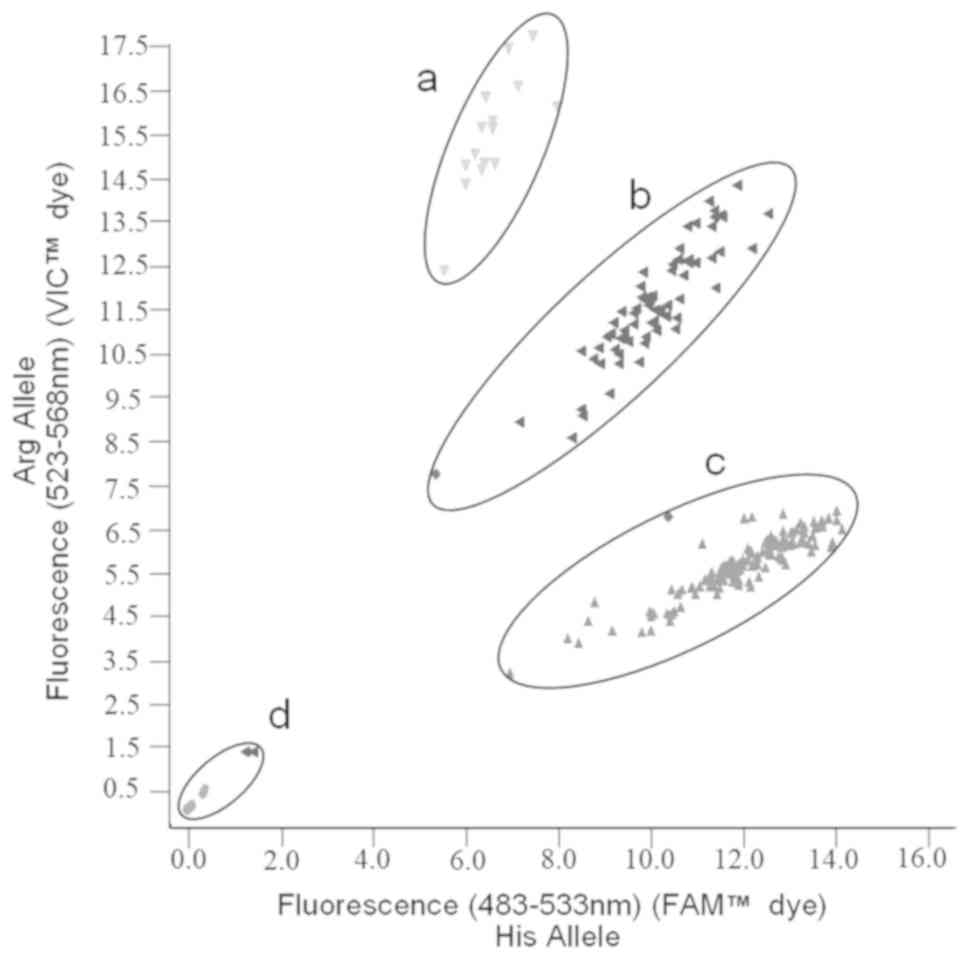
A typical TaqMan assay scatter plot for SNP
rs1229984 is shown in Fig. 1 as
obtained from the LightCycler after RT-PCR. Each dot represents one
sample (an individual participant), and the X and Y axes equate to
the levels of fluorescence of two dyes FAM and VIC for His allele
(SNP presence) and Arg allele (wild-type), respectively.
Identification of both alleles for each sample was calculated based
on fluorescence levels from the LightCycler. Samples with only a
substantial increase in FAM dye fluorescence conferred homozygosity
for His allele. A substantial increase in only VIC dye fluorescence
conferred homozygosity for Arg allele and substantial increase in
both fluorescence signals conferred heterozygosity for alleles
Arg/His. Thus, region A represents samples of alleles Arg/Arg
(homozygous wild-type), region B of alleles Arg/His (heterozygous
SNP) and region C of alleles His/His (homozygous SNP). Ambiguous
samples or those that did not sufficiently fluoresce were repeated
until clear separation was obtained, as represented by region
D.
Information regarding overall genotyping results and
allelic counts is provided in Table
II. As expected, genotypes His/His and Arg/His were most
prevalent in this Japanese population with overall genotype
frequencies as follows-His/His: 799 (59%); Arg/His: 486 (36%);
Arg/Arg: 74 (5%). Allele frequencies for His allele and Arg allele
were calculated using the Hardy-Weinberg equations: p² + 2pq +
q²=1; p + q=1, where p is defined as the frequency of His allele
and q as the frequency of Arg allele (since SNP rs1229984 is
controlled by a pair of alleles). This showed that 1042 (77%)
individuals had His allele and 317 (23%) had Arg allele. Our
results matched allele frequency data in Japanese SNP database HGVD
(Human Genetic Variation Database) (19). Results also suggest the study
population is in Hardy-Weinberg Equilibrium (HWE) since the P-value
was >0.05 so observed allele frequencies in this study
population do not greatly differ from the expected frequencies.
This suggests the distribution of alleles in the population is
unlikely due to chance so further association analysis are
interpretable (20). Overall,
results were in accordance with previous data on allelic
distributions of SNP rs1229984 that suggest reference His allele is
more prevalent than alternate Arg allele in Japanese populations,
differing from the worldwide distribution (12).
 | Table IIOverall genotyping results and
allelic counts for SNP rs1229984 in all participants. |
Table II
Overall genotyping results and
allelic counts for SNP rs1229984 in all participants.
| SNP | Alleles | Genotypes | P-value
(HWE)a |
|---|
| rs1229984 | His | Arg | His/His | Arg/His | Arg/Arg | 0.993 |
| No (%) | 1,042(77) | 317(23) | 799(59) | 486(36) | 74(5) | |
Association analysis using genetic
models
Results of multinomial logistic regression analysis
exploring the association between genotype status and odds of
cancer are shown in Tables III and
IV. Association analyses were
performed for both overall cancer and all specific cancer
phenotypes.
 | Table IIIMultinomial logistic regression
association analysis using dominant inheritance genetic model
(Arg/Arg: Arg/His+His/His). |
Table III
Multinomial logistic regression
association analysis using dominant inheritance genetic model
(Arg/Arg: Arg/His+His/His).
| | Crude
ORa | Adjusted
ORb |
|---|
| Cancer
phenotype | Genotype | OR |
P-valuec | OR |
P-valuec |
|---|
| | Arg/Arg | 1.00
(reference) | N/A | 1.00
(reference) | N/A |
| Total Cancer |
Arg/His+His/His | 0.7
(0.42-1.15) | 0.129 | 0.66
(0.39-1.13) | 0.129 |
| Gastric |
Arg/His+His/His | 1.78
(0.7-4.51) | 0.231 | 1.77 (0.7-4.5) | 0.231 |
| Lung |
Arg/His+His/His | 0.96
(0.43-2.15) | 0.708 | 0.85
(0.38-1.94) | 0.708 |
| Colon |
Arg/His+His/His | 0.62
(0.29-1.34) | 0.480 | 0.74
(0.33-1.69) | 0.480 |
| Pancreatic |
Arg/His+His/His | 0.65
(0.23-1.87) | 0.321 | 0.58
(0.2-1.69) | 0.321 |
| Liver |
Arg/His+His/His | 1.04
(0.31-3.43) | 0.975 | 1.02
(0.31-3.4) | 0.975 |
 | Table IVMultinomial logistic regression
association analysis using recessive inheritance genetic model
(Arg/Arg+Arg/His: His/His). |
Table IV
Multinomial logistic regression
association analysis using recessive inheritance genetic model
(Arg/Arg+Arg/His: His/His).
| | Crude
ORa | Adjusted
ORb |
|---|
| Cancer
phenotype | Genotype | OR, (95% CI) |
P-valuec | OR, (95% CI) |
P-valuec |
|---|
| |
Arg/Arg+Arg/His | 1.00
(reference) | N/A | 1.00
(reference) | N/A |
| Total Cancer | His/His | 0.99
(0.79-1.23) | 0.901 | 0.95
(0.75-1.20) | 0.657 |
| Gastric | His/His | 1.21
(0.86-1.71) | 0.264 | 1.16
(0.82-1.64) | 0.399 |
| Lung | His/His | 0.7
(0.48-1.00) | 0.051 | 0.64
(0.44-0.93) | 0.020 |
| Colon | His/His | 1.1
(0.72-1.66) | 0.662 | 1.08
(0.71-1.65) | 0.724 |
| Pancreatic | His/His | 0.88
(0.49-1.59) | 0.678 | 0.95
(0.51-1.77) | 0.877 |
| Liver | His/His | 1.12
(0.63-1.99) | 0.705 | 1.07
(0.6-1.91) | 0.814 |
Results were calculated using dominant inheritance
and recessive inheritance genetic models to account for different
modes SNP of inheritance. Confounders were adjusted for in odds
ratio (OR) calculations with adjustment for only age at death and
sex in crude OR, and adjustment for age at death; sex; smoking
status and drinking status in adjusted OR. Cancer phenotypes
reported are shown in order of decreasing sample size. Further
association analysis for cancer phenotypes with low prevalence have
not been reported in detail due to low power.
Genotype homozygous Arg/Arg was used as the
reference for comparison to other genotypes (Arg/His, His/His) in
the dominant genetic model. Results show there was no significant
difference in odds of overall cancer when comparing other genotypes
(Arg/His, His/His) to reference Arg/Arg since Arg/Arg:
Arg/His+His/His adjusted OR=0.66 (95% CI=0.39-1.13, P=0.129).
Results for individual cancer risk in all cancer phenotypes
similarly suggested there was no significant difference in odds of
specific cancer when comparing Arg/Arg: Arg/His+His/His using a
dominant model. The results for stomach, lung, colon, pancreatic
and liver cancer are presented in further detail below for
reference (Tables III and IV).
The genotypes Arg/Arg+Arg/His were used as the
reference for comparison to genotype His/His in the recessive
genetic model. Results show there is no significant difference in
odds of cancer overall when comparing genotype His/His to reference
Arg/Arg+Arg/His since Arg/Arg+Arg/His: His/His OR=0.95 (95%
CI=0.75-1.20, P=0.657). However, adjusted results show there is a
significant decrease in odds of lung cancer when comparing genotype
His/His to reference Arg/Arg+Arg/His since Arg/Arg+Arg/His: His/His
OR=0.64 (95% CI=0.44-0.93, P=0.020). This suggests that individuals
who are genotype His/His (homozygous for rs1229984) may be at
decreased risk of lung cancer compared to those of genotypes
Arg/His or Arg/Arg. However, this result must be interpreted with
caution due to a low number of lung cancer cases. No other
significant associations with specific cancer phenotypes were found
across all models (Tables III and
IV).
In all examined models there was no significant
association between rs1229984 and overall odds of cancer nor odds
of specific cancer phenotypes, except in the case of lung cancer in
a recessive genetic model where a significant decrease in odds was
found in those homozygous for the SNP (genotype His/His).
Discussion
This study primarily aimed to clarify the
association between single nucleotide polymorphism rs1229984 in the
ADH1B gene and both overall and specific cancer risk in a
Japanese population.
Detailed results for the following cancers were
presented: total cancer, stomach, lung, colon, pancreatic and
liver. Analyses of liver, pancreas and colon cancer were presented
due to evidence that these are alcohol-related cancers (21). Stomach cancer is presented due to
previous investigations into its association with rs229984
(22-25).
Lung cancer is presented due to adequate sample size and
significant findings. However, results for individual lung cancer
sub-types have not been presented due to inadequate sample size and
reduced power.
Overall, results showed no statistically significant
association between SNP rs1229984 in both overall and specific
cancer risk, except for lung cancer in a recessive genetic model
where results suggested homozygous SNP presence (genotype His/His)
may decrease cancer risk.
A recent meta-analysis found no significant
association between rs1229984, colorectal, hepatocellular, stomach
nor pancreatic cancer supporting our findings of no association
between rs1229984 and specific cancer phenotypes in this study
(26). However, lung cancer was not
explored in this meta-analysis due to low reporting in studies
suggesting research into associations with lung cancer require
larger sample sizes (26). The true
effect of ADH gene SNPs on alcohol metabolism is debated.
Birley et al suggest SNPs in the ADH region have
lower effects on alcohol metabolism than previously expected
(27) so the overall effect of
rs1229984 on cancer risk may be minimal or non-existent. This
evidence supports our finding of a lack of association between
rs1229984 and overall and specific cancer risk but is rarely
replicated in the literature perhaps due to publication bias.
Alternatively, this lack of association may be
masked by the fact that single SNPs have little effect in the
predisposition to complex traits, such as cancer, and that hundreds
to thousands of loci are likely involved (8). Combining results for multiple loci may
reveal the synergistic effect of multiple SNPs in alcohol
metabolism and cancer risk. Indeed, the role of other ADH1B
and ALDH2 variants remains an area of interest (28). For example, the HapMap project
estimates there are at least 449 polymorphic variants in the
ADH region which may be involved in ADH gene
expression, many of which have not been studied (29). Another author suggests rs1229984 may
be in linkage disequilibrium with other unidentified ADH
regions since it has previously been shown to be in linkage
disequilibrium with ADH1C sites (30). This suggests multiple variants may
influence the expression of rs1229984 and consequently affect
cancer risk.
Evidence regarding the effect of rs1229984 on
specific cancer risk has been contradictory. SNP presence may
confer increased risk, decreased risk or no association cancer risk
(31), whilst others argue the
variant confers increased risk (32-34).
As mentioned prior, the mechanism of how this variant may affect
cancer risk is poorly understood but may be due to altered
enzymatic function of ADH1B in individuals with
rs1229984(6). Some studies suggest
that rs1229984 presence may confer a protective effect against only
alcohol-related cancers, such as oesophageal cancer. In one study,
Arg/Arg individuals (homozygous wild type) had a 3.99-fold
increased risk of developing oesophageal cancer compared with
His/His individuals (homozygous rs1229984) (35). This reduced risk may be explained by
the theory that those with rs1229984 are more likely to be alcohol
adverse or abstinent due to unpleasant side effects of acetaldehyde
(36). Some suggest the interplay of
ADH1B genotypes with levels of alcohol consumption may
modulate oesophageal cancer risk (37). Despite evidence of this protective
effect of rs1229984, findings have not been widely reproducible and
meta-analyses are conflicting perhaps due to few robust studies and
publication bias in the field (38).
The relationship between levels of alcohol
consumption and cancer risk is complex. Epidemiological evidence
has widely associated alcohol consumption with specific cancer risk
with the International Agency for Research on Cancer (IARC)
categorising the secondary metabolite acetaldehyde as ‘carcinogenic
to humans’ in Group 1(39). A
dose-response relationship between alcohol consumption and cancer
risk has been well established (21). However, in this study few samples had
complete alcohol data and so drinking status was stratified only as
‘drinkers’ or ‘non-drinkers’, disallowing association analysis
between levels of alcohol consumption, genotype and cancer risk.
Thus, associations may have been missed as quantitative amount of
drinking were not used in analyses potentially leading to exposure
misclassification of participants.
Our finding that rs1229984 presence may confer a
protective effect against lung cancer may be influenced by unknown
factors such as levels of alcohol consumption. Indeed, results
showed a significant difference in cancer prevalence between
drinkers and non-drinkers suggesting levels of alcohol intake may
be important (Table I). Evidence
suggests that heavy drinkers may be at increased risk of lung
cancer than moderate or never drinkers (21). The proposed mechanism is that high
concentrations of ethanol exposure may cause decreased NK cell
activity and immune suppression, leading to increased lung
metastases (40). However, research
into the association between levels of alcohol consumption and
specifically lung cancer risk is limited. Overall, the effect of
SNPs on specific cancer risk must be further researched before
drawing conclusions.
This study had strengths and limitations. Selection
bias may have been introduced as participants were consecutive
autopsy cases meaning cause of admission or death may not be
randomised. The contribution of any gene to cancer risk is
modulated by many other social and environmental factors.
Therefore, there may be uncontrolled confounders such as diet,
levels of alcohol consumption, occupational status, socioeconomic
status or ethnicity that may affect both the expression of
rs1229984 and overall cancer risk. Indeed, rs1229984 has shown
reduced cancer risk amongst Asians and mixed ethnicity groups, but
increased risk amongst Caucasians suggesting there may be ethnic
differences in alcohol metabolism and cancer risk (26). Measurement bias was reduced through
machine genotype identification; however, autopsy practice may have
differed between pathologists leading to potential
misclassification of cancer outcomes. Lifestyle data was collected
through a self-reported questionnaire from medical records leading
to potential recall bias and inaccuracy of data. Cases were
unmatched to controls meaning unknown confounders may not have been
controlled for. Results may not be generalizable to other global
populations or age at death groups, as this study focused on
elderly Japanese population. Results may also not be generalizable
to any association between rs1229984 and other diseases such as as
alcohol use disorders (11) which
has been extensively studied, or to non-alcohol-related disease
(41). To our knowledge this is the
first study looking more specifically at associations between
rs1229984 in elderly Japanese population and cancer risk.
In conclusion, this study suggested there was no
significant association between SNP rs1229984 and overall or
specific cancer risk, except in the case of lung cancer where
results suggested homozygous SNP presence (genotype His/His)
decreased overall lung cancer risk.
Acknowledgements
The authors would like to thank Professor Graham
Cooke of Public Health at Imperial College London who contributed
in the writing and editing of the manuscript.
Funding
The present study was funded by a grant from
GMEXT/JSPS (grant no. 17K09081) and The Joint Usage/Research
Program of the Medical Research Institute, TMDU.
Availability of data and materials
All datasets used or analysed for this study are
available from the corresponding author upon reasonable
request.
Authors' contributions
The present study was designed by MM. Sample
collection and preparation was performed by MS and TA. MS and TA
conducted the autopsy and pathological analyses for all samples
used. Experiments and statistical analyses were performed by PG and
SP. The manuscript was written by PG with contributions from SP and
MM. All authors read and approved of the final manuscript.
Ethics approval and consent to
participate
Informed consent for the use of autopsy samples and
patient clinical data for this study was obtained from the family
of study participants at the time of autopsy. Clinical data such as
drinking and smoking habit were extracted from patient medical
records with informed consent. The study protocol was approved by
The Ethics Committees of Tokyo Geriatric Hospital and Tokyo Medical
and Dental University (Tokyo, Japan) and authorized by TMDU
Research Ethics Committee (approval no. 2016-011-02).
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
World Health Organisation: Global status
report on alcohol and health, 2018.
|
|
2
|
International Association of Cancer
Registries GI for CRD: Cancer Tomorrow-IARC, 2018.
|
|
3
|
World Health Organisation: Cancer Country
Profiles 2014, 2014. http://www9.who.int/cancer/country-profiles/JPN_2020.pdf.
Accesed April 18, 2019.
|
|
4
|
Bray F, Ferlay J, Soerjomataram I, Siegel
RL, Torre LA and Jemal A: Global cancer statistics 2018: GLOBOCAN
estimates of incidence and mortality worldwide for 36 cancers in
185 countries. CA Cancer J Clin. 68:394–424. 2018.PubMed/NCBI View Article : Google Scholar
|
|
5
|
World Health Organisation: Human Genomics
in Global Health, Genes and human diseases. Genes and
noncommunicable diseases, 2019. https://www.who.int/genomics/public/geneticdiseases/en/index3.html.
|
|
6
|
Edenberg HJ: The genetics of alcohol
metabolism: Role of alcohol dehydrogenase and aldehyde
dehydrogenase variants. Alcohol Res Health. 30:5–13.
2007.PubMed/NCBI
|
|
7
|
Seitz HK and Becker P: Alcohol metabolism
and cancer risk. Alcohol Res Health. 38:38–47. 2007.PubMed/NCBI
|
|
8
|
Polimanti R and Gelernter J: ADH1B: From
alcoholism, natural selection, and cancer to the human phenome. Am
J Med Genet B Neuropsychiatr Genet. 177:113–125. 2018.PubMed/NCBI View Article : Google Scholar
|
|
9
|
Bierut LJ, Goate AM, Breslau N, Johnson
EO, Bertelsen S, Fox L, Agrawal A, Bucholz KK, Grucza R,
Hesselbrock V, et al: ADH1B is associated with alcohol dependence
and alcohol consumption in populations of European and African
ancestry. Mol Psychiatry. 17:445–450. 2012.PubMed/NCBI View Article : Google Scholar
|
|
10
|
Edenberg HJ and McClintick JN: Alcohol
dehydrogenases, aldehyde dehydrogenases, and alcohol use disorders:
A critical review. Alcohol Clin Exp Res. 42:2281–2297.
2018.PubMed/NCBI View Article : Google Scholar
|
|
11
|
Tawa EA, Hall SD and Lohoff FW: Overview
of the genetics of alcohol use disorder. Alcohol Alcohol.
51:507–514. 2016.PubMed/NCBI View Article : Google Scholar
|
|
12
|
U.S National Library of Medicine NC for
BI: dbSNP Short Genetic Variations, Reference SNP (rs) Report,
rs1229984, 2018.
|
|
13
|
Sawabe M, Arai T, Kasahara I, Esaki Y,
Nakahara K, Hosoi T, Orimo H, Takubo K, Murayama S and Tanaka N:
Developments of geriatric autopsy database and internet-based
database of Japanese single nucleotide polymorphisms for geriatric
research (JG-SNP). Mech Ageing Dev. 125:547–552. 2004.PubMed/NCBI View Article : Google Scholar
|
|
14
|
Parlayan C, Ikeda S, Sato N, Sawabe M,
Muramatsu M and Arai T: Association analysis of single nucleotide
polymorphisms in miR-146a and miR-196a2 on the prevalence of cancer
in elderly Japanese: A case-control study. Asian Pac J Cancer Prev.
15:2101–2107. 2014.PubMed/NCBI View Article : Google Scholar
|
|
15
|
Oda K, Tanaka N, Arai T, Araki J, Song Y,
Zhang L, Kuchiba A, Hosoi T, Shirasawa T, Muramatsu M and Sawabe M:
Polymorphisms in pro- and anti-inflammatory cytokine genes and
susceptibility to atherosclerosis: A pathological study of 1503
consecutive autopsy cases. Hum Mol Genet. 16:592–599.
2007.PubMed/NCBI View Article : Google Scholar : Access date: April
18, 2019.
|
|
16
|
Life Technologies Corporation:
TaqMan® SNP Genotyping Assays, 2014. http://http://tools.thermofisher.com/content/sfs/manuals/TaqMan_SNP_Genotyping_Assays_man.pdf.
|
|
17
|
Thermo Fisher Scientific Inc: TaqMan
multiplex real-time PCR, 2016. https://www.thermofisher.com/content/dam/LifeTech/Documents/PDFs/PG1600-PJT1731-COL12035-TaqManMultiplex-qPCR-ProdBulletin-Global-FHR.pdf.
|
|
18
|
Green J and Britten N: Qualitative
research and evidence based medicine. BMJ. 316:1230–1232.
1998.PubMed/NCBI View Article : Google Scholar
|
|
19
|
Higasa K, Miyake N, Yoshimura J, Okamura
K, Niihori T, Saitsu H, Doi K, Shimizu M, Nakabayashi K, Aoki Y, et
al: Human genetic variation database, a reference database of
genetic variations in the Japanese population. J Hum Genet.
61:547–553. 2016.PubMed/NCBI View Article : Google Scholar
|
|
20
|
Wigginton JE, Cutler DJ and Abecasis GR: A
note on exact tests of Hardy-Weinberg equilibrium. Am J Hum Genet.
76:887–893. 2005.PubMed/NCBI View
Article : Google Scholar
|
|
21
|
Bagnardi V, Rota M, Botteri E, Tramacere
I, Islami F, Fedirko V, Scotti L, Jenab M, Turati F, Pasquali E, et
al: Alcohol consumption and site-specific cancer risk: A
comprehensive dose-response meta-analysis. Br J Cancer.
112:580–593. 2015.PubMed/NCBI View Article : Google Scholar
|
|
22
|
Wang HL, Zhou PY, Liu P and Zhang Y: ALDH2
and ADH1 genetic polymorphisms may contribute to the risk of
gastric cancer: A meta-analysis. PLoS One. 9(e88779)2014.PubMed/NCBI View Article : Google Scholar
|
|
23
|
Jelski W, Chrostek L, Zalewski B and
Szmitkowski M: Alcohol dehydrogenase (ADH) isoenzymes and aldehyde
dehydrogenase (ALDH) activity in the sera of patients with gastric
cancer. Dig Dis Sci. 53:2101–2105. 2008.PubMed/NCBI View Article : Google Scholar
|
|
24
|
Ghosh S, Bankura B, Ghosh S, Saha ML,
Pattanayak AK, Ghatak S, Guha M, Nachimuthu SK, Panda CK, Maji S,
et al: Polymorphisms in ADH1B and ALDH2 genes associated with the
increased risk of gastric cancer in West Bengal, India. BMC Cancer.
17(782)2017.PubMed/NCBI View Article : Google Scholar
|
|
25
|
Hidaka A, Sasazuki S, Matsuo K, Ito H,
Sawada N, Shimazu T, Yamaji T, Iwasaki M, Inoue M and Tsugane S:
JPHC Study Group. Genetic polymorphisms of ADH1B, ADH1C and ALDH2,
alcohol consumption, and the risk of gastric cancer: The Japan
Public Health Center-based prospective study. Carcinogenesis.
36:223–231. 2015.PubMed/NCBI View Article : Google Scholar
|
|
26
|
Tan B and Ning N: Association of
ADH1BArg47His polymorphism with the risk of cancer: A
meta-analysis. Biosci Rep. 39: pii(BSR20181915)2019.PubMed/NCBI View Article : Google Scholar
|
|
27
|
Birley AJ, James MR, Dickson PA,
Montgomery GW, Heath AC, Martin NG and Whitfield JB: ADH single
nucleotide polymorphism associations with alcohol metabolism in
vivo. Hum Mol Genet. 18:1533–1542. 2009.PubMed/NCBI View Article : Google Scholar
|
|
28
|
Crabb DW, Matsumoto M, Chang D and You M:
Overview of the role of alcohol dehydrogenase and aldehyde
dehydrogenase and their variants in the genesis of alcohol-related
pathology. Proc Nutr Soc. 63:49–63. 2004.PubMed/NCBI View Article : Google Scholar
|
|
29
|
Edenberg HJ and Bosron WF: Alcohol
dehydrogenases. Comprehensive Toxicol. 111–130. 2010.
|
|
30
|
Osier MV, Pakstis AJ, Soodyall H, Comas D,
Goldman D, Odunsi A, Okonofua F, Parnas J, Schulz LO, Bertranpetit
J, et al: A global perspective on genetic variation at the ADH
genes reveals unusual patterns of linkage disequilibrium and
diversity. Am J Hum Genet. 71:84–99. 2002.PubMed/NCBI View
Article : Google Scholar
|
|
31
|
Mao N, Nie S, Hong B, Li C, Shen X and
Xiong T: Association between alcohol dehydrogenase-2 gene
polymorphism and esophageal cancer risk: A meta-analysis. World J
Surg Oncol. 14(191)2016.PubMed/NCBI View Article : Google Scholar
|
|
32
|
Zhang G, Mai R and Huang B: ADH1B Arg47His
polymorphism is associated with esophageal cancer risk in
high-incidence Asian population: Evidence from a meta-analysis.
PLoS One. 5(e13679)2010.PubMed/NCBI View Article : Google Scholar
|
|
33
|
Guo H, Zhang G and Mai R: Alcohol
dehydrogenase-1B Arg47His polymorphism and upper aerodigestive
tract cancer risk: A meta-analysis including 24,252 subjects.
Alcohol Clin Exp Res. 36:272–278. 2012.PubMed/NCBI View Article : Google Scholar
|
|
34
|
McKay JD, Truong T, Gaborieau V, Chabrier
A, Chuang SC, Byrnes G, Zaridze D, Shangina O, Szeszenia-Dabrowska
N, Lissowska J, et al: A genome-wide association study of upper
aerodigestive tract cancers conducted within the INHANCE
consortium. PLoS Genet. 7(e1001333)2011.PubMed/NCBI View Article : Google Scholar
|
|
35
|
Chen YJ, Chen C, Wu DC, Lee CH, Wu CI, Lee
JM, Goan YG, Huang SP, Lin CC, Li TC, et al: Interactive effects of
lifetime alcohol consumption and alcohol and aldehyde dehydrogenase
polymorphisms on esophageal cancer risks. Int J Cancer.
119:2827–2831. 2006.PubMed/NCBI View Article : Google Scholar
|
|
36
|
Li D, Zhao H and Gelernter J: Strong
association of the alcohol dehydrogenase 1B gene (ADH1B) with
alcohol dependence and alcohol-induced medical diseases. Biol
Psychiatry. 70:504–512. 2011.PubMed/NCBI View Article : Google Scholar
|
|
37
|
Lee CH, Lee JM, Wu DC, Goan YG, Chou SH,
Wu IC, Kao EL, Chan TF, Huang MC, Chen PS, et al: Carcinogenetic
impact of ADH1B and ALDH2 genes on squamous cell carcinoma risk of
the esophagus with regard to the consumption of alcohol, tobacco
and betel quid. Int J Cancer. 122:1347–1356. 2008.PubMed/NCBI View Article : Google Scholar
|
|
38
|
Lee YH: Meta-analysis of genetic
association studies. Ann Lab Med. 35:283–287. 2015.PubMed/NCBI View Article : Google Scholar
|
|
39
|
International Agency for Research on
Cancer, World Health Organisation: IARC Monographs on the
Identification of Carcinogenic Hazards to Humans, 2019. https://monographs.iarc.fr/list-of-classifications.
Accessed April 18. 2019.
|
|
40
|
Ben-Eliyahu S, Page GG, Yirmiya R and
Taylor AN: Acute alcohol intoxication suppresses natural killer
cell activity and promotes tumor metastasis. Nat Med. 2:457–460.
1996.PubMed/NCBI View Article : Google Scholar
|
|
41
|
Jiménez-Jiménez FJ, Gómez-Tabales J,
Alonso-Navarro H, Zurdo M, Turpín-Fenoll L, Millán-Pascual J,
Adeva-Bartolomé T, Cubo E, Navacerrada F, Rojo-Sebastián A, et al:
Association between the rs1229984 polymorphism in the alcohol
dehydrogenase 1B gene and risk for restless legs syndrome. Sleep.
40(zsx174)2017.PubMed/NCBI View Article : Google Scholar
|