Introduction
Hepatocellular carcinoma (HCC) is one of the most
common types of cancer in the world and is the second most common
cause of cancer-associated mortality (1). More than 70% of HCC cases develop from a
background of chronic liver disease, such as chronic hepatitis or
liver cirrhosis, caused by viral infection or alcohol abuse
(2,3). In
Japan, the hepatitis C virus (HCV) is the most common cause of HCC.
Although the incidence of non-B non-C-associated HCC is increasing,
~60% of HCC cases are associated with HCV (4).
For early-stage HCC with a limited number of
cancerous lesions, smaller size, and without vascular invasion,
liver resection is the best treatment; however, the cumulative
5-year recurrence rate remained as high as ~80% following curative
hepatectomy (5,6). Chemotherapy is generally applied in
advanced cases with multiple cancerous and/or metastatic lesions;
however, only a limited number of molecular targeting drugs, such
as sorafenib, have demonstrated significant effects during clinical
testing (7,8).
It has been reported that the HCV infection induces
DNA damage via oxidative stress produced by chronic inflammation.
HCV core protein also increases reactive oxygen species by inducing
altered mitochondrial function (9,10). The DNA
damage generated by these processes is hypothesized to cause
mutations in tumor-suppressor genes and oncogenes observed in HCC
(11–13). These mutations are thought to be
important in the development of HCC; however, there are many HCC
cases without mutations in known tumor-associated genes. In
addition, a previous study demonstrated that the HCV core protein
induces cell transformation directly (14) and this may be mediated by increased
c-MYC protooncogene expression (15).
This pathogenetic complexity makes it difficult to understand the
mechanism of HCC development following HCV infection.
To determine the molecular mechanism underlying HCC
development and progression, the current study attempted to
identify novel cancer-associated genes in a mouse model of skin
cancer. By screening for aberrantly methylated genome regions or
genes showing aberrant expression levels in mouse squamous cell
carcinoma (SCC) tissues, novel genes involved in the development
and progression of human cancers, including HCC were identified.
For example, zygote arrest 1 (Zar1) was aberrantly methylated in
mouse SCC genomic DNA and its human ortholog, Zar1, was shown to be
aberrantly methylated in the genome of human HCC (16).
Previously, the ubiquitin-conjugating enzyme, CDC34,
was found to be highly expressed and hypomethylated in SCC tissue
samples (17). In the
ubiquitin-proteasome system (UPS), E2 enzymes, including CDC34,
conjugate ubiquitin (which is activated by E1 enzymes) onto target
molecules in conjunction with E3 enzymes (18). As UPS regulates degradation of various
types of protein, it is crucial in a variety of cellular processes.
UPS aberrations have been observed in numerous types of human tumor
(19,20). However, limited information is
available regarding aberrations of CDC34 in HCC. To determine the
role of CDC34 in the development or progression of HCC, the current
study investigated the expression patterns of CDC34 in human HCC
surgical specimens.
Materials and methods
Patients and tissue samples
Fifty-eight HCV-positive HCCs [classified as grade I
or II according to the Union for International Cancer Control TNM
classification (21)] and paired
non-tumorous tissue samples were obtained from the Nihon University
Hospital (Tokyo, Japan). These tissues and data were collected from
August 1995 to April 2010 after obtaining written informed consent
from each subject. The present study complied with the tenets of
the Declaration of Helsinki and was approved by the Institutional
Review Board of Nihon University School of Medicine (approval no.
214-0). All surgical specimens were frozen in liquid nitrogen
immediately following resection and stored at −80°C.
RNA extraction
RNA was extracted from HCC and non-tumorous tissue
samples using Invitrogen TRIzol reagent (Thermo Fisher Scientific,
Inc., Waltham, MA, USA) according to the manufacturers protocol.
All samples showed RNA integrity ≥7.0, as determined using an
Agilent 2100 Bioanalyzer (Agilent Technologies, Inc., Santa Clara,
CA, USA).
Reverse transcription-quantitative
polymerase chain reaction (RT-qPCR)
Total RNA was isolated from the tissue samples
obtained from the HCV-positive patients. Aliquots of total RNA (500
ng) were reverse-transcribed using iScript (Bio-Rad Laboratories,
Inc., Hercules, CA, USA) and the generated cDNAs were used for qPCR
using SYBR®Premix Ex TaqTM II (Takara Bio, Inc., Otsu,
Japan). CDC34 and GAPDH expression levels served as the target and
endogenous control, respectively. The amplification conditions were
as follows: 95°C for 30 sec, followed by 40 cycles of denaturation
at 95°C for 5 sec, annealing at 60°C for 10 sec and polymerization
at 72°C for 30 sec. To obtain standard curve for each genes,
RT-qPCR was done by using a dilution series of cDNA mixture as a
template. The experiments were performed in triplicate and the
amounts of genes were worked out by extrapolation from the standard
curves. The primer sequences were as follows: forward,
5′-CTACGAGGGCGGCTACTTC-3′ and reverse, 5′-TCTTGGTCAGGAACCGAAAG-3′,
for CDC34; and forward, 5′-GCACCGTCAAGGCTGAGAAC-3′ and reverse,
5′-TGGTGAAGACGCCAGTGGA-3′ for GAPDH.
Immunofluorescence staining
Sections (3 µm) were deparaffinized in xylene,
dehydrated through a graded alcohol series, washed in
phosphate-buffered saline and subjected to hematoxylin-eosin
staining or immunofluorescence (IF) staining. Prior to IF staining,
sections were heated in 0.05% citrate buffer solution (pH 6.0) at
121°C for 15 min, for antigen retrieval. To prevent non-specific
staining, sections were incubated with 3%
H2O2 and Protein Block (Dako; Agilent
Technologies, Inc.) at room temperature for 10 min. The sections
were then incubated with anti-CDC34 rabbit polyclonal primary
antibody (dilution, 1:100; cat. no. HPA002382; Sigma-Aldrich; Merck
KGaA, Darmstadt, Germany) at 4°C overnight. For detection of
primary antibodies, biotinylated anti-rabbit IgG goat secondary
antibody (dilution, 1:2,000; cat. no. E0432, Dako; Agilent
Technologies, Inc.) at room temperature for 30 min and
streptavidin-conjugated fluorescein isothiocyanate (Dako; Agilent
Technologies, Inc.) at room temperature for 10 min. All sections
were examined under a fluorescence microscope (DP71 microscope with
U-RFL-T fluorescence power supply unit; Olympus Corp., Tokyo,
Japan).
Statistical analysis
Continuous data are expressed as the median (range).
The χ2 test was used for categorical variables.
Statistical analyses were performed using JMP 10 software (SAS
Institute Japan Ltd., Tokyo, Japan) and P<0.05 was considered to
indicate a statistically significant difference.
Results
Clinicopathological features of the
patients
The 58 patients (48 men and 10 women) had
HCV-derived HCC. The median age of the patients was 68.5 years
(range 48–78 years). A total of 31 cases (53.4%) had non-liver
cirrhosis (LC) background and 27 cases (46.6%) had LC background.
The tumor size was ≥3 cm in 24 cases (41.3%), and >2 tumors were
found in 6 cases (10.3%). The histological grade was
well-differentiated (W/D) in 24 cases (41.3%) and moderately
differentiated (M/D) or poorly differentiated (P/D) in 34 cases
(58.6%). Details of the pathological parameters are presented in
Table I.
 | Table I.Comparison of clinical backgrounds and
expression of CDC34. |
Table I.
Comparison of clinical backgrounds and
expression of CDC34.
| Variable | Low tumor expression
(n=29) | High tumor expression
(n=29) | P-value | T/N ratio <2
(n=47) | T/N ratio ≥2
(n=11) | P-value |
|---|
| Sex |
|
| 0.16 |
|
| 0.33 |
| M | 26 | 22 |
| 40 | 8 |
|
| F | 3 | 7 |
| 7 | 3 |
| Age
(years)a | 68 (48–78) | 70 (50–73) | 0.36 | 67.3 (48–78) | 66.5 (50–73) | 0.74 |
| ICG15R (%) |
|
| 0.10 |
|
| 0.03 |
|
<15 | 15 | 21 |
| 26 | 10 |
|
| ≥15 | 14 | 8 |
| 21 | 1 |
|
| Alb (g/dl) |
|
| 1.00 |
|
| 0.93 |
|
<3.5 | 5 | 5 |
| 8 | 2 |
|
| ≥3.5 | 24 | 24 |
| 39 | 9 |
|
| AST (U/l) |
|
| 0.37 |
|
| 0.91 |
|
<40 | 6 | 9 |
| 12 | 3 |
|
| ≥40 | 23 | 20 |
| 35 | 8 |
|
| ALT (U/l) |
|
| 0.03 |
|
| 0.21 |
| <
40 | 7 | 15 |
| 16 | 6 |
|
| ≥40 | 22 | 14 |
| 31 | 5 |
|
| T.Bil (mg/dl) |
|
| 0.69 |
|
| 0.14 |
|
<1.2 | 25 | 26 |
| 39 | 11 |
|
|
≥1.2 | 4 | 3 |
| 8 | 0 |
|
| AFP (ng/ml) |
|
| 0.06 |
|
| 0.03 |
| <
20 | 9 | 16 |
| 17 | 8 |
|
|
≥20 | 20 | 30 |
| 30 | 3 |
|
| Background |
|
| 0.45 |
|
| 0.45 |
|
non-LC | 24 | 7 |
| 24 | 7 |
|
| LC | 23 | 4 |
| 23 | 4 |
|
| Vascular
invasion |
|
| 0.60 |
|
| 0.96 |
|
(−) | 15 | 17 |
| 26 | 6 |
|
|
(+) | 14 | 12 |
| 21 | 5 |
|
| Tumor size
(cm) |
|
| 0.11 |
|
| 0.02 |
|
<3 | 14 | 20 |
| 24 | 10 |
|
| ≥3 | 15 | 9 |
| 23 | 1 |
|
| Tumor number |
|
| 0.39 |
|
| 0.34 |
|
<1 | 25 | 27 |
| 43 | 9 |
|
| ≥2 | 4 | 2 |
| 4 | 2 |
|
| Histological
grade |
|
| 0.03 |
|
| 0.10 |
|
W/D | 8 | 16 |
| 17 | 7 |
|
| M/D,
P/D | 21 | 13 |
| 30 | 4 |
|
Expression in HCV-positive HCC and
non-tumorous tissue samples
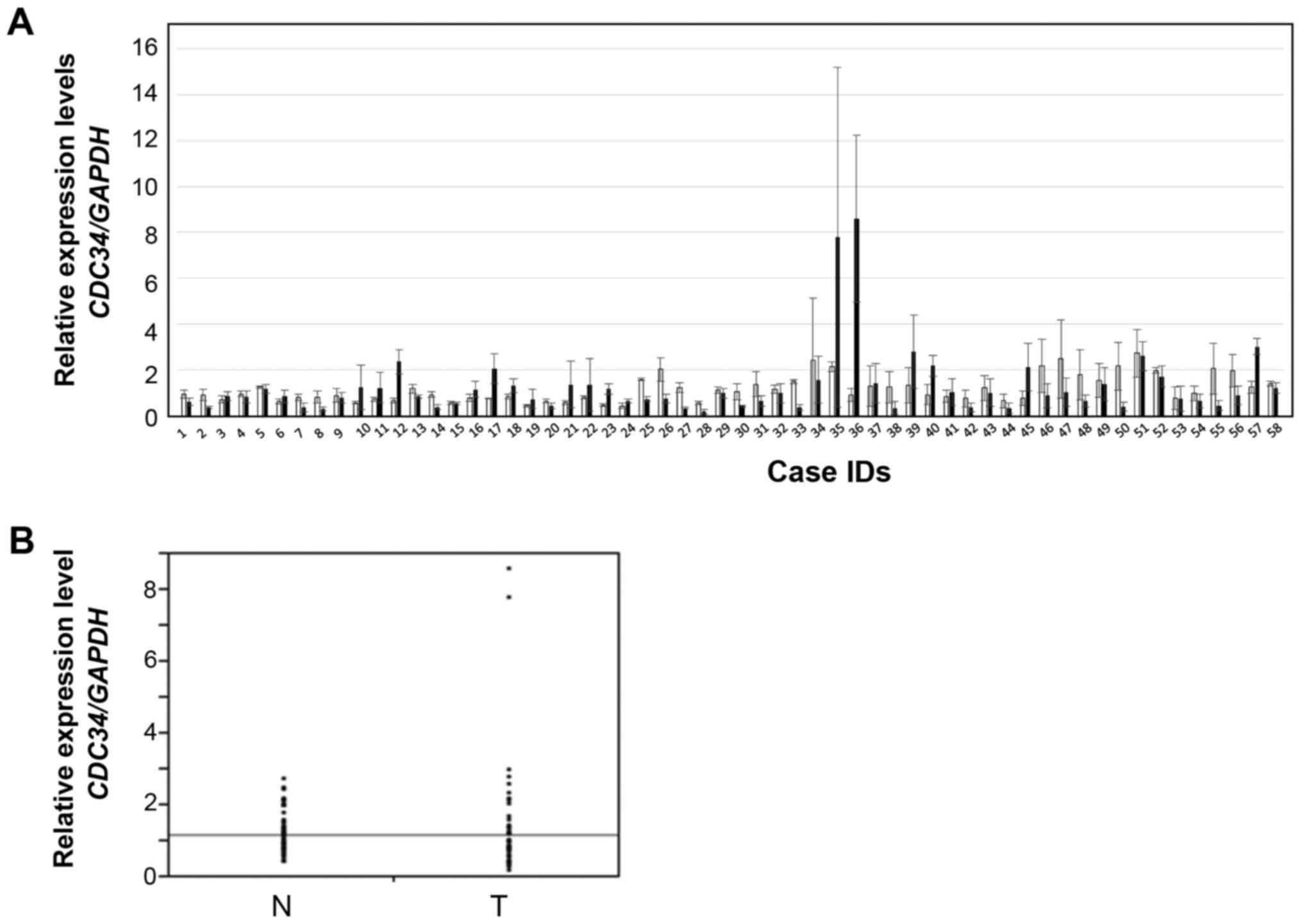
To examine the role of CDC34 in HCC, the level of
CDC34 mRNA expression in 58 HCV-positive HCC and non-tumorous
tissue samples were examined by RT-qPCR. The expression patterns of
CDC34 in HCC and non-tumorous tissue samples varied between the
cases (Fig. 1A). No significant
differences were identified in the expression levels of CDC34
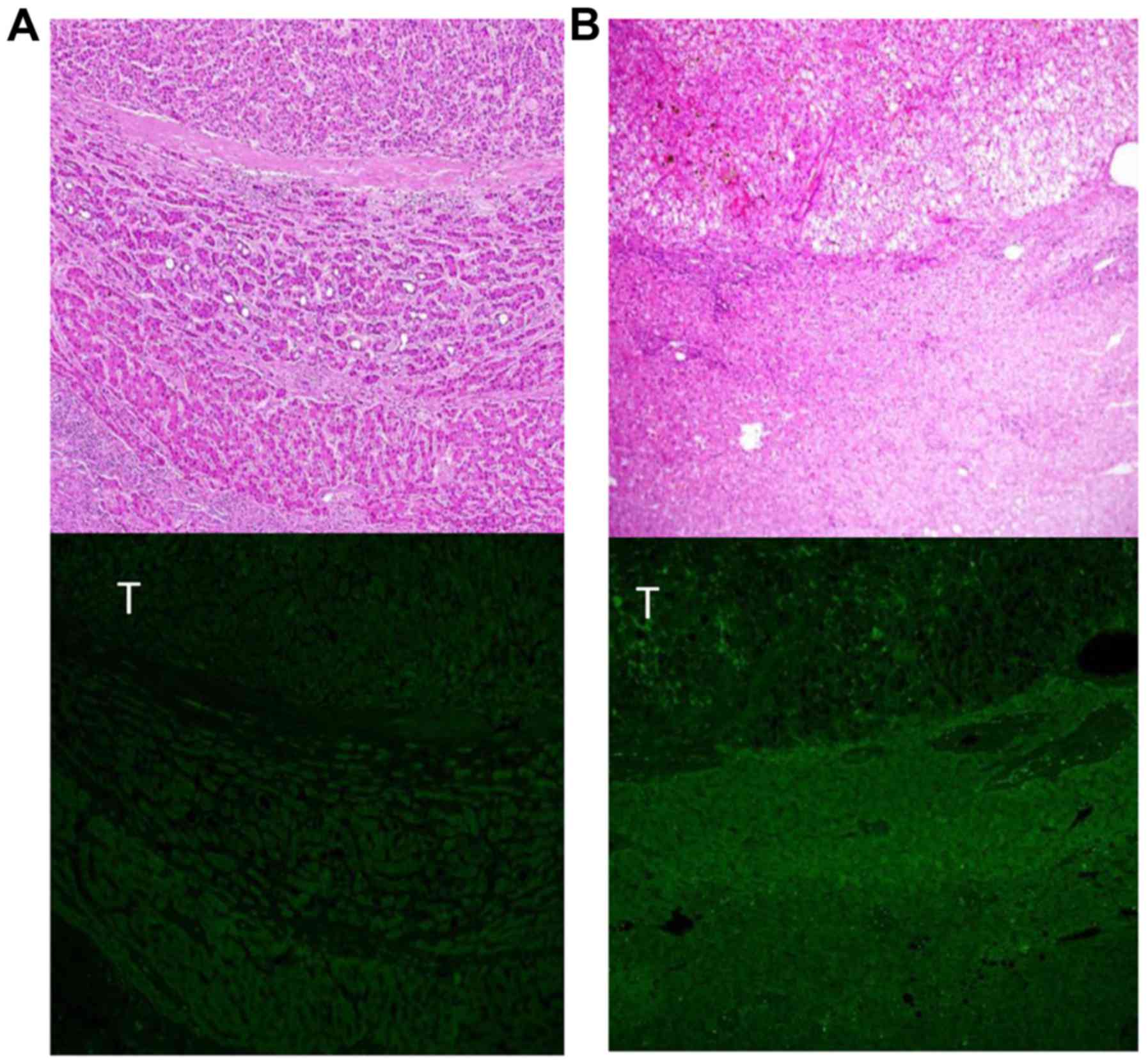
between HCC and non-tumorous tissue samples (Fig. 1B). The representative fluorescence
staining images using anti-CDC34 antibody are presented in Fig. 2. In case 37 (Fig. 2A), the strength of the staining was
similar between cancerous and non-cancerous regions. By contrast,
the fluorescence signal indicating CDC34 protein expression was
stronger in the cancerous area than the non-cancerous area in case
39 (Fig. 2B). These staining patterns
were similar to the expression patterns of CDC34 mRNA in cases 37
and 39, respectively. In the two cases, staining patterns of CDC34
were inhomogeneous; however, no obvious morphological features or
other characteristics were observed in the CDC34-positive
cells.
Correlation between CDC34 expression
and clinicopathological features
Table I presents the
correlations between CDC34 expression in the HCC tissue samples and
clinicopathological features. It was clearly demonstrated that
reduced ALT levels (<40 U/l) and lower histological grade (W/D)
were significantly associated with higher CDC34 expression
levels.
The correlations between the T/N ratio, which is the
ratio of CDC34 expression in HCCs to that in non-tumorous tissues,
and clinicopathological features are presented in Table I. Among the 58 cases, CDC34 expression
in HCC was at least 2x higher than that in the non-tumorous tissue
samples in 11 cases (19.0%; Table I).
In the present study, a smaller tumor size (<3.0 cm) was
significantly associated with a higher T/N ratio (T/N ≥2.0;
P<0.02). In addition, reduced levels of indocyanine green (ICG;
<15%) and α-fetoprotein (AFP; <20 ng/ml) were associated with
T/N ≥2.0 (P<0.03).
Discussion
As protein ubiquitination affects protein stability,
UPS is crucial for protein catabolism in the cytosol and nucleus.
As UPS regulates numerous aspects of cellular function, defects in
this system may result in the pathogenesis of various significant
human diseases, including tumors (19,20).
Ubiquitin may be added to a substrate protein by the concerted
action of ubiquitin-activating enzyme (E1), ubiquitin-conjugating
enzyme (E2), and ubiquitin protein ligase (E3) (18).
CDC34, which encodes an E2 enzyme, was originally
identified in yeast as a gene essential for G1/S
transition (22). It has been
implicated in cell cycle progression by ubiquitination of
cyclin-dependent kinase inhibitors, such as P27kip1, and
the G2/M checkpoint protein, WEE1 G2 checkpoint kinase
(23). In addition, CDC34 was shown to
be a critical target of let-7 microRNA, which exerts
tumor-suppressive functions (24).
These studies indicated that CDC34 performs oncogenic or
tumor-promoting functions; however, there have been studies
regarding the aberrance of CDC34 in human cancer. To date,
upregulated expression levels of CDC34 have been observed in acute
lymphoblastic leukemia (25) and HCC
(26), in which the quantity of CDC34
mRNA was shown to be significantly higher in surgically resected
HCC than in non-tumorous liver tissue samples. In addition, it was
demonstrated that the expression level of CDC34 mRNAs was higher in
P/D tumors than in W/D and M/D tumors (26). Inconsistent with this previous study
(26), the present study indicated
that increased CDC34 expression levels in HCC tissue samples were
associated with favorable clinicopathological features, including
lower ALT levels and histological grade. Furthermore, a higher T/N
ratio of CDC34 was associated with favorable features, such as
smaller tumor size, lower ICG levels and lower AFP levels. Although
not significant, there was a notable trend for W/D histological
grade to be associated with an increased T/N ratio of CDC34. ALT is
an enzyme produced predominantly by hepatocytes, and a higher blood
level of ATL indicates liver damage or disease. ICG retention rate
after 15 min is a useful preoperative liver function evaluation
factor and serves as an effective postoperative mortality
predictor. AFP, which is abundant in fetal plasma, but not in the
plasma of adults (27), is a specific
marker for HCC. Thus, reduced ALT, ICG and AFP levels are
associated with favorable outcome.
The discrepancy between the present results and
those reported previously (26) may be
attributable to the differences in sample size and viral status of
HCC between the two studies. The number of cases in the previous
study was limited (26), and the viral
status varied among the cases in the previous study. However, all
tissue specimens analyzed in the present study were exclusively
HCV-positive HCCs.
Numerous studies have indicated that CDC34 exerts
oncogenic properties, and there have been studies to develop drugs
targeting CDC34 (18); however, the
present study indicated the possibility that CDC34 performs
tumor-suppressive functions in HCC. This result suggests that CDC34
targets known tumor-suppressive genes, as well as oncogenes.
In conclusion, the present results indicated that
increased CDC34 expression levels or T/N ratio in HCC tissue
samples are associated with favorable phenotypes of HCC, such as a
small tumor size, and reduced levels of ICG and AFP. These findings
suggest that there may be genes that perform oncogenic functions
among the CDC34 targets in HCC. Therefore, CDC34 knockdown or
overexpressing cells may be useful tools to search new cancer
related genes. Furthermore, CDC34 expression levels in HCC may
serve as an indicator of HCC prognosis in patients.
Acknowledgements
The present study was supported by a Grant-in-Aid
for Scientific Research (grant no. 24249068) from the Ministry of
Education, Culture, Sports, Science and Technology (MEXT), Japan
and by the MEXT-Supported Program for the Strategic Research
Foundation at Private Universities (2011–2015). The authors would
like to thank their colleagues from the Department of Digestive
Surgery at the Nihon University School of Medicine (Tokyo,
Japan).
References
|
1
|
Globocan 2012: Estimated Cancer Incidence,
Mortality and Prevalence Worldwide in 2012. Lyon, France: 2012,
http://globocan.iarc.fr/Pages/fact_sheets_cancer.aspx
|
|
2
|
Forner A, Llovet JM and Bruix J:
Hepatocellular carcinoma. Lancet. 379:1245–1255. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
van Malenstein H, van Pelt J and Verslype
C: Molecular classification of hepatocellular carcinoma anno 2011.
Eur J Cancer. 47:1789–1797. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Lavanchy D: The global burden of hepatitis
C. Liver Int. 29 Suppl 1:74–81. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Grazi GL, Ercolani G, Pierangeli F, Del
Gaudio M, Cescon M, Cavallari A and Mazziotti A: Improved results
of liver resection for hepatocellular carcinoma on cirrhosis give
the procedure added value. Ann Surg. 234:71–78. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Imamura H, Matsuyama Y, Tanaka E, Ohkubo
T, Hasegawa K, Miyagawa S, Sugawara Y, Minagawa M, Takayama T,
Kawasaki S and Makuuchi M: Risk factors contributing to early and
late phase intrahepatic recurrence of hepatocellular carcinoma
after hepatectomy. J Hepatol. 38:200–207. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Llovet JM, Ricci S, Mazzaferro V, Hilgard
P, Gane E, Blanc JF, De Oliveira AC, Santoro A, Raoul JL, Forner A,
et al SHARP Investigators Study Group, : Sorafenib in advanced
hepatocellular carcinoma. N Engl J Med. 359:378–390. 2008.
View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Cheng AL, Guan Z, Chen Z, Tsao CJ, Qin S,
Kim JS, Yang TS, Tak WY, Pan H, Yu S, et al: Efficacy and safety of
sorafenib in patients with advanced hepatocellular carcinoma
according to baseline status: Subset analyses of the phase III
Sorafenib Asia-Pacific trial. Eur J Cancer. 48:1452–1465. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Ivanov AV, Bartosch B, Smirnova OA,
Isaguliants MG and Kochetkov SN: HCV and oxidative stress in the
liver. Viruses. 5:439–469. 2013. View
Article : Google Scholar : PubMed/NCBI
|
|
10
|
Korenaga M, Wang T, Li Y, Showalter LA,
Chan T, Sun J and Weinman SA: Hepatitis C virus core protein
inhibits mitochondrial electron transport and increases reactive
oxygen species (ROS) production. J Biol Chem. 280:37481–37488.
2005. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Minouchi K, Kaneko S and Kobayashi K:
Mutation of p53 gene in regenerative nodules in cirrhotic liver. J
Hepatol. 37:231–239. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Laurent-Puig P and Zucman-Rossi J:
Genetics of hepatocellular tumors. Oncogene. 25:3778–3786. 2006.
View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Li CW, Chang PY and Chen BS: Investigating
the mechanism of hepatocellular carcinoma progression by
constructing genetic and epigenetic networks using NGS data
identification and big database mining method. Oncotarget.
7:79453–79473. 2016.PubMed/NCBI
|
|
14
|
Lemon SM and McGivern DR: Is hepatitis C
virus carcinogenic? Gastroenterology. 142:1274–1278. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Higgs MR, Lerat H and Pawlotsky JM:
Hepatitis C virus-induced activation of β-catenin promotes c-Myc
expression and a cascade of pro-carcinogenetic events. Oncogene.
32:4683–4693. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Takagi K, Fujiwara K, Takayama T, Mamiya
T, Soma M and Nagase H: DNA hypermethylation of zygote arrest 1
(ZAR1) in hepatitis C virus positive related hepatocellular
carcinoma. Springerplus. 2:1502013. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Fujiwara K, Ghosh S, Liang P, Morien E,
Soma M and Nagase H: Genome-wide screening of aberrant DNA
methylation which associated with gene expression in mouse skin
cancers. Mol Carcinog. 54:178–188. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Ceccarelli DF, Tang X, Pelletier B,
Orlicky S, Xie W, Plantevin V, Neculai D, Chou YC, Ogunjimi A,
Al-Hakim A, et al: An allosteric inhibitor of the human Cdc34
ubiquitin-conjugating enzyme. Cell. 145:1075–1087. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Micel LN, Tentler JJ, Smith PG and
Eckhardt GS: Role of ubiquitin ligases and the proteasome in
oncogenesis: Novel targets for anticancer therapies. J Clin Oncol.
31:1231–1238. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Morrow JK, Lin HK, Sun SC and Zhang S:
Targeting ubiquitination for cancer therapies. Future Med Chem.
7:2333–2350. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Wittekind C, Compton CC, Greene FL and
Sobin LH: TNM residual tumor classification revisited. Cancer.
94:2511–2516. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Schwob E, Böhm T, Mendenhall MD and
Nasmyth K: The B-type cyclin kinase inhibitor p40SIC1 controls the
G1 to S transition in S. cerevisiae. Cell. 79:233–244. 1994.
View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Tyers M and Jorgensen P: Proteolysis and
the cell cycle: With this RING I do thee destroy. Curr Opin Genet
Dev. 10:54–64. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Legesse-Miller A, Elemento O, Pfau SJ,
Forman JJ and Tavazoie S: Coller HA let-7 Overexpression leads to
an increased fraction of cells in G2/M, direct down-regulation of
Cdc34, and stabilization of Wee1 kinase in primary fibroblasts. J
Biol Chem. 284:6605–6609. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Eliseeva E, Pati D, Diccinanni MB, Yu AL,
Mohsin SK, Margolin JF and Plon SE: Expression and localization of
the CDC34 ubiquitin-conjugating enzyme in pediatric acute
lymphoblastic leukemia. Cell Growth Differ. 12:427–433.
2001.PubMed/NCBI
|
|
26
|
Tanaka K, Kondoh N, Shuda M, Matsubara O,
Imazeki N, Ryo A, Wakatsuki T, Hada A, Goseki N, Igari T, et al:
Enhanced expression of mRNAs of antisecretory factor-1, gp96, DAD1
and CDC34 in human hepatocellular carcinomas. Biochim Biophys Acta.
1536:1–12. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Butterfield LH, Ribas A, Meng WS, Dissette
VB, Amarnani S, Vu HT, Seja E, Todd K, Glaspy JA, McBride WH, et
al: T-cell responses to HLA-A*0201 immunodominant peptides derived
from α-fetoprotein in patients with hepatocellular cancer. Clin
Cancer Res. 9:5902–5908. 2003.PubMed/NCBI
|