Introduction
Hepatitis B infection is a life-threatening liver
disease resulting from the hepatitis B virus (HBV) (1). There is high inter-individual
variability in the clinical presentation of HBV infection, ranging
from self-limited to acute fulminant hepatitis. This can cause
chronic liver inflammation leading to cirrhosis and hepatocellular
carcinoma (HCC) (2). Despite the
presence of effective antiviral therapies and vaccines (3), the mortality rates have increased from
0.8 to 1.4 million from 1990 to 2013(4). Therefore, it is crucial to consider
the molecular aspects affecting HBV.
Non-coding RNAs (ncRNAs) are RNA molecules that are
not translated into proteins (5).
They regulate diverse cellular functions and processes by
controlling gene expression (6).
High-throughput DNA sequencing and array-based technologies have
revolutionized the classification of ncRNAs (7). Among several types of ncRNAs, short
ncRNAs, including microRNAs (miRNAs/miRs) and long ncRNAs
(lncRNAs), have been considered a standpoint (8).
miRNAs are ~19 to 22 nucleotides (nt) in length
(9). To date, >2,000 miRNAs have
been registered in the ‘miRbase’ database; however, the functional
role of the majority of miRNAs remains unclear; they have emerged
as eminent players in human pathophysiological processes (10). miRNAs affect gene expression through
various mechanisms, such as de-adenylation, targeting mRNA cleavage
and suppression of translation, supporting the evidence that a
single miRNA can regulate hundreds of genes; hundreds of miRNAs
(11) can also regulate a single
gene. An example of miRNAs that have been highly expressed in the
liver is miR-100, which is located on chromosome 11 at
11q24.1(12). It promotes HBV
protein production, DNA replication and progeny secretion (13).
lncRNAs, the most recent acknowledged class of
ncRNAs, are transcripts with lengths >200 nt without
protein-coding capacity (14).
lncRNAs are messenger RNA (mRNA)-like transcripts, but without
stable open reading frames (ORFs). The majority of lncRNAs can
regulate gene expression through chromatin modification,
transcription and post-transcriptional processing (15). An increasing number of lncRNAs have
been characterized in studies, focusing on their roles in
regulating gene expression (16,17).
Metastasis-associated lung adenocarcinoma transcript
1 (MALAT1) is one of the most abundant lncRNAs in normal tissues
and is highly conserved between humans, also known as
nuclear-enriched abundant transcript 2 (NEAT2) (18). It consists of >8,000 nt and is
coded by chromosome 11q13. MALAT1 has been reported to regulate
gene expression; there is substantial evidence to suggest the vital
role of MALAT1 in liver cell proliferation (19,20).
Recently, MALAT1 has gained considerable attention due to its
association with a number of diseases, also acting as a potential
biomarker for the diagnosis, prediction and therapeutic target for
numerous types of cancer (21).
Single nucleotide polymorphisms (SNPs) have been
suggested to be biological markers for revealing the evolutionary
history and common genetic polymorphisms that explain the heritable
risk for common diseases (22).
SNPs in ncRNAs have been reported to alter their secondary
structure or modify expression levels, thereby influencing their
regulatory function, contributing to disease development (23). In a previous study, the authors
examined the presence of SNP (rs1834306 T/C) in miR-100 in
HBV-infected patients and its effect on gene expression (24). In continuation of this, the present
study focused on examining SNP rs619586 (A/G) in MALAT1, and its
expression level in HBV-infected Egyptian patients compared to
other healthy controls. Possible correlations between both miR-100
and MALAT1 in HBV infection were also investigated.
Subjects and methods
Ethics approval
All subjects provided written informed consent for
genetic analysis in the present observational prospective
case-control study. All methods and analyses were carried out
following the guidelines of the Ministry of Health and approved by
the Research Ethics Committee for Experimental and Clinical Studies
at the Faculty of Pharmacy, University of Cairo, Egypt [BC
(1837)].
Patients and study design
A total of 200 subjects; 100 outpatients (70 males
and 30 females; under the medical supervision of the National Liver
Institute, Menoufia University, Menoufia, Egypt) with an approved
diagnosis of HBV infection by enzyme-linked immunosorbent assay
(ELISA) and polymerase chain reaction (PCR) were included in
parallel to 100 individuals (64 males and 36 females) with normal
liver function test results, no history of hepatic diseases and
negative for HBV and hepatitis C virus (HCV) serology, which served
as controls. As previously described by Motawi et al
(24), viral assessment in all
subjects was performed. Patients with HCV infection or other viral
or hepatic disorders were excluded from the study. All biochemical
investigations included alanine aminotransferase (ALT) and
aspartate aminotransferase (AST) activities. According to the
manufacturer's instructions, total bilirubin, albumin and
creatinine levels were measured in blood for all subjects using a
Cobas 6000 analyzer (Roche Diagnostics GmbH).
SNP selection
Based on the data from the HapMap (http://www.hapmap.org); NCBI dbSNP (http://www.ncbi.nlm.nih.gov/SNP/) and miRNAs
(http://microrna.sanger.ac.uk) databases,
miR100 (rs1834306 T/C) and MALAT1 (rs619586 A/G) SNPs were selected
for analyses in the present study.
DNA extraction and SNP genotyping
Genomic DNA was extracted from 5 ml of venous blood
samples [collected in ethylene-diamine-tetra-acetic acid (EDTA)
sterile vacutainer] from each participant using the GentraPuregene
Blood kit (Qiagen GmbH) according to the manufacturer's
instructions. A Nanodrop™ 2000/2000c spectrophotometer (Thermo
Fisher Scientific, Inc.) was used to assess the purity and the
concentration of the extracted DNA. The extracted DNA was applied
to 1% agarose gel electrophoresis to confirm its integrity.
miR-100 rs1834306 T/C was analyzed using
PCR-sequence-specific primers (PCR-SSP), as previously described by
Motawi et al (24). However,
MALAT1 polymorphism rs619586 A/G was genotyped by restriction
fragment length polymorphism-PCR (RFLP-PCR). The primer sequences
of both miR-100 andMALAT1 were designed using Primo SNP 3.4: SNP
PCR Primer Design (https://www.changbioscience.com/primo/primosnp.html)
and secondly checked using primer blast (https://www.ncbi.nlm.nih.gov/tools/primer-blast/)
(Table I). A 25 µl PCR reaction
mixture contained MyTaq™ Red Mastermix (2X; Meridian Life Science,
Inc.), 10 pmoles of each primer and 150 ng DNA. The PCR reaction
conditions were as follows: 95˚C for 10 min (one cycle) followed by
35 cycles of 94˚C for the 30 sec, 59˚C for 30 sec, and 72˚C for 1
min, then a final extension step at 72˚C for 7 min. All PCR
reactions were performed in a 2720 thermal cycler (Applied
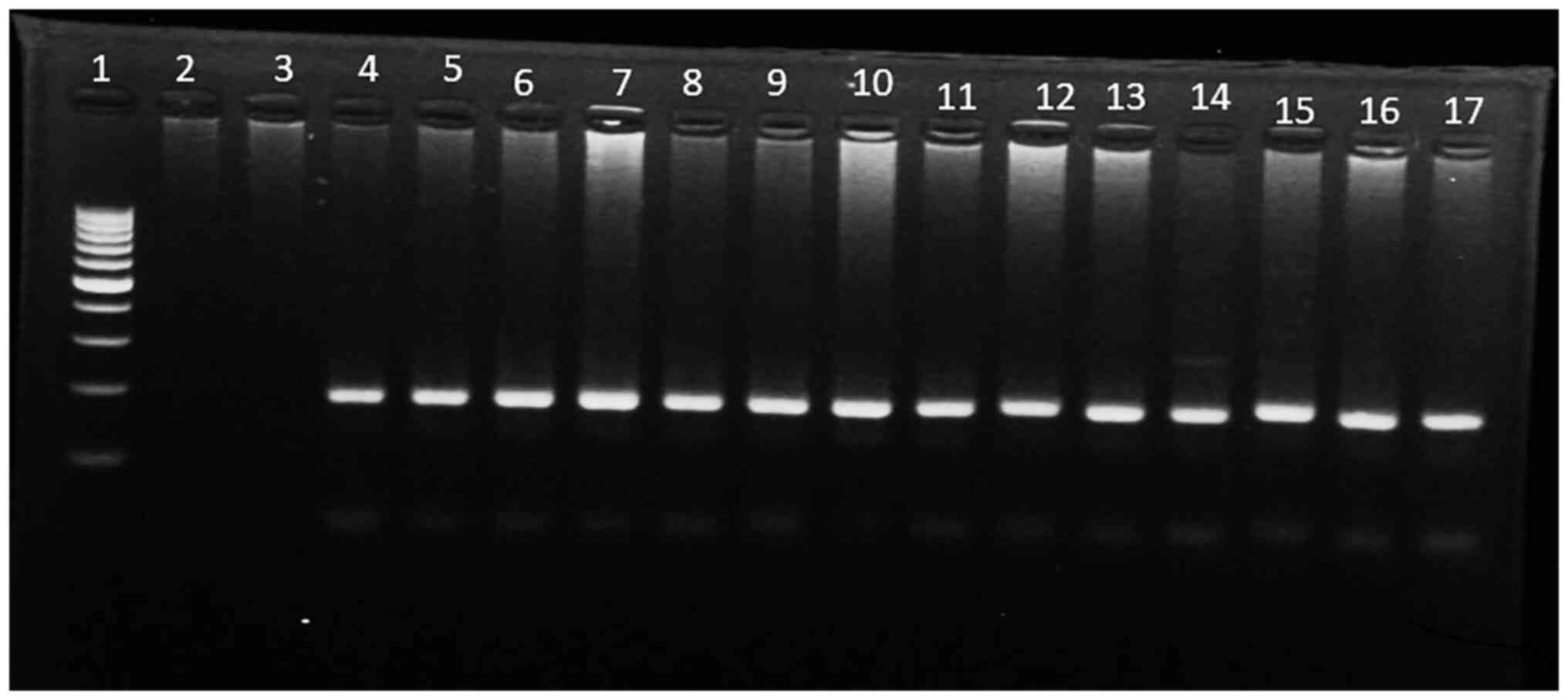
Biosystems; Thermo Fisher Scientific, Inc.). A 2% agarose
electrophoresis stained with ethidium bromide (10 mg/ml) was used
to visualize the PCR product (188 bp) in comparison to the 100 bp
DNA ladder (Fermentas; Thermo Fisher Scientific, Inc.) (Fig. 1). The PCR product was digested by
the addition of BveI (BspMI) restriction enzymes
(Fermentas; Thermo Fisher Scientific, Inc.). The restriction
product size was 188bp for the A/A genotype, 121/67 bp for G/G and
188/121/67 bp for A/G. The digestion products were visualized by 3%
agarose gel electrophoresis and estimated by comparing with the 50
bp DNA Ladder (Fermentas; Thermo Fisher Scientific, Inc.). In
total, 10% of samples were randomly selected to be sequenced to
control genotyping quality and validate the results.
 | Table IPCR primers sequences used for the
amplification of miR-100 and MALAT-1 in patients with HBV and the
controls. |
Table I
PCR primers sequences used for the
amplification of miR-100 and MALAT-1 in patients with HBV and the
controls.
| SNP | Primers | PCR Product
(bp) | Enzyme | Restriction
product |
|---|
| miR-100 | Forward T:
5'-GTGGAAACCAAGGGAAGCACGT-3' | 301 | - | - |
| rs1834306 | Forward C:
5'-TGGAAACCAAGGGAAGCACGC-3' | | | |
| T/C | Reverse:
5'-ATAAGCAAAGCCCCAGGTCC-3' | | | |
| MALAT-1 |
Forward:5'-AAAGTCCGCCATTTTGCCAC-3' | 188 | BspMI | AA:188 |
| rs619586 | Reverse:
5’-CACAAAACCCCCGGAACTT-3’ | | | AG:188/121/67 |
| A/G | | | | GG:121/67 |
RNA isolation and reverse
transcription-quantitative PCR (RT-qPCR)
Total RNA (from 200 µl plasma samples) was purified
using the miRNeasy Mini kit (cat. no. 217004; Qiagen, Inc.)
according to the manufacturer's instructions. For the miR-100
expression level, cDNA was prepared using the miScriptII RT kit
(cat. no. 218061; Qiagen, Inc.). qPCR was performed using the
miScript SYBR-Green PCR kit (cat. no. 218073; Qiagen, Inc.), as
previously described in the study by Motawi et al (24).
To determine the transcripts of the gene of interest
(MALAT1), RNA was reverse-transcribed using the High-Capacity DNA
Reverse Transcription kit (cat no. 4368814; Applied Biosystems;
Thermo Fisher Scientific, Inc.). The target cDNA was then amplified
using the TaqMan™ Universal Master Mix II (cat no. 4440043; Applied
Biosystems; Thermo Fisher Scientific, Inc.) according to the
manufacturer's protocol. The expression of glyceraldehyde
3-phosphate dehydrogenase (GAPDH) was used as an internal control
for quantitative normalization. RT-qPCR amplification began with an
initial holding period at 50˚C for 2 min, 95˚C for 10 min followed
by a PCR program consisting of 40 cycles of 95˚C for 15 sec and
60˚C for 1 min. Differences in the Ct values (Ct) between MALAT1
and GADPH were calculated using the formula ΔΔCt=ΔCt (tested
sample)-ΔCt (control sample) to determine the relative expression
levels; the fold change in MALAT1 was calculated using the
2-ΔΔCq method (25).
Statistical analysis
All statistical analyses were performed using the
clinical Statistical Package for Social Science (SPSS) version 19
(SPSS, Inc.). Data are presented as the mean ± standard
deviation/error (SD/SE). An independent paired t-test was
applied to compare numerical variables between the patients with
HBV and controls for quantitative variables. SNP/STAT was performed
using the online tool (http://bioinfo.iconcologia.net/SNP stats). The
receiver operating characteristic (ROC) curve analyzed the
sensitivity versus specificity of the scoring system. Pearson's
correlation analysis was used for correlation analysis. All
P-values were two-tailed; A P-value <0.05 was considered to
indicate a statistically significant difference.
Results
Demographic and biochemical
characteristics of patients with HBV vs. the control subjects
All patients with HBV were found positive for
hepatitis B surface antigen (HBsAg) and HBV-DNA. The demographic
and biochemical characteristics of the patients with HBV vs. the
control subjects have been previously described in the study by
Motawi et al (24)
Genetic variation of MALAT1 rs619586
A/G and miR-100 rs1834306 T/C
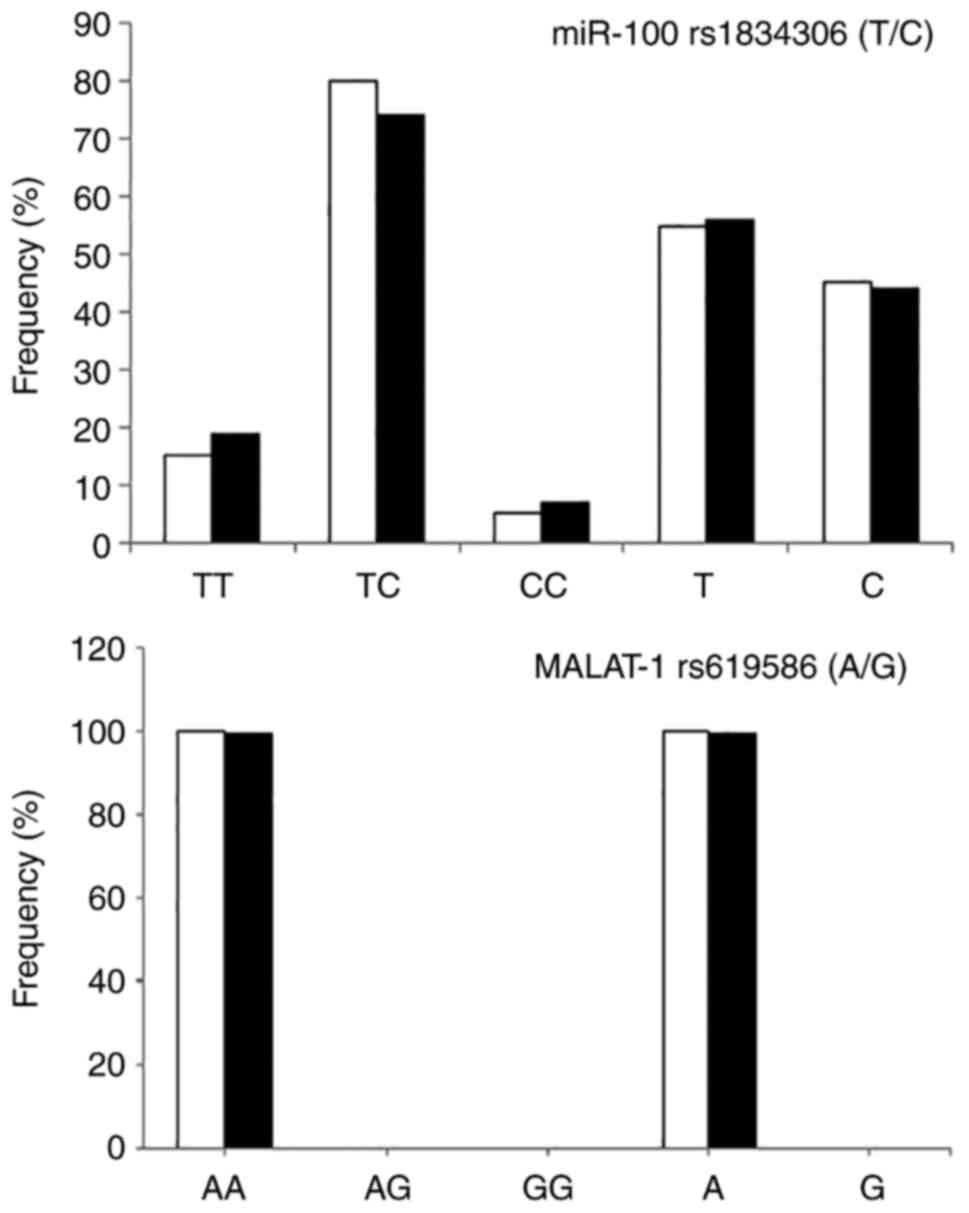
The study of the genotypes of MALAT1 rs619586 A/G
revealed the presence of only one genotype: The dominant AA
genotype, with a complete disappearance of other genotypes in the
two studied groups. On the contrary, all genotypes of miR-100
rs1834306 (T/C) were found, although no statistically significant
difference in the genotype distribution between patients with HBV
and the normal controls was demonstrated. The genotype and allelic
frequency of MALAT1 rs619586 (A/G) and miR-100 rs1834306 (T/C) are
presented in Fig. 2.
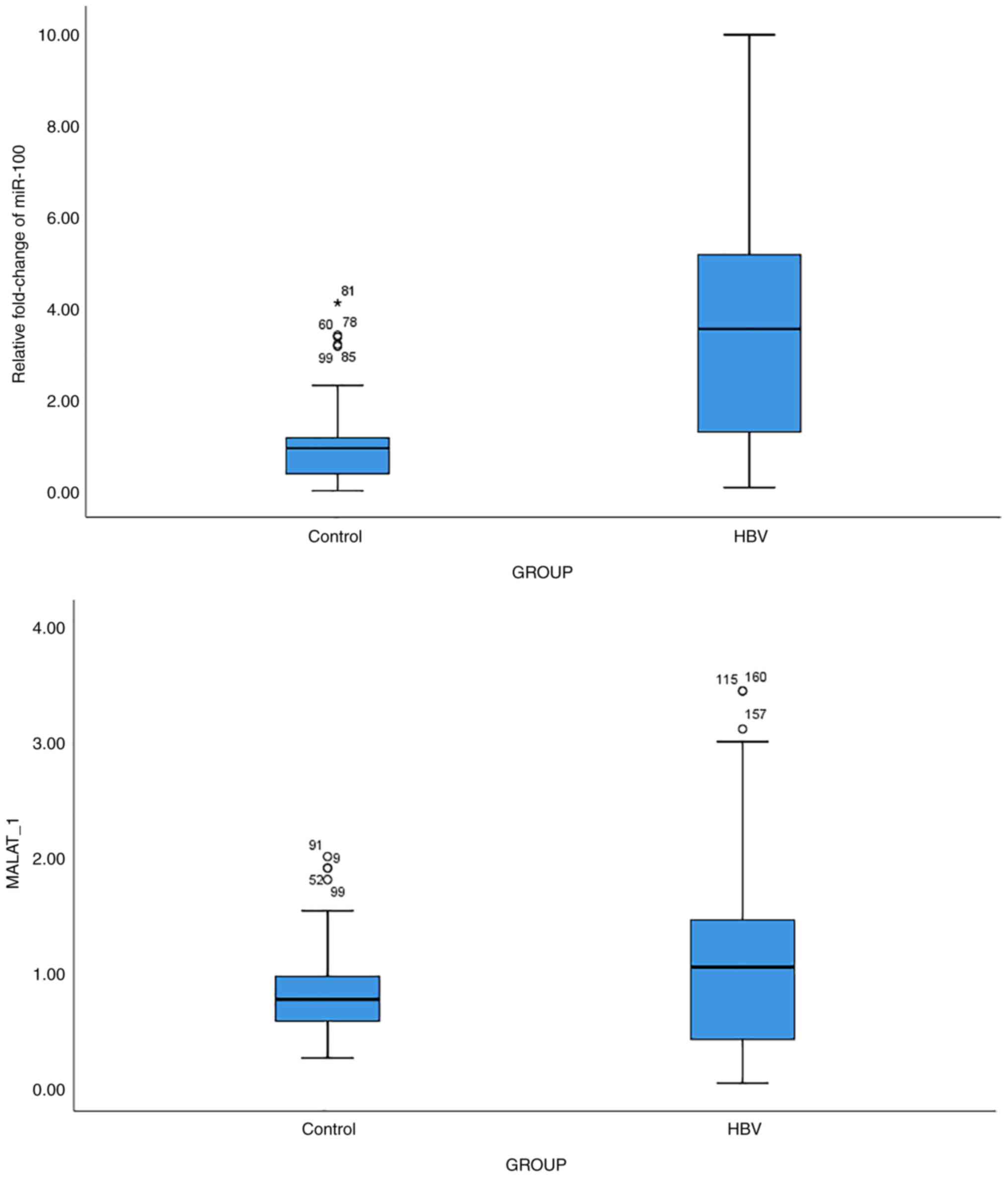
Expression of miR-100 and MALAT1
The analysis of the miR-100 and MALAT1 expression
levels revealed a significant upregulation ofmiR-100 (P<0.001)
and MALAT1 (P<0.001) expression in patients with HBV vs. the
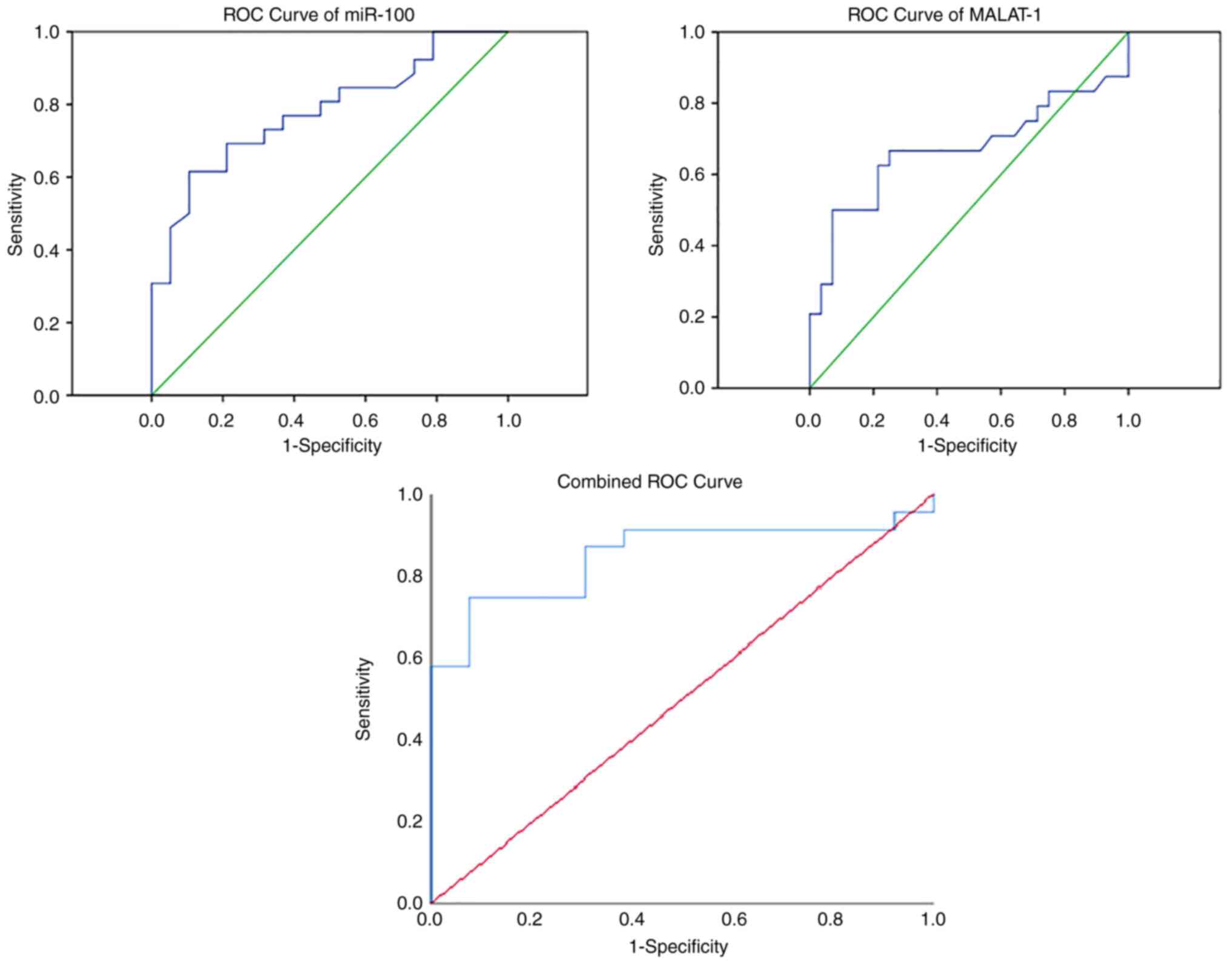
controls (Fig. 3). The results of
the analysis of the ROC curve for both miR-100 and MALAT1 are
summarized in Table II. Combined
ROC curve analysis resulted in an improvement in the diagnostic
potential of both ncRNAs, leading to 75% sensitivity and 92%
specificity (Fig. 4).
 | Table IIROC curve of miR-100 and MALAT-1. |
Table II
ROC curve of miR-100 and MALAT-1.
| Parameters | Cut-off | AUC | Sensitivity | Specificity | 95% CI | P-value |
|---|
| miR-100 | 1.017 | 0.778 | 76.9%, | 63.2% | 0.644-0.913 | P<0.01 |
| MALAT1 | 0.830 | 0.670 | 66.7% | 75% | 0.510-0.831 | P<0.05 |
| Combined | | 0.853 | | | 0.727-0.978 | P<0.001 |
Correlations between increased miR-100
and MALAT1 expression levels with some biochemical tests and the
viral load in patients HBV vs. the controls
No statistically significant correlation was found
between MALAT1 and miR-100 concerning their genotyping or
expression level in patients with HBV. A positive correlation
between the viral load of HBV, and both MALAT1 expression and
(r=0.282 and r2 =0.079; P<0.05) and miR-100
expression (r=0.489 and r2=0.239; P<0.001) was found.
In view of the biochemical tests, a positive correlation (r=0.316
and r2=0.099; P<0.05) between MALAT1 expression and
the ALT level was detected (Table
III).
 | Table IIICorrelation between biochemical tests
and the viral load with miR-100 and MALAT1 expression level. |
Table III
Correlation between biochemical tests
and the viral load with miR-100 and MALAT1 expression level.
| | r/r2
value and P-value | Group | AST | ALT | Albumin | Total
bilirubin | PCR | miR-100 | MALAT1 |
|---|
| Group | r | 1 | | | | | | | |
| AST | r | 0.365b | 1 | | | | | | |
| | r2 | 0.133 | | | | | | | |
| | p | <0.001 | | | | | | | |
| ALT | r | 0.368b | 0.865b | 1 | | | | | |
| | r2 | 0.135 | 0.748 | | | | | | |
| | p | <0.001 | <0.001 | | | | | | |
| Albumin | r | -0.718b | -0.069 | -0.078 | 1 | | | | |
| | r2 | 0.515 | 0.004 | 0.006 | | | | | |
| | p | <0.001 | NS | NS | | | | | |
| Total
bilirubin | r | 0.353b | 0.724b | 0.781b | -0.051 | 1 | | | |
| | r2 | 0.124 | 0.524 | 0.609 | | | | | |
| | p | <0.001 | <0.001 | <0.001 | NS | | | | |
| PCR | r | 0.598b | 0.698b | 0.722b | -0.280b | 0.732b | 1 | | |
| | r2 | 0.357 | 0.487 | 0.521 | 0.078 | 0.535 | | | |
| | p | <0.001 | <0.001 | <0.001 | <0.001 | <0.001 | | | |
| miR-100 | r | 0.478b | 0.356a | 0.334a | -0.302a | 0.401b | 0.489b | 1 | |
| | r2 | 0.228 | 0.126 | 0.111 | 0.091 | 0.160 | 0.239 | | |
| | p | <0.001 | <0.05 | <0.05 | <0.05 | <0.01 | <0.001 | | |
| MALAT1 | r | 0.294a | 0.236 | 0.316a | -0.151 | 0.248 | 0.282a | 0.102 | 1 |
| | r2 | 0.086 | 0.055 | 0.099 | 0.022 | 0.061 | 0.079 | 0.010 | |
| | p | <0.05 | NS | <0.05 | NS | NS | <0.05 | NS | |
Discussion
Viral infections are of global public health
concern; HBV is one of the leading causes of mortality (26), and the main obstacle to its
treatment is the inability to achieve a full cure for HBV (27). Thus, it is urgent to consider the
molecular field affecting HBV. Recent research has devoted ample
attention to genetic alterations (28). In this respect, diverse classes of
ncRNAs, ranging from miRNAs to lncRNAs, play a crucial role in the
epigenetic regulation of gene expression; genome stability also
acts as a defense against foreign genetic elements (29).
Over the past decade, an increasing number of
scientific studies and research have focused on the pivotal
biological functions of one of the key lncRNAs, MALAT1, which was
originally discovered as a prognostic marker for lung cancer
metastasis, and has been linked to several other human tumor
entities (30). Previous studies
have revealed that MALAT1 gene polymorphisms are associated with
disease susceptibility; for example, the MALAT1 rs619586 has been
found to be associated with a decreased risk of developing HCC and
colorectal cancer (31-32).
With respect to an SNP of MALAT1 rs619586 A/G in
HBV-infected patients vs. the control group, the genotyping results
of the present study revealed the dominant appearance of AA
genotype in both groups. By contrast, the AG and GG genotypes were
lacking. In agreement with these results, Motawi et al
(33) demonstrated that the AA
genotype was more frequent than AG or GG genotypes in HBV-infected
Egyptian patients. An earlier study on the Chinese population
reported that the AA genotype was the most frequent with a lack of
significance, apart from reporting no significant association
between MALAT1 rs619586 SNP and HBV clearance (31). In the Taiwanese population, Yuan
et al (34) pointed to an
insignificant association between MALAT1 rs619586 and the risk of
developing HCC in the HBV-positive subgroup, with higher frequency
persistence of the AA genotype over the AG and GG genotypes.
However, Wang et al (35)
found that tge MALAT1 rs619586 polymorphism decreased the risk of
developing HCC under a dominant model, indicating that this SNP has
the potential to be a biomarker for HCC risk and prognosis. In
general, only a limited number of studies have focused on the
genetic variation of MALAT1 rs619586 A/G in HBV infection. In other
diseases, the AA genotype is the most dominant genotype, such as in
lung cancer (36), congenital heart
disease (37), ischemic stroke
(38), thyroid carcinoma (39) and recurrent miscarriage (40).
It is not surprising that disease-associated SNPs
can alter their gene expression levels (23). Supporting this issue, recent studies
have focused (41-46)
on the dysregulation of MALAT1 in various diseases. The present
study demonstrated a significant upregulation of MALAT1 expression
in the plasma of patients with HBV compared to healthy controls.
Generally, only a limited number of studies are available to date
which quantify MALAT1 expression in patients with HBV. Consistent
with the results obtained herein, Konishi et al (41) reported that MALAT1 plasma levels
were progressively and significantly elevated in both hepatic
disease and patients with HCC. Evidence from other recent studies
has reported an increase in the MALAT1 level in HCC tissues
(42,43). Multiple lines of evidence have
reported the prognostic usefulness of MALAT1 across various types
of cancer (44-45), such as being a putative non-invasive
biomarker in HCV-induced HCC (46).
In addition to MALAT1, miR-100 is another type of
non-protein-coding transcript, miRNA. Previously, it was reported
by Motawi et al (25) that
miR-100 was significantly upregulated inpatients with HBV, and this
elevation is synchronized with the presence of the T allele,
suggesting that miR-100 may be considered a potential molecular
marker to appraise the prognosis of patients with HBV. As lncRNAs
may interact with miRNAs and modulate each other's expression
(47), MALAT1 has been described to
regulate several miRNAs (48).
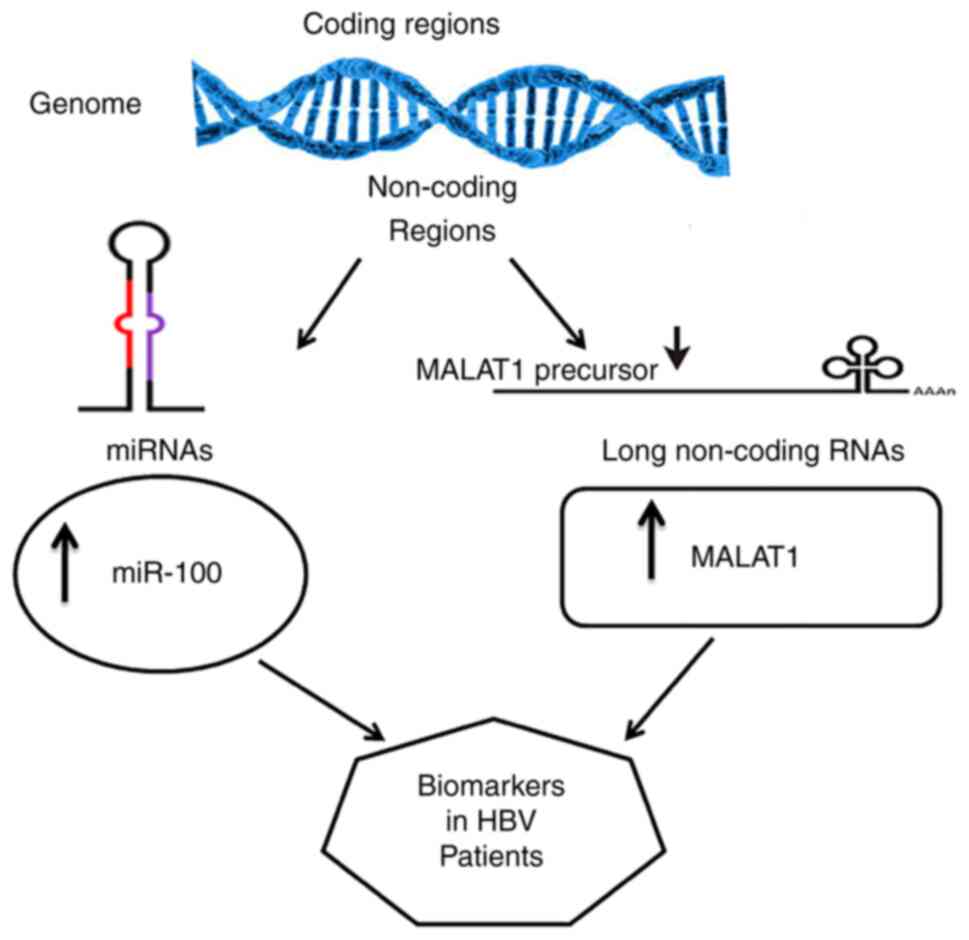
The present study hypothesized the presence of
correlations between miR-100 and MALAT1. Although no significant
correlation was observed, miR-100 and MALAT1 were significantly
upregulated in the HBV-infected patients. A positive correlation
between the viral load of HBV and both miR-100 and MALAT1 was
detected. Both miR-100 and MALAT1 may be regarded as non-invasive
molecular markers in HBV infection in the Egyptian population
(Fig. 5). Therefore, the present
preliminary study focused on two major classes of ncRNAs; miR-100
and MALAT1. Both ncRNAs were upregulated in patients with HBV, and
both were found to be positively correlated with the HBV viral
load. Accordingly, they may be considered a molecular biomarker in
HBV infection. To confirm these findings, further studies with
larger sample sizes with other SNPs in both genes are required to
clarify the associations between SNPs and their susceptibility to
HBV infection in the Egyptian population.
Acknowledgements
Not applicable.
Funding
Funding: No funding was received.
Availability of data and materials
The datasets generated during and/or analyzed in the
current study are available from the corresponding author on
reasonable request.
Authors' contributions
The present study was carried out in collaboration
between all authors. RMT designed the research and contributed new
reagents/analytic tools. SZE provided the patient samples and
clinical data. AEM and EAEM performed the experiments. RMT and EAEM
analyzed and interpreted the data. SMR and TKM participated in the
experimental design and in the writing of the manuscript. RMT, AEM
and EAEM confirm the authenticity of the raw data. All authors
wrote, and have read and approved the final manuscript.
Ethics approval and consent to
participate
All subjects provided written informed consent for
genetic analysis in the present observational prospective
case-control study. All methods and analyses were carried out
followingtheguidelines of the Ministry of Health and approved by
the Research Ethics Committee for Experimental and Clinical Studies
at the Faculty of Pharmacy, University of Cairo, Egypt [BC
(1837)].
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Ezbarami ZT, Hassani P, Tafreshi MZ and
Majd HA: A qualitative study on individual experiences of chronic
hepatitis B patients. Nurs Open. 4:310–318. 2017.PubMed/NCBI View
Article : Google Scholar
|
|
2
|
Asgari S, Chaturvedi N, Scepanovic P,
Hammer C, Semmo N, Giostra E, Müllhaupt B, Angus P, Thompson AJ,
Moradpour D and Fellay J: Human genomics of acute liver failure due
to hepatitis B virus infection: An exome sequencing study in liver
transplant recipients. J Viral Hepat. 26:271–277. 2019.PubMed/NCBI View Article : Google Scholar
|
|
3
|
Revill P, Testoni B, Locarnini S and
Zoulim F: Global strategies are required to cure and eliminate HBV
infection. Nat Rev Gastroenterol Hepatol. 13:239–248.
2016.PubMed/NCBI View Article : Google Scholar
|
|
4
|
Alavi M, Grebely J, Hajarizadeh B, Amin J,
Larney S, Law MG, George J, Degenhardt L and Dore GJ: Mortality
trends among people with hepatitis B and C: A population-based
linkage study, 1993-2012. BMC Infect Dis. 18(215)2018.PubMed/NCBI View Article : Google Scholar
|
|
5
|
Cao MX, Jiang YP, Tang YL and Liang X: The
crosstalk between lncRNA and microRNA in cancer metastasis:
Orchestrating the epithelial-mesenchymal plasticity. Oncotarget.
8:12472–12483. 2017.PubMed/NCBI View Article : Google Scholar
|
|
6
|
Yamamura S, Imai-Sumida M, Tanaka Y and
Dahiya R: Interaction and cross-talk between non-coding RNAs. Cell
Mol Life Sci. 75:467–484. 2018.PubMed/NCBI View Article : Google Scholar
|
|
7
|
Tripathi R, Chakraborty P and Varadwaj PK:
Unraveling long non-coding RNAs through analysis of high-throughput
RNA-sequencing data. Noncoding RNA Res. 2:111–118. 2017.PubMed/NCBI View Article : Google Scholar
|
|
8
|
Bayoumi AS, Sayed A, Broskova Z, Teoh JP,
Wilson J, Su H, Tang YL and Kim IM: Crosstalk between long
non-coding RNAs and microRNAs in health and disease. Int J Mol Sci.
17(356)2016.PubMed/NCBI View Article : Google Scholar
|
|
9
|
Kohlhapp FJ, Mitra AK, Lengyel E and Peter
ME: MicroRNAs as mediators and communicators between cancer cells
and the tumor microenvironment. Oncogene. 34:5857–5868.
2015.PubMed/NCBI View Article : Google Scholar
|
|
10
|
Deng K, Wang H, Guo X and Xia J: The
crosstalk between long, non-coding RNAs and microRNAs in gastric
cancer. Acta Biochim Biophys Sin (Shanghai). 48:111–116.
2015.PubMed/NCBI View Article : Google Scholar
|
|
11
|
Paneru B, Ali A, Al-Tobasei R, Kenney B
and Salem M: Crosstalk among lncRNAs, microRNAs and mRNAs in the
muscle ‘degradome’ of rainbow trout. Sci Rep.
8(8416)2018.PubMed/NCBI View Article : Google Scholar
|
|
12
|
Qin C, Huang RY and Wang ZX: Potential
role of miR-100 in cancer diagnosis, prognosis, and therapy. Tumor
Biol. 36:1403–1409. 2015.PubMed/NCBI View Article : Google Scholar
|
|
13
|
Lin Y, Deng W, Pang J, Kemper T, Hu J, Yin
J, Zhang J and Lu M: The microRNA-99 family modulates hepatitis B
virus replication by promoting IGF-1R/PI3K/Akt/mTOR/ULK1
signaling-induced autophagy. Cell Microbiol. 19:1–15.
2017.PubMed/NCBI View Article : Google Scholar
|
|
14
|
Tan JY and Marques AC: miRNA-mediated
crosstalk between transcripts: The missing ‘linc’? Bioessays.
38:295–301. 2016.PubMed/NCBI View Article : Google Scholar
|
|
15
|
He RZ, Luo DX and Mo YY: Emerging roles of
lncRNAs in the post-transcriptional regulation in cancer. Genes
Dis. 6:6–15. 2019.PubMed/NCBI View Article : Google Scholar
|
|
16
|
Bhat SA, Ahmad SM, Mumtaz PT, Malik AA,
Dar MA, Urwat U, Shah RA and Ganai NA: Long non-coding RNAs:
Mechanism of action and functional utility. Noncoding RNA Res.
1:43–50. 2016.PubMed/NCBI View Article : Google Scholar
|
|
17
|
Fernandes JCR, Acuña SM, Aoki JI,
Floeter-Winter LM and Muxel SM: Long non-coding RNAs in the
regulation of gene expression: Physiology and disease. Noncoding
RNA. 5(17)2019.PubMed/NCBI View Article : Google Scholar
|
|
18
|
Amodio N, Raimondi L, Juli G, Stamato MA,
Caracciolo D, Tagliaferri P and Tassone P: MALAT1: A druggable long
non-coding RNA for targeted anti-cancer approaches. J Hematol
Oncol. 11(63)2018.PubMed/NCBI View Article : Google Scholar
|
|
19
|
Ren D, Li H, Li R, Sun J, Guo P, Han H,
Yang Y and Li J: Novel insight into MALAT-1 in cancer: Therapeutic
targets and clinical applications. Oncol Lett. 11:1621–1630.
2016.PubMed/NCBI View Article : Google Scholar
|
|
20
|
Sun Y and Ma L: New insights into long
non-coding RNA malat1 in cancer and metastasis. Cancers (Basel).
11(216)2019.PubMed/NCBI View Article : Google Scholar
|
|
21
|
Zhao M, Wang S, Li Q, Ji Q, Guo P and Liu
X: Malat1: A long non-coding RNA highly associated with human
cancers. Oncol Lett. 16:19–26. 2018.PubMed/NCBI View Article : Google Scholar
|
|
22
|
Fareed M and Afzal M: Single nucleotide
polymorphism in genome-wide association of human population: A tool
for broad spectrum service. Egypt J Med Hum Genet. 14:123–134.
2013.
|
|
23
|
Castellanos-Rubio A and Ghosh S:
Disease-associated SNPs in inflammation-related lncRNAs. Front
Immunol. 10(420)2019.PubMed/NCBI View Article : Google Scholar
|
|
24
|
Motawi TK, Mady AE, Shaheen S, Elshenawy
SZ, Talaat RM and Rizk SM: Genetic variation in microRNA-100
(miR-100) rs1834306 T/C associated with Hepatitis B virus (HBV)
infection: Correlation with expression level. Infect Genet Evol.
73:444–449. 2019.PubMed/NCBI View Article : Google Scholar
|
|
25
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta DeltaC(T)) method. Methods. 25:402–408.
2001.PubMed/NCBI View Article : Google Scholar
|
|
26
|
Fedeli U, Grande E, Grippo F and Frova L:
Mortality associated with hepatitis C and hepatitis B virus
infection: A nationwide study on multiple causes of death data.
World J Gastroenterol. 23:1866–1871. 2017.PubMed/NCBI View Article : Google Scholar
|
|
27
|
Lok AS, Zoulim F, Dusheiko G and Ghany MG:
Hepatitis B cure: From discovery to regulatory approval.
Hepatology. 66:1296–1313. 2017.PubMed/NCBI View Article : Google Scholar
|
|
28
|
Ahmadi A, Kaviani S, Yaghmaie M,
Pashaiefar H, Ahmadvand M, Jalili M, Alimoghaddam K, Eslamijouybari
M and Ghavamzadeh A: Altered expression of MALAT 1 lncRNA in
chronic lymphocytic leukemia patients, correlation with cytogenetic
findings. Blood Res. 53:320–324. 2018.PubMed/NCBI View Article : Google Scholar
|
|
29
|
Frías-Lasserre D and Villagra CA: The
importance of ncRNAs as epigenetic mechanisms in phenotypic
variation and organic evolution. Front Microbiol.
8(2483)2017.PubMed/NCBI View Article : Google Scholar
|
|
30
|
Wu M, Lin Z, Li X, Xin X, An J, Zheng Q,
Yang Y and Lu D: HULC cooperates with MALAT1 to aggravate liver
cancer stem cells growth through telomere repeat-binding factor 2.
Sci Rep. 6(36045)2016.PubMed/NCBI View Article : Google Scholar
|
|
31
|
Liu Y, Pan S, Liu L, Zhai X, Liu J, Wen J,
Zhang Y, Chen J, Shen H and Hu Z: A genetic variant in long
non-coding RNA HULC contributes to risk of HBV-related
hepatocellular carcinoma in a Chinese population. PLoS One.
7(e35145)2012.PubMed/NCBI View Article : Google Scholar
|
|
32
|
Zhao K, Jin S, Wei B, Cao S and Xiong Z:
Association study of genetic variation of lncRNA MALAT1 with
carcinogenesis of colorectal cancer. Cancer Manag Res.
10:6257–6261. 2018.PubMed/NCBI View Article : Google Scholar
|
|
33
|
Motawi TMK, El-Maraghy SA, Sabry D and
Mehana NA: The expression of long non-coding RNA genes is
associated with expression with polymorphisms of HULC rs7763881 and
MALAT1 rs619586 in hepatocellular carcinoma and HBV Egyptian
patients. J Cell Biochem. 120:14645–14656. 2019.PubMed/NCBI View Article : Google Scholar
|
|
34
|
Yuan LT, Chang JH, Lee HL, Yang YC, Su SC,
Lin CL, Yang SF and Chien MH: Genetic variants of lncRNA MALAT1
exert diverse impacts on the risk and clinicopathologic
characteristics of patients with hepatocellular carcinoma. J Clin
Med. 8(1406)2019.PubMed/NCBI View Article : Google Scholar
|
|
35
|
Wang BG, Xu Q, Lv Z, Fang XX, Ding HX, Wen
J and Yuan Y: Association of twelve polymorphisms in three
onco-lncRNa genes with hepatocellular cancer risk and prognosis: A
case-control study. World J Gastroenterol. 24:2482–2490.
2018.PubMed/NCBI View Article : Google Scholar
|
|
36
|
Gong WJ, Peng JB, Yin JY, Li XP, Zheng W,
Xiao L, Tan LM, Xiao D, Chen YX, Li X, et al: Association between
well-characterized lung cancer lncRNA polymorphisms and
platinum-based chemotherapy toxicity in Chinese patients with lung
cancer. Acta Pharmacol Sin. 38:581–590. 2017.PubMed/NCBI View Article : Google Scholar
|
|
37
|
Li Q, Zhu W, Zhang B, Wu Y, Yan S, Yuan Y,
Zhang H, Li J, Sun K, Wang H and Yu T: The MALAT1 gene polymorphism
and its relationship with the onset of congenital heart disease in
Chinese. Biosci Rep. 38(BSR20171381)2018.PubMed/NCBI View Article : Google Scholar
|
|
38
|
Zhu R, Liu X and He Z: Long non-coding RNA
H19 and MALAT1 gene variants in patients with ischemic stroke in a
northern Chinese Han population. Mol Brain. 11(58)2018.PubMed/NCBI View Article : Google Scholar
|
|
39
|
Wang ML and Liu JX: MALAT1 rs619586
polymorphism functions as a prognostic biomarker in the management
of differentiated thyroid carcinoma. J Cell Physiol. 235:1700–1710.
2020.PubMed/NCBI View Article : Google Scholar
|
|
40
|
Che D, Yang Y, Xu Y, Fang Z, Pi L, Fu L,
Zhou H, Tan Y, Lu Z, Li L, et al: The lncRNA MALAT1 rs619586 G
variant confers decreased susceptibility to recurrent miscarriage.
Front Physiol. 10(385)2019.PubMed/NCBI View Article : Google Scholar
|
|
41
|
Konishi H, Ichikawa D, Yamamoto Y, Arita
T, Shoda K, Hiramoto H, Hamada J, Itoh H, Fujita Y, Komatsu S, et
al: Plasma level of metastasis-associated lung adenocarcinoma
transcript 1 is associated with liver damage and predicts
development of hepatocellular carcinoma. Cancer Sci. 107:149–154.
2016.PubMed/NCBI View Article : Google Scholar
|
|
42
|
Pan Y, Tong S, Cui R, Fan J, Liu C, Lin Y,
Tang J, Xie H, Lin P, Zheng T and Yu X: Long non-coding MALAT1
functions as a competing endogenous RNA to regulate vimentin
expression by sponging miR-30a-5p in hepatocellular carcinoma. Cell
Physiol Biochem. 50:108–120. 2018.PubMed/NCBI View Article : Google Scholar
|
|
43
|
Zhao ZB, Chen F and Bai XF: Long
non-coding RNA MALAT1 regulates hepatocellular carcinoma growth
under hypoxia via sponging microRNA-200a. Yonsei Med J. 60:727–734.
2019.PubMed/NCBI View Article : Google Scholar
|
|
44
|
Wang Z, Katsaros D, Biglia N, Shen Y, Fu
Y, Loo LWM, Jia W, Obata Y and Yu H: High expression of long
non-coding RNA MALAT1 in breast cancer is associated with poor
relapse-free survival. Breast Cancer Res Treat. 171:261–271.
2018.PubMed/NCBI View Article : Google Scholar
|
|
45
|
Zhu K, Ren Q and Zhao Y: . LncRNA MALAT1
overexpression promotes proliferation, migration and invasion of
gastric cancer by activating the PI3K/AKT pathway. Oncol Lett.
17:5335–5342. 2019.PubMed/NCBI View Article : Google Scholar
|
|
46
|
Toraih EA, Ellawindy A, Fala SY, Al Ageeli
E, Gouda NS, Fawzy MS and Hosny S: Oncogenic long non-coding RNA
MALAT1 and HCV-related hepatocellular carcinoma. Biomed
Pharmacother. 102:653–669. 2018.PubMed/NCBI View Article : Google Scholar
|
|
47
|
Wang X, Li M, Wang Z, Han S, Tang X, Ge Y,
Zhou L, Zhou C, Yuan Q and Yang M: Silencing of long non-coding RNA
MALAT1 by miR-101 and miR-217 inhibits proliferation, migration,
and invasion of esophageal squamous cell carcinoma cells. J Biol
Chem. 290:3925–3935. 2015.PubMed/NCBI View Article : Google Scholar
|
|
48
|
Liu D, Zhu Y, Pang J, Weng X, Feng X and
Guo Y: Knockdown of long non-coding RNA MALAT1 inhibits growth and
motility of human hepatoma cells via modulation of miR-195. J Cell
Biochem. 119:1368–1380. 2018.PubMed/NCBI View Article : Google Scholar
|