Introduction
Breast cancer is the most common malignancy in
women, accounting for 627,000 deaths worldwide in 2018(1). Lymph node involvement is one of the
most important prognostic factors in breast cancer (2). Axillary lymph node dissection (ALND)
significantly reduces recurrence and improves regional control and
nodal staging, which is important for the selection of adjuvant
therapy (3) and prognostic
evaluation (4). However, ALND is
associated with various adverse side effects, such as lymphedema,
numbness, chronic pain, seroma or infection, and its impact on
survival and recurrence is subject to controversy (5,6).
Sentinel lymph node biopsy (SLNB) is a common
procedure used to detect the presence of metastatic cells and to
decide whether ALND is required. SLNB may also help with breast
cancer staging (7). Histological
examination of step section or serial section slides of SLNs is the
most widely used method. One-step nucleic acid amplification (OSNA;
Sysmex Corporation) is an alternative loop-mediated isothermal
amplification (LAMP)-based semi-quantitative assay that quantifies
copies of cytokeratin (CK)19 mRNA, which is expressed in most
breast cancer cells (8). The
determination of CK19 mRNA copy number can predict the presence of
micro- or macro-metastases in the SNL (9). Several studies have shown that OSNA
is more sensitive and objective compared with histological
examination (10). It is also a
cost-effective strategy (11,12)
that is widely used in Europe and Japan (13,14).
OSNA is an intraoperative procedure; therefore, ALND
can be performed during the same surgery, thereby avoiding a second
surgery. The OSNA procedure also makes it possible to commence
adjuvant treatment earlier (15).
It is also important to identify patients in whom ALND can safely
be avoided, without increasing the risk of recurrence (16,17).
The aim of this retrospective study was to determine
whether CK19 mRNA copy number in the SNL could predict positivity
of ALND.
Materials and methods
Study population
A total of 812 patients with early-stage invasive
breast cancer underwent breast surgery, SNL biopsy and OSNA
analysis between January 2010 and August 2014 at the Institut de
Cancérologie de Lorraine (ICL; Vandoeuvre-lès-Nancy, France).
Surgery and OSNA procedures were decided for all patients with
clinically or ultrasonographically node-negative cT1-2 breast
cancer. All patients provided informed oral consent and a signed a
non-opposition form and the study was approved by the Ethics
Committee of the ICL (CAV-2009-osna).
The exclusion criteria were as follows: i) Patients
who had received neoadjuvant treatment, had undergone previous
ipsilateral breast or axillary surgery, had cT3-T4 tumors, and
clinically or ultrasonographically positive axilla confirmed by
fine-needle aspiration biopsy; ii) a total of 20 patients were
excluded due to positive inhibition status (+I), corresponding to
values greater than the highest point of the calibration curve,
making the determination of exact number of copies not possible;
and iii) a total of 49 patients were also excluded as one central
slice of their SLNs had been investigated by histology, potentially
decreasing the number of copies of CK19 mRNA detected. Data
collected from each patient are listed in Table I. Data for the current study were
obtained from the prospective breast cancer database at the ICL.
All data were anonymized prior to analysis to protect patient
confidentiality.
 | Table IComparison of clinicopathological
characteristics between the included and the excluded patients. |
Table I
Comparison of clinicopathological
characteristics between the included and the excluded patients.
|
Characteristics | All patients
(n=812), n (%) | Included (n=197), n
(%) | Excluded (n=615), n
(%) | P-value |
|---|
| Age, years (mean ±
SD) | 60±11 | 59±11 | 61±12 | 0.264 |
| Body mass index ≥30
kg/m2 | 170 (20.9) | 42 (21.3) | 128 (20.8) | 0.879 |
| Tumor size ≥13
mm | 377 (46.4) | 111 (56.3) | 266 (43.2) | 0.001 |
| Bloom-Richardson
histological grade | | | | |
|
1 | 229 (29.6) | 59 (30.3) | 170 (29.4) | 0.787 |
|
2 | 397 (51.3) | 102 (52.3) | 295 (50.9) | |
|
3 | 148 (19.1) | 34 (17.4) | 114 (19.7) | |
| Tumor
localization | | | | |
|
Outer or
lower-outer quadrant | 165 (20.7) | 35 (17.8) | 130 (21.6) | 0.245 |
|
Other | 633 (79.3) | 162 (82.2) | 471 (78.4) | |
| Histological
type | | | | |
|
Ductal | 598 (73.6) | 153 (77.7) | 445 (72.4) | 0.070 |
|
Lobular | 76 (9.4) | 21 (10.7) | 55 (8.9) | |
|
Other | 138 (17.0) | 23 (11.7) | 115 (18.7) | |
| Positive ER
status | 742 (91.4) | 183 (92.9) | 559 (90.9) | 0.384 |
| Positive PR
status | 635 (78.2) | 156 (79.2) | 479 (77.9) | 0.700 |
| Positive ER and/or
PR status | 751 (92.5) | 183 (92.3) | 568 (92.5) | 0.804 |
| Positive HER2
receptor status | 52 (6.4) | 13 (6.6) | 39 (6.3) | 0.898 |
| Triple-negative
breast cancer | 47 (5.8) | 12 (6.1) | 35 (5.7) | 0.834 |
| Sentinel lymph
nodes removed, median (range) | 3 (2-4) | 2 (2-4) | 3 (2-4) | 0.081 |
SLN biopsy procedure
SLNs were localized using the isotope method, alone
or combined with the dye procedure. The isotope method consisted of
99mTc-labeled rhenium sulfur (Amersham; Cytiva)
periareolar injection the day before surgery, followed by
lymphoscintigraphy 1-3 h later. The dye procedure consisted of 2 ml
of patent blue dye (Guerbet) administered by subareolar injection
at surgery. SLNs were identified using a hand-held gamma-probe
(Europrobe 3; Euromedical Instruments), isolated, and perinodal fat
was removed. All suspicious lymph nodes identified during surgery
were sent for analysis. Data on the SLNs included their color (blue
or not), localization, signal intensity and size.
Lymph nodes with a weight of >0.6 g were
subdivided into two or more samples and processed separately, as
recommended by the manufacturer of the OSNA assay (Sysmex
Corporation). A maximum of 4 samples were assessed per run, for a
total running time of 15-60 min for 1-4 samples, respectively.
Histopathology
Each non-SNL was measured, cut longitudinally into
2-mm sections, fixed in formalin for 8 h at room temperature and
embedded in paraffin. The sections were then prepared for
hematoxylin and eosin staining.
Breast tumors were examined by hematoxylin and eosin
staining. CK19 (clone RCK 108; cat. no. M0888; Agilent
Technologies, Inc.), hormonal receptors, including estrogen
receptor (ER; clone SP1; cat. no. 790-4325) and progesterone
receptor (PR; clone 1E2; cat. no. 790-4296; Ventana Medical
Systems, Inc.; Roche Diagnostics), HER2 (clone 4B5; cat. no.
790-4493; Ventana Medical Systems, Inc.; Roche Diagnostics) and
Ki-67 (clone MIB-1; cat. no. M7240; Agilent Technologies, Inc.)
expression were determined using immunohistochemistry. All assays
were automated using Benchmark (Roche Diagnostics) according to the
manufacturer's protocols. Histopathological categories were defined
according to the sixth edition of the TNM classification (18).
OSNA analysis and mRNA CK19 copy
determination
The OSNA assay was processed as previously described
using the OSNA BC System (Sysmex Corporation) (9). Briefly, whole SLNs were homogenized
in 4 ml Lynorhag lysis buffer (Sysmex Corporation). The homogenate
was centrifuged at 10,000 x g for 1 min at room temperature and
directly used as a template for amplification. CK19 mRNA detection
was assessed using reverse transcription-LAMP with the RD-100i
analyzer (Sysmex Corporation).
Results for each sample were presented on the
RD-100i instrument in qualitative categories along with the CK19
mRNA copy number/µl. The (-), (+), (++) and (+I) symbols were used
by the OSNA instrument to indicate copy numbers of <250,
250-5,000, >5,000 and greater than the highest point of the
standard curve, respectively.
According to the cut-off levels defined by Tsujimoto
et al (9), a copy number
between 250 and 5,000 copies/µl (+) was considered as predictive of
the presence of SLN micrometastases in the analyzed lymph node, and
a copy number >5,000/µl (++) was considered as predictive of the
presence of SLN macrometastases. A copy number <250 copies/µl
was considered as predictive of the absence of tumor cells.
The number of copies was then estimated using the
number of copies measured in a 1/10 dilution of the sample. The
node total copy number was estimated by adding CK19 mRNA copies of
each piece of the sample, in nodes weighing >0.6 g. Tubes
containing more than one node for the same patient were excluded
from the analysis. Only the SLN with the highest number of copies
was considered for each patient.
Statistical analysis
Statistical analysis was performed using SAS
software version 9.4 (SAS Institute Inc.). The significance level
was set at 0.05. Qualitative variables are described as number and
percentage, and quantitative variables as mean ± standard
deviation, or median and interquartile range (IQR), according to
the normality test (Kolmogorov-Smirnov test). Predictive factors of
positive ALND were investigated using bivariate logistic regression
and the results are expressed as ORs and 95% CIs. The log-linearity
assumption of the logistic model was checked by categorizing each
variable in 10 groups (corresponding to deciles) and by examining
the plots of the logit of observed percentages of positive ALND in
each class. Quantitative variables were transformed into binary
variables if the log-linearity assumption was violated, using the
threshold maximizing sensitivity and specificity (Youden index).
All variables with a P-value <0.10 in bivariate logistic
regression were included in a multivariate logistic regression
model with backward selection at P=0.10. The results of the final
multivariate model are presented as adjusted ORs (95% CIs). The
stability of the selected model was investigated using the
bootstrap resampling method (19).
Results
Study population
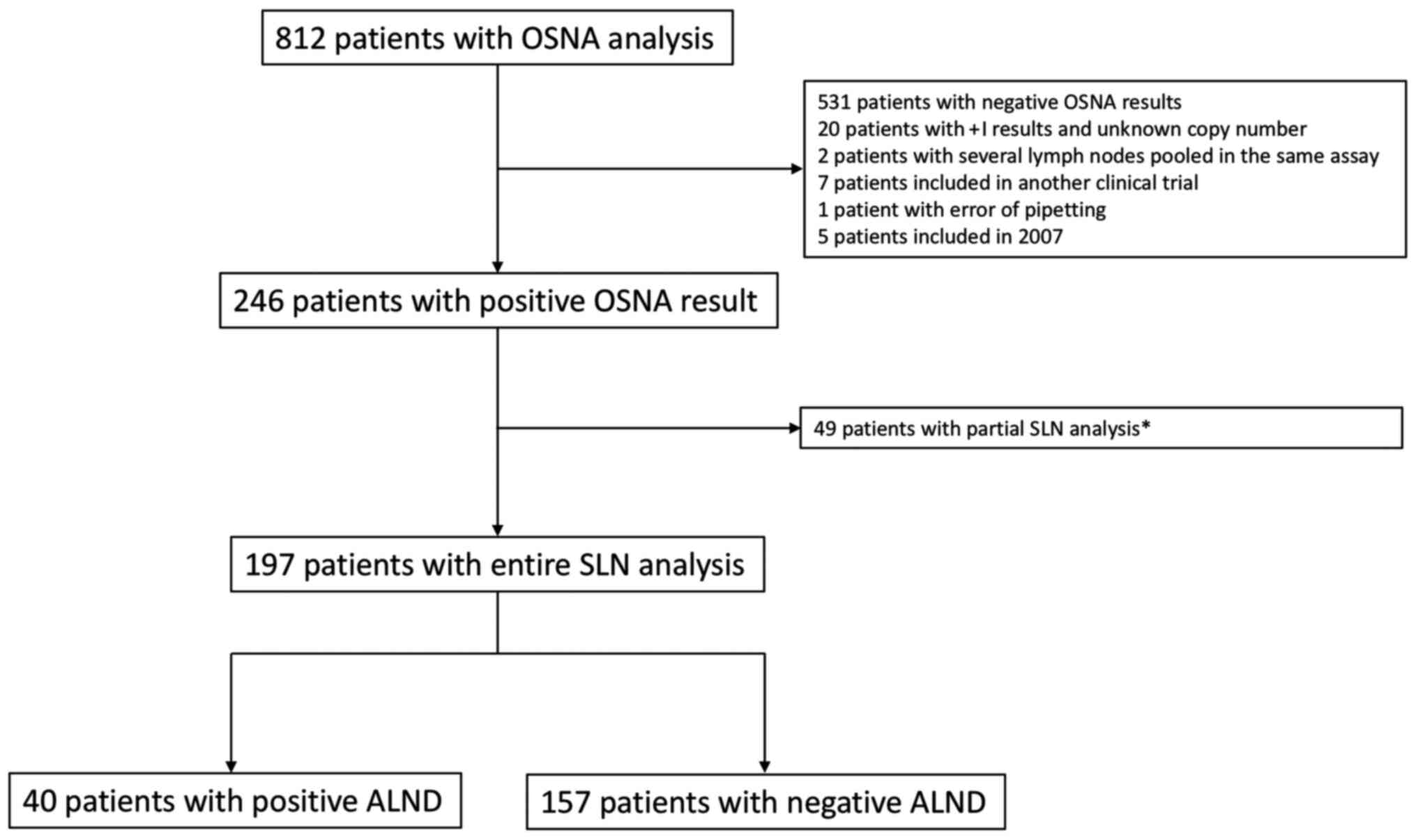
Among the 812 patients who underwent OSNA analysis,
246 patients had at least one positive SLN. Among these, a total of
197 patients with positive OSNA analysis were included in this
retrospective study (Fig. 1). A
comparison of the characteristics of included (n=197) vs. excluded
(n=615) patients is presented in Table
I. Patient and disease characteristics are summarized in
Table II. Patients with SLN
micro- or macrometastases as determined by OSNA underwent ALND.
Patients with a negative OSNA result did not undergo further
ALND.
 | Table IIClinicopathological characteristics
for the 197 patients and according to the positivity of ALND. |
Table II
Clinicopathological characteristics
for the 197 patients and according to the positivity of ALND.
|
Characteristics | All patients
(n=197), n (%) | Negative ALND
(n=157), n (%) | Positive ALND
(n=40), n (%) | P-value |
|---|
| Age, years (mean ±
SD) | 59±11 | 60±11 | 58±13 | 0.296 |
| Body mass index ≥30
kg/m2 | 42 (21.3) | 29 (18.5) | 13 (32.5) | 0.053 |
| Tumor size ≥13
mma | 111 (56.3) | 82 (52.2) | 29 (72.5) | 0.021 |
| Bloom-Richardson
histological grade | | | | 0.548 |
|
1 | 59 (30.3) | 49 (31.6) | 10 (25.0) | |
|
2 | 102 (52.3) | 78 (50.3) | 24 (60.0) | |
|
3 | 34 (17.4) | 28 (18.1) | 6 (15.0) | |
| Tumor
localization | | | | 0.023 |
|
Outer or
lower-outer quadrant | 35 (17.8) | 23 (14.6) | 12 (30.0) | |
|
Other | 162 (82.2) | 134 (85.4) | 28(70) | |
| Histological
type | | | | |
|
Ductal | 153 (77.7) | 118 (75.2) | 35 (87.5) | 0.144 |
|
Lobular | 21 (10.7) | 20 (12.7) | 1 (2.5) | |
|
Other | 23 (11.7) | 19 (12.1) | 4 (10.0) | |
| Positive ER
status | 183 (92.9) | 145 (92.4) | 38 (95.0) | 0.739 |
| Positive PR
status | 156 (79.2) | 121 (77.1) | 35 (87.5) | 0.147 |
| Positive ER and/or
PR status | 183 (92.3) | 145 (92.4) | 38 (95.0) | 0.739 |
| Positive HER2
receptor status | 13 (6.6) | 11 (7.0) | 2 (5.0) | 1 |
| Triple-negative
breast cancer | 12 (6.1) | 10 (6.4) | 2 (5.0) | 1 |
| SLNs removed,
median (range) | 2 (2-4) | 2 (2-3) | 2.5 (2-4) | 0.38 |
| Positive SLNs,
median (range) | 1 (1-1) | 1 (1-1) | 1 (1-2) | 0.23 |
|
1 | 147 (74.8) | 121 (77.1) | 26 (65.0) | |
|
2 | 39 (19.4) | 30 (19.1) | 9 (22.5) | |
|
3 | 10 (5.2) | 6 (3.8) | 4 (10.0) | |
|
4 | 1 (0.6) | 0 | 1 (2.5) | |
| SLN maximal copy
number/µl, median (range) | 2,100
(540-38,000) | 1,300
(530-6,900) | 60,830
(5,000-695,000) | <0.001 |
|
≥4,700a | 77 (39.1) | 46 (29.3) | 31 (77.5) | <0.001 |
|
≥5,000 | 74 (37.6) | 44 (28.0) | 30 (75.0) | <0.001 |
OSNA analysis
The median number of SLNs removed was 2 (IQR, 2-4).
A total of 123 patients (62%) had SLN micrometastases, while 74 had
SLN macrometastases (38%). The median number of ALNs removed was 15
(IQR, 13-19.5) with a median of 2 (IQR, 1-3) positive ALNs. A total
of 40 patients (20%) had non-SNL metastases. The patient
characteristics according to lymph node status are presented in
Table II. A tumor size >13 mm
localized in the outer quadrant (OQ) or lower-outer quadrant (LOQ)
of the breast was more frequent in the group with positive ALND.
Two or more positive SLNs were found in all patients with positive
ALND. The CK19 mRNA copy number was also higher in the positive
ALND group [median, 60,830 (IQR, 5,000-695,000) vs. 1,300 (IQR,
530-6,900)]. The threshold of 4,700 CK19 mRNA copies was found to
be the optimal cut-off for distinguishing patients with vs. those
without non-SNLs. The value of 5,000 CK19 mRNA copies commonly used
to differentiate micrometastases and macrometastases was retained,
since only 3 patients had values between 4,700 and 5,000. A total
of 30 patients of the group with positive ALND (75%) had SLN
macrometastases vs. only 28% in the group with negative ALND,
corresponding to a specificity of 72% (113/157). The positive
predictive value was 40.5% (30/74) and the negative predictive
value (NPV) was 92% (113/123). The factors predictive of positive
ALND are presented in Table III,
whereas a comparison of the OSNA cut-off of the present study with
other alternatives from the literature for prediction of non-SLN
metastasis is presented in Table
IV.
 | Table IIIPredictive factors for positive
axillary lymph node dissection by bivariate and multivariate
logistic regression analysis. |
Table III
Predictive factors for positive
axillary lymph node dissection by bivariate and multivariate
logistic regression analysis.
| | Bivariate
analyses | Multivariate
analysis |
|---|
| Factors | OR (95% CI) | P-value | OR (95% CI) | P-value |
|---|
| Age (per 1-year
increase) | 0.98
(0.95-1.01) | 0.295 | | |
| Body mass index ≥30
kg/m2 | 2.12
(0.98-4.61) | 0.056 | | |
| Tumor size >13
mm | 2.41
(1.13-5.16) | 0.023 | | |
| Bloom-Richardson
histological grade | | 0.551 | | |
|
1 | 1.0
(reference) | | | |
|
2 | 1.51
(0.66-3.42) | | | |
|
3 | 1.05
(0.34-3.20) | | | |
| Tumor localization
in the outer or lower-outer quadrant | 2.50
(1.11-5.60) | 0.026 | 2.84
(1.14-7.05) | 0.025 |
| Ductal
carcinoma | 2.31
(0.85-6.32) | 0.102 | | |
| Copy number
≥5,000/µl | 7.70
(3.48-17.08) | <0.001 | 8.07
(3.58-18.23) | <0.001 |
| Positive ER
status | 1.57
(0.34-7.33) | 0.564 | | |
| Positive PR
status | 2.08
(0.76-5.71) | 0.154 | | |
| Positive ER or PR
status | 1.57
(0.34-7.38) | 0.564 | | |
| Positive HER2
status | 0.70
(0.15-3.29) | 0.650 | | |
| Triple-negative
breast cancer | 0.77
(0.16-3.68) | 0.747 | | |
 | Table IVComparison of our OSNA cut-off with
other alternatives from literature for prediction of non-SLN
metastasis. |
Table IV
Comparison of our OSNA cut-off with
other alternatives from literature for prediction of non-SLN
metastasis.
| Study (Refs.) | Number of
patients | Threshold
(copies/µl) | Se | Sp | PPV | NPV | FN, n (%) | FP, n (%) | Method | AUC | Problems |
|---|
| Present study | 812 patients 197
OSNA+ | 5,000 | 75.0 | 72.0 | 40.5 | 91.9 | 10 (8.1) | 44 (59.5) | Maximal copy
number | 0.77 | |
| Peg et al
(37) | 697 patients
OSNA+ | 15,000 | 76.7 | 55.2 | 41.1 | 85.5 | 14.7% | | TTL | 0.709 | T1-T3 breast
tumors. |
| Deambrogio et
al (38) | 1,080 patients 194
OSNA+ | 7,700 | 78 | 57 | 50 | 83 | 15 (17.4) | 54(50) | | | T1-T3 breast
tumors. 46 patients with OSNA+ analysis did not undergo further
surgery. |
| Heilmann et
al (39) | 143 patients 39
OSNA+ | 7,900 | 91 | 61 | | | | | | | T1-T3 breast
tumors. Part of the lymph node analyzed by histology. |
| Terrenato et
al (20) | 1,140 patients 318
OSNA+ | 2,150 | 94.9 | 51.4 | 46.5 | 95.8 | 5 (4.2) | 107 (53.5) | TTL | 0.765 | No description of
+I case management. Lack of representation of cancers other than
ductal or lobular carcinomas. |
| Nabais et al
(40) | 598 patients 58
OSNA+ | 190,000 | 73.3 | 74.4 | | 88.9 | | | TTL | 0.805 | T1-T3 breast
tumors. |
| Banerjee et
al (41) | 170 patients 49
OSNA+ | 1,400 | | | | | | | | | T1-T3 breast
tumors. 50% of the lymph node analyzed by OSNA. |
| Espinosa-Bravo
et al (42) | 306 patients 108
OSNA+ | 120,000 | 47 | 85.3 | 56 | 80 | | | TTL | | |
| Buglioni et
al (43) | 709 patients 179
OSNA+ | 2,000 | | | | | | | | | 50% of the lymph
node analyzed by OSNA. |
By multivariate analysis, two parameters remained
significantly associated with positive ALND, namely SLN
macrometastases (OR=8.07, 95% CI: 3.58-18.23) and tumor
localization in the OQ [centered around the 3 o'clock position
(left breast) or 9 o'clock position (right breast)] or LOQ
(OR=2.84; 95% CI: 1.14-7.05). The AUC was 0.77 (95% CI: 0.69-0.85).
Considering that 20% of the patients had non-SLN metastases in our
population (40/197), the estimated probability from the
multivariate model was 36% for patients with SLN macrometastases
and tumor outside of the OQ or LOQ, and 61% for patients with SLN
macrometastases and tumor localization within the OQ or LOQ. The
estimated probability of positive ALND for patients with
micrometastases was 6% in case of tumors located outside of the OQ
or LOQ and 16% in case of tumors within the OQ or LOQ.
Discussion
Intraoperative SLN evaluation has limited ability to
detect metastases due to the partial evaluation of the node
(10,11). SLN histopathological examination is
thus performed postoperatively in several centers. Since 2007, the
OSNA assay has been used as an objective, simple and automated tool
for the intraoperative assessment of whole SLNs. The high
concordance of OSNA with histological techniques has been shown in
several studies (20-23),
as has its high sensitivity and specificity (24). OSNA avoids sampling errors and
second-stage surgeries due to false-negative results, without
increasing operative time, except in breast-conserving surgery
(25).
However, the need for ALND in SLN-positive early
breast cancer remains controversial. ALND is a possible cause of
morbidity, incurs greater costs and is associated with lower
quality of life (26).
Furthermore, the selection of adjuvant therapy currently relies
more on the characteristics of the primary tumor rather than on the
number of affected lymph nodes. Several studies have shown poorer
prognosis and a higher recurrence rate in cases with SLN
micrometastases without adjuvant therapy (6,27,28).
In line with the findings of the IBCSG 23-01 trial (5), the American College of Surgeons
Oncology Group (ACOSOG) Z0011 randomized trial (16) stated that ALND could be avoided in
patients with T1-T2 N0 breast cancer and 1-2 SLN metastases
undergoing breast-conserving surgery and receiving adjuvant
whole-breast irradiation and adjuvant systemic therapy. In the
AMAROS trial (17), the authors
demonstrated the non-inferiority of axillary radiotherapy vs. ALND
in patients with SLN micro- or macrometastases.
Due to certain limitations in these trials,
including a high rate of loss to follow-up (18.6% in the ACOSOG
study), and imbalances in several prognostic characteristics
between groups (29,16), several trials are still ongoing to
confirm these results (30). The
2015 National Comprehensive Cancer Network guidelines (14) already stated that ALND was not
necessary in the population of the ACOSOG trial. Current
recommendations in France indicate that ALND is necessary if SLN
macro-metastases are present, whereas multidisciplinary discussion
is recommended in case of micrometastases (31).
The use of objective tools capable of predicting
non-SLN axillary involvement could therefore be useful, at least
for patients who do not meet the Z0011 criteria, or who were
underrepresented in that trial (for example, patients with invasive
lobular carcinoma, estrogen receptor-negative status, or age <50
years). An optimal negative cut-off would also help to identify
patients who would not benefit from ALND, given that it can safely
be omitted if SLN is negative (32).
Many available prediction models for positive ALND
are based on factors that cannot be determined preoperatively and
are therefore not clinically relevant. The OSNA procedure can be
performed during surgery and is an independent predictive factor of
potential further axillary metastasis progression, with a good
diagnostic capacity [area under the receiver operating
characteristics curve (AUC) = 0.77 in the present study].
Predictive cut-offs of CK19 mRNA copy number have
already been investigated in other studies (20,37-43).
Some authors (43) considered the
maximal copy number, whereas others (20,37,40,42)
considered the total tumor load (TTL), defined by the number of
CK19 mRNA copies in all positive SLNs. TTL could be considered as
more representative of the tumor cell load, but is linked to the
number of SLNs analyzed during the procedure, which depends on the
highly variable standard practices in each center. This variability
in practices may explain the differences between the cut-offs
across published studies.
In some studies (39,41),
SLN sections were used for histological evaluation. Our
supplementary analysis including the 49 patients with histological
analysis of SLN sections confirmed that this practice may lead to
possible underestimation of CK19 mRNA total copies. This analysis
yielded a cut-off of 3,500 copies [AUC=0.741 (95% CI: 0.657-0.825),
data not shown]. A false-negative result would prevent some
patients from undergoing ALND. The undervaluation of copy number
could also explain the varying cut-offs reported in published
studies (40,41). There is potential for bias in our
study due to the exclusion of 20 +I cases the total copy number of
which was not available (33) and
because of the 49 cases with histological analysis of the central
section of the SLN. Some disadvantages of the OSNA assay must also
be considered, such as te inability to conduct further histological
analysis.
Shimazu et al (34) proposed an intraoperative nomogram
based on tumor size and TTL, but their NPV and AUC were lower
compared with those in the present study. Furthermore, a central
section was removed for histological examination in one
institution.
In our cohort of patients, when the copy number was
<5,000, 113 patients had no further axillary involvement (92%)
and only 10 patients (8%) had positive ALND. These results indicate
that ALND can safely be avoided when the tumor is localized outside
of the OQ or LOQ of the breast, and the copy number is <5,000. A
total of 30 ALNDs were positive when CK19 mRNA was >5,000 copies
(41%). These results support the concept that ALND must be
considered in this case, particularly when the tumor is in the OQ.
We believe that the high cut-offs described by Heilmann et
al (39), Deambrogio et
al (38) or Peg et al
(37) may result in a very high
false-negative rate. We herein confirmed that ALND can safely be
avoided in patients with tumors in the other quadrants if the CK19
mRNA copy number is <5,000. These results are almost in line
with previous published studies (20,38),
and the copy threshold for OSNA was confirmed based on a large
cohort. The present study may also help to overcome certain
drawbacks of previous studies (39,41),
such as partial evaluation of the node.
Predictive thresholds for non-SLN positivity should
be assessed in other cancers, such as cervical cancer, in which
pelvic lymphadenectomy results are negative in >80% of cases
(35). The OSNA assay may also
contribute to prognostic evaluation (36).
In conclusion, a cut-off of 5,000 copies for CK19
mRNA combined with tumor localization may represent an
intraoperative objective and useful tool for predicting further
non-SLN axillary involvement and the need for completion ALND in
patients with breast cancer.
Acknowledgements
Not applicable.
Funding
Funding: No funding was received.
Availability of data and materials
The datasets used and/or analyzed during the current
study are available from the corresponding author on reasonable
request.
Authors' contributions
Conceptualization, JLM, PR and FM; methodology, AH
and JLM; validation and interpretation of raw data, AH, FM and JLM;
formal analysis, JS; data collection, MK, MR, MH, PG and HP;
resources, PR, FM and JLM; data curation, MK and AH; original
manuscript draft preparation, HP; manuscript review and editing,
AH; visualization, HP and AH; supervision, AH; project
administration, JLM, AL and AH. All the authors have read and
approved the final version of the manuscript for publication.
Ethics approval and consent to
participate
All patients provided informed oral consent and a
signed a non-opposition form and the study was approved by the
Ethics Committee of the ICL (CAV-2009-osna).
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Bray F, Ferlay J, Soerjomataram I, Siegel
RL, Torre LA and Jemal A: Global cancer statistics 2018: GLOBOCAN
estimates of incidence and mortality worldwide for 36 cancers in
185 countries. CA Cancer J Clin. 68:394–424. 2018.PubMed/NCBI View Article : Google Scholar
|
|
2
|
Fisher B, Bauer M, Wickerham DL, Redmond
CK, Fisher ER, Cruz AB, Foster R, Gardner B, Lerner H, Margolese R,
et al: Relation of number of positive axillary nodes to the
prognosis of patients with primary breast cancer. An NSABP update.
Cancer. 52:1551–1557. 1983.PubMed/NCBI View Article : Google Scholar
|
|
3
|
Moore MP and Kinne DW: Axillary
lymphadenectomy: A diagnostic and therapeutic procedure. J Surg
Oncol. 66:2–6. 1997.PubMed/NCBI View Article : Google Scholar
|
|
4
|
Jakub JW, Bryant K, Huebner M, Hoskin T,
Boughey JC, Reynolds C and Degnim AC: The number of axillary lymph
nodes involved with metastatic breast cancer does not affect
outcome as long as all disease is confined to the sentinel lymph
nodes. Ann Surg Oncol. 18:86–93. 2011.PubMed/NCBI View Article : Google Scholar
|
|
5
|
Galimberti V, Cole BF, Zurrida S, Viale G,
Luini A, Veronesi P, Baratella P, Chifu C, Sargenti M, Intra M, et
al: International Breast Cancer Study Group Trial 23-01
investigators: Axillary dissection versus no axillary dissection in
patients with sentinel-node micrometastases (IBCSG 23-01): A phase
3 randomised controlled trial. Lancet Oncol. 14:297–305.
2013.PubMed/NCBI View Article : Google Scholar
|
|
6
|
Reed J, Rosman M, Verbanac KM, Mannie A,
Cheng Z and Tafra L: Prognostic implications of isolated tumor
cells and micrometastases in sentinel nodes of patients with
invasive breast cancer: 10-year analysis of patients enrolled in
the prospective East Carolina University/Anne Arundel Medical
Center Sentinel Node Multicenter Study. J Am Coll Surg.
208:333–340. 2009.PubMed/NCBI View Article : Google Scholar
|
|
7
|
Carlo JT, Grant MD, Knox SM, Jones RC,
Hamilton CS, Livingston SA and Kuhn JA: Survival analysis following
sentinel lymph node biopsy: A validation trial demonstrating its
accuracy in staging early breast cancer. Proc Bayl Univ Med Cent.
18:103–107. 2005.PubMed/NCBI View Article : Google Scholar
|
|
8
|
Chu PG and Weiss LM: Keratin expression in
human tissues and neoplasms. Histopathology. 40:403–439.
2002.PubMed/NCBI View Article : Google Scholar
|
|
9
|
Tsujimoto M, Nakabayashi K, Yoshidome K,
Kaneko T, Iwase T, Akiyama F, Kato Y, Tsuda H, Ueda S, Sato K, et
al: One-step nucleic acid amplification for intraoperative
detection of lymph node metastasis in breast cancer patients. Clin
Cancer Res. 13:4807–4816. 2007.PubMed/NCBI View Article : Google Scholar
|
|
10
|
Li D, Xu X, Chen J, Chen J, Yang B, Yang
W, Xu W, Wu J and Shi D: Utility of one-step nucleic acid
amplification (OSNA) assay in detecting breast cancer metastases of
sentinel lymph nodes in a Chinese population. Breast Cancer.
22:135–140. 2015.PubMed/NCBI View Article : Google Scholar
|
|
11
|
Leidenius MHK, Krogerus LA, Toivonen TS
and Von Smitten KJ: The feasibility of intraoperative diagnosis of
sentinel lymph node metastases in breast cancer. J Surg Oncol.
84:68–73. 2003.PubMed/NCBI View Article : Google Scholar
|
|
12
|
Raia-Barjat T, Trombert B, Khaddage A,
Douchet C, Seffert P, Peoc'h M, Falk AT, Magné N and Chauleur C:
OSNA (one-step nucleic acid amplification) sentinel lymph node
intraoperative molecular analysis in breast cancer: A cost-benefit
analysis. Med Oncol. 31(322)2014.PubMed/NCBI View Article : Google Scholar
|
|
13
|
Castellano I, Macrì L, Deambrogio C,
Balmativola D, Bussone R, Ala A, Coluccia C and Sapino A:
Reliability of whole sentinel lymph node analysis by one-step
nucleic acid amplification for intraoperative diagnosis of breast
cancer metastases. Ann Surg. 255:334–342. 2012.PubMed/NCBI View Article : Google Scholar
|
|
14
|
Saruta Y and Puig-Junoy J: Cost and budget
impact analysis of an accurate intraoperative sentinel lymph node
diagnosis for breast cancer metastasis. Appl Health Econ Health
Policy. 14:323–335. 2016.PubMed/NCBI View Article : Google Scholar
|
|
15
|
Klingler S, Marchal F, Rauch P, Kenouchi
O, Chrétien AS, Genin P, Leroux A and Merlin JL: Using one-step
nucleic acid amplification (OSNA) for intraoperative detection of
lymph node metastasis in breast cancer patients avoids second
surgery and accelerates initiation of adjuvant therapy. Ann Oncol.
24:2305–2309. 2013.PubMed/NCBI View Article : Google Scholar
|
|
16
|
Giuliano AE, Ballman KV, McCall L, Beitsch
PD, Brennan MB, Kelemen PR, Ollila DW, Hansen NM, Whitworth PW,
Blumencranz PW, et al: Effect of axillary dissection vs no axillary
dissection on 10-year overall survival among women with invasive
breast cancer and sentinel node metastasis: The ACOSOG Z0011
(Alliance) Randomized Clinical Trial. JAMA. 318:918–926.
2017.PubMed/NCBI View Article : Google Scholar
|
|
17
|
Donker M, van Tienhoven G, Straver ME,
Meijnen P, van de Velde CJ, Mansel RE, Cataliotti L, Westenberg AH,
Klinkenbijl JH, Orzalesi L, et al: Radiotherapy or surgery of the
axilla after a positive sentinel node in breast cancer (EORTC
10981-22023 AMAROS): A randomised, multicentre, open-label, phase 3
non-inferiority trial. Lancet Oncol. 15:1303–1310. 2014.PubMed/NCBI View Article : Google Scholar
|
|
18
|
Sobin LH and Wittekind CH (eds): UICC: TNM
Classification of Malignant Tumors. 6th edition. Wiley-Liss, New
York, NY, 2002.
|
|
19
|
Sauerbrei W and Schumacher M: A bootstrap
resampling procedure for model building: Application to the Cox
regression model. Stat Med. 11:2093–2109. 1992.PubMed/NCBI View Article : Google Scholar
|
|
20
|
Terrenato I, D'Alicandro V, Casini B,
Perracchio L, Rollo F, De Salvo L, Di Filippo S, Di Filippo F,
Pescarmona E, Maugeri-Saccà M, et al: A cut-off of 2150 cytokeratin
19 mRNA copy number in sentinel lymph node may be a powerful
predictor of non-sentinel lymph node status in breast cancer
patients. PLoS One. 12(e0171517)2017.PubMed/NCBI View Article : Google Scholar
|
|
21
|
Tamaki Y: One-step nucleic acid
amplification (OSNA): Where do we go with it? Int J Clin Oncol.
22:3–10. 2017.PubMed/NCBI View Article : Google Scholar
|
|
22
|
Snook KL, Layer GT, Jackson PA, de Vries
CS, Shousha S, Sinnett HD, Nigar E, Singhal H, Chia Y, Cunnick G,
et al: OSNA Study Group: Multicentre evaluation of intraoperative
molecular analysis of sentinel lymph nodes in breast carcinoma. Br
J Surg. 98:527–535. 2011.PubMed/NCBI View
Article : Google Scholar
|
|
23
|
Cserni G: Intraoperative analysis of
sentinel lymph nodes in breast cancer by one-step nucleic acid
amplification. J Clin Pathol. 65:193–199. 2012.PubMed/NCBI View Article : Google Scholar
|
|
24
|
Shi F, Liang Z, Zhang Q, Wang C and Liu X:
The performance of one-step nucleic acid amplification assay for
intraoperative detection of sentinel lymph node macrometastasis in
breast cancer: An updated meta-analysis. Breast. 39:39–45.
2018.PubMed/NCBI View Article : Google Scholar
|
|
25
|
Chaudhry A, Williams S, Cook J, Jenkins M,
Sohail M, Calder C, Winters ZE and Rayter Z: The real-time
intra-operative evaluation of sentinel lymph nodes in breast cancer
patients using One Step Nucleic Acid Amplification (OSNA) and
implications for clinical decision-making. Eur J Surg Oncol.
40:150–157. 2014.PubMed/NCBI View Article : Google Scholar
|
|
26
|
Mansel RE, Fallowfield L, Kissin M, Goyal
A, Newcombe RG, Dixon JM, Yiangou C, Horgan K, Bundred N, Monypenny
I, et al: Randomized multicenter trial of sentinel node biopsy
versus standard axillary treatment in operable breast cancer: The
ALMANAC Trial. J Natl Cancer Inst. 98:599–609. 2006.PubMed/NCBI View Article : Google Scholar
|
|
27
|
Tjan-Heijnen VC, Pepels MJ, de Boer M,
Borm GF, van Dijck JA, van Deurzen CH, Adang EM, Menke-Pluymers MB,
van Diest PJ and Bult P: Impact of omission of completion axillary
lymph node dissection (cALND) or axillary radiotherapy (ax RT) in
breast cancer patients with micrometastases (pN1mi) or isolated
tumor cells (pN0(i+)) in the sentinel lymph node (SN): Results from
the MIRROR study. J Clin Oncol. 27 (Suppl 15):CRA506. 2009.
|
|
28
|
Pepels MJ, de Boer M, Bult P, van Dijck
JA, van Deurzen CH, Menke-Pluymers MB, van Diest PJ, Borm GF and
Tjan-Heijnen VC: Regional recurrence in breast cancer patients with
sentinel node micrometastases and isolated tumor cells. Ann Surg.
255:116–121. 2012.PubMed/NCBI View Article : Google Scholar
|
|
29
|
Huang T-W, Kuo KN, Chen K-H, Chen C, Hou
WH, Lee WH, Chao TY, Tsai JT, Su CM, Huang MT, et al:
Recommendation for axillary lymph node dissection in women with
early breast cancer and sentinel node metastasis: A systematic
review and meta-analysis of randomized controlled trials using the
GRADE system. Int J Surg. 34:73–80. 2016.PubMed/NCBI View Article : Google Scholar
|
|
30
|
Houvenaeghel G, Cohen M, Raro P, De Troyer
J, de Lara CT, Gimbergues P, Gauthier T, Faure-Virelizier C,
Vaini-Cowen V, Lantheaume S, et al: Others investigators (SERC
trial group): Overview of the pathological results and treatment
characteristics in the first 1000 patients randomized in the SERC
trial: Axillary dissection versus no axillary dissection in
patients with involved sentinel node. BMC Cancer.
18(1153)2018.PubMed/NCBI View Article : Google Scholar
|
|
31
|
Dauplat MM, E. Barranger E, Classe JM,
Toledano A and Houvenaeghel G: L'exploration et le traitement de la
région axillaire des tumeurs infiltrantes du sein. Oncologie.
15:589–592. 2013.
|
|
32
|
Veronesi U, Paganelli G, Galimberti V,
Viale G, Zurrida S, Bedoni M, Costa A, de Cicco C, Geraghty JG,
Luini A, et al: Sentinel-node biopsy to avoid axillary dissection
in breast cancer with clinically negative lymph-nodes. Lancet.
349:1864–1867. 1997.PubMed/NCBI View Article : Google Scholar
|
|
33
|
Horimoto Y, Tanabe M, Kazuno S, Miura Y,
Mogushi K, Sonoue H, Arakawa A, Kajino K, Kobayashi T and Saito M:
Elucidation of inhibitory effects on metastatic sentinel lymph
nodes of breast cancer during One-Step Nucleic Acid Amplification.
Sci Rep. 8(7563)2018.PubMed/NCBI View Article : Google Scholar
|
|
34
|
Shimazu K, Sato N, Ogiya A, Sota Y,
Yotsumoto D, Ishikawa T, Nakamura S, Kinoshita T, Tsuda H, Ohi Y,
et al: Intraoperative nomograms, based on one-step nucleic acid
amplification, for prediction of non-sentinel node metastasis and
four or more axillary node metastases in breast cancer patients
with sentinel node metastasis. Ann Surg Oncol. 25:2603–2611.
2018.PubMed/NCBI View Article : Google Scholar
|
|
35
|
Bizzarri N, Pedone Anchora L, Zannoni GF,
Santoro A, Valente M, Inzani F, Gallotta V, Conte C, Chiantera V,
Fanfani F, et al: Role of one-step nucleic acid amplification
(OSNA) to detect sentinel lymph node low-volume metastasis in
early-stage cervical cancer. Int J Gynecol Cancer. 30:364–371.
2020.PubMed/NCBI View Article : Google Scholar
|
|
36
|
Shimazu K, Miyake T, Okuno J, Naoi Y,
Tanei T, Shimoda M, Kagara N, Kim SJ and Noguchi S: One-step
Nucleic Acid Amplification can identify sentinel node-negative
breast cancer patients with excellent prognosis. Anticancer Res.
39:1447–1454. 2019.PubMed/NCBI View Article : Google Scholar
|
|
37
|
Peg V, Espinosa-Bravo M, Vieites B,
Vilardell F, Antúnez JR, de Salas MS, Delgado-Sánchez JJ, Pinto W,
Gozalbo F, Petit A, et al: Intraoperative molecular analysis of
total tumor load in sentinel lymph node: A new predictor of
axillary status in early breast cancer patients. Breast Cancer Res
Treat. 139:87–93. 2013.PubMed/NCBI View Article : Google Scholar
|
|
38
|
Deambrogio C, Castellano I, Paganotti A,
Zorini EO, Corsi F, Bussone R, Franchini R, Antona J, Miglio U,
Sapino A, et al: A new clinical cut-off of cytokeratin 19 mRNA copy
number in sentinel lymph node better identifies patients eligible
for axillary lymph node dissection in breast cancer. J Clin Pathol.
67:702–706. 2014.PubMed/NCBI View Article : Google Scholar
|
|
39
|
Heilmann T, Mathiak M, Hofmann J,
Mundhenke C, van Mackelenbergh M, Alkatout I, Wenners A,
Eckmann-Scholz C and Schem C: Intra-operative use of one-step
nucleic acid amplification (OSNA) for detection of the tumor load
of sentinel lymph nodes in breast cancer patients. J Cancer Res
Clin Oncol. 139:1649–1655. 2013.PubMed/NCBI View Article : Google Scholar
|
|
40
|
Nabais C, Figueiredo J, Lopes P, Martins M
and Araújo A: Total tumor load assessed by one-step nucleic acid
amplification assay as an intraoperative predictor for non-sentinel
lymph node metastasis in breast cancer. Breast. 32:33–36.
2017.PubMed/NCBI View Article : Google Scholar
|
|
41
|
Banerjee SM, Michalopoulos NV, Williams
NR, Davidson T, El Sheikh S, McDermott N, Tran-Dang MA, Davison S
and Keshtgar MR: Detailed evaluation of one step nucleic acid
(OSNA) molecular assay for intra-operative diagnosis of sentinel
lymph node metastasis and prediction of non-sentinel nodal
involvement: Experience from a London teaching hospital. Breast.
23:378–384. 2014.PubMed/NCBI View Article : Google Scholar
|
|
42
|
Espinosa-Bravo M, Sansano I, Pérez-Hoyos
S, Ramos M, Sancho M, Xercavins J, Rubio IT and Peg V: Prediction
of non-sentinel lymph node metastasis in early breast cancer by
assessing total tumoral load in the sentinel lymph node by
molecular assay. Eur J Surg Oncol. 39:766–773. 2013.PubMed/NCBI View Article : Google Scholar
|
|
43
|
Buglioni S, Di Filippo F, Terrenato I,
Casini B, Gallo E, Marandino F, Maini CL, Pasqualoni R, Botti C, Di
Filippo S, et al: Quantitative molecular analysis of sentinel lymph
node may be predictive of axillary node status in breast cancer
classified by molecular subtypes. PLoS One.
8(e58823)2013.PubMed/NCBI View Article : Google Scholar
|