Introduction
Mucoepidermoid carcinoma (MEC), representing 5% of
all salivary gland tumors and 26% of malignant salivary gland
tumors registered for the last 39 years in Hiroshima, Japan, is the
most common malignant tumor of the major and minor salivary glands
(1,2). MEC is characterized by its cellular
heterogeneity and consists of mucin-producing, epidermoid and
intermediate cells. Clinical and pathological parameters (age,
tumor size, presence of cervical lymphadenopathy, distant spread,
perineural invasion and histological grade) of MEC have been
associated with tumor biological behavior and patient management
(3). Pathological classification of
MEC is graded as low-, intermediate- or high-grade based on adverse
features, such as perineural invasion, angiolymphatic invasion,
coagulative necrosis, infiltrative growth, high mitotic rate,
anaplasia and cystic components of <20% (4).
An important genetic abnormality in MEC is the
translocation between chromosomes 11q and 19p, which has been
hypothesized to be an early event in the pathogenesis of MEC
(5,6), and has been reported in >50% of MEC
tumors (7). Low-grade tumors have a
higher incidence rate of this fusion compared with that in
high-grade tumors (8) and patients
with fusion-positive cancer tend to have improved survival time,
with significantly lower risks of recurrence, metastases or
cancer-related mortality (9). The
majority of fusion genes in MEC are associated with a specific
chromosomal t(11;19) (q14-21;p12-13) translocation that joins exon
1 of the cAMP response element-binding (CREB) protein-binding
domain of CREB-regulated transcription coactivator 1 (CRTC1)
gene to exons 2-5 of the Notch coactivator mastermind-like gene 2
(MAML2) gene, resulting in the expression of a new
CRTC1-MAML2 fusion gene (10). This translocation generates a fusion
protein comprised of CRTC1 (also called MECT1,
TORC1 or WAMP1) at 19q21 and the C-terminal
transcriptional activation domain of MAML2 at 11q21
(11-14).
Previous analysis suggested that another member of the CRTC
family, at 15q26, CRTC3, also fused with MAML2
(15). Okabe et al (16) and Nakayama et al (17) showed that CRTC1-MAML2 or
CRTC3-MAML2 fusions occurred in 40-80% of primary salivary
gland MECs, and was associated with a distinct tumor subset that
had favorable clinicopathological features and an indolent clinical
course.
Previously, amphiregulin (AREG), a member of the
epidermal growth factor (EGF) family, was identified as a target of
the CRTC1-MAML2 fusion gene and secreted AREG was shown to
activate EGF receptor (EGFR) signaling in an autocrine manner
(18). Furthermore, mutations in
EGFR itself are rare in salivary gland carcinomas (19), while copy number alternations in
EGFR are frequently found in high-grade MEC, regardless of
fusion gene positivity (20). The
molecular pathology and oncology of MEC are still poorly
understood. Established authentic cell lines are essential to
determine the biological characteristics of MEC, and a number of
cell cultures and models have emerged; however, the cell line
usability is limited (21). The
present study reports the establishment of a MEC cell line
(HCM-MEC010) carrying the CRTC1-MAML2 fusion gene and
activated EGFR. The potential uses for this cell line will also be
discussed to understand the biological characteristics of MEC.
Materials and methods
Cell line generation and cell
culture
A patient with MEC provided consent in accordance
with Hyogo College of Medicine (Hyogo, Japan) institutional
policies. Tumor samples were obtained according to an approved
Institutional Review Board protocol of Hyogo College of Medicine
(approval no. 276; Hyogo, Japan). The present study was also
conducted in accordance with the Declaration of Helsinki. Clinical
and pathological data were collected from the medical records of
the patient. Tumor tissues were minced into 1-2-mm pieces with a
disposable scalpel and placed in primary culture. To separate the
stromal cells from the mass culture, a magnetic-activated cell
sorting (MACS) system was used. Briefly, MACS buffer, containing 1X
PBS, 0.5% BSA, 2 mM EDTA (pH 7.2) (cat. no. 130-042-901; Miltenyi
Biotec Inc.), was pre-cooled to 4˚C. To remove the fibroblasts, the
single cell suspension was centrifuged at 300 x g for 10 min at
room temperature. and positive selection was performed using CD326
(EpCAM) MicroBeads and a MidiMACSTM Separator (Miltenyi Biotic
GmbH), according to the manufacturer's instructions. The obtained
primary human MEC cells were seeded in F-medium (22) with 10 µM Y-27632 (FUJIFILM Wako Pure
Chemical Corporation). After 1 week, the culture medium was
replaced with fresh medium, which was changed every 4 days
thereafter. At the same time, the fibroblasts derived from the
tumor tissue of the same patient, were obtained and grown in
F-medium. Once cells reached confluence (80%), they were washed
with PBS (Mg2+ and Ca2+ free) (23) and detached with 0.05% EDTA/trypsin
for 5 min at 38˚C (24). After
centrifugation at 167 x g for 5 min at 4˚C, the MEC cells were
resuspended in F-medium, containing Y-27632 and seeded
(0.3x106 cells) in 60 mm dishes. An epithelial cell line
was successfully established from the sample of the patient and was
termed HCM-MEC010. The morphology of the exponentially
proliferating cells in a monolayer was reviewed and documented
using inverted phase contrast microscopy. The cells were also
tested for mycoplasma infection using the MycoAlert®
Assay (Lonza Group, Ltd.) and the cell culture growth medium and
with fluorescent microscopy using the Mycoplasma Hoechst Stain
Assay (MP Biomedicals, LLC).
Short tandem repeat (STR)
authentication of the MEC cell line
To verify the identity of the cell line, genomic DNA
was extracted from the blood of the patient, whose tumor sample was
used to generate the HCM-MEC010 cell line, as well as from the cell
line using the QIAamp DNA Mini kit (Qiagen, Inc.) according to the
manufacturer's protocol. DNA genotyping using STR profiling was
performed using the GenePrint 10 System (Promega Corporation) and
the Applied Biosystems 3130xl Analyzer (Applied Biosystems; Thermo
Fisher Scientific, Inc.) and analyzed by BEX Co., Ltd. The
evaluation value (EV) was determined using the following equation:
EV=(number of coincidental peaks) x 2/total number of peaks in cell
A and total number of peaks in cell B.
Reverse transcription (RT)-PCR of the
CRTC1-MAML2 fusion oncogene
The HCM-MEC010 cell line was plated in 100-mm dishes
and cultured to 90% confluence. RNA was extracted using
TRIzol® (Invitrogen; Thermo Fisher Scientific, Inc.) and
RT-PCR was performed using the PrimeScript RT-PCR kit (Takara Bio,
Inc.) according to the manufacturer's instructions. The following
primers were used: CRTC1 forward 1, 5'-TTCGAGGAGGTCATGAAGGA-3' and
2, 5'-ATGGCGACTTCGAACAATCCGCGGAA-3'; MAML2 reverse 1,
5'-TTGCTGTTGGCAGGAGATAG-3' and 2, 5'-GGGTCGCTTGCTGTTGGCAGGAG-3'
(18), which amplified 101 and 194
bp fragments, respectively. Amplification of the GAPDH gene
(forward, 5'-CAATGACCCCTTCATTGACC-3' and reverse,
5'-GACAAGCTTCCCGTTCTCAG-3') was performed as a control.
Successfully amplified RT-PCR products of the CRTC1-MAML2
fusion gene were purified and sequenced (24) using BigDye™ Terminator
v3.1 Cycle Sequencing kit (Thermo Fisher Scientific, Inc.) and 2%
agarose gel electrophoresis.
Western blot analysis
The culture medium was removed and the cells were
washed with PBS (Mg2+ and Ca2+ free). RIPA
buffer was added (cat. no. sc-24948; Santa Cruz, Inc.) and the
cells were incubated at 4˚C for 60 min, then centrifuged at 12,000
x g for 20 min 4˚C. The supernatant was the total cell lysate.
Proteins were extracted from the HCM-MEC010 and human tongue
squamous cell carcinoma (SAS; purchased from the Japanese
Collection of Research Bioresources Cell Bank) cell lines as
previously described (25). Protein
concentration was measured using a Bradford assay (26) Western blot analysis was performed as
previously described (25). The
primary and secondary antibodies are listed in Table I. The protein expression ratio,
compared with that in SAS cells, was measured using ImageJ v1.53e
software (National Institutes of Health). The data are presented as
the mean ± SD. The experiment was repeated three times.
 | Table IPrimary and secondary antibodies used
for western blot analysis and immunofluorescence. |
Table I
Primary and secondary antibodies used
for western blot analysis and immunofluorescence.
| A, Primary
antibodies |
|---|
| | Dilution | |
|---|
| Name | Cat. no. | Western blot |
Immunofluorescence | Supplier |
|---|
| Rabbit monoclonal
anti-EGFR | 4267 | 1/1000 | 1/60 | CST |
| Rabbit monoclonal
anti-p-EGFR | 3777 | 1/1000 | | CST |
| Rabbit monoclonal
anti-AKT | 4691 | 1/1000 | | CST |
| Rabbit monoclonal
anti-p-AKT | 4060 | 1/1000 | | CST |
| Rabbit monoclonal
anti-AREG | 16036-1-AP | 1/1000 | | ProteinTech Group,
Inc. |
| Rabbit monoclonal
anti-E-cadherin | 3195 | 1/1000 | 1/100 | CST |
| Rabbit monoclonal
anti-N-cadherin | 13116 | 1/1000 | 1/100 | CST |
| Rabbit monoclonal
anti-vimentin | 5741 | 1/1000 | 1/100 | CST |
| Rabbit monoclonal
anti-tubulin | 2148 | 1/1000 | | CST |
| Mouse monoclonal
anti-actin | 47778 | 1/1000 | | |
| B, Secondary
antibodies |
| | Dilution | |
| Name | Cat. no. | Western blot |
Immunofluorescence | Supplier |
| Alexa Flur 488 goat
anti-rabbit IgG (H+L) | A-11008 | | 1/400 | Molecular Probes;
Thermo Fisher Scientific, Inc. |
| Anti-mouse IgG,
HRP-linked | 7076 | 1/1000 | | CST |
| Anti-IgG (H+L
chain) rabbit pAb-HRP | 458 | 1/10000 | | Molecular and
Biological Laboratories Co., Ltd. |
| Goat anti-mouse
HRP | ab97023 | 1/1000 | | Abcam |
Immunofluorescence staining
The cultured HCM-MEC010 and SAS cell lines were
fixed in 3.7% formaldehyde for 20 min at room temperature. After
permeabilization with 0.2% Triton-X/PBS for 5 min at room
temperature, the cells were blocked with 2% (w/v) BSA (Nacalai
Tesque, Inc.)/PBS, then washed with PBS (Mg2+ and
Ca2+ free) and incubated with the primary antibodies
overnight at 4˚C. The cells were washed with PBS (Mg2+
and Ca2+ free), then incubated with the secondary
antibody and Rhodamine phalloidin (Cytoskeleton, Inc.) for 2 h at
room temperature. The samples were mounted in Vecta shield
containing DAPI (Vector Laboratories). Fluorescent images were
captured using a confocal laser-scanning microscope (LSM780; Zeiss
AG). The primary and secondary antibodies are listed in Table I.
RNA analysis
RNA-Sequencing (RNA-Seq) libraries were generated
using RNA extracted from the HCM-MEC010 cell line, as previously
described (27), with the TruSeq
Stranded mRNA Library Prep kit for Illumina, Inc., following the
manufacturer's instructions, then sequenced on a NovaSeq 6000
System (Illumina, Inc.). The analysis was performed by Takara Bio,
Inc.
Hematoxylin and eosin-staining
A section of the hard palate was fixed in 10%
formalin solution at room temperature for 24 h and embedded in
paraffin. Sections (5-µm thick) were cut from the paraffin blocks
and stained with hematoxylin (0.09%) for 5 min and eosin (0.13%)
for 9 min at room temperature according to standard methods
(28). The images were captured
using a light microscope (BX51; Olympus Corporation).
Patient
A 45-year-old Japanese female noticed spontaneous
dull pain and swelling in her hard palate for 1 month and was
referred to Hyogo College of Medicine, Nishinomiya, Hyogo, Japan on
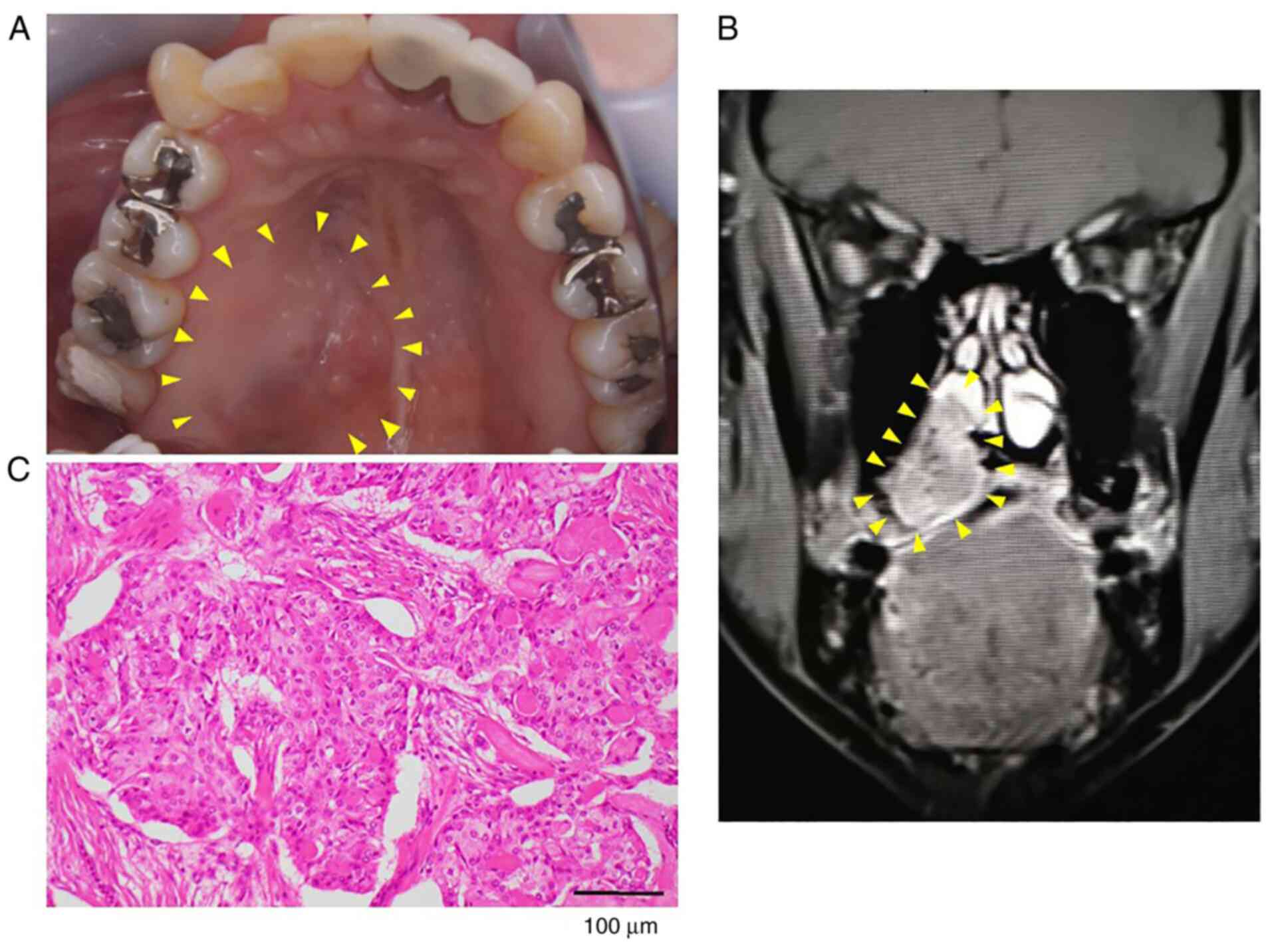
January, 2019. On examination, diffuse swelling was observed in the
right hard palate. There was no trismus. The surface of the mass
was smooth and was soft on palpation (Fig. 1A). Bilateral cervical lymph nodes
were palpable, but painless and mobile. Magnetic resonance imaging
showed an irregular mass measuring 30x20x18 mm in the right hard
palate, and resorption in the nasal septum and posterior wall of
the maxillary sinus (Fig. 1B). The
clinical diagnosis was a malignant tumor of the hard palate. A
biopsy was performed intraorally and the lesion was pathologically
diagnosed as low-grade MEC using Armed Forces Institute of
Pathology (29).
Results
The patient was admitted to Hyogo College of
Medicine, Nishinomiya, Hyogo, Japan and treated by partial
resection of the hard palate, supraomohyoid neck dissection and
reconstruction using an anterolateral thigh flap under general
anesthesia. Hematoxylin and eosin-stained tumor tissue
microscopically showed an overlying stratified squamous epithelium,
mucous cells and squamous cells that were polygonal-to-ovoid in
shape with eosinophilic cytoplasms (Fig. 1C). The mucous cells were cuboidal or
goblet-like and tended to line the cysts. The squamous cells formed
solid sheets. The tumor was diagnosed as mucoepidermoid carcinoma,
low-grade type, pT4aN0M0 MEC of the hard palate. All dissected
cervical lymph nodes showed no metastatic cells. At the 30-month
follow up, the patient's prognosis was excellent and she had
maintained a disease-free status.
Establishment of a MEC cell line from
a patient tumor
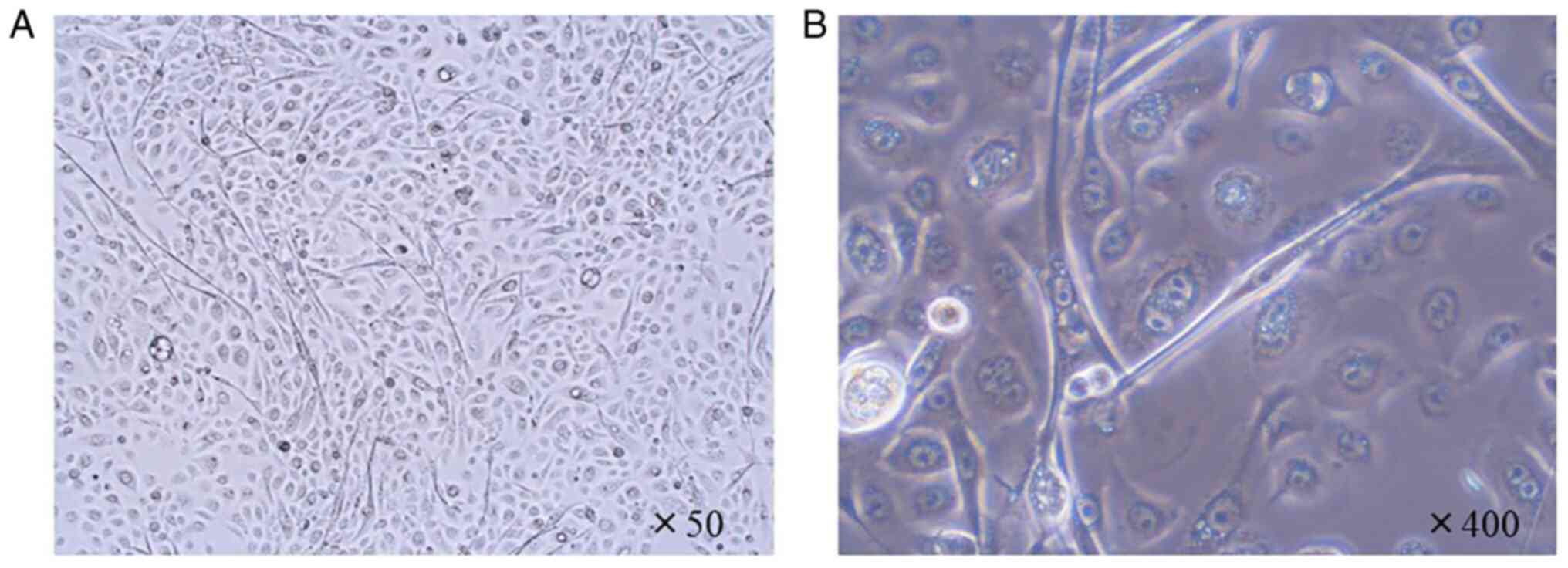
A new MEC cell line, termed HCM-MEC010 was
established, which maintained a cobblestone epithelial-like
morphology for at least 30 passages (Fig. 2A and B). To confirm that the HCM-MEC010 cell
line was derived from the tumor sample of the patient, STR
profiling was performed using the DNA extracted from the
high-passage HCM-MEC010 cell line and the blood from the patient.
Genotypic analysis confirmed that the cell line was derived from
the tumor and no contamination with other cell types was detected
(EV, 1.0). (Table SI; Figs. S1 and S2).
RT-PCR analysis reveals that
HCM-MEC010 cells express the CRTC1-MAML2 fusion gene
As the CRTC1-MAML2 gene fusion is common in
MEC (9), the fusion event was
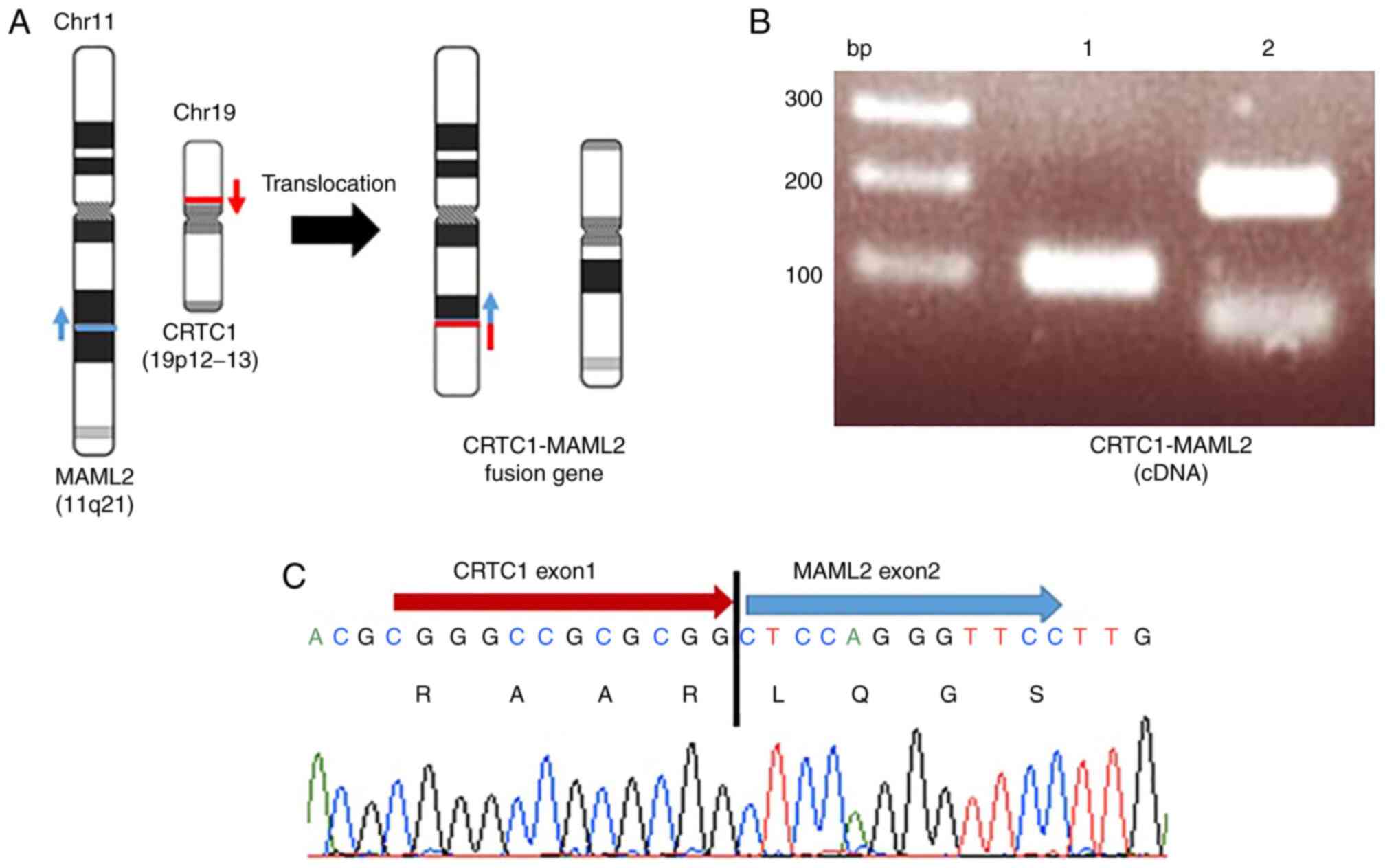
analyzed in the HCM-MEC010 cell line using RT-PCR. Fig. 3A shows the translocation event
between chromosomes 11 and 19, while Fig. 3B shows the RT-PCR amplified
fragments (lane 1, 101 bp and lane 2, 196 bp) using primer sets 1
or 2, respectively. The fusion transcript of CRTC1 and
MAML2 genes was confirmed using Sanger sequencing (Fig. 3C). This revealed the fusion products
of CRTC1 exon 1 and MAML2 exon 2 with the predicted
splicing event, indicating that a translocation event had occurred
between the first introns of CRTC1 and MAML2.
Protein expression in the HCM-MEC010
cell line
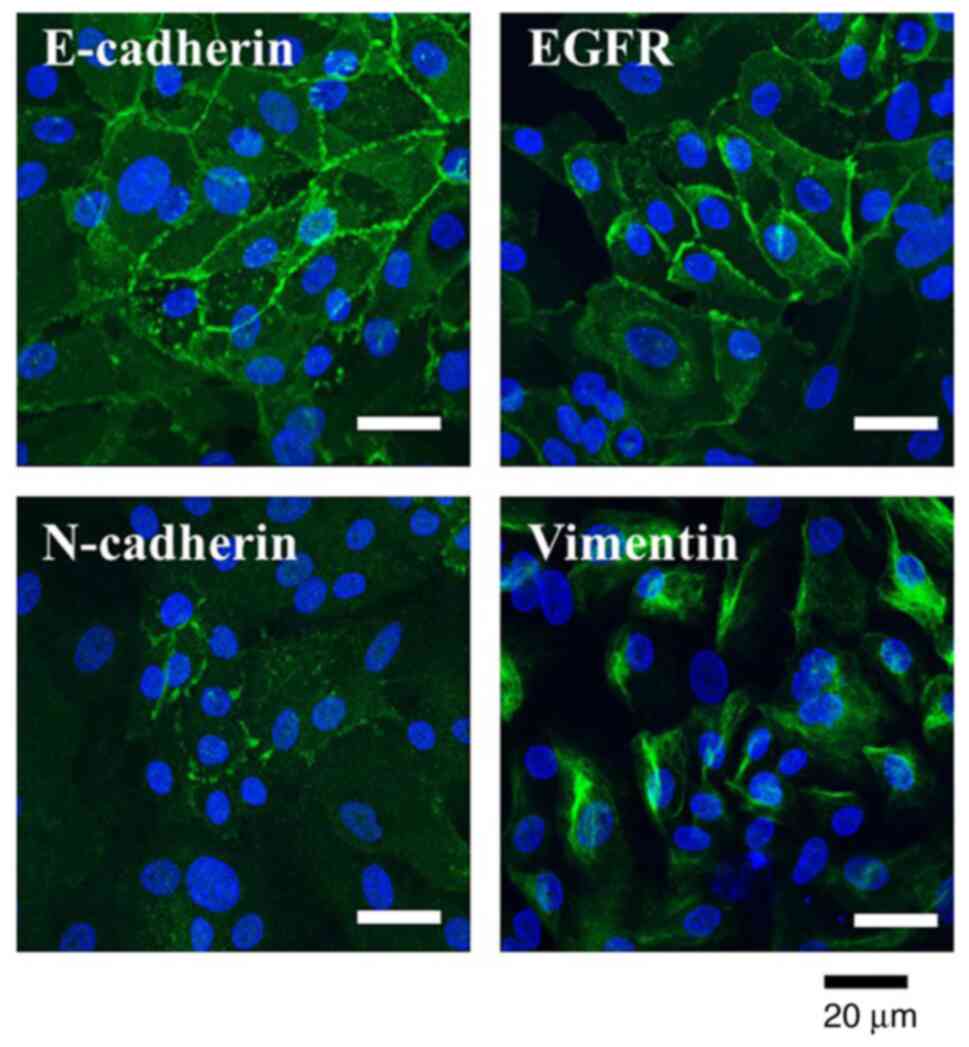
Next, the protein expression of the epithelial and
mesenchymal markers in the HCM-MEC010 cell line was confirmed using
immunofluorescent staining. EGFR and E-cadherin were expressed on
the cell membrane in the HCM-MEC010 cells, while N-cadherin
expression was only faintly detected. Vimentin expression was also
detected in HCM-MEC010 cells (Fig.
4).
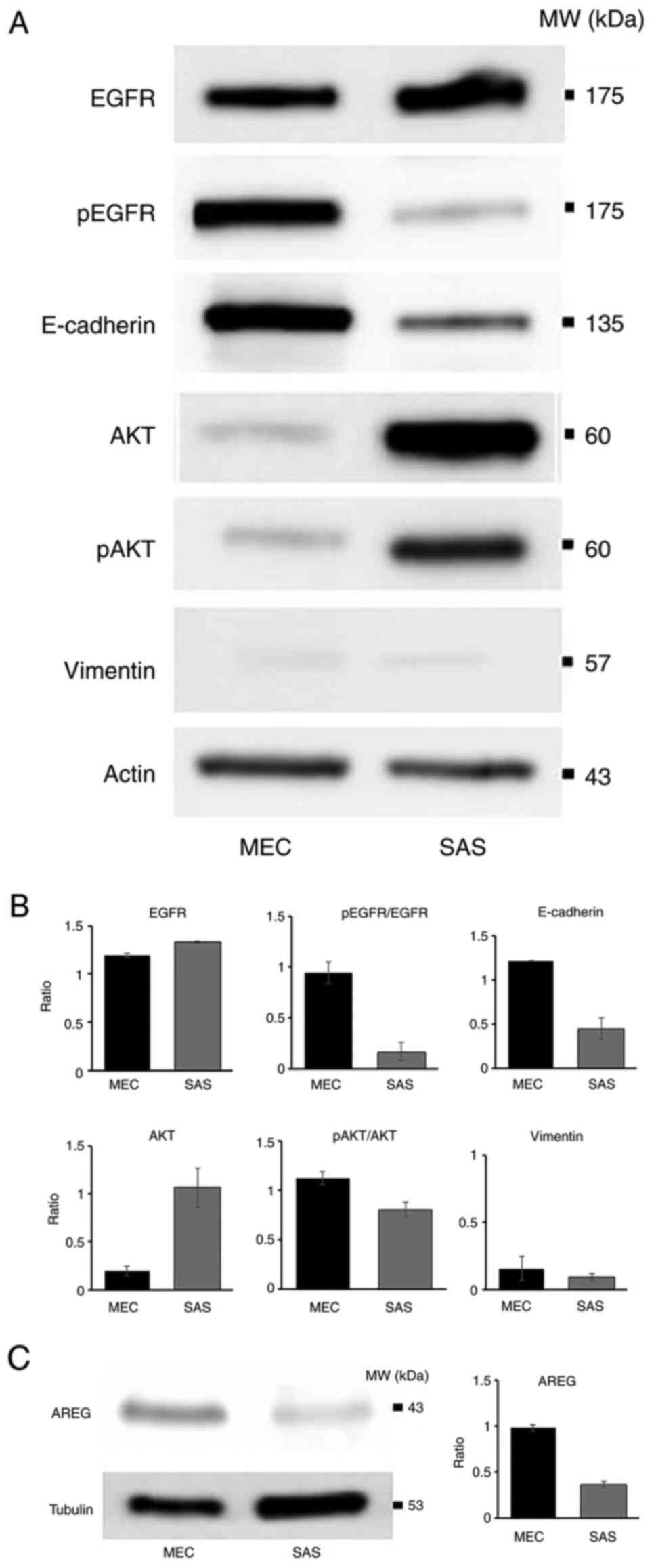
HCM-MEC010 cells express AREG and show
EGFR activation
As the AREG-EGFR signaling cascade has been
identified as a CRTC1-MAML2 fusion gene target (18), AREG expression and the status of the
EGFR cascade was analyzed in the HCM-MEC010 cell line. The human
tongue SAS cell line was used as a comparison as the SAS cell line
contains a mutation in the HER4 gene, which encodes one of
the other types of human EGFR, and the authentic EGFR
pathway is not involved in cell proliferation (30). EGFR was expressed in both cell
types, but the AREG expression level was much higher in the
HCM-MEC010 cell line compared with that in the SAS cell line
(Fig. 5). Furthermore, EGFR was
phosphorylated (p) in the HCM-MEC010 cell line compared with that
in the SAS cell line, indicating the activation of the EGFR
pathway. In addition, the expression level of AKT and p-AKT was
lower in the HCM-MEC010 cell line compared with that in the SAS
cell line. In the SAS cell line, AKT can be phosphorylated by both
the AREG-EGFR and HER4 pathways (31,32),
and high levels of AKT phosphorylation in the SAS cell line must
represent an additive effect of HER4 pathway activation (33). E-cadherin was expressed at higher
levels in the HCM-MEC010 cell line compared with that in the SAS
cell line. Vimentin expression was detected in small amounts in
both the HCM-MEC010 and SAS cell lines (Fig. 5).
RNA-seq analysis of the HCM-MEC010
cell line revealed epidermoid characteristics
To further characterize the HCM-MEC010 cell line,
RNA-Seq analysis was performed. MEC is known to be composed of a
mixture of mucous, epidermoid, and intermediate cells (34). RNA-Seq analysis revealed the high
expression level of genes in the keratin family, including
KRT5, KRT14, KRT6A, KRT17, and
KRT7. Table II lists the
top 200 expressed genes. However, expression of the mucous cell
marker MUC was not detected. These results, together with
the cell morphology results, suggest that the HCM-MEC010 cell line
is considered to be of epidermoid, but not mucinous, origin.
 | Table IIRNA-Sequencing data for the MEC cell
line. |
Table II
RNA-Sequencing data for the MEC cell
line.
| Entrez gene ID | Gene symbol | Description | TPM |
|---|
| 6280 | S100A9 | S100 calcium
binding protein A9 | 25012.1582 |
| - | RNR2 | - | 19355.6582 |
| 6277 | S100A6 | S100 calcium
binding protein A6 | 15991.2832 |
| 1915 | EEF1A1 | Eukaryotic
translation elongation factor 1 α 1 | 13189.91406 |
| 9168 | TMSB10 | Thymosin β 10 | 11790.02637 |
| 3852 | KRT5 | Keratin 5 | 10268.93652 |
| 6590 | SLPI | Secretory leukocyte
peptidase inhibitor | 9873.083008 |
| 6222 | RPS18 | Ribosomal protein
S18 | 9865.65332 |
| 3861 | KRT14 | Keratin 14 | 9010.899414 |
| 301 | ANXA1 | Annexin A1 | 8724.607422 |
| 302 | ANXA2 | Annexin A2 | 8183.348633 |
| 6130 | RPL7A | Ribosomal protein
l7a | 7233.334961 |
| 3853 | KRT6A | Keratin 6A | 7038.019531 |
| 6205 | RPS11 | Ribosomal protein
S11 | 6782.036133 |
| 3872 | KRT17 | Keratin 17 | 6779 |
| 6282 | S100A11 | S100 calcium
binding protein A11 | 6758.137695 |
| 57402 | S100A14 | S100 calcium
binding protein A14 | 6577.536133 |
| 6136 | RPL12 | Ribosomal protein
L12 | 6571.3125 |
| 6202 | RPS8 | Ribosomal protein
S8 | 5322.165039 |
| 23521 | RPL13A | Ribosomal protein
l13a | 5300.688477 |
| 6175 | RPLP0 | Ribosomal protein
lateral stalk subunit P0 | 5200.693848 |
| 6144 | RPL21(2) | Ribosomal protein
L21 | 5167.824219 |
| 7114 | TMSB4X | Thymosin β 4
X-linked | 4858.939453 |
| 6201 | RPS7 | Ribosomal protein
S7 | 4807.380371 |
| 6281 | S100A10 | S100 calcium
binding protein A10 | 4705.583984 |
| 6206 | RPS12 | Ribosomal protein
S12 | 4461.196777 |
| 6230 | RPS25 | Ribosomal protein
S25 | 4362.669922 |
| 6122 | RPL3 | Ribosomal protein
L3 | 4190.952148 |
| 2597 | GAPDH |
Glyceraldehyde-3-phosphate
dehydrogenase | 4084.755371 |
| 4502 | MT2A | Metallothionein
2A | 4024.866699 |
| 3855 | KRT7 | Keratin 7 | 3954.171631 |
| 6194 | RPS6 | Ribosomal protein
S6 | 3893.395996 |
| 6152 | RPL24 | Ribosomal protein
L24 | 3876.068848 |
| 6142 | RPL18A | Ribosomal protein
l18a | 3788.035645 |
| 60 | ACTB | Actin β | 3734.336914 |
| 6156 | RPL30 | Ribosomal protein
L30 | 3719.687988 |
| 6279 | S100A8 | S100 calcium
binding protein A8 | 3664.906982 |
| 10399 | RACK1 | Receptor for
activated C kinase 1 | 3650.605225 |
| 100133941 | CD24 | CD24 molecule | 3593.078369 |
| 6191 | RPS4X | Ribosomal protein
S4 X-linked | 3478.193848 |
| - | RNR1 | - | 3421.098877 |
| 2950 | GSTP1 | Glutathione
S-transferase pi 1 | 3338.100586 |
| 6187 | RPS2 | Ribosomal protein
S2 | 3280.663086 |
| 6207 | RPS13 | Ribosomal protein
S13 | 3170.464111 |
| 11224 | RPL35 | Ribosomal protein
L35 | 3153.293701 |
| 1937 | EEF1G | Eukaryotic
translation elongation factor 1 γ | 3140.122559 |
| 6125 | RPL5 | Ribosomal protein
L5 | 3137.963623 |
| 6170 | RPL39 | Ribosomal protein
L39 | 3100.071045 |
| 4637 | MYL6 | Myosin light chain
6 | 3067.146484 |
| 3868 | KRT16 | Keratin 16 | 3044.359619 |
| 4736 | RPL10A | Ribosomal protein
l10a | 2961.195801 |
| 6141 | RPL18 | Ribosomal protein
L18 | 2929.649658 |
| 1476 | CSTB | Cystatin B | 2910.897217 |
| 6124 | RPL4 | Ribosomal protein
L4 | 2865.387207 |
| 4070 | TACSTD2 | Tumor associated
calcium signal transducer 2 | 2787.421387 |
| 6147 | RPL23A | Ribosomal protein
l23a | 2730.734131 |
| 71 | ACTG1 | Actin γ 1 | 2705.085693 |
| 220 | ALDH1A3 | Aldehyde
dehydrogenase 1 family member A3 | 2588.035156 |
| 6135 | RPL11 | Ribosomal protein
L11 | 2561.839844 |
| 3880 | KRT19 | Keratin 19 | 2536.187012 |
| 6132 | RPL8 | Ribosomal protein
L8 | 2522.498047 |
| 6181 | RPLP2 | Ribosomal protein
lateral stalk subunit P2 | 2490.632813 |
| 3866 | KRT15 | Keratin 15 | 2464.382324 |
| 6699 | SPRR1B | Small proline rich
protein 1B | 2444.126465 |
| 6159 | RPL29 | Ribosomal protein
L29 | 2439.016113 |
| 2512 | FTL | Ferritin light
chain | 2432.441895 |
| 6193 | RPS5 | Ribosomal protein
S5 | 2432.29126 |
| 6233 | RPS27A | Ribosomal protein
s27a | 2403.434326 |
| 6129 | RPL7 | Ribosomal protein
L7 | 2332.271973 |
| 6273 | S100A2 | S100 calcium
binding protein A2 | 2289.59375 |
| 6133 | RPL9 | Ribosomal protein
L9 | 2237.880371 |
| 1475 | CSTA | Cystatin A | 2159.565186 |
| 6128 | RPL6 | Ribosomal protein
L6 | 2119.131592 |
| 2495 | FTH1 | Ferritin heavy
chain 1 | 2094.474121 |
| 3921 | RPSA | Ribosomal protein
SA | 2085.400391 |
| 5266 | PI3 | Peptidase inhibitor
3 | 2079.049805 |
| 2171 | FABP5 | Fatty acid binding
protein 5 | 2073.613281 |
| 5052 | PRDX1 | Peroxiredoxin
1 | 2053.132568 |
| 3956 | LGALS1 | Galectin 1 | 2031.178833 |
| 6143 | RPL19 | Ribosomal protein
L19 | 2021.314087 |
| 25818 | KLK5 | Kallikrein related
peptidase 5 | 1822.719238 |
| 3939 | LDHA | Lactate
dehydrogenase A | 1803.661499 |
| 6176 | RPLP1 | Ribosomal protein
lateral stalk subunit P1 | 1802.172241 |
| 51458 | RHCG | Rh family C
glycoprotein | 1785.147339 |
| 6303 | SAT1 | Spermidine/spermine
N1-acetyltransferase 1 | 1763.299316 |
| 9982 | FGFBP1 | Fibroblast growth
factor binding protein 1 | 1742.557251 |
| 7178 | TPT1 | Tumor protein,
translationally-controlled 1 | 1741.52832 |
| 6227 | RPS21 | Ribosomal protein
S21 | 1726.189087 |
| 3934 | LCN2 | Lipocalin 2 | 1720.297241 |
| 3315 | HSPB1 | Heat shock protein
family B (small) member 1 | 1668.157104 |
| 1973 | EIF4A1 | Eukaryotic
translation initiation factor 4A1 | 1623.40625 |
| 1938 | EEF2 | Eukaryotic
translation elongation factor 2 | 1612.361694 |
| 5055 | SERPINB2 | Serpin family B
member 2 | 1610.25 |
| 2810 | SFN | Stratifin | 1591.703979 |
| 6703 | SPRR2D | Small proline rich
protein 2D | 1568.223389 |
| 26986 | PABPC1 | Poly(A) binding
protein cytoplasmic 1 | 1534.452637 |
| 6204 | RPS10 | Ribosomal protein
S10 | 1532.445679 |
| 10410 | IFITM3 | Interferon induced
transmembrane protein 3 | 1529.12146 |
| 6189 | RPS3A | Ribosomal protein
S3A | 1509.361816 |
| 6154 | RPL26 | Ribosomal protein
L26 | 1432.493286 |
| 3918 | LAMC2 | Laminin subunit γ
2 | 1429.380249 |
| 83442 | SH3BGRL3 | SH3 domain binding
glutamate rich protein like 3 | 1395.721313 |
| 6139 | RPL17 | Ribosomal protein
L17 | 1375.231934 |
| 1933 | EEF1B2 | Eukaryotic
translation elongation factor 1 β 2 | 1351.424194 |
| 10974 | ADIRF | Adipogenesis
regulatory factor | 1348.772461 |
| 6134 | RPL10 | Ribosomal protein
L10 | 1336.026611 |
| 5268 | SERPINB5 | Serpin family B
member 5 | 1335.237183 |
| 6700 | SPRR2A | Small proline rich
protein 2A | 1285.784912 |
| 10094 | ARPC3 | Actin related
protein 2/3 complex subunit 3 | 1270.268311 |
| 2152 | F3 | Coagulation factor
III, tissue factor | 1268.36792 |
| 2197 | FAU | FAU ubiquitin like
and ribosomal protein S30 fusion | 1255.56189 |
| 9124 | PDLIM1 | PDZ and LIM domain
1 | 1252.652954 |
| 64065 | PERP | P53 apoptosis
effector related to PMP22 | 1252.282227 |
| 4869 | NPM1 | Nucleophosmin
1 | 1247.643188 |
| 7295 | TXN | Thioredoxin | 1169.833984 |
| 3553 | IL1B | Interleukin 1
β | 1166.45752 |
| 5054 | SERPINE1 | Serpin family E
member 1 | 1154.025146 |
| 6171 | RPL41 | Ribosomal protein
L41 | 1152.395996 |
| 25824 | PRDX5 | Peroxiredoxin
5 | 1133.30603 |
| 6173 | RPL36A | Ribosomal protein
l36a | 1111.359619 |
| 5315 | PKM | Pyruvate kinase
M1/2 | 1092.81897 |
| 1072 | CFL1 | Cofilin 1 | 1085.361328 |
| 6289 | SAA2 | Serum amyloid
A2 | 1073.526978 |
| 4071 | TM4SF1 | Transmembrane 4 L
six family member 1 | 1063.068237 |
| 506 | ATP5F1B | ATP synthase F1
subunit β | 1047.457275 |
| 5834 | PYGB | Glycogen
phosphorylase B | 1047.218994 |
| 928 | CD9 | CD9 molecule | 1021.081299 |
| 10628 | TXNIP | Thioredoxin
interacting protein | 1021.076111 |
| 103910 | MYL12B | Myosin light chain
12B | 1012.033325 |
| 3854 | KRT6B | Keratin 6B | 1011.945374 |
| 3688 | ITGB1 | Integrin subunit β
1 | 1004.073792 |
| 3312 | HSPA8 | Heat shock protein
family A (Hsp70) member 8 | 1000.302063 |
| 6288 | SAA1 | Serum amyloid
A1 | 999.111145 |
| 1382 | CRABP2 | Cellular retinoic
acid binding protein 2 | 986.4415283 |
| 6224 | RPS20 | Ribosomal protein
S20 | 975.680481 |
| 10109 | ARPC2 | Actin related
protein 2/3 complex subunit 2 | 966.9124146 |
| 1992 | SERPINB1 | Serpin family B
member 1 | 952.090332 |
| 306 | ANXA3 | Annexin A3 | 951.5344238 |
| 4501 | MT1X | Metallothionein
1X | 939.5453491 |
| 5660 | PSAP | Prosaposin | 936.7683105 |
| 6286 | S100P | S100 calcium
binding protein P | 924.9679565 |
| 567 | B2M |
β-2-microglobulin | 919.1690674 |
| 3914 | LAMB3 | Laminin subunit β
3 | 918.9204102 |
| 1308 | COL17A1 | Collagen type XVII
α 1 chain | 916.5231323 |
| 824 | CAPN2 | Calpain 2 | 912.717041 |
| 2706 | GJB2 | Gap junction
protein β 2 | 904.8463745 |
| 3860 | KRT13 | Keratin 13 | 894.9153442 |
| 3646 | EIF3E | Eukaryotic
translation initiation factor 3 subunit E | 893.5683594 |
| 5479 | PPIB | Peptidylprolyl
isomerase B | 883.137207 |
| 7316 | UBC | Ubiquitin C | 875.6885986 |
| 3326 | HSP90AB1 | Heat shock protein
90 α family class B member 1 | 871.9744263 |
| 642587 | MIR205HG | MIR205 host
gene | 864.2874146 |
| 468 | ATF4 | Activating
transcription factor 4 | 850.9224243 |
| 140576 | S100A16 | S100 calcium
binding protein A16 | 849.9338989 |
| 6155 | RPL27 | Ribosomal protein
L27 | 841.65802 |
| 6228 | RPS23 | Ribosomal protein
S23 | 837.4863281 |
| 25984 | KRT23 | Keratin 23 | 837.0656738 |
| 54541 | DDIT4 | DNA damage
inducible transcript 4 | 831.8173218 |
| 112694756 | LOC112694756 | Uncharaterized
LOC112694756 | 831.1845093 |
| 9349 | RPL23 | Ribosomal protein
L23 | 826.6482544 |
| 7184 | HSP90B1 | Heat shock protein
90 β family member 1 | 826.4506836 |
| 1337 | COX6A1 | Cytochrome c
oxidase subunit 6A1 | 820.6051025 |
| 1974 | EIF4A2 | Eukaryotic
translation initiation factor 4A2 | 800.7364502 |
| 6188 | RPS3 | Ribosomal protein
S3 | 796.1228638 |
| 6157 | RPL27A | Ribosomal protein
l27a | 790.3303833 |
| 5757 | PTMA | Prothymosin α | 790.0863037 |
| 826 | CAPNS1 | Calpain small
subunit 1 | 783.6133423 |
| 5328 | PLAU | Plasminogen
activator, urokinase | 780.4100342 |
| 2023 | ENO1 | Enolase 1 | 778.8522949 |
| 1509 | CTSD | Cathepsin D | 771.4251709 |
| 10476 | ATP5PD | ATP synthase
peripheral stalk subunit d | 768.3088989 |
| 7534 | YWHAZ | Tyrosine
3-monooxygenase/tryptophan 5-monooxygenase activation protein
ζ | 767.7701416 |
| 292 | SLC25A5 | Solute carrier
family 25 member 5 | 758.4469604 |
| 5216 | PFN1 | Profilin 1 | 753.312439 |
| 1340 | COX6B1 | Cytochrome c
oxidase subunit 6B1 | 751.3442383 |
| 8407 | TAGLN2 | Transgelin 2 | 741.7597046 |
| 689 | BTF3 | Basic transcription
factor 3 | 738.1211548 |
| 374 | AREG | Amphiregulin | 735.1116333 |
| 10376 | TUBA1B | Tubulin α 1b | 732.8063965 |
| 6210 | RPS15A | Ribosomal protein
s15a | 728.9209595 |
| 3909 | LAMA3 | Laminin subunit α
3 | 723.6885986 |
| 7086 | TKT | Transketolase | 713.4926147 |
| 5650 | KLK7 | Kallikrein related
peptidase 7 | 708.7366333 |
| 4323 | MMP14 | Matrix
metallopeptidase 14 | 702.4146118 |
| 4312 | MMP1 | Matrix
metallopeptidase 1 | 700.8983154 |
| 6229 | RPS24 | Ribosomal protein
S24 | 700.0944824 |
| 10653 | SPINT2 | Serine peptidase
inhibitor, Kunitz type 2 | 695.8338623 |
| 4831 | NME2 | NME/NM23 nucleoside
diphosphate kinase 2 | 694.8643799 |
| 10971 | YWHAQ | Tyrosine
3-monooxygenase/tryptophan 5-monooxygenase activation protein
τ | 692.3873291 |
| 5478 | PPIA | Peptidylprolyl
isomerase A | 682.8765869 |
| 7980 | TFPI2 | Tissue factor
pathway inhibitor 2 | 679.0671997 |
| 6146 | RPL22 | Ribosomal protein
L22 | 678.4135132 |
| 3945 | LDHB | Lactate
dehydrogenase B | 671.2799683 |
| 351 | APP | Amyloid β precursor
protein | 665.9901733 |
| 1508 | CTSB | Cathepsin B | 665.0159302 |
| 10209 | EIF1 | Eukaryotic
translation initiation factor 1 | 664.9918213 |
| 8673 | VAMP8 | Vesicle associated
membrane protein 8 | 659.6922607 |
| 7416 | VDAC1 | Voltage dependent
anion channel 1 | 659.1289063 |
| 4946 | OAZ1 | Ornithine
decarboxylase antizyme 1 | 656.2600098 |
| 6168 | RPL37A | Ribosomal protein
l37a | 649.401123 |
Discussion
The isolation of primary tumor cells from patient
samples is the first step for several genetic, biochemical and
pharmacological experiments relevant to personalized cancer
treatment (35). However, such
studies are limited due to cell availability. The establishment of
a cancer cell line is a traditional, but still powerful and
informative method of studying human cancer. The present study
reports the establishment of a MEC cell line with a
CRTC1-MAML2 fusion gene.
Several studies have shown that the presence of the
CRTC1/3-MAML2 fusion gene confers an improved prognosis,
with improved disease-free survival and fewer distant metastasis in
MEC (36,37). There are rare exceptions to this
rule, including fusion-positive high-grade MEC with multiple
additional genetic variations, such as mutations in CDKN2A,
that have been associated with a poor prognosis (38).
The function of the CRTC1-MAML2 fusion gene
has been intensively studied. Its transformation ability was
identified using the RK3E cell line (39) and its importance for tumor state
maintenance has also been demonstrated. Initially, it was
hypothesized to cause tumor growth by the constitutive activation
of Notch signaling via the MAML2 gene portion. Furthermore,
the N terminus CRTC1 domain-mediated aberrant activation of
cAMP/CREB signaling has also been identified as a cause of tumor
formation (14,40). The interaction between AP-1 and MYC
oncoprotein with CRTC1–MAML2 fusion proteins has been reported
(41), suggesting that the
CRTC1-MAML2 fusion gene regulates several different
signaling pathways. AREG is a known cAMP/CREB-regulated
gene, whose expression positively correlates with that of
CRTC1-MAML2 in MEC (42). As
AREG-EGFR signaling was identified as one of the CRTC1-MAML2
fusion gene targets, EGFR signaling could represent the mechanism
of action by which the fusion gene promotes carcinogenesis.
These observations suggest an overall role for EGFR
in the pathogenesis of MEC and the EGFR pathway could be a possible
therapeutic target. As several drugs target this pathway, AREG–EGFR
signaling was analyzed in the HCM-MEC010 cell line in the present
study. The HCM-MEC010 cell line was found to express AREG and
phosphorylate EGFR. Immunofluorescence analysis localized EGFR
expression to the HCM-MEC010 cell membrane. These data suggest that
the EGFR ligand, AREG, activated EGFR in an autocrine manner;
therefore, antibodies that block AREG-EGFR binding or drugs that
interfere with EGFR activation could be used for CRTC1-MAML2
fusion-positive MEC treatment. However, further analysis is
required to identify suitable therapies.
MECs are composed of mucin-producing, epidermoid,
and intermediate cells; however, RNA-Seq analysis of the HCM-MEC010
cell line detected little expression of MUC genes in the
mucous cell marker family, indicating that mucin-producing cells
and intermediate cells may have been removed during culture. MECs
develop in excretory duct cells (43) and the mixture of three different
cell types in MECs predicts their common origin. Duct and acinar
cell differentiation are typically lineage-restricted; however,
after irradiation, both duct and acinar cells can differentiate
into different cell types (44). It
is conceivable that established epidermoid-like cells are competent
to differentiate into acinar cells, which is a predicted
characteristic of injured duct stem cells. Further analysis will
assist in the clarification into the origin of MECs. Cancer stem
cells have been hypothesized to be involved in tumor formation
(43). The results of the present
study potentially indicate these cells may be of the same
origin.
In conclusion, a MEC cell line, HCM-MEC010, with a
CRTC1-MAML2 gene fusion was established. This cell line
showed typical MEC characteristics, including AREG expression and
EGFR activation; therefore, it could be used to assist in the
identification of EGFR-targeted drugs for the treatment of
CRTC1-MAML2 fusion gene-harboring MEC.
Supplementary Material
Results of STR test in cells. STR,
short tandem repeat.
Result of STR test in blood. STR,
short tandem repeat.
Resultsfrom short tandem repeat
analysis.
Acknowledgements
The authors would like to thank Ms. Shinobu Osawa
(Department of Oral and Maxillofacial Surgery, Hyogo College of
Medicine, Nishinomiya, Japan) for preparation of the experiments
and Ms. Takako Nanba (Department of Oral and Maxillofacial Surgery,
Hyogo College of Medicine, Nishinomiya, Japan) for the management
of the grants. The authors would also like to thank Nikki March and
Sarah Williams for editing a draft version of the manuscript.
Funding
Funding: This study was supported by JSPS Grants-in-Aid for
Scientific Research (grant nos. 16H11737 and 19H 10277), a
Grant-in-Aid for Graduate Students, and a Hyogo College of Medicine
and Hyogo Health Foundation Cancer Research Award.
Availability of data and materials
The datasets generated and/or analyzed during the
current study are not publicly available due to a pending patent
application, but are available from the corresponding author on
reasonable request.
Authors' contributions
KN, SK, KaY, KT, HK and YN conceived and designed
the present study. KN, SK, KaY, YF, KyY and YN performed the
experiments. KN, SK, KoY and YN analyzed the data. KN, SK and YN
wrote, reviewed, and revised the manuscript. All authors read and
approved the final manuscript. KN and SK confirm the authenticity
of all the raw data.
Ethics approval and consent to
participate
The current study was approved by the Institutional
Review Board of Hyogo College of Medicine (Hyogo, Japan) and was
conducted in accordance with the Declaration of Helsinki. The
patient provided written informed consent to participate.
Patient consent for publication
The patient provided written informed consent for
the publication of their case study.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Sentani K, Ogawa I, Ozawa K, Sadakane A,
Utada M, Tsuya T, Kajihara H, Yonehara S, Takeshima Y and Yasui W:
Characteristics of 5015 salivary gland neoplasms registered in the
Hiroshima tumor tissue registry over a period of 39 years. J Clin
Med. 8(566)2019.PubMed/NCBI View Article : Google Scholar
|
|
2
|
Behboudi A, Enlund F, Winnes M, Andrén Y,
Nordkvist A, Leivo I, Flaberg E, Szekely L, Mäkitie A, Grenman R,
et al: Molecular classification of mucoepidermoid
carcinomas-prognostic significance of the MECT1-MAML2 fusion
oncogene. Genes Chromosomes Cancer. 45:470–481. 2006.PubMed/NCBI View Article : Google Scholar
|
|
3
|
Ettl T, Schwarz-Furlan S, Gosau M and
Reichert TE: Salivary gland carcinomas. Oral Maxillofac Surg.
16:267–283. 2012.PubMed/NCBI View Article : Google Scholar
|
|
4
|
Katabi N, Ghossein R, Ali S, Dogan S,
Klimstra D and Ganly I: Prognostic features in mucoepidermoid
carcinoma of major salivary glands with emphasis on tumour
histologic grading. Histopathology. 65:793–804. 2014.PubMed/NCBI View Article : Google Scholar
|
|
5
|
El-Naggar AK, Lovell M, Killary AM,
Clayman GL and Batsakis JG: A mucoepidermoid carcinoma of minor
salivary gland with t(11;19)(q21;p13.1) as the only karyotypic
abnormality. Cancer Genet Cytogenet. 87:29–33. 1996.PubMed/NCBI View Article : Google Scholar
|
|
6
|
Saade RE, Bell D, Garcia J, Roberts D and
Weber R: Role of CRTC1/MAML2 translocation in the prognosis and
clinical outcomes of mucoepidermoid carcinoma. JAMA Otolaryngol
Head Neck Surg. 142:234–240. 2016.PubMed/NCBI View Article : Google Scholar
|
|
7
|
Bell D and El-Naggar AK: Molecular
heterogeneity in mucoepidermoid carcinoma: Conceptual and practical
implications. Head Neck Pathol. 7:23–27. 2013.PubMed/NCBI View Article : Google Scholar
|
|
8
|
Jee KJ, Persson M, Heikinheimo K,
Passador-Santos F, Aro K, Knuutila S, Odell EW, Mäkitie A, Sundelin
K, Stenman G and Leivo I: Genomic profiles and CRTC1-MAML2 fusion
distinguish different subtypes of mucoepidermoid carcinoma. Mod
Pathol. 26:213–222. 2013.PubMed/NCBI View Article : Google Scholar
|
|
9
|
O'Neill ID: t(11;19) translocation and
CRTC1-MAML2 fusion oncogene in mucoepidermoid carcinoma. Oral
Oncol. 45:2–9. 2009.PubMed/NCBI View Article : Google Scholar
|
|
10
|
Seethala RR, Dacic S, Cieply K, Kelly LM
and Nikiforova MN: A reappraisal of the MECT1/MAML2 translocation
in salivary mucoepidermoid carcinomas. Am J Surg Pathol.
34:1106–1121. 2010.PubMed/NCBI View Article : Google Scholar
|
|
11
|
Tonon G, Modi S, Wu L, Kubo A, Coxon AB,
Komiya T, O'Neil K, Stover K, El-Naggar A, Griffin JD, et al:
t(11;19)(q21;p13) translocation in mucoepidermoid carcinoma creates
a novel fusion product that disrupts a Notch signaling pathway. Nat
Genet. 33:208–213. 2003.PubMed/NCBI View
Article : Google Scholar
|
|
12
|
Enlund F, Behboudi A, Andrén Y, Oberg C,
Lendahl U, Mark J and Stenman G: Altered Notch signaling resulting
from expression of a WAMTP1-MAML2 gene fusion in mucoepidermoid
carcinomas and benign Warthin's tumors. Exp Cell Res. 292:21–28.
2004.PubMed/NCBI View Article : Google Scholar
|
|
13
|
Wu L, Liu J, Gao P, Nakamura M, Cao Y,
Shen H and Griffin JD: Transforming activity of MECT1-MAML2 fusion
oncoprotein is mediated by constitutive CREB activation. EMBO J.
24:2391–2402. 2005.PubMed/NCBI View Article : Google Scholar
|
|
14
|
Coxon A, Rozenblum E, Park YS, Joshi N,
Tsurutani J, Dennis PA, Kisch IR and Kaye FJ: Mect1-Maml2 fusion
oncogene linked to the aberrant activation of cyclic AMP/CREB
regulated genes. Cancer Res. 65:7137–7144. 2005.PubMed/NCBI View Article : Google Scholar
|
|
15
|
Fehr A, Röser K, Heidorn K, Hallas C,
Löning T and Bullerdiek J: A new type of MAML2 fusion in
mucoepidermoid carcinoma. Genes Chromosomes Cancer. 47:203–206.
2008.PubMed/NCBI View Article : Google Scholar
|
|
16
|
Okabe M, Miyabe S, Nagatsuka H, Terada A,
Hanai N, Yokoi M, Shimozato K, Eimoto T, Nakamura S, Nagai N, et
al: MECT1-MAML2 fusion transcript defines a favorable subset of
mucoepidermoid carcinoma. Clin Cancer Res. 12:3902–3907.
2006.PubMed/NCBI View Article : Google Scholar
|
|
17
|
Nakayama T, Miyabe S, Okabe M, Sakuma H,
Ijichi K, Hasegawa Y, Nagatsuka H, Shimozato K and Inagaki H:
Clinicopathological significance of the CRTC3-MAML2 fusion
transcript in mucoepidermoid carcinoma. Mod Pathol. 22:1575–1581.
2009.PubMed/NCBI View Article : Google Scholar
|
|
18
|
Chen Z, Chen J, Gu Y, Hu C, Li JL, Lin S,
Shen H, Cao C, Gao R, Li J, et al: Aberrantly activated AREG-EGFR
signaling is required for the growth and survival of CRTC1-MAML2
fusion-positive mucoepidermoid carcinoma cells. Oncogene.
33:3869–3877. 2014.PubMed/NCBI View Article : Google Scholar
|
|
19
|
Dahse R, Driemel O, Schwartz S, Dahse J,
Kromeyer-Hauschild K, Berndt A and Kosmehl H: Epidermal growth
factor receptor kinase domain mutations are rare in salivary gland
carcinomas. Br J Cancer. 100:623–625. 2009.PubMed/NCBI View Article : Google Scholar
|
|
20
|
Nakano T, Yamamoto H, Hashimoto K, Tamiya
S, Shiratsuchi H, Nakashima T, Nishiyama K, Higaki Y, Komune S and
Oda Y: HER2 and EGFR gene copy number alterations are predominant
in high-grade salivary mucoepidermoid carcinoma irrespective of
MAML2 fusion status. Histopathology. 63:378–392. 2013.PubMed/NCBI View Article : Google Scholar
|
|
21
|
Warner KA, Adams A, Bernardi L, Nor C,
Finkel KA, Zhang Z, McLean SA, Helman J, Wolf GT, Divi V, et al:
Characterization of tumorigenic cell lines from the recurrence and
lymph node metastasis of a human salivary mucoepidermoid carcinoma.
Oral Onco. 49:1059–1066. 2013.PubMed/NCBI View Article : Google Scholar
|
|
22
|
Liu X, Ory V, Chapman S, Yuan H, Albanese
C, Kallakury B, Timofeeva OA, Nealon C, Dakic A, Simic V, et al:
ROCK inhibitor and feeder cells induce the conditional
reprogramming of epithelial cells. Am J Pathol. 180:599–607.
2012.PubMed/NCBI View Article : Google Scholar
|
|
23
|
Dulbecco R and Vogt M: Plaque formation
and isolation of pure lines with poliomyelitis viruses. J Exp Med.
99:167–182. 1954.PubMed/NCBI View Article : Google Scholar
|
|
24
|
Noguchi K, Wakai K, Kiyono T, Kawabe M,
Yoshikawa K, Hashimoto-Tamaoki T, Kishimoto H and Nakano Y:
Molecular analysis of keratocystic odontogenic tumor cell lines
derived from sporadic and basal cell nevus syndrome patients. Int J
Oncol. 51:1731–1738. 2017.PubMed/NCBI View Article : Google Scholar
|
|
25
|
Hiromoto T, Noguchi K, Yamamura M, Zushi
Y, Segawa E, Takaoka K, Moridera K, Kishimoto H and Urade M:
Up-regulation of neutrophil gelatinase-associated lipocalin in oral
squamous cell carcinoma: Relation to cell differentiation. Oncol
Rep. 26:1415–1421. 2011.PubMed/NCBI View Article : Google Scholar
|
|
26
|
Gupta R, Kalita P, Patil O and Mohanty S:
An investigation of folic acid-protein association sites and the
effect of this association on folic acid self-assembly. J Mol
Model. 21(308)2015.PubMed/NCBI View Article : Google Scholar
|
|
27
|
Rio DC, Ares M Jr, Hannon GJ and Nilsen
TW: Purification of RNA using TRIzol (TRI reagent). Cold Spring
Harb Protoc. 2010(pdb.prot5439)2010.PubMed/NCBI View Article : Google Scholar
|
|
28
|
Prophet EB, Mills B, Arrington JB and
Sobin LH: Laboratory methods in histotechnology (Armed Forces
Institute of Phatology). American Registry of Pathology,
Washington, DC, 1992.
|
|
29
|
Seethala RR: An update on grading of
salivary gland carcinomas. Head Neck Pathol. 3:69–77.
2009.PubMed/NCBI View Article : Google Scholar
|
|
30
|
Ohnishi Y, Minamino Y, Kakudo K and Nozaki
M: Resistance of oral squamous cell carcinoma cells to cetuximab is
associated with EGFR insensitivity and enhanced stem cell-like
potency. Oncol Rep. 32:780–786. 2014.PubMed/NCBI View Article : Google Scholar
|
|
31
|
Meng C, Wang S and Wang X, Lv J, Zeng W,
Chang R, Li Q and Wang X: Amphiregulin inhibits TNF-α-induced
alveolar epithelial cell death through EGFR signaling pathway.
Biomed Pharmacother. 125(109995)2020.PubMed/NCBI View Article : Google Scholar
|
|
32
|
Telesco SE, Vadigepalli R and
Radhakrishnan R: Molecular modeling of ErbB4/HER4 kinase in the
context of the HER4 signaling network helps rationalize the effects
of clinically identified HER4 somatic mutations on the cell
phenotype. Biotechnol J. 8:1452–1464. 2013.PubMed/NCBI View Article : Google Scholar
|
|
33
|
Li X, Huang Q, Wang S, Huang Z, Yu F and
Lin J: HER4 promotes the growth and metastasis of osteosarcoma via
the PI3K/AKT pathway. Acta Biochim Biophys Sin (Shanghai).
52:345–362. 2020.PubMed/NCBI View Article : Google Scholar
|
|
34
|
Luna MA: Salivary mucoepidermoid
carcinoma: Revisited. Adv Anat Pathol. 13:293–307. 2006.PubMed/NCBI View Article : Google Scholar
|
|
35
|
Mitra A, Mishra L and Li S: Technologies
for deriving primary tumor cells for use in personalized cancer
therapy. Trends Biotechnol. 31:347–354. 2013.PubMed/NCBI View Article : Google Scholar
|
|
36
|
Okumura Y, Miyabe S, Nakayama T, Fujiyoshi
Y, Hattori H, Shimozato K and Inagaki H: Impact of CRTC1/3-MAML2
fusions on histological classification and prognosis of
mucoepidermoid carcinoma. Histopathology. 59:90–97. 2011.PubMed/NCBI View Article : Google Scholar
|
|
37
|
Tirado Y, Williams MD, Hanna EY, Kaye FJ,
Batsakis JG and El-Naggar AK: CRTC1/MAML2 fusion transcript in high
grade mucoepidermoid carcinomas of salivary and thyroid glands and
Warthin's tumors: Implications for histogenesis and biologic
behavior. Genes Chromosomes Cancer. 46:708–715. 2007.PubMed/NCBI View Article : Google Scholar
|
|
38
|
Anzick SL, Chen WD, Park Y, Meltzer P,
Bell D, El-Naggar AK and Kaye FJ: Unfavorable prognosis of
CRTC1-MAML2 positive mucoepidermoid tumors with CDKN2A deletions.
Genes Chromosomes Cancer. 49:59–69. 2010.PubMed/NCBI View Article : Google Scholar
|
|
39
|
Komiya T, Park Y, Modi S, Coxon AB, Oh H
and Kaye FJ: Sustained expression of Mect1-Maml2 is essential for
tumor cell growth in salivary gland cancers carrying the t(11;19)
translocation. Oncogene. 25:6128–6132. 2006.PubMed/NCBI View Article : Google Scholar
|
|
40
|
Wu J, Wang N, Yang Y, Jiang G, Zhan H and
Li F: LINC01152 upregulates MAML2 expression to modulate the
progression of glioblastoma multiforme via Notch signaling pathway.
Cell Death Dis. 12(115)2021.PubMed/NCBI View Article : Google Scholar
|
|
41
|
Amelio AL, Fallahi M, Schaub FX, Zhang M,
Lawani MB, Alperstein AS, Southern MR, Young BM, Wu L, Zajac-Kaye
M, et al: CRTC1/MAML2 gain-of-function interactions with MYC create
a gene signature predictive of cancers with CREB-MYC involvement.
Proc Natl Acad Sci USA. 111:3260–3268. 2014.PubMed/NCBI View Article : Google Scholar
|
|
42
|
Shinomiya H, Ito Y, Kubo M, Yonezawa K,
Otsuki N, Iwae S, Inagaki H and Nibu KI: Expression of amphiregulin
in mucoepidermoid carcinoma of the major salivary glands: A
molecular and clinicopathological study. Hum Pathol. 57:37–44.
2016.PubMed/NCBI View Article : Google Scholar
|
|
43
|
Porcheri C and Mitsiadis TA: Physiology,
pathology and regeneration of salivary glands. Cells.
8(976)2019.PubMed/NCBI View Article : Google Scholar
|
|
44
|
Weng PL, Aure MH, Maruyama T and Ovitt CE:
Limited regeneration of adult salivary glands after severe injury
involves cellular plasticity. Cell Rep. 24:1464–1470.e3.
2018.PubMed/NCBI View Article : Google Scholar
|