Introduction
Lung cancer is the main type of cancer-related
fatalities worldwide, and there is an extremely strong association
between smoking and the formation of lung cancer (1,2). Tobacco is
a chemical compound that contains >5,000 types of chemicals
(3), of which 73 types were identified
as carcinogens by the International Agency for Research on Cancer.
There are >20 types that are associated with lung cancer,
including polycyclic aromatic hydrocarbons, such as benzopyrene and
4-(methylnitrosamino)-1-(3-pyridyl)-1-butanone (NNK) (4,5).
Studies have investigated the mechanism of
smoking-induced lung cancer for decades, and its mechanisms include
genetic alterations such as p53, KRAS mutations caused by forming
adducts with DNA replication, activation of cell surface receptors,
AKT, PKA-induced apoptotic genes, and the direct inhibition of the
activity of tumor suppressor genes by tobacco, undermining the
balance of the activity between oncogenes and tumor suppressor
genes. However, the main mechanism is the formation of DNA adducts
(6).
Benzopyrene is the most studied carcinogen in the
tobacco-related carcinogen, which causes DNA damage by DNA adduct
formation, and the specific DNA adducts have been detected in the
lungs of smokers (7). DNA adducts can
be cleared by DNA repair systems [such as nucleotide excision
repair (NER)] (8), and intracellular
DNA adducts can be rapidly cleared by effective intracellular DNA
repair. While DNA is replicated, if the damaged DNA can not be
repaired, DNA polymerase will terminate the copying, thereby
preventing replication of damaged DNA or cell death. When there is
more DNA polymerase than DNA adducts, this leads to gene mutations
caused by mismatches. The clearance of DNA adducts mainly depend on
NER, in which DNA polymerase has an irreplaceable role. DNA
polymerase δ (Pol δ) is essential for DNA replication, which
consists of four subunits, which are p125, p50, p68 and p12. The
expression of the smallest DNA Pol δ subunit, POLD4, was low in
certain non-small cell lung cancers and small cell lung cancers in
our previous study, and the lack of POLD4 protein expression or
downregulation of its expression by siRNA weakened DNA replication
and the NER capability (9).
The p12 subunit is rapidly degraded in cultured
human cells by DNA damage or replication stress induced by
treatments with ultraviolet (UV) radiation, methyl
methanesulfonate, hydroxyurea and aphidicolin (10,11). It is
well known that there is a close association between smoking and
lung cancer, and we hypothesize that smoking promotes the cancer
risk by decreasing POLD4 expression.
4-Nitroquinoline 1-oxide (4NQO) is a quinoline
derivative and a tumorigenic compound used in the assessment of the
efficacy of diets, drugs and procedures in the prevention and
treatment of cancer in animal models. It induces DNA lesions
usually corrected by nucleotide excision repair. In the present
study, 4NQO protein expression was observed in lung cancer cell
line A549, which was treated by the major carcinogen in tobacco,
benzopyrene analogs, in order to explore the mechanism of POLD4 in
smoking-induced lung cancer.
Materials and methods
Cell culture
The human lung cancer A549 cells were obtained from
the Cell Bank of Shanghai Institutes for Biological Sciences,
Chinese Academy of Sciences (Shanghai, China) and maintained in
RPMI-1640 medium supplemented with 10% heat-inactivated fetal
bovine serum. All the cells were incubated in a humidified
atmosphere with 5% CO2 at 37°C.
Transfection
Transfection was carried out using 50 nmol/l of a
siRNA (POLD4 siRNA: Sense, 5-GCAUCUCUAUCCCCUAUGATT-3 and antisense,
5-UCAUAGGGGAUAGAGAUGCTT-3). Duplex (Sigma-Aldrich, St. Louis, MO,
USA) targeting POLD4 mRNA or negative control 1 (control: Sense,
5-CUUUAAGCUCCCUGACGUUU-3 and antisense, 5-ACGCUCAGGGAGCUUAAAGUG-3,
Ambion, Carlsbad, CA, USA) were assessed with Lipofectamine® 2000
(Invitrogen Life Technologies, Carlsbad, CA, USA).
4NQO and MG132 treatments
The A549 cells were treated with different
concentrations of 4NQO (0, 0.1 and 0.4 µmol/l) for 48 h, and were
subsequently washed three times with phosphate-buffered saline
prior to protein analysis.
The degradation of p12 is due to an accelerated rate
of proteolysis that is inhibited by the proteasome inhibitors,
MG132 and lactacystin. At 3 h before 4NQO treatment, the proteasome
inhibitor MG132 (10 mmol/l) was added to block the activation of
calpain, and the cells were harvested for protein analysis.
MTT
The siRNA-treated A549 cells or non-treated cells
were separately cultured in 200 µl of culture medium in 96-well
plates at a density of 8,000 cells/well. The following day, the
medium was replaced with medium containing 4NQO or Taxol, followed
by incubation for 48 h. Viable cells were measured in triplicate
using TetraColor One (Seikagaku Co., Tokyo, Japan) with reference
to the viability of mock-treated cells.
Western blot analysis
The total cell lysates of treated cells were
separated by electrophoresis on 12.5% SDS-PAGE and transferred onto
a nitrocellulose membrane. The membrane was subsequently stained
with Ponceau S and cut into several pieces according to the
molecular weight of each designed protein. The pieces of membrane
were blocked with 5% w/v non-fat dry milk in Tris-buffered saline
and Tween-20 (TBST) buffer [20 mmol/l Tris-HCl (pH 7.4), 150 mmol/l
NaCl and 0.05% Tween-20] for 1 h at room temperature. The blots
were subsequently incubated with individual primary antibodies
(POLD4, 1:1,000; cat. no. H00057804-M10; Novus Biologicals, LCC,
Littleton, CO, USA; α-tubulin, 1:5,000; cat. no. sc-8035; Santa
Cruz Biotechnology, Inc., Dallas, TX, USA), corresponding to each
designed protein for 1 h at room temperature. After three 15-min
washes in TBST, the blots were incubated with alkaline
phosphatase-conjugated goat anti-mouse or anti-rabbit
immunoglobulin G (Pierce, Rockford, IL, USA) for 1 h and washed
with TBST three times for 10 min. The Perfect Protein Western Blot
kit (Novagen, Madison, WI, USA) was used for signal generation.
Statistical analysis
The statistical software SPSS 17.0 (SPSS, Inc.,
Chicago, IL, USA) was used to calculate the significance according
to Student's t-test. P<0.05 was considered to indicate a
statistically significant difference.
Results
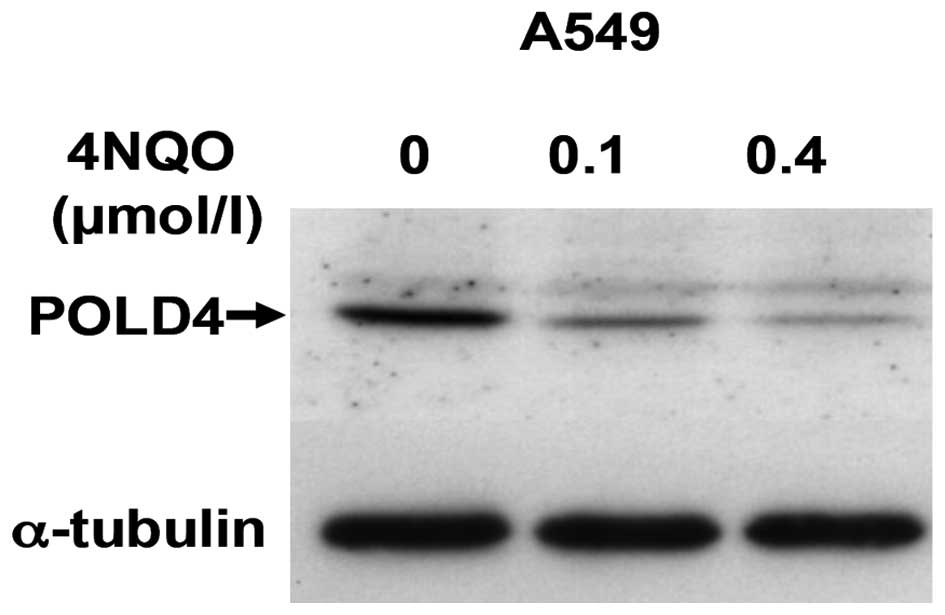
4NQO reduces POLD4 protein levels in
the lung cancer A549 cell line
Certain studies have shown that UV radiation or
hydroxyurea treatment converts Pol δ in vivo to the
three-subunit form lacking p12. In the present experiment, after 48
h treatment of A549 cells by 4NQO, the POLD4 protein expression was
evaluated by western blotting and the expression of POLD4 in
4NQO-treated A549 cells significantly decreased. There was a
positive correlation between the decrease and the 4NQO
concentration (Fig. 1).
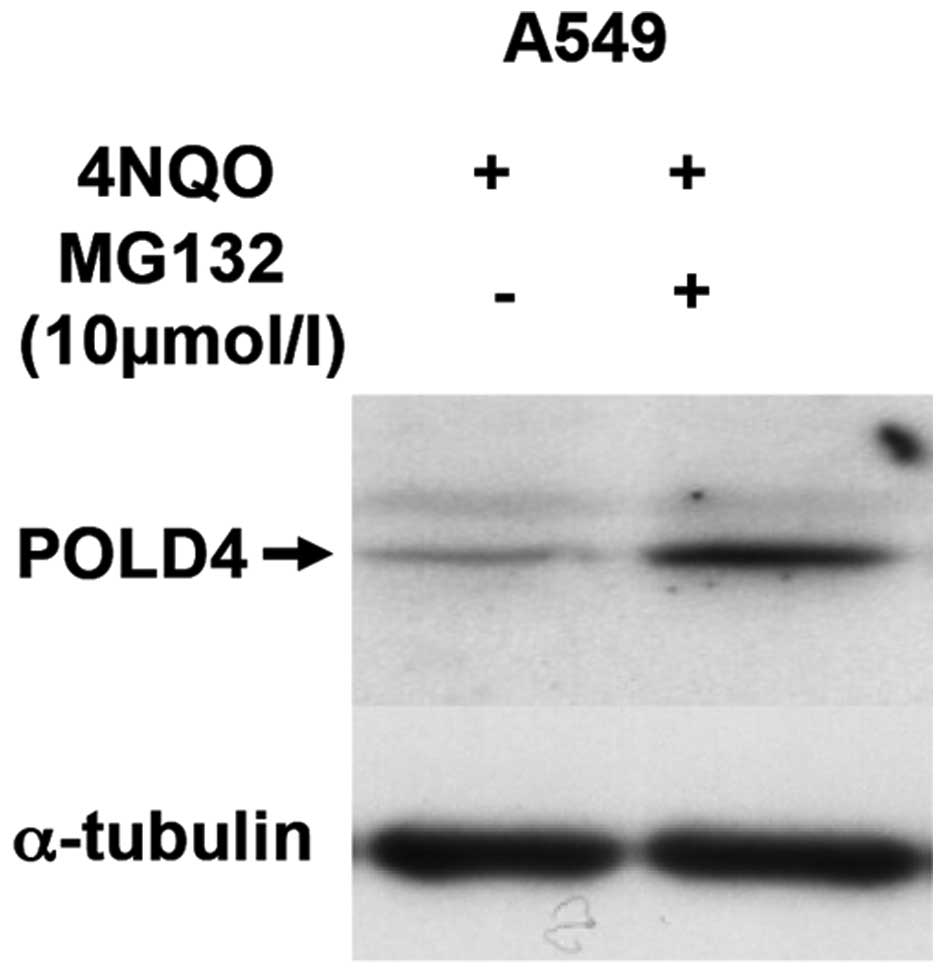
Low expression of POLD4 induced by
4NQO may be regulated by ubiquitination
POLD4 is degraded in response to DNA damage through
the ubiquitin-proteasome pathway (12). In order to clarify whether POLD4
degradation is caused by 4NQO through the ubiquitin-proteasome
pathway, cells were treated prior to 4NQO treatment by protease
activity inhibitor MG132, and it was observed that protease
activity inhibitors can reverse the decreased POLD4 protein levels
caused by 4NQO (Fig. 2), indicating
that the low POLD4 expression due caused by 4NQO may be via the
ubiquitin pathway.
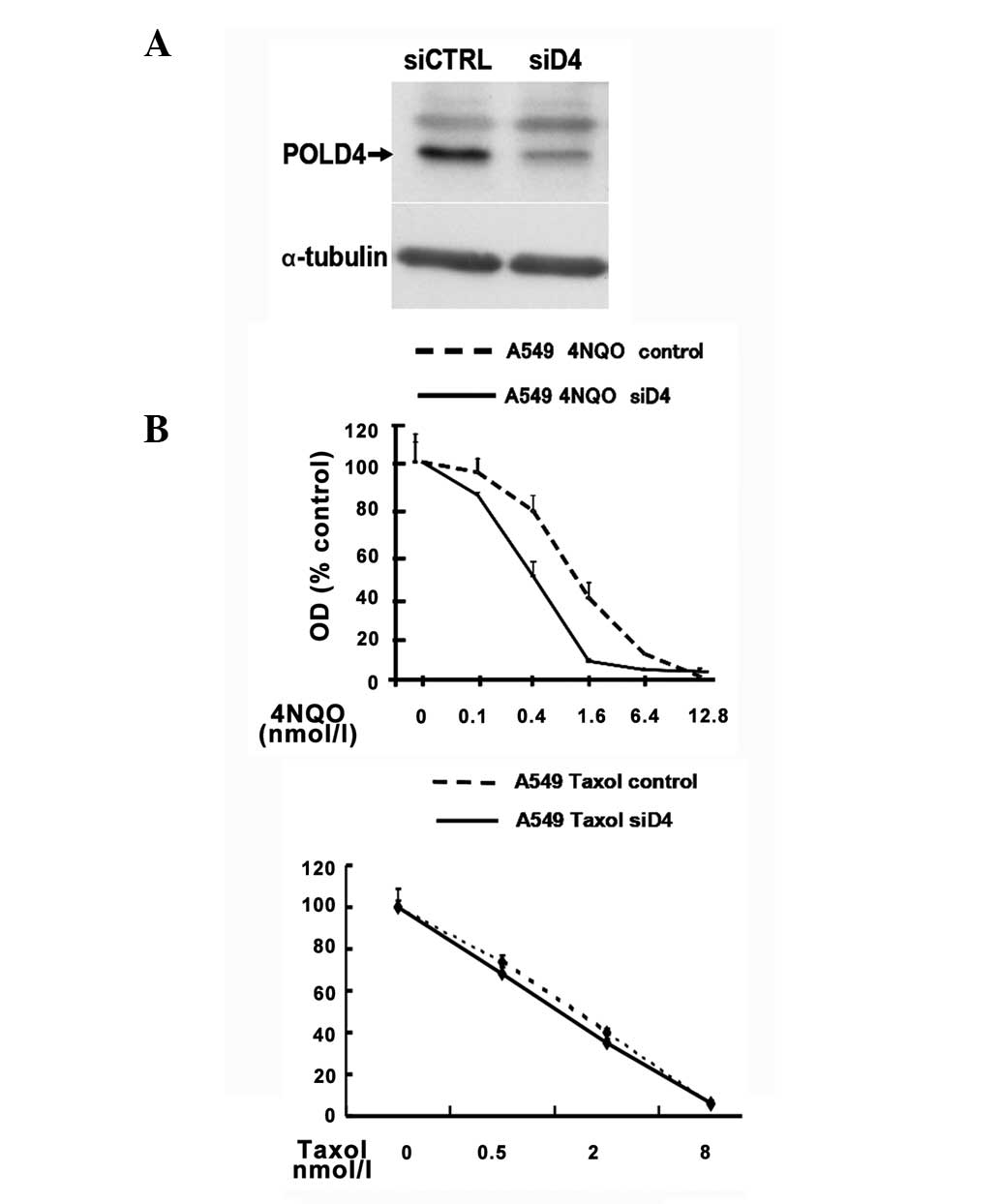
Low expression of POLD4 weakens the
DNA NER ability
Exposure to tobacco smoke and UV radiation can
result in various types of DNA damage and subsequently lead to
cancer formation. 4NQO is the UV radiation-mimetic chemical, which
has been thought to cause bulky DNA adducts and chromosomal
aberrations in exposed cells, and 4NQO-induced DNA damage can be
repaired by a global repair mechanism (13). Therefore, 4NQO-induced mutagen
sensitivity assays have been used to study susceptibility in
4NQO-treated cells in the present study. siRNA interference was
used to regulate the expression levels of POLD4 in A549 to observe
sensitivity of siRNA-treated cells and non-treated cells to 4NQO
and Taxol. The low POLD4 expression cells had a more enhanced
sensitivity to 4NQO toxicity compared to the siRNA non-treated
cells, indicating that low POLD4 expression weakened DNA NER
capacity. However, there was no difference in the sensitivity to
Taxol between the siRNA-treated cells and non-treated cells,
indicating that POLD4 expression is not involved in the sensitivity
to Taxol (P>0.05; Fig. 3 and
Table I).
 | Table I.Taxol and 4NQO-induced mutagen
sensitivity assays. |
Table I.
Taxol and 4NQO-induced mutagen
sensitivity assays.
| Treatments | Control | SiD4 | F | P-value |
|---|
| Taxol, nmol/l |
|
|
|
| 0.5 | 70.0±2.0 | 75.0±2.6 | 6.9 | 0.059 |
| 2 | 35.7±2.1 | 40.3±2.5 | 6.1 | 0.069 |
| 8 |
6.3±1.5 |
6.3±0.6 | 0.0 | 1.000 |
| 4NQO, µmol/l |
|
|
|
| 0.1 | 95.7±2.5 | 85.0±2.6 | 25.6 | 0.007 |
| 0.4 | 79.3±2.1 | 53.0±3.6 | 145.1 | 0.000 |
| 1.6 | 39.7±2.5 | 10.0±1.7 | 282.9 | 0.000 |
| 6.4 |
9.3±1.2 |
4.3±0.6 | 45.0 | 0.003 |
| 12.8 |
3.0±1.0 |
3.0±0.8 |
0.0 | 1.000 |
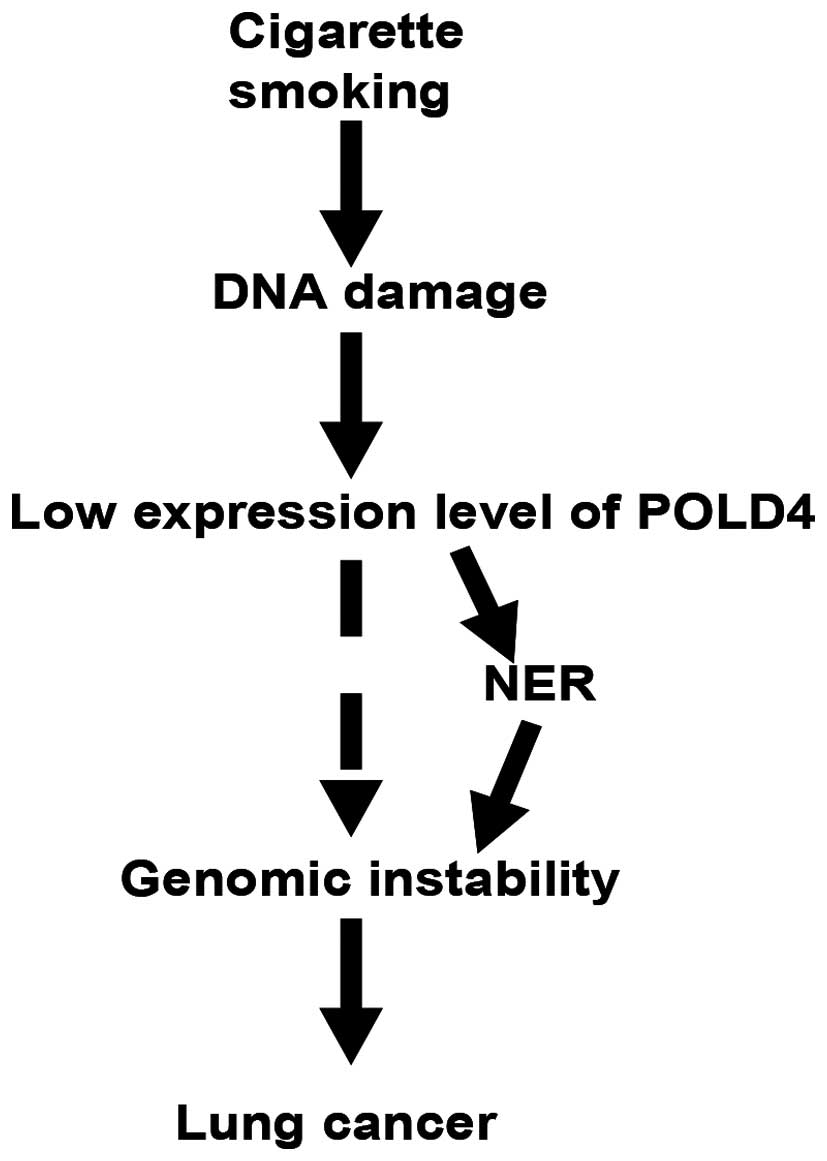
Possible mechanisms of POLD4 in the
effects of 4NQO on A549 cells
The formation of DNA damage caused by tobacco
inhalation decreased POLD4 expression levels. Low POLD4 expression
can cause genomic instability through different mechanisms; by
contrast, the repair of DNA damage caused by 3–4 benzopyrene in
tobacco depends on the NER capacity of Pol δ, while low POLD4
expression of the risk may weaken the NER capacity, thereby
increasing the instability of the genome. These are two reasons for
the cause of genomic instability and ultimately the increase in the
risk (Fig. 4) of lung cancer
formation.
Discussion
In previous studies, POLD4 expression in small cell
lung cancer and a small section of non-small cell lung cancers is
lower when compared to normal samples (9). Clinical data showed that almost all small
cell lung cancers have a smoking history, indicating the
possibility of a correlation between smoking-induced lung cancer
and POLD4.
There are >20 types of lung cancer carcinogens in
tobacco, of which the most significant carcinogens are polycyclic
aromatic hydrocarbons such as benzopyrene and NNK. The most studied
polycyclic aromatic hydrocarbon is benzopyrene. In the present
study, the benzopyrene analogue 4NQO was used. 4NQO is a synthetic
water-soluble carcinogen that is commonly used as a carcinogen in
murine models for investigating the various stages of oral
carcinogenesis. 4NQO exerts potent intracellular oxidative stress
and its metabolic product binds to DNA predominantly at the guanine
residues. These insults appear similar to the damage induced by
other carcinogens that are present in tobacco (14).
DNA Pol δ is involved in numerous DNA damage
responses, and the Pol δ4 holoenzyme consists of four subunits,
which are p125, p50, p68 and p12. The p12 subunit is known to be
rapidly degraded in response to DNA damage by UV radiation,
hydroxyurea or DNA replication stress, leading to the in
vivo conversion of Pol δ4 to Pol δ3, a trimeric form lacking
the p12 subunit (15), via the
ubiquitin-proteasome pathway (10,11).
In our previous study (9), the low expression of POLD4 in small cell
and non-small cell lung cancer suggested that there is a possible
association between smoking and POLD4 genes. In the present study,
4NQO decreased the POLD4 protein expression in the lung cancer cell
line A549, and the effect could be reversed by the proteasome
inhibitor MG132.
4NQO produces a DNA adduct that is removed through
Pol δ-dependent NER DNA. The effects of the decreased POLD4
expression were explored in the NER capacity in A549 cells, and the
results showed that a low POLD4 expression enhanced the cell
sensitivity to 4NQO, indirectly indicating that the POLD4 decrease
weakened the NER capacity. In the present study, another type of
cytotoxic drug, Taxol, was selected as a control to cause
cytotoxicity, however, the repair of DNA damage did not depend on
the NER capacity.
The study by Meng et al (11) showed that genotoxic agents, including
UV and alkylating chemicals, induce a DNA damage response in which
Pol δ4 is converted to a trimer (Pol δ3) by degradation of p12, and
Pol δ3 exhibits an enhanced ability for the detection of errors in
primers and templates in comparison to its parent enzyme, whereas
in the present study, a low expression of Pol δ4 decreased the NER
capacity.
In our previous study, we proved that regulation of
DNA polymerase POLD4 influenced genomic instability in lung cancer,
and found that downregulation of POLD4 in Calu6 cells results in
G1-S blockage through suppression of the AKT-Skp2-p27
pathway (9,16,17).
Our previous study also reported that the
shRNA-mediated reduction of POLD4 resulted in a marked decrease in
colony formation activity in Calu6, ACC-LC-319 and PC-10 cells, and
POLD4 reduction was also associated with an increased population of
karyomere-like cells, which may be an indication of DNA replication
stress and/or DNA damage. siRNA-mediated reduction of POLD4 in
cells with an abundant expression resulted in a cell cycle delay,
checkpoint activation and an elevated frequency of chromosomal
gap/break formation.
In conclusion, the present study reported that a
significant tobacco carcinogen, benzopyrene analogue 4NQO,
downregulated the POLD4 expression level, and thereby caused the
decreased NER, further resulting in genomic instability, and
ultimately an increased risk of lung cancer formation. Combined
with the results of our previous study, we have proposed the
mechanisms of action for POLD4 in smoking-induced lung cancer.
Acknowledgements
The present study was supported partly by grants
from the National Natural Science Foundation of China (no.
81141093), the Science Foundation of the Fujian Province, China
(no. 2013J01290), Technology Foundation for Selected Overseas
Chinese Scholar, Ministry of Personnel of China (2012) and the
Nursery Research Fund of Second Affiliated Hospital of Fujian
Medical University (no. 2012MP73).
References
|
1
|
Hecht SS: Tobacco carcinogens, their
biomarkers and tobacco-induced cancer. Nat Rev Cancer. 3:733–744.
2003. View
Article : Google Scholar : PubMed/NCBI
|
|
2
|
Osada H and Takahashi T: Genetic
alterations of multiple tumor suppressors and oncogenes in the
carcinogenesis and progression of lung cancer. Oncogene.
21:7421–7434. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Yovel Y, Franz MO, Stilz P and Schnitzler
HU: Plant classification from bat-like echolocation signals. PLoS
Comput Biol. 4:e10000322008. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Hecht SS: Chemoprevention of cancer by
isothiocyanates, modifiers of carcinogen metabolism. J Nutr.
129:768S–774S. 1999.PubMed/NCBI
|
|
5
|
Hecht SS, Yuan JM and Hatsukami D:
Applying tobacco carcinogen and toxicant biomarkers in product
regulation and cancer prevention. Chem Res Toxicol. 23:1001–1008.
2010. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Hecht SS: Lung carcinogenesis by tobacco
smoke. Int J Cancer. 131:2724–2732. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Pfeifer GP, Denissenko MF, Olivier M,
Tretyakova N, Hecht SS and Hainaut P: Tobacco smoke carcinogens,
DNA damage and p53 mutations in smoking-associated cancers.
Oncogene. 21:7435–7451. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Hoeijmakers JH: Genome maintenance
mechanisms for preventing cancer. Nature. 411:366–374. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Huang QM, Tomida S, Masuda Y, Arima C, Cao
K, Kasahara TA, Osada H, Yatabe Y, Akashi T, Kamiya K, et al:
Regulation of DNA polymerase POLD4 influences genomic instability
in lung cancer. Cancer Res. 70:8407–8416. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Zhang S, Zhou Y, Trusa S, Meng X, Lee EY
and Lee MY: A novel DNA damage response: Rapid degradation of the
p12 subunit of dna polymerase delta. J Biol Chem. 282:15330–15340.
2007. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Meng X, Zhou Y, Zhang S, Lee EY, Frick DN
and Lee MY: DNA damage alters DNA polymerase delta to a form that
exhibits increased discrimination against modified template bases
and mismatched primers. Nucleic Acids Res. 37:647–657. 2009.
View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Zhang S, Zhou Y, Sarkeshik A, Yates JR
III, Thomson TM, Zhang Z, Lee EY and Lee MY: Identification of RNF8
as a ubiquitin ligase involved in targeting the p12 subunit of DNA
polymerase δ for degradation in response to DNA damage. J Biol
Chem. 288:2941–2950. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Li C, Wang LE and Wei Q: DNA repair
phenotype and cancer susceptibility - a mini review. Int J Cancer.
124:999–1007. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Kanojia D and Vaidya MM:
4-nitroquinoline-1-oxide induced experimental oral carcinogenesis.
Oral Oncol. 42:655–667. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Chea J, Zhang S, Zhao H, Zhang Z, Lee EY,
Darzynkiewicz Z and Lee MY: Spatiotemporal recruitment of human DNA
polymerase delta to sites of UV damage. Cell Cycle. 11:2885–2895.
2012. View
Article : Google Scholar : PubMed/NCBI
|
|
16
|
Huang QM, Akashi T, Masuda Y, Kamiya K,
Takahashi T and Suzuki M: Roles of POLD4, smallest subunit of DNA
polymerase delta, in nuclear structures and genomic stability of
human cells. Biochem Biophys Res Commun. 391:542–546. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Huang Q, Suzuki M, Zeng Y, Zhang H, Yang D
and Lin H: Downregulation of POLD4 in Calu6 cells results in G1-S
blockage through suppression of the Akt-Skp2-p27 pathway. Bioorg
Med Chem Lett. 24:1780–1783. 2014. View Article : Google Scholar : PubMed/NCBI
|