Introduction
According to the GLOBOCAN 2012 statistics,
colorectal cancer (CRC) is the third most common cancer in the
world. Every year there are ~0.7 million CRC-related mortalities
worldwide (1). The usual treatments
for CRC are surgery, radiotherapy, chemotherapy and drugs; however,
the therapeutic effects of these strategies are insufficient due to
the occurrence of invasion and metastasis (2). Tumor metastasis is a complicated
process involving multiple steps, one of which is known as the
epithelial-to-mesenchymal transition (EMT). As a result of the EMT,
epithelial cells gain a mesenchymal phenotype, enabling them to
migrate to and invade a distant organ (3). An enhanced understanding of the
molecular mechanisms regulating the EMT in CRC is therefore
critical for more effective treatments.
microRNAs (miRNAs) are small, single-stranded RNA
molecules of 20–24 bases in length that belong to the non-coding
RNA species. miRNAs regulate gene expression by binding to the
3′-untranslated region (UTR) of their target mRNA, inhibiting
translation or degrading the mRNA. To date, >1,400 miRNAs have
been identified, and >5,300 human genes have been predicted to
be targets of miRNAs (4). miRNAs are
involved in numerous biological processes, including embryonic
development, cell differentiation and metabolism (5–7).
Aberrant miRNA expression in cancer has been found to participate
in several pathological processes, such as EMT and other events
contributing to tumorigenesis (8,9).
miR-20a is a member of the miR-17-92 cluster, which
includes miR-17-5p, miR-17-3p, miR-18a, miR-19a, miR-20a, miR-19b-l
and miR-92-1, and is located on chromosome 13q31 (10). miR-20a has been reported to be
associated with EMT in various types of cancers. In hepatocellular
carcinoma (HCC), miR-20a restoration has been shown to inhibit cell
proliferation and induce apoptosis by targeting the 3′-UTR of
Mcl-1, while decreased expression of miR-20a has been reported to
correlate significantly with the aggressive characteristics of HCC
cells (11). In addition, the
upregulated expression of miR-20a has been found to decrease the
migratory ability of oral squamous cell carcinoma (OSCC) cell lines
(12), and the ectopic expression of
miR-20a has been demonstrated to induce the EMT and enhance the
metastasis of gallbladder carcinoma (GBC) cells in vitro and
in vivo (13). In CRC,
however, the role of miR-20a in EMT remains largely unknown. The
aim of the present study, therefore, was to characterize the
expression profile of miR-20a in CRC tissues and analyze its
correlation with the clinicopathological characteristics of the
condition, and then to clarify the role of miR-20a in EMT in CRC
cells lines.
Materials and methods
Tissue samples and cell culture
The present study was approved by the Committee for
the Ethical Review of Research at the Provincial Hospital
affiliated to Shandong University (Jinan, China). A total of 30
samples of tumor and adjacent non-tumor tissue were obtained from
the Provincial Hospital, and signed consent forms were obtained
from all patients. The human SW620, SW480, LS174T and HCT116 CRC
cell lines and the normal human colonic epithelial cell line FHC
were purchased from the Cell Bank of Chinese Academy of Sciences
(Shanghai, China). The cells were cultured in RPMI-1640 and
Dulbecco's modified Eagle's medium (Gibco-BRL, Grand Island, NY,
USA) supplemented with 10% fetal bovine serum (Gibco-BRL) at 37°C
in a humidified atmosphere containing 5% CO2.
Reverse transcription-quantitative
polymerase chain reaction (RT-qPCR)
Total RNA was extracted using TRIzol® reagent
(Invitrogen Life Technologies, Carlsbad, CA, USA) according to the
manufacturer's instructions, and the miRNA was reverse-transcribed
into miRNA using a One-Step PrimeScript® miRNA cDNA Synthesis kit
(Takara Bio, Inc., Shiga, Japan). RT-qPCR was performed using a
SYBR® Premix Ex Taq kit (Takara Bio, Inc.) on an ABI 7300 qPCR
system (Applied Biosystems, Foster City, CA, USA). The primers for
miR-20a and the reference gene U6 were purchased from RiboBio Co.,
Ltd. (Guangzhou, China) and were as follows: miR-20a, F 5-TAC GAT
AAA GTG CTT ATA GTG CAG GTA G-3; U6, F 5-AAA GAC CTG TAC GCC AAC
AC-3. The PCR cycle conditions consisted of an initial denaturation
step at 95°C for 30 sec, followed by 40 cycles at 95°C for 30 sec
and 60°C for 1 min, and a final extension step of 60°C for 1 min.
The relative expression ratio of miR-20a was quantified using the
2−ΔΔCT method.
Establishment of CRC cell lines with
knocked down and overexpressed miR-20a
Plasmids containing miR-20a inhibitor, miR-20a
mimics, inhibitor negative control and mimic negative control were
purchased from GenePharma (Shanghai, China). The SW620 and LS174T
cell lines were selected for knockdown and overexpression
modifications, respectively. A total of 2×105 cells were
seeded into a six-well plate. Twenty-four hours later, when the
cells were 70–90% confluent, 2 µg plasmids were transfected into
the cells using Lipofectamine® 2000 (Invitrogen Life Technologies).
Stably transfected cell lines were selected using 400 µg/µl G418
(Sigma, St. Louis, MO, USA) for 1 month.
Cell proliferation assay
Cell proliferation was measured using a cell
counting kit-8 (CCK-8) assay (Dojindo Molecular Technologies,
Kumamoto, Japan) according to the manufacturer's instructions.
Transwell® assay
Cell invasion was measured using Transwell chambers
(Corning, Inc., Corning, NY, USA). Cells were suspended in
serum-free medium and seeded in the upper chamber; 500 µl complete
RPMI-1640 was added to the lower well. The cells were incubated for
24 h and the cells that invaded the lower surface were fixed with
methanol for 10 min and stained with crystal violet (Beyotime Co.,
Shanghai, China) for 30 min. The cells were counted under a
microscope (Nikon, Tokyo, Japan) at a magnification of x400.
Wound healing assay
A total of 5×105 cells were seeded in a
six-well plate and incubated overnight until the cells had grown to
a confluent monolayer. The wound was then made by scratching with a
200-µl Eppendorf tip, and the wounded monolayers were washed with
phosphate-buffered saline twice to remove any non-adherent cells,
prior to being incubated in serum-free medium. Images of the wound
areas were captured under a microscope (Nikon).
Western blot analysis
Radioimmunoprecipitation assay lysis buffer and
phenylmethylsulfonyl fluoride (Beyotime Co.) were added to the
cells. After lysis for 30 min, the cellular extracts were
centrifuged at 13,000 × g for 10 min at 4°C, and the supernatant
was frozen at −80°C. The total extracted protein was quantified
using a bicinchoninic acid protein assay kit (Beyotime Co.). The
proteins were separated using 10% sodium dodecyl
sulfate-polyacrylamide gel electrophoresis, and then transferred to
polyvinylidene difluoride membranes and blocked in 5% non-fat milk
in Tris buffered saline/Tween 20 buffer. The membranes were
incubated overnight at 4°C with primary antibodies against
E-cadherin (#14472, mouse monoclonal, 1:500), vimentin (#3878,
rabbit polyclonal, 1:500), matrix metalloproteinase-2 (MMP-2)
(#4022, rabbit polyclonal, 1:1,000) and MMP-9 (#13667, rabbit
monoclonal, 1:500), purchased from Cell Signaling Technology Inc.
(Beverly, MA, USA), and mouse monoclonal primary antibodies against
TIMP-2 (#MAB3310, mouse monoclonal, 1:500, Chemicon International,
Inc., Temecula, CA, USA) and β-actin (#BM0627, 1:1,000, Wuhan
Boster Biological Technology, Ltd., Wuhan, China). After the
membranes were washed with TBST 6 × 10 min, the membranes were
incubated with secondary antibodies (#BA1054, goat anti-rabbit and
#BA1051, goat anti-mouse 1:2,000; Wuhan Boster Biological
Technology, Ltd.). The protein bands were detected using an
enhanced chemiluminescence detection system (Beyotime Co.). β-actin
was used as a loading control.
Statistical analysis
Data are presented as the mean ± standard deviation
from at least three separate experiments. The correlations between
miR-20a expression levels and pathological features were analyzed
with the χ2 test. All statistical analyses were
conducted using SPSS version 17.0 statistical software (SPSS, Inc.,
Chicago, IL, USA). P<0.05 was considered to indicate a
statistically significant difference.
Results
Increased miR-20a expression in CRC
tissues and cell lines
RT-qPCR analysis was carried out to detect the level
of miR-20a in 30 tumor and matched non-tumor tissues, four CRC cell
lines (SW620, SW480, LS174T and HCT116) and one normal colon
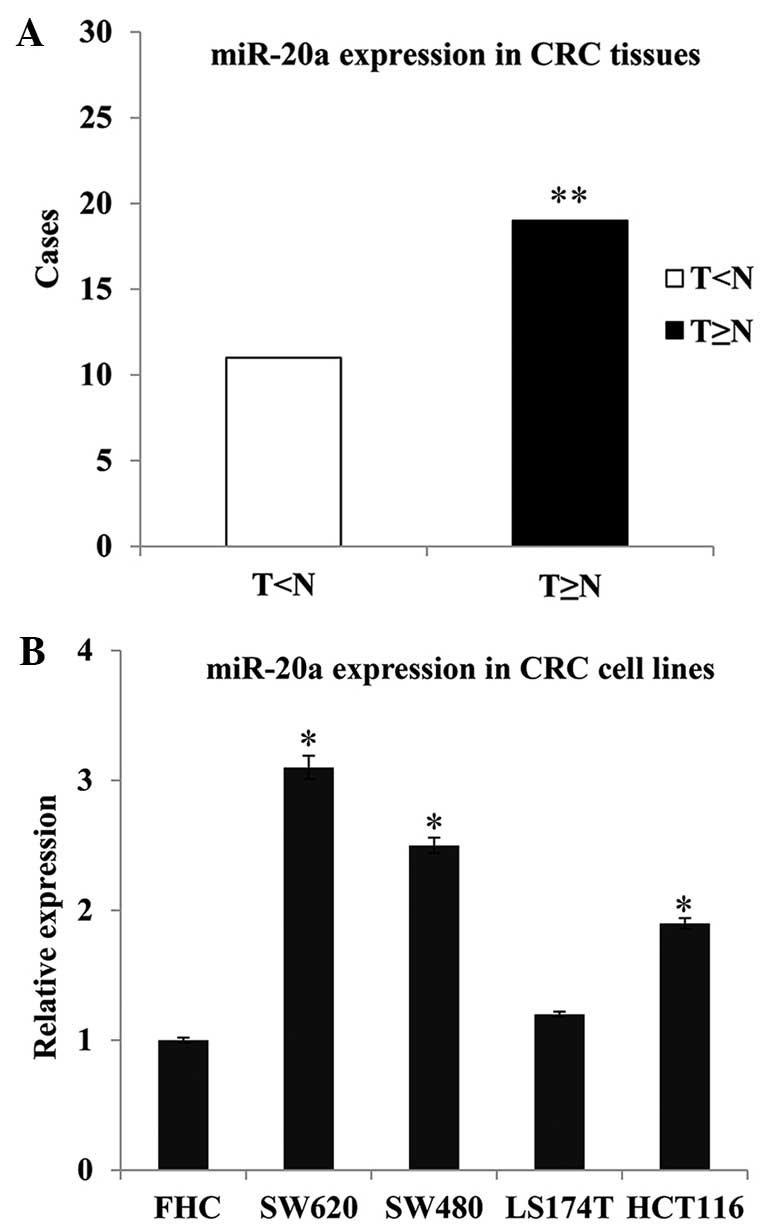
epithelium cell line (FHC). The results showed an increased
expression of miR-20a in the tumor tissues compared with the
non-tumor tissues; the frequency of miR-20a upregulation was 63.33%
(19/30; P=0.008) (Fig. 1A). The four
CRC cell lines also exhibited higher miR-20a expression levels than
the normal colon epithelium cells (SW620 vs. FHC, P=0.016; SW480
vs. FHC, P=0.021; and HCT116 vs. FHC, P=0.024) (Fig. 1B).
In order to explore the association between the
expression pattern of miR-20a and the clinicopathological
characteristics of the patients with CRC, 30 patients were
categorized into low (<1.5-fold) and high (≥1.5-fold) miR-20a
expression groups. The results are summarized in Table I. Statistical analyses revealed that
the high expression of miR-20a was associated with tumor invasion
(P=0.015) and lymph node metastasis (P=0.047), but not with age,
gender, differentiation or distant metastasis. These data indicate
that increased miR-20a expression plays an important role in CRC
metastasis.
 | Table I.Association between miR-20a expression
and the clinicopathological characteristics of patients with
colorectal cancer. |
Table I.
Association between miR-20a expression
and the clinicopathological characteristics of patients with
colorectal cancer.
|
| Relative miR-20a
expression |
|---|
|
|
|
|---|
| Characteristic | Low (n) | High (n) | P-value |
|---|
| Gender |
|
|
|
| Male | 8 | 11 | 0.466 |
|
Female | 3 | 8 |
|
| Age in years |
|
|
|
|
<60 | 2 | 7 | 0.419 |
| ≥60 | 9 | 12 |
|
| Differentiation |
|
|
|
|
Poorly | 3 | 7 | 0.702 |
|
Moderately/well | 8 | 12 |
|
| Local invasion |
|
|
|
| Yes | 7 | 3 | 0.015 |
| No | 4 | 16 |
|
| Lymph node
metastasis |
|
|
|
| Yes | 8 | 5 | 0.047 |
| No | 3 | 14 |
|
| Distant
metastasis |
|
|
|
| Yes | 6 | 4 | 0.108 |
| No | 5 | 15 |
|
Levels of miR-20a in CRC cell lines
with knocked down and overexpressed miR-20a
SW620 cells transfected with miR-20a inhibitor and
inhibitor negative control plasmids were termed SW620-Si20a and
SW620-NC cells, respectively. LS174T cells transfected with miR-20a
mimic and mimic negative control plasmids were termed LS174T-OV20a
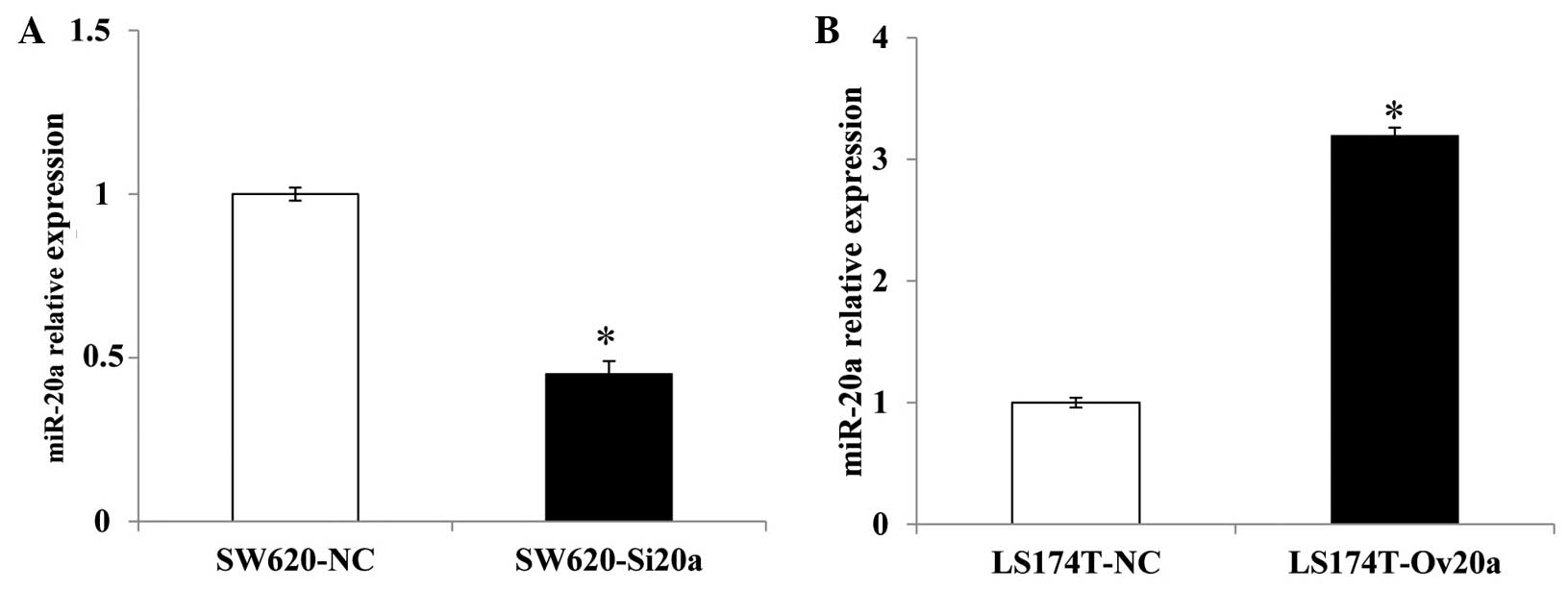
and LS174T-NC cells, respectively. RT-qPCR was conducted to confirm
the expression levels of miR-20a, and the results showed that,
compared with the SW620-NC cells, the expression of miR-20a in the
SW620-Si20a cells was decreased (P=0.031). By comparison, the
LS174T-OV20a cells showed an increased level of miR-20a when
compared with the LS174T-NC cells (P=0.015) (Fig. 2).
miR-20a promotes the proliferative
ability of colon cells
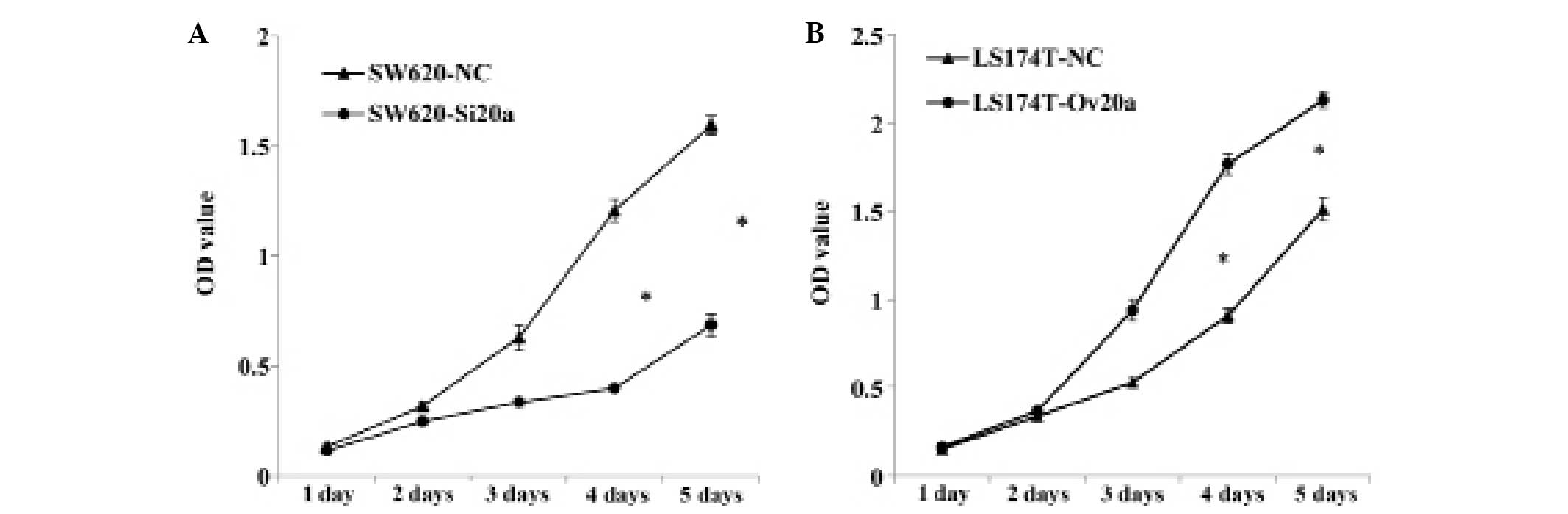
The CCK-8 assay was employed to detect the
proliferative ability of cells with knocked down or overexpressed
miR-20a. Suppression of miR-20a significantly inhibited the growth
of SW620 cells at the fourth day (P<0.05), while miR-20a
overexpression enhanced LS174T cells proliferation at the same
time-point (P<0.05) (Fig. 3A and
B). These results indicate that miR-20a promotes the
proliferative ability of CRC cell lines.
miR-20a enhances the invasive and
migratory ability of CRC cells
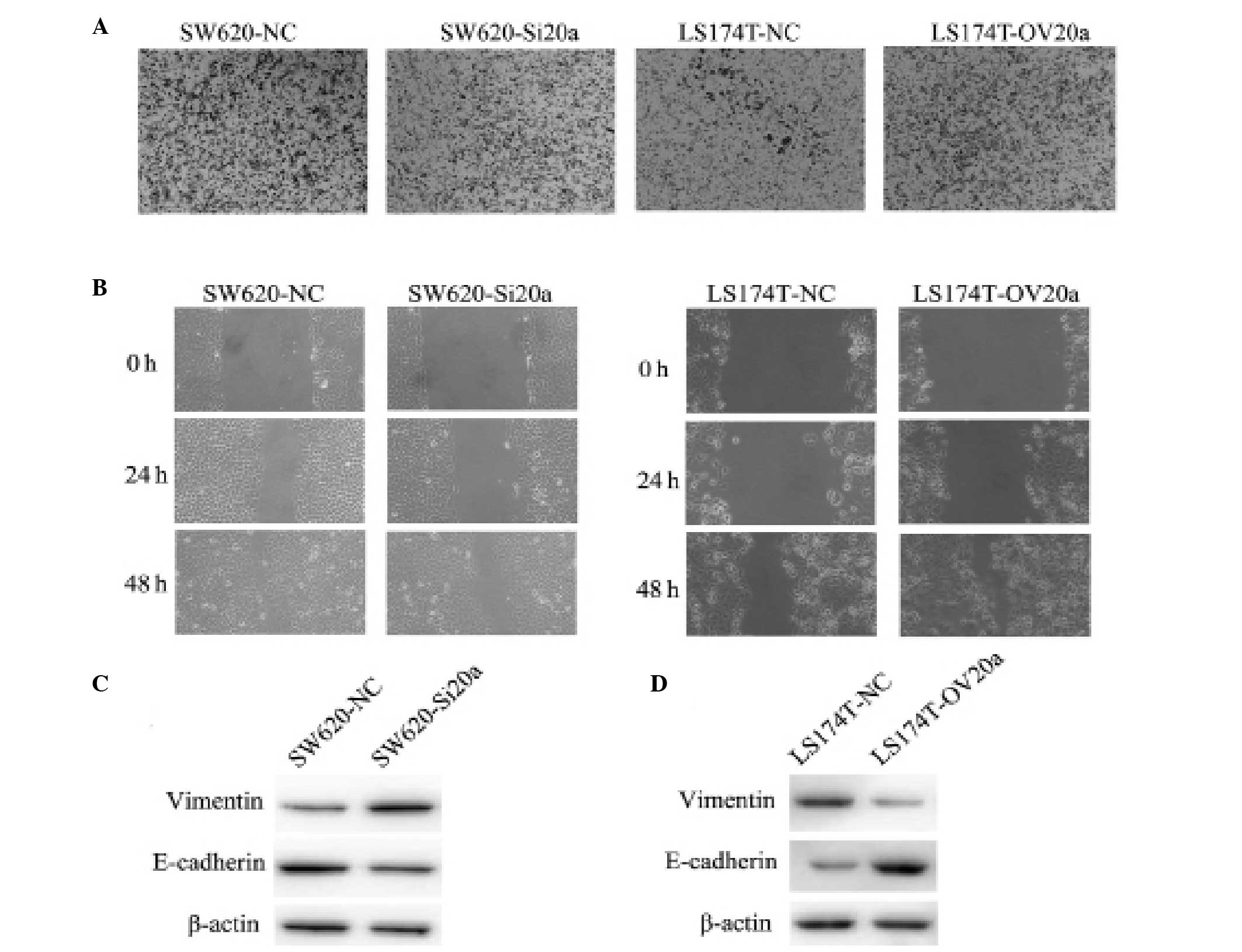
To evaluate the role of miR-20a in invasion and
migration, Transwell and wound healing assays were performed using
the stably transfected CRC cell lines. Compared with the control,
SW620-Si20a cells exhibited a significantly inhibited invasive
ability. By contrast, overexpression of miR-20a increased the
invasion of LS174T-OV20a cells as compared with the LS174T-NC cells
(Fig. 4A). Consistent with these
results, the wound healing assay indicated that miR-20a-knockdown
slowed the rate of wound closure, while miR-20a overexpression
increased the migratory ability (Fig.
4B). Western blot analyses showed that the loss of miR-20a
expression enhanced the level of vimentin and reduced the level of
E-cadherin in SW620 cells; by contrast, ectopic expression of
miR-20a in the LS174T cells led to an increased level of E-cadherin
and a decreased level of vimentin (Fig.
4C and D). These findings demonstrate that miR-20a enhances the
EMT of CRC cells.
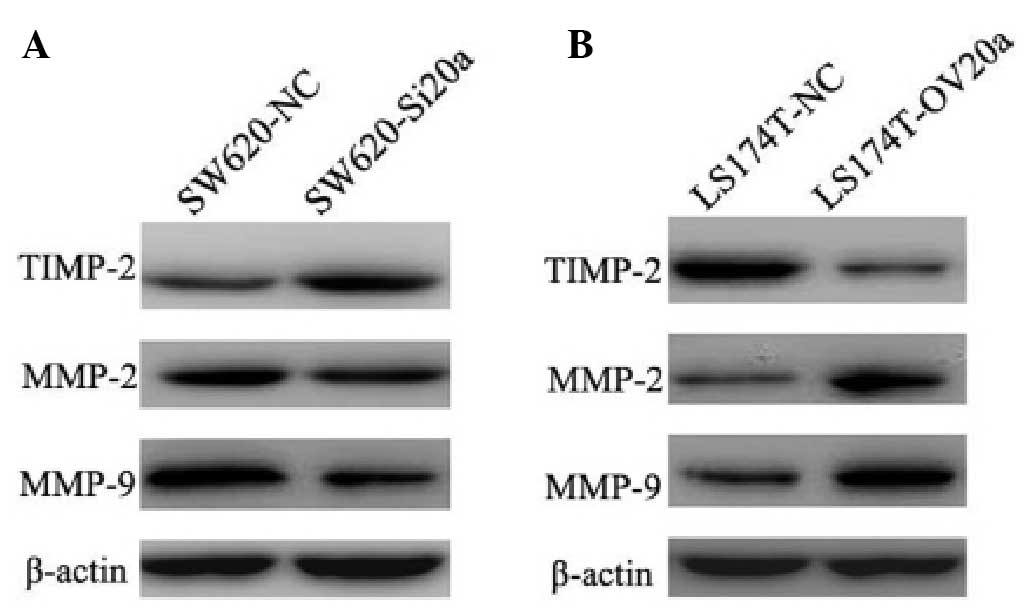
miR-20a enhances EMT by modulating the
expression of TIMP-2 and MMPs
The finding that miR-20a is involved in the
promotion of cell invasion and migration suggested that miR-20a is
additionally involved in the EMT process. To further explore the
mechanism by which miR-20a modulated the EMT, western blot analysis
was performed to examine the expression of EMT-related markers. In
the SW620-Si20a cells, knockdown of miR-20a upregulated the TIMP-2
and downregulated the MMP-2 and MMP-9 protein levels (Fig. 5A). In the LS174T-OV20a cells,
overexpression of miR-20a inhibited the expression of TIMP-2 and
induced the expression of MMP-2 and MMP-9 (Fig. 5B). These data indicate that miR-20a
enhances EMT by regulating the expression of TIMP-2, MMP-2 and
MMP-9.
Discussion
In CRC, the high mortality rates are primarily a
result of tumor metastasis. At diagnosis, most patients are found
to exhibit distant metastasis; even following surgery, ~50% of
these patients are likely to undergo tumor relapse and succumb
(14). The inhibition of tumor
metastasis, therefore, is a promising therapeutic approach.
Increasing numbers of miRNAs have been demonstrated to play
significant roles in cell proliferation and the migration and
invasion of various types of tumor, suggesting that these molecules
represent a novel class of therapeutic target in cancer (8).
Studies have shown that miR-20a exhibits different
expression profiles in different types of cancer (11–13,15–18).
Upregulation of miR-20a has been found in GBC (13), colon adenocarcinoma (15) and gliomas (16), while miR-20a downregulation has been
reported in primary HCC (11), OSCC
(12), breast cancer (17) and pancreatic carcinoma (18). This suggests that the function of
miR-20a may vary among different types of tumor. The present data
showed an elevated expression profile of miR-20a in 30 CRC cases,
which is consistent with the data from a previous study (19). Further analysis revealed that the
high expression of miR-20a in patients with CRC was associated with
tumor invasion and lymph node metastasis. It should be noted,
however, that the number of clinical samples in this study was
relatively small; therefore, further research with larger samples
is necessary to verify the findings. By investigating the
biological function of miR-20a in CRC cells, the present study
showed that miR-20a enhanced the proliferation, invasion and
migration of the cells.
EMT is an biological process that can be classified
into one of three subtypes: Embryogenesis and organ development,
tissue regeneration and organ fibrosis or cancer progression and
metastasis (20). During cancer
progression, cancer cells detach from the basement membrane,
migrate to and invade surrounding tissues and eventually colonize
remote sites (21). A hallmark of
cells that have undergone EMT is the loss of the epithelial cell
marker E-cadherin and the gain of the mesenchymal cell markers
N-cadherin and vimentin (22). Given
that the present results revealed that miR-20a increased the
EMT-related cell invasion and migration, the next step was to
examine the changes in the markers of EMT. The epithelial marker
E-cadherin was upregulated following transfection with the miR-20a
mimics and decreased following transfection with the inhibitor of
miR-20a. By comparison, the mesenchymal marker vimentin was
downregulated following transfection with the miR-20a mimics and
increased following transfection with the inhibitor of miR-20a.
These results further confirmed that miR-20a was involved in
modulating the EMT.
To further investigate the mechanism of miR-20a in
enhancing EMT, the expression of TIMP-2, MMP-2 and MMP-9 was
examined. MMP-2 and MMP-9 are members of the MMP family that
degrade the basement membrane and extracellular matrix,
facilitating the invasion and migration of cancer cells (23,24).
Aberrant expression of MMP-2 and MMP-9 has been confirmed to be
associated with proliferation, invasion and EMT in a variety of
types of cancer (25,26). TIMP-2 is the key endogenous regulator
of MMPs; by inhibiting the activity of MMPs, it thus inhibits tumor
migration (27). The present data
showed that, with the downregulation of miR-20a in SW620 cells, the
expression of TIMP-2 was upregulated and that of MMP-2 and MMP-9
was downregulated. The opposite results were obtained in LS174T
cells overexpressing miR-20a. Wang et al (28) found that TIMP-2 is the direct target
of miR-20a; in the present study, therefore, miR-20a may have
increased the expression of MMP-2 and MMP-9 by suppressing the
expression of TIMP-2.
In conclusion, the results of the present study have
demonstrated that miR-20a is upregulated in CRC tissues and that
this upregulation is associated with tumor invasion and lymph node
metastases (P<0.05). Furthermore, miR-20a enhances the EMT of
CRC cells by modulating the expression of TIMP-2, MMP-2 and
MMP-9.
Acknowledgements
This study was supported by Natural Science
Foundation of Shandong Province (grant no. 2013ZRB14250).
References
|
1
|
World Health Organization (WHO), .
GLOBOCAN 2012: Estimated Cancer Incidence, Mortality and Prevalence
Worldwide in 2012. http://globocan.iarc.fr/Pages/fact_sheets_cancer.aspxAccessed.
April 30–2015
|
|
2
|
Gupta GP and Massagué J: Cancer
metastasis: Building a framework. Cell. 127:679–695. 2006.
View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Lamouille S, Subramanyam D, Blelloch R and
Derynck R: Regulation of epithelial-mesenchymal and
mesenchymal-epithelial transitions by microRNAs. Curr Opin Cell
Biol. 25:200–207. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Schetter AJ, Okayama H and Harris CC: The
role of microRNAs in colorectal cancer. Cancer J. 18:244–252. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Melton C, Judson RL and Blelloch R:
Opposing microRNA families regulate self-renewal in mouse embryonic
stem cells. Nature. 463:621–626. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Yoo AS, Sun AX, Li L, et al:
MicroRNA-mediated conversion of human fibroblasts to neurons.
Nature. 476:228–231. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Rottiers V and Näär AM: MicroRNAs in
metabolism and metabolic disorders. Nat Rev Mol Cell Biol.
13:239–250. 2012. View
Article : Google Scholar : PubMed/NCBI
|
|
8
|
Rossi S, Di Narzo AF, Mestdagh P, et al:
microRNAs in colon cancer: A roadmap for discovery. FEBS Lett.
586:3000–3007. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Bartels CL and Tsongalis GJ: MicroRNAs:
Novel biomarkers for human cancer. Ann Biol Clin (Paris).
68:263–272. 2010.(In French). PubMed/NCBI
|
|
10
|
Tsuchida A, Ohno S, Wu W, et al: miR-92 is
a key oncogenic component of the miR-17-92 cluster in colon cancer.
Cancer Sci. 102:2264–2271. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Fan MQ, Huang CB, Gu Y, Xiao Y, Sheng JX
and Zhong L: Decrease expression of microRNA-20a promotes cancer
cell proliferation and predicts poor survival of hepatocellular
carcinoma. J Exp Clin Cancer Res. 32:212013. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Chang CC, Yang YJ, Li YJ, et al:
MicroRNA-17/20a functions to inhibit cell migration and can be used
a prognostic marker in oral squamous cell carcinoma. Oral Oncol.
49:923–931. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Chang Y, Liu C, Yang J, et al: MiR-20a
triggers metastasis of gallbladder carcinoma. J Hepatol.
59:518–527. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Jeong WJ, Cha PH and Choi KY: Strategies
to overcome resistance to epidermal growth factor receptor
monoclonal antibody therapy in metastatic colorectal cancer. World
J Gastroenterol. 20:9862–9871. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Schetter AJ, Leung SY, Sohn JJ, et al:
MicroRNA expression profiles associated with prognosis and
therapeutic outcome in colon adenocarcinoma. JAMA. 299:425–436.
2008. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Malzkorn B, Wolter M, Liesenberg F, et al:
Identification and functional characterization of microRNAs
involved in the malignant progression of gliomas. Brain Pathol.
20:539–550. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Volinia S, Calin GA, Liu CG, et al: A
microRNA expression signature of human solid tumors defines cancer
gene targets. Proc Natl Acad Sci USA. 103:2257–2261. 2006.
View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Yan H, Wu J, Liu W, Zuo Y, Chen S, Zhang
S, Zeng M and Huang W: MicroRNA-20a overexpression inhibited
proliferation and metastasis of pancreatic carcinoma cells. Hum
Gene Ther. 21:1723–1734. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Diosdado B, van de Wiel MA, Terhaar Sive
Droste JS, et al: MiR-17-92 cluster is associated with 13q gain and
c-myc expression during colorectal adenoma to adenocarcinoma
progression. Br J Cancer. 101:707–714. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Kalluri R and Weinberg RA: The basics of
epithelial-mesenchymal transition. J Clin Invest. 119:1420–1428.
2009. View
Article : Google Scholar : PubMed/NCBI
|
|
21
|
Zhu QC, Gao RY, Wu W and Qin HL:
Epithelial-mesenchymal transition and its role in the pathogenesis
of colorectal cancer. Asian Pac J Cancer Prev. 14:2689–2698. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Sun XJ, Zhang P, Li HH, Jiang ZW, Jiang CC
and Liu H: Cisplatin combined with metformin inhibits migration and
invasion of human nasopharyngeal carcinoma cells by regulating
E-cadherin and MMP-9. Asian Pac J Cancer Prev. 15:4019–4023. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Kessenbrock K, Plaks V and Werb Z: Matrix
metalloproteinases: Regulators of the tumor microenvironment. Cell.
141:52–67. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Levi E, Fridman R, Miao HQ, Ma YS, Yayon A
and Vlodavsky I: Matrix metalloproteinase 2 releases active soluble
ectodomain of fibroblast growth factor receptor 1. Proc Natl Acad
Sci USA. 93:7069–7074. 1996. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Chen R, Cui J, Xu C, et al: The
significance of MMP-9 over MMP-2 in HCC invasiveness and recurrence
of hepatocellular carcinoma after curative resection. Ann Surg
Oncol. 19 (Suppl 3):S375–S384. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Liu Y, Bi T, Shen G, et al: Lupeol induces
apoptosis and inhibits invasion in gallbladder carcinoma GBC-SD
cells by suppression of EGFR/MMP-9 signaling pathway.
Cytotechnology. Jul 19–2014.(Epub ahead of print). View Article : Google Scholar
|
|
27
|
Kallakury BV, Karikehalli S, Haholu A,
Sheehan CE, Azumi N and Ross JS: Increased expression of matrix
metalloproteinases 2 and 9 and tissue inhibitors of
metalloproteinases 1 and 2 correlate with poor prognostic variables
in renal cell carcinoma. Clin Cancer Res. 7:3113–3119.
2001.PubMed/NCBI
|
|
28
|
Wang Z, Wang B, Shi Y, et al: Oncogenic
miR-20a and miR-106a enhance the invasiveness of human glioma stem
cells by directly targeting TIMP-2. Oncogene. 34:1407–1419. 2015.
View Article : Google Scholar : PubMed/NCBI
|