Introduction
Diabetes mellitus, as one of the most frequently
diagnosed metabolic disorders, affects ~7% of the population
worldwide (1). The high-glucose
environment in diabetic patients affects the normal function of
major organs, leading to the occurrence of a series of diabetic
complications (2). Diabetes mellitus
may be classified into three major types, including type I, type II
and gestational diabetes. In has been reported that type II
diabetic patients have a 2–6-fold increased risk of death from
cardiovascular complications compared with that of healthy people
(3). The prevention of
cardiovascular disease in patients with type II diabetes is
critical for their survival (4).
A high blood glucose concentration globally affects
the expression of genes, including a large set of long non-coding
RNAs (lncRNAs) (5), which are
critical factors in physiological and pathological processes
(6). Those lncRNAs exhibit
upregulated or downregulated expression during the development of
diabetes to promote or inhibit the progression of diabetes and its
associated complications (7–9). The lncRNA steroid receptor RNA
activator (SRA) has been proven to participate in multiple human
diseases (10,11). Genetic variants of lncRNA SRA are
closely correlated with the risk of breast cancer (10). Furthermore, overexpression of lncRNA
SRA was indicated to promote hepatic steatosis through the
inhibition of adipose triglyceride lipase (11). lncRNA-SRA was recently proved to
promote the proliferation of vascular smooth muscle cells (VSMCs)
(12), which have pivotal roles in
the pathogenesis of cardiovascular diseases (13). The present study was performed to
assess the role of lncRNA-SRA in cardiovascular disease in patients
with type II diabetes mellitus and in VSMCs under high-glucose
conditions in vitro. The results may indicate that
lncRNA-SRA is implicated in the pathogenesis of diabetic
cardiovascular disease by regulating the viability of VSMCs.
Materials and methods
Patient groups and clinical
samples
A total of 108 patients with type II diabetic
cardiovascular diseases were diagnosed by laboratory tests and
stress perfusion cardiovascular magnetic resonance imaging at
Nanning Second People's Hospital (Nanning, China) between January
2012 and January 2013. Among those patients, 34 cases were included
in the present study according to strict inclusion and exclusion
criteria to serve as the diabetic cardiovascular disease group (DCD
group). The inclusion criteria were as follows: i) The patients
were diagnosed for the first time; ii) no treatment prior to
admission; iii) informed consent. The exclusion criteria were i)
other diabetic complications; ii) other severe diseases; iii)
chronic diseases; iv) patient age of >70 years. At the same
time, 178 type II diabetic patients without any obvious
complications were included to serve as the diabetic group (D
group) and 44 age- and gender-matched healthy controls were
included as the control group (C group). The DCD group included 19
males and 15 females, with an age range of 29–69 years and a mean
age of 47.2±5.1 years. The D group included 93 males and 85
females, with an age range of 25–69 years and a mean age of
46.2±6.2 years. The C group included 27 males and 17 females, with
an age range of 26–68 years and a mean age of 46.7±5.5 years. Whole
blood (10 ml) was extracted from each participant on the day of
admission, and was used to isolate the plasma using a routine
method. The plasma was stored in liquid nitrogen until analysis.
The present study was approved by the Ethics Committee of Nanning
Second People's Hospital (Nanning, China), and all participants
provided written informed consent.
Follow-up
The 178 type II diabetic patients without any
obvious complications were followed up every 2 months for 5 years.
The occurrence of cardiovascular disease was recorded. A total of
172 patients completed the follow-up procedure. A total of 6
patients died during the follow-up.
Cell culture and transfection
Human VSMCs were purchased from Lonza Group Ltd.
(cat. no. CC-2583; Basel, Switzerland). VSMCs were cultured in
medium 231 (cat. no. M231500; Gibco; Thermo Fisher Scientific,
Inc., Waltham, MA, USA) supplemented with smooth muscle growth
supplement (cat. no. S00725; Gibco; Thermo Fisher Scientific, Inc.)
and maintained at 37°C in a humidified atmosphere containing 5%
CO2. Full-length lncRNA-SRA cDNA was amplified via
polymerase chain reaction (PCR) using primers with a NheI
restriction site at the 5′end. The cDNA used in PCR reaction was
synthesized using total RNA extracted from plasma samples obtained
from patients, which was mentioned in next section. Full-length
lncRNA-SRA cDNA was cloned into NheI linearized pEGFPC3
vector (Clontech, Palo Alto, CA, USA) to generate the lncRNA-SRA
expression vector. VSMCs (5×105) were transfected with
10 nM lncRNA-SRA expression vector using Lipofectamine
2000® reagent (Invitrogen; Thermo Fisher Scientific,
Inc.), according to the manufacturer's protocol. Untransfected
cells were used as the control group and cells transfected with
empty vector were used the negative control group. For D-glucose
treatment, 0, 10, 30 or 50 mM D-glucose was added to the culture
medium and VSMCs were cultured for 6, 12 or 18 h following
transfection prior to any subsequent experimentation.
Reverse transcription-quantitative PCR
(RT-qPCR)
Total RNA was extracted from VSMCs or plasma samples
using TRIzol® reagent (Invitrogen; Thermo Fisher
Scientific, Inc.), accoridng to the manufacturer's protocol. The
RNA concentration was measured using a NanoDrop™ 2000
spectrophotometer (Thermo Fisher Scientific, Inc.). Total RNA was
reverse transcribed into cDNA using SuperScript III Reverse
Transcriptase (Thermo Fisher Scientific, Inc.) using the following
reaction conditions: 54°C for 30 min and 75°C for 5 min. qPCR was
subsequently performed using the PowerUp SYBR™ Green Master mix
(cat. no. A25743; Applied Biosystems; Thermo Fisher Scientific,
Inc.). The following primer pairs were used for qPCR: lncRNA-SRA
forward, 5′-GCTAGGGCACTAGGTTGTCGC-3′ and reverse,
5′-CGCCTGGCACTGCTGCAGGAAC-3′; β-actin forward,
5′-GACCTCTATGCCAACACAGT-3′ and reverse, 5′-AGTACTTGCGCTCAGGAGGA-3′
and 1·S rRNA forward, 5′-GACCTCTATGCCAACACAGT-3′ and reverse,
5′-AGTACTTGCGCTCAGGAGGA-3′. The following thermocycling conditions
were used for qPCR: Initial denaturation at 95°C for 50 sec; 40
cycles of 95°C for 15 sec and 57°C for 40 sec. lncRNA-SRA was
quantified using the 2−ΔΔCq method (14) and normalized to β-actin and 18S
rRNA.
MTT assay
After transfection, the expression of lncRNA-SRA in
VSMCs was detected by RT-qPCR. Subsequent experiments were
performed only in case of the overexpression rate of lncRNA-SRA
reaching 200%. VSMCs were collected and cell suspensions were
prepared with a final density of 5×104 cells per ml. A
total of 0.1 ml cell suspension was added to each well of a 96-well
plate, followed by the addition of D-glucose at a final
concentration of 10, 30 or 50 mM. The plate was incubated at 37°C
with 5% CO2 for 6 h. Following incubation, 10 µl MTT was
added to each well and cells were cultured for a further 4 h at
37°C. Following MTT incubation, DMSO (10 µl/well) was added to
dissolve the purple formazan crystals. The optical density (OD) was
measured at 570 nm using a microplate reader (BioTek™ 800™ TS;
BioTek Instruments, Inc., Winooski, VT, USA). The OD value of
control cells treated with 0 mM D-glucose was set as 100% and the
viability of the cells in the treated groups was expressed as the
percentage of their OD value vs. that in the control group.
Statistical analysis
GraphPad Prism 6 (GraphPad Software Inc., La Jolla,
CA, USA) was used for all statistical analyses. Gene expression and
cell viability data were reported as the mean ± standard deviation
and compared by one-way analysis of variance followed by Fisher's
least-significant difference test. The incidence of diabetic
cardiovascular disease was compared using the Student's t-test.
Correlation analyses were performed by determining Pearson's
correlation coefficient. Receiver operating characteristic (ROC)
curves were drawn and the area under the curve (AUC) was determined
to assess the ability of the lncRNA to distinguish between the
different groups. P<0.05 was considered to indicate a
statistically significant difference.
Results
LncRNA-SRA is significantly
downregulated in patients with type II diabetic cardiovascular
disease
The expression level of lncRNA-SRA was detected by
RT-qPCR in the plasma of patients in the DCD, D and C groups. The
clinicopathological features of these patients are summarized in
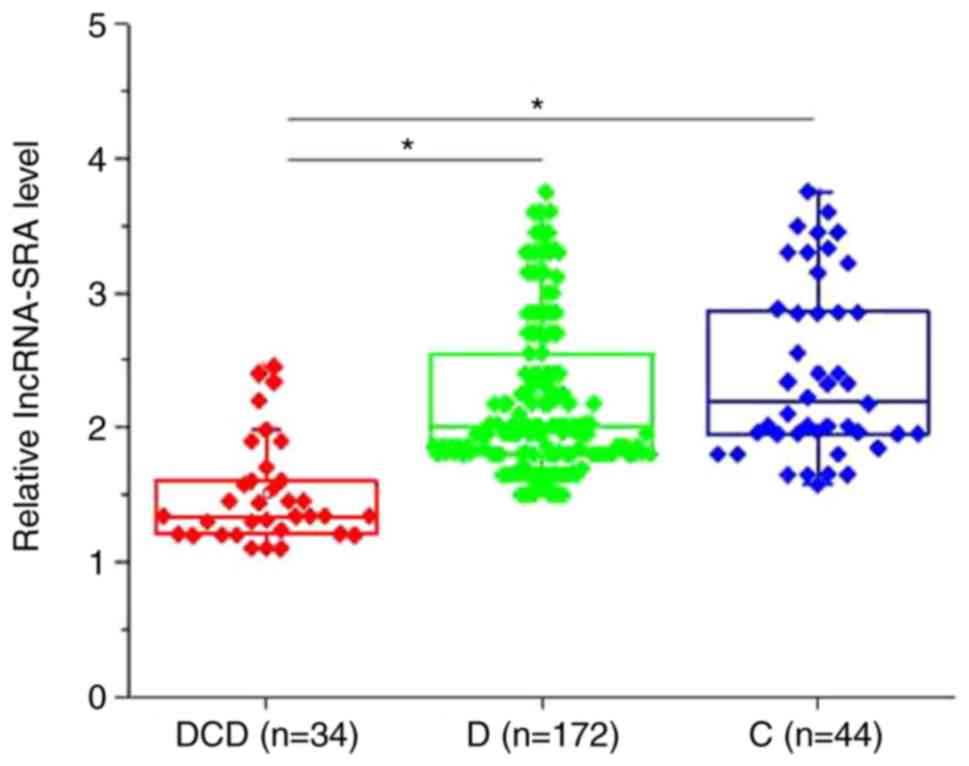
Table I. As presented in Fig. 1, the plasma levels of lncRNA-SRA were
significantly decreased in patients with type II diabetic
cardiovascular disease compared with type II diabetic patients
without any obvious complications and healthy controls (P<0.05).
No significant differences in the plasma levels of lncRNA-SRA were
obtained between type II diabetic patients without any obvious
complications and healthy controls (P>0.05). Of note, the plasma
levels of lncRNA-SRA in the DCD group were significantly and
inversely correlated with the systolic blood pressure (r=−0.82,
P<0.0001), the levels of low-density lipoprotein (r=−0.79,
P<0.0001) and the levels of triglycerides (r=−0.75,
P<0.0001), and were significantly and positively correlated with
the levels of high-density lipoprotein (r=0.71, P<0.0001) (data
not shown).
 | Table I.Clinicopathological parameters of
patients within the 3 groups. |
Table I.
Clinicopathological parameters of
patients within the 3 groups.
| Clinical
parameter | Group C (n=44) | Group D (n=178) | Group DCD (n=34) |
|---|
| Age (years) |
46.7±5.5 | 46.2±6.2 | 47.2±5.1 |
| Sex (n, %) |
|
|
|
| Male | 27 (61.4) | 93 (52.2) | 19 (55.9) |
|
Female | 17 (38.6) | 85 (47.8) | 15 (44.1) |
| BMI
(kg/m2) | 21.3±1.9 | 24.4±2.6 | 24.7±2.3 |
| Systolic blood
pressure (mmHg) | 116.7±3.4 | 117.1±3.7 | 135.5±7.2 |
| Diastolic blood
pressure (mmHg) | 76.7±5.9 | 77.2±6.7 | 79.2±9.9 |
| Oral glucose
tolerance test blood glucose level (mmol/l) | 6.2±1.8 | 14.4±2.7 | 14.9±2.6 |
| LDL (mg/dl) | 77.5±22.1 | 84.5±19.7 | 88.7±22.2 |
| HDL (mg/dl) | 57.4±9.2 | 49.2±8.8 | 44.5±11.8 |
| Triglycerides
(mg/dl) | 137.8±89.2 | 150.2±78.4 | 177.5±100.8 |
Downregulation of lncRNA-SRA
distinguishes patients with type II diabetic cardiovascular disease
from healthy controls and type II diabetic patients without any
obvious complications
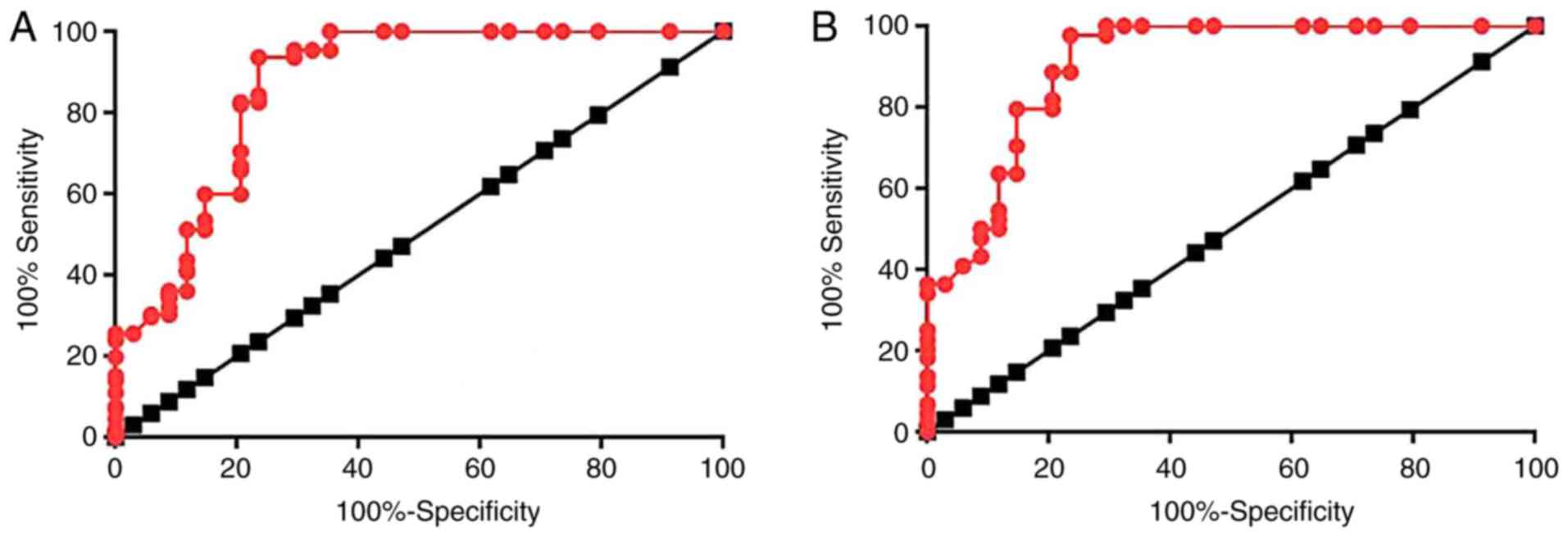
ROC curve analysis was performed to evaluate the
diagnostic value of lncRNA-SRA for diabetic cardiovascular disease.
With the healthy controls as a reference, the area under the curve
(AUC) was 0.9041, with a standard error of 0.03625 and 95%
confidence interval of 0.8830–0.9752 (Fig. 2A). With the type II diabetic patients
without any obvious complications as a reference, the AUC was
0.8679, with a standard error of 0.04222 and a 95% confidence
interval of 0.7851–0.9507 (Fig.
2B).
Low plasma levels of lncRNA-SRA are
associated with a high incidence of cardiovascular disease in type
II diabetic patients
A total of 172 type II diabetic patients without any
obvious complications on the day of admission completed the 5-year
follow-up. According to the median relative plasma level of
lncRNA-SRA (1.44), these patients were divided into a high (n=86)
and a low (n=86) expression group. During the follow-up,
cardiovascular disease occurred in 41 cases, including 29 cases in
the low expression group and 12 cases in the high expression group.
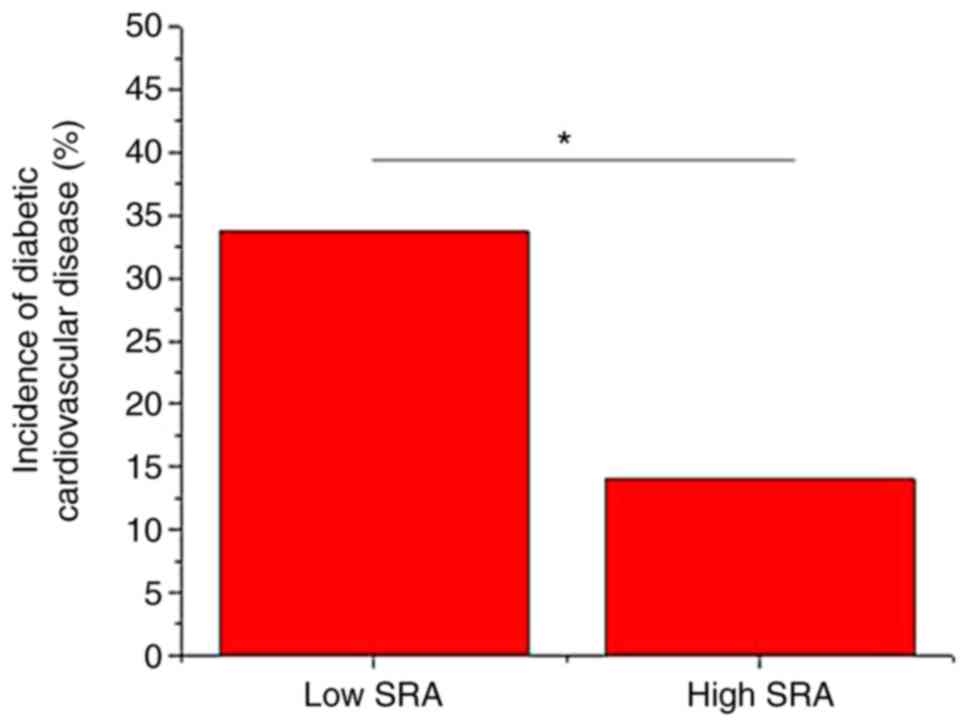
As presented in Fig. 3, the
incidence of cardiovascular disease was significantly higher in the
low expression group compared with that in the high expression
group (P<0.05).
A high-glucose environment has no
significant effect on lncRNA-SRA expression in VSMCs
VSMCs were cultured with 0 mM (control), or with 10,
30 or 50 mM D-glucose in culture medium for 6, 12 or 18 h.
Subsequently, the expression of lncRNA-SRA in the VSMCs was
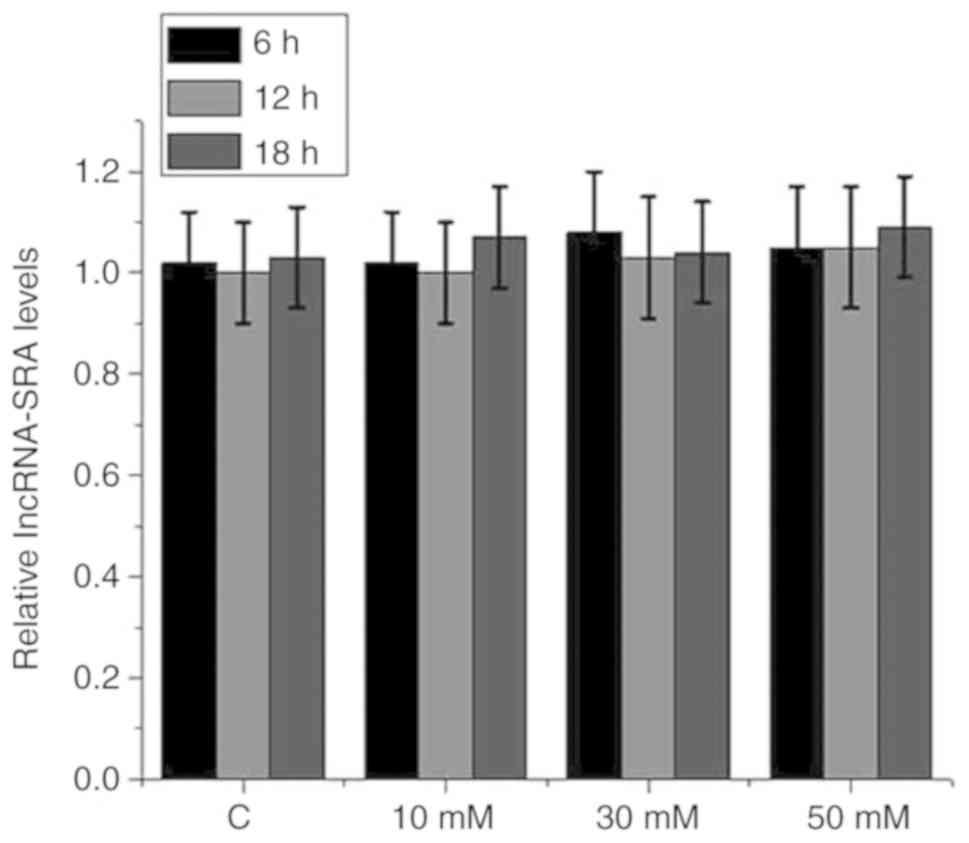
detected by RT-qPCR. As presented in Fig. 4, treatment with high glucose at
different concentrations for different durations did not
significantly affect lncRNA-SRA expression in VSMCs
(P>0.05).
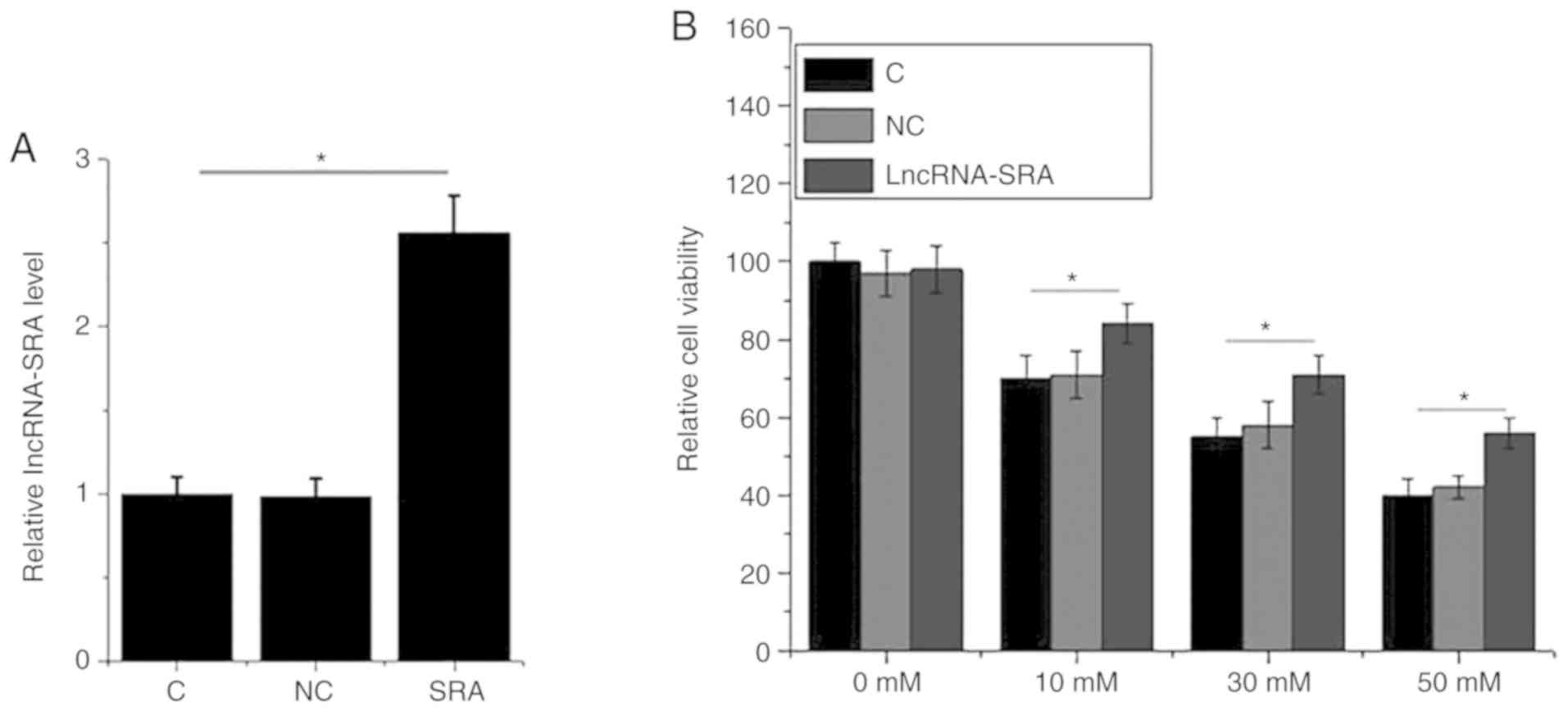
lncRNA-SRA overexpression improves the
viability of VSMCs under high-glucose treatment
An MTT assay was performed to assess the impact of
lncRNA-SRA on the viability of VSMCs under high-glucose treatment
for 12 h. lncRNA-SRA overexpression was achieved by plasmid
transfection (Fig. 5A). High glucose
(10, 30 or 50 mM D-glucose) significantly reduced the viability of
VSMCs compared with the control group (P<0.05; Fig. 5A). In addition, lncRNA-SRA
overexpression significantly increased the viability of VSMCs
compared with the untransfected control cells and negative
control-transfected cells in the presence of D-glucose, but not in
the absence of glucose (P<0.05; Fig.
5B).
Discussion
lncRNA-SRA is involved in regulating the
proliferation of VSMCs (13),
indicating its potential involvement in the pathogenesis of
cardiovascular diseases (15). The
key result of the present study is that lncRNA-SRA is specifically
downregulated in diabetic patients with cardiovascular disease, and
the downregulation of lncRNA-SRA may serve as a potential
diagnostic marker for this disease.
The development of diabetes is associated with the
occurrence of a series of complications (16,17).
Previous studies suggested that lncRNAs are key factors in the
development of diabetic complications. Sun et al (16) reported that lncRNA Erb-b2 receptor
tyrosine kinase 4-intron region (Erbb4-IR) was upregulated in a
mouse model of diabetic kidney injury, and that the upregulation of
lncRNA Erbb4-IR is closely associated with disease progression. A
study by Zhuo et al (17)
revealed that lncRNA H19 is downregulated in diabetic
cardiomyopathy and overexpression of lncRNA H19 inhibits autophagy
by epigenetically silencing DIRAS family GTPase 3. However, most
lncRNAs involved in diabetic complications are regulated by high
glucose and have no diagnostic value to distinguish patients with a
specific diabetic complication from diabetic patients without this
complication (16,17). lncRNA-SRA has pivotal roles in
different types of human disease. Genetic variants of lncRNA SRA
have been proved to be closely associated with the risk of breast
cancer (10). lncRNA-SRA also
inhibits the expression of adipose triglyceride lipase, thereby
promoting hepatic steatosis (11).
However, the involvement of lncRNA-SRA in diabetic cardiovascular
disease has remained elusive. It has been reported that lncRNA-SRA
promotes the proliferation of VSMCs, which have pivotal roles in
the pathogenesis of cardiovascular diseases (12,13). The
present study demonstrated that the plasma levels of lncRNA-SRA may
serve as a potential diagnostic biomarker for type II diabetic
cardiovascular disease. However, the expression of lncRNA-SRA has
been reported to be affected by multiple diseases. Therefore, the
diagnostic specificity should be further investigated.
A high-glucose environment affects the expression of
certain lncRNAs, and altered expression of those lncRNAs
participates in diabetes-associated pathological processes
(18,19). Normal blood glucose levels are ~5 mM
(20). In the present study, a
high-glucose environment was created by the addition of D-glucose
(10, 30 or 50 mM) to the cell culture medium. However, high-glucose
treatment had no significant effect on lncRNA-SRA expression in
VSMCs. Therefore, lncRNA-SRA may not participate in the initiation
of cardiovascular disease in diabetic patients, while indirect
effects cannot be excluded. lncRNA-SRA expression may be altered by
the presence of cardiovascular disorders during the development of
diabetes. Of note, ectopic overexpression of lncRNA-SRA increased
the viability of VSMCs in a high-glucose environment. Therefore,
lncRNA-SRA overexpression may serve as a potential therapeutic
strategy for the treatment of diabetic cardiovascular disease.
Various factors contribute to the development of
diabetic complications (21,22). The present 5-year follow-up study
revealed that low plasma levels of lncRNA-SRA were associated with
a significantly increased incidence of cardiovascular disease in
patients with type II diabetes. Therefore, detection of plasma
lncRNA-SRA may provide guidance for the prevention of diabetic
cardiovascular disease. Of note, the present study has certain
limitations. For instance, the molecular mechanisms of the role of
lncRNA-SRA in diabetic cardiovascular disease were not elucidated.
In addition, no in vivo experimental validation was
performed and therefore this will be examined in future studies.
Furthermore, future studies may be required to examine the
expression on lncRNA-SRA in patients with cardiovascular disease
without diabetes.
In conclusion, the present study suggested that
downregulation of lncRNA-SRA is involved in the pathogenesis of
diabetic cardiovascular disease. Plasma lncRNA-SRA may serve as a
potential diagnostic biomarker for this disease.
Acknowledgements
Not applicable.
Funding
Not applicable.
Availability of data and materials
The datasets used and/or analyzed in the present
study are available from the corresponding author on reasonable
request.
Authors' contributions
SR and YQ designed the experiments. SR, YZ and BL
performed experiments. KB, LW, YL and YYL prepared the materials
and analyzed the data. YQ interpreted the data and drafted the
manuscript. All authors read and approved the final version of the
manuscript.
Ethics approval and consent to
participate
This study passed the review of the Ethics Committee
of Nanning Second People's Hospital (Nanning, China) and all
participants provided written informed consent.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Wild S, Roglic G, Green A, et al: Global
prevalence of diabetes: estimates for the year 2000 and projections
for 2030. Diabetes care. 27:1047–1053. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
van Dieren S, Beulens JW, van der Schouw
YT, Grobbee DE and Neal B: The global burden of diabetes and its
complications: An emerging pandemic. Eur J Cardiovasc Prev Rehabil.
17 (Suppl 1):S3–S8. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Gæde P, Vedel P, Larsen N, Jensen GV,
Parving HH and Pedersen O: Multifactorial intervention and
cardiovascular disease in patients with type 2 diabetes. N Engl J
Med. 348:383–393. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Colhoun HM, Betteridge DJ, Durrington PN,
Hitman GA, Neil HA, Livingstone SJ, Thomason MJ, Mackness MI,
Charlton-Menys V and Fuller JH; CARDS investigators, : Primary
prevention of cardiovascular disease with atorvastatin in type 2
diabetes in the Collaborative Atorvastatin Diabetes Study (CARDS):
Multicentre randomised placebo-controlled trial. Lancet.
364:685–696. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Ruan Y, Lin N, Ma Q, Chen R, Zhang Z, Wen
W, Chen H and Sun J: Circulating LncRNAs analysis in patients with
type 2 diabetes reveals novel genes influencing glucose metabolism
and Islet β-cell function. Cell Physiol Biochem. 46:335–350. 2018.
View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Esteller M: Non-coding RNAs in human
disease. Nat Rev Genet. 12:861–869. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Liu JY, Yao J, Li XM, Song YC, Wang XQ, Li
YJ, Yan B and Jiang Q: Pathogenic role of lncRNA-MALAT1 in
endothelial cell dysfunction in diabetes mellitus. Cell Death Dis.
5:e15062014. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Li X, Wang H, Yao B, Xu W, Chen J and Zhou
X: lncRNA H19/miR-675 axis regulates cardiomyocyte apoptosis by
targeting VDAC1 in diabetic cardiomyopathy. Sci Rep. 6:363402016.
View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Wang S, Xu H, Zou L, Xie J, Wu H, Wu B, Yi
Z, Lv Q, Zhang X, Ying M, et al: LncRNA uc.48+ is involved in
diabetic neuropathic pain mediated by the P2X3 receptor in the
dorsal root ganglia. Purinergic Signal. 12:139–148. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Yan R, Wang K, Peng R, Wang S, Cao J, Wang
P and Song C: Genetic variants in lncRNA SRA and risk of breast
cancer. Oncotarget. 7:22486–22496. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Chen G, Yu D, Nian X, Liu J, Koenig RJ, Xu
B and Sheng L: LncRNA SRA promotes hepatic steatosis through
repressing the expression of adipose triglyceride lipase (ATGL).
Sci Rep. 6:355312016. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Long J, Zhu N, Zhang CJ, Liu C, Tuo QH,
Liao DF and Qin L: LncRNA-SRA promotes vascular smooth muscle cells
proliferation via MEK/ERK/CREB signaling pathway. Atheroscler
Suppl. 32:113–121. 2018. View Article : Google Scholar
|
|
13
|
Rivard A and Andrés V: Vascular smooth
muscle cell proliferation in the pathogenesis of atherosclerotic
cardiovascular diseases. Histol Histopathol. 15:557–571.
2000.PubMed/NCBI
|
|
14
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Ahlqvist E, Van Zuydam NR, Groop LC and
McCarthy MI: The genetics of diabetic complications. Nat Rev
Nephrol. 11:277–287. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Sun SF, Tang PMK, Feng M, Xiao J, Huang
XR, Li P, Ma RCW and Lan HY: Novel lncRNA Erbb4-IR promotes
diabetic kidney injury in db/db mice by targeting miR-29b.
Diabetes. 67:731–744. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Zhuo C, Jiang R, Lin X and Shao M: LncRNA
H19 inhibits autophagy by epigenetically silencing of DIRAS3 in
diabetic cardiomyopathy. Oncotarget. 8:1429–1437. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Cao B, Liu N and Wang W: High glucose
prevents osteogenic differentiation of mesenchymal stem cells via
lncRNA AK028326/CXCL13 pathway. Biomed Pharmacother. 84:544–551.
2016. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Gong Y, Zhu Y, Zhu B, Si X, Heng D, Tang
Y, Sun X and Lin L: LncRNA MALAT1 is up-regulated in diabetic
gastroparesis and involved in high-glucose-induced cellular
processes in human gastric smooth muscle cells. Biochem Biophys Res
Commun. 496:401–406. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Güemes M, Rahman SA and Hussain K: What is
a normal blood glucose? Arch Dis Child. 101:569–574. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Brownlee M and Hirsch IB: Glycemic
variability: A hemoglobin A1c-independent risk factor for diabetic
complications. JAMA. 295:1707–1708. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Liebl A, Mata M and Eschwege E: Evaluation
of risk factors for development of complications in type II
diabetes in Europe. Diabetologia. 45 (Suppl 1):S23–S28. 2002.
View Article : Google Scholar
|