Introduction
Hand, foot and mouth disease (HFMD) is a common
infectious disease in children. The clinical symptoms are mainly
lesions of varying severity in the palms, feet and mouth, which may
be accompanied by fever, sore throat and diarrhea. Enterovirus 71
(EV71) and Coxsackie virus A16 (CA16) are the most common pathogens
in pediatric patients with HFMD (1,2).
Infection with EV71 frequently results in severe central nervous
system complications, including acute encephalitis, paralytic
paralysis, encephalomyelitis and other associated neurological
complications that may be fatal, particularly in children aged
<5 years. Therefore, HFMD currently poses a serious public
health threat (3-5).
EV71 belongs to the enterovirus family of
picornaviruses and is composed of a single strand of RNA (~7.4 KB).
The VP1 gene encodes the peripheral immunogenic protein, which is
the major pathogenic factor of the virus. It has an important role
in the adsorption and desorption of viral infection (4,6).
Furthermore, this protein includes the major neutralization epitope
that may be used for virus identification and evolutionary analysis
(7). In the present study, the VP1
gene of EV71, the most important pathogen causing HFMD, was
analyzed with the aim of enriching the genetic database of HFMD in
Yunnan, China. The results of the present study may provide novel
clues to elucidate the molecular genetic pathogenesis of HFMD
epidemics and provide a scientific basis for the clinical
assessment of HFMD risk and prognosis, early clinical intervention
and prevention, as well as the development of vaccines or
therapeutic targets.
Materials and methods
Subjects and specimen collection
A total of 590 patients with suspected HFMD were
recruited at Yan'an Hospital Affiliated to Kunming Medical
University (Kunming, China) between January 2015 and December 2016.
The patients included 355 males and 235 females aged 0-36 years. A
total of 470 patients were ≥5 years old, 118 patients were between
5-18 years old and 2 patients were ≤18 years old. The patients were
primarily treated at the dermatology, pediatric and other clinical
departments with the first symptoms being lesions in the palms,
feet and oral cavity, accompanied by fever and diarrhea. All
patients provided oral informed consent. The study was approved by
Kunming Yan'an Hospital/Kunming Medical University Affiliated
Yan'an Hospital Medical Ethics Committee (document no.
2017-035-01). According to the Hand-foot-mouth Disease Prevention
and Control Guidelines (2008 edition) issued by the Ministry of
Health of the P.R. China (8), fecal
specimens were collected within 7 days of disease onset.
Fluorescence PCR detection of
EV71
EV71 nucleic acid was extracted from the clinical
stool samples of suspected cases using a EV71 nucleic acid
detection kit (Jiangsu Mole Bioscience Co., Ltd.) based on the
centrifuge column method, according to the manufacturer's protocol.
Fluorescence PCR was used to detect the EV71 pathogen using an ABI
StepOne Plus system (Applied Biosystems; Thermo Fisher Scientific,
Inc.) and the parameters presented in Table I.
 | Table ICycling parameters of fluorescence PCR
detection of EV71. |
Table I
Cycling parameters of fluorescence PCR
detection of EV71.
| Step | Temperature (˚C) | Time (min:sec) | Cycles (n) |
|---|
| 1 | 45 | 15:00 | 1 |
| 2 | 95 | 02:00 | 1 |
| 3 | 94 | 00:10 | 40 |
| | 60 | 00:40 | |
EV71 viral RNA extraction from
EV71-positive specimens
The EV71 nucleic acid detection kit (Jiangsu Mole
Bioscience Co., Ltd.) was used to extract viral RNA from 50 samples
tested positive for EV71, according to the manufacturer's
protocol.
Reverse transcription-PCR
(RT-PCR)
The extracted EV71 viral RNA was used as the
template for RT-PCR. A reverse transcription-PCR kit (Takara
Biotechnology Co., Ltd.) was used for RT-PCR according to the
manufacturer's instructions. The components of the reaction system
are presented in Table II. The
reaction mixture was placed in the PCR instrument, reacted at 65˚C
for 5 min and placed on ice. The components used for the next
reaction are presented in Table
III. The thermocycling conditions were as follows: 37˚C for 15
min, 50˚C for 15 min and 65˚C for 10 min. The cDNA obtained was
used as a template for the PCR amplification of the VP1 gene.
 | Table IIFirst step of reverse transcription of
EV71. |
Table II
First step of reverse transcription of
EV71.
| Component | Volume (µl) |
|---|
| Oligo dT primer (2.5
µM) | 1 |
| Random 6mers (20
µM) | 1 |
| Template RNA | 2 |
| dNTP mixture (10 mM
each) | 1 |
| RNase-free
H2O | 5 |
| Total | 10 |
 | Table IIISecond step of reverse transcription
of EV71. |
Table III
Second step of reverse transcription
of EV71.
| Component | Volume (µl) |
|---|
| Fist-step reaction
mixture | 10.0 |
| PrimeScript reverse
transcriptase (200 U/µl) | 1.0 |
| 5x PrimeScript
buffer | 4.0 |
| RNase inhibitor (40
U/µl) | 0.5 |
| RNase-free
H2O | 4.5 |
| Total | 20.0 |
PCR amplification of the VP1 gene
VP1 gene primers were designed using Primer Premier
software (version 5.0; www.PremierBiosoft.com; Table IV). Primers were synthesized by
Beijing Biomed Technology Development Co., Ltd (http://www.biomed168.com/). The product cDNA obtained
from the above RT-PCR reactions serves as a template for the PCR
amplification of VP1 gene. A 50-µl PCR mix consisting of 2 µl
complementary (c)DNA template, 1 µl 30 pmol forward and reverse
primers, 5 µl 10XeasyTaq DNA buffer, 0.5 µl 5 U/µl EasyTaq DNA
polymerase, 5 µl 2.5 mmol/l deoxyribose nucleoside triphosphate
mixture and 35.5 µl RNA-free pure water. The PCR cycling parameters
were as follows: 1 cycle at 94˚C for 3 min, 35 cycles at 94˚C for 3
min, 56˚C for 30 sec and 72˚C for 1 min, and 1 cycle at 72˚C and 3
min. The products were subjected to 1% agarose gel electrophoresis,
stained with Gelview nucleic acid dye (Beijing Bioteke
Biotechnology Co., Ltd) and observed using a 6000 gel imaging
system (Bio-Rad Laboratories, Inc). PCR products were compared with
a DL5000 DNA marker (Takara Biotechnology Co., Ltd.) and the
presence of target bands at the required length was considered to
indicate a positive result.
 | Table IVPCR primer sequences and product
fragment size. |
Table IV
PCR primer sequences and product
fragment size.
| Site | PCR product (bp) | Primer sequence |
|---|
| Enterovirus 71
VP1 | 1,200 | P1:
5'-GGCTGCAATCGTCTGTTACC-3' |
| | | P2:
5'-AAGTCCCGAGAGCTGTCTTCAAATTATGGGAGAAATCGTC-3' |
VP1 gene sequencing and analysis
After purification, VP1 gene amplification products
were sent to Beijing Biomed Technology Development Co., Ltd
(http://www.biomed168.com/) for sequence
determination using an ABI3730XL automatic DNA sequence analyzer
(Applied Biosystems; Thermo Fisher Scientific, Inc.). Sequence
comparison was performed using the Basic Local Alignment Search
Tool (National Center for Biotechnology Information).
Gene sequence homology comparison and
phylogenetic tree analysis
Gene sequence comparison and homology analysis were
performed with DNAMAN software (version 5.1.0.0; Lynnon Biosoft),
according to the software instructions. MEGA software (version 4.0;
https://www.megasoftware.net/) was used
to construct phylogenetic trees with neighbor joining and a Kimura
2-parameter model. The bootstrap value was 1,000. All of the
reference strain sequences were derived from GeneBank (https://www.ncbi.nlm.nih.gov/pubmed).
Homologous modeling analysis
The protein structure database (PDB; http://www.rcsb.org/) was searched to select
appropriate protein structure templates (PDB ID: 3J23). Pymol
software (version 2.3; https://pymol.org/) was used to simulate the protein
spatial structure of the VP1-coding amino acid sequence
variation.
Results
Clinical features
Among the 590 patients with suspected HFMD admitted
to Yan'an Hospital Affiliated to Kunming Medical University
(Kunming, China) between January 2015 and December 2016, the
positive rate of an enterovirus nucleic acid etiology was 60.8%
(359/590). The pathogen composition of 359 enterovirus nucleic
acid-positive cases is shown in Table
V. The age and sex composition of 50 EV71-positive cases is
shown in Table VI.
 | Table VPathogen composition of 359 EV nucleic
acid-positive cases. |
Table V
Pathogen composition of 359 EV nucleic
acid-positive cases.
| Year | EV nucleic
acid-positive cases | EV71 | CA16 | Other EV types |
|---|
| 2015 | 203 | 28 (13.8) | 32 (15.8) | 143 (70.4) |
| 2016 | 156 | 22 (14.1) | 26 (16.7) | 108 (69.2) |
| Total | 359 | 50 (13.9) | 58 (16.2) | 251 (69.9) |
 | Table VIAge and sex composition of 50
EV71-positive cases. |
Table VI
Age and sex composition of 50
EV71-positive cases.
| Age | EV71-positive
cases | Male | Female |
|---|
| ≤5 | 35 | 23 (65.7) | 12 (34.3) |
| 5-18 | 13 | 9 (69.2) | 4 (30.8) |
| ≥18 | 2 | 0 (0) | 2 (100.0) |
| Total | 50 | 32 (64.0) | 18 (36.0) |
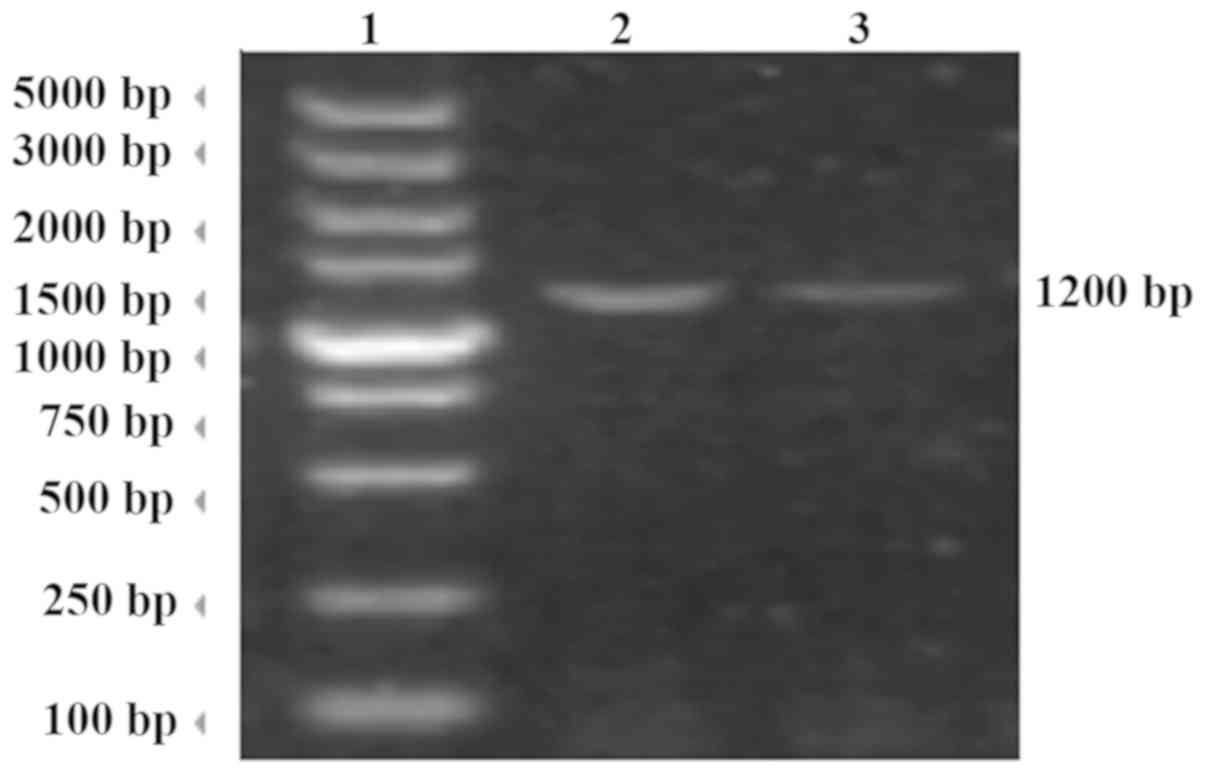
Electrophoresis of VP1 gene
amplicon
The viral RNA was extracted from EV71-positive
samples and the VP1 gene was amplified by RT-PCR. The products were
identified by agarose gel electrophoresis and the length of the
target gene was 1,200 bp (Fig.
1).
Sequencing analysis of VP1
gene-positive amplification products
The VP1 gene amplification products of the 50
EV71-positive samples were sent to Beijing Biomed Technology
Development Co., Ltd (http://www.biomed168.com/) for gene sequencing. 50
sequencing results were compared with the gene sequences of the
EV71 reference strain (KU936121.1) in GenBank, and 50 sequence
alignment diagrams in GenBank show homology values between
97.96-99.43%. An example of sequence alignment diagrams is shown
(Fig. 2).
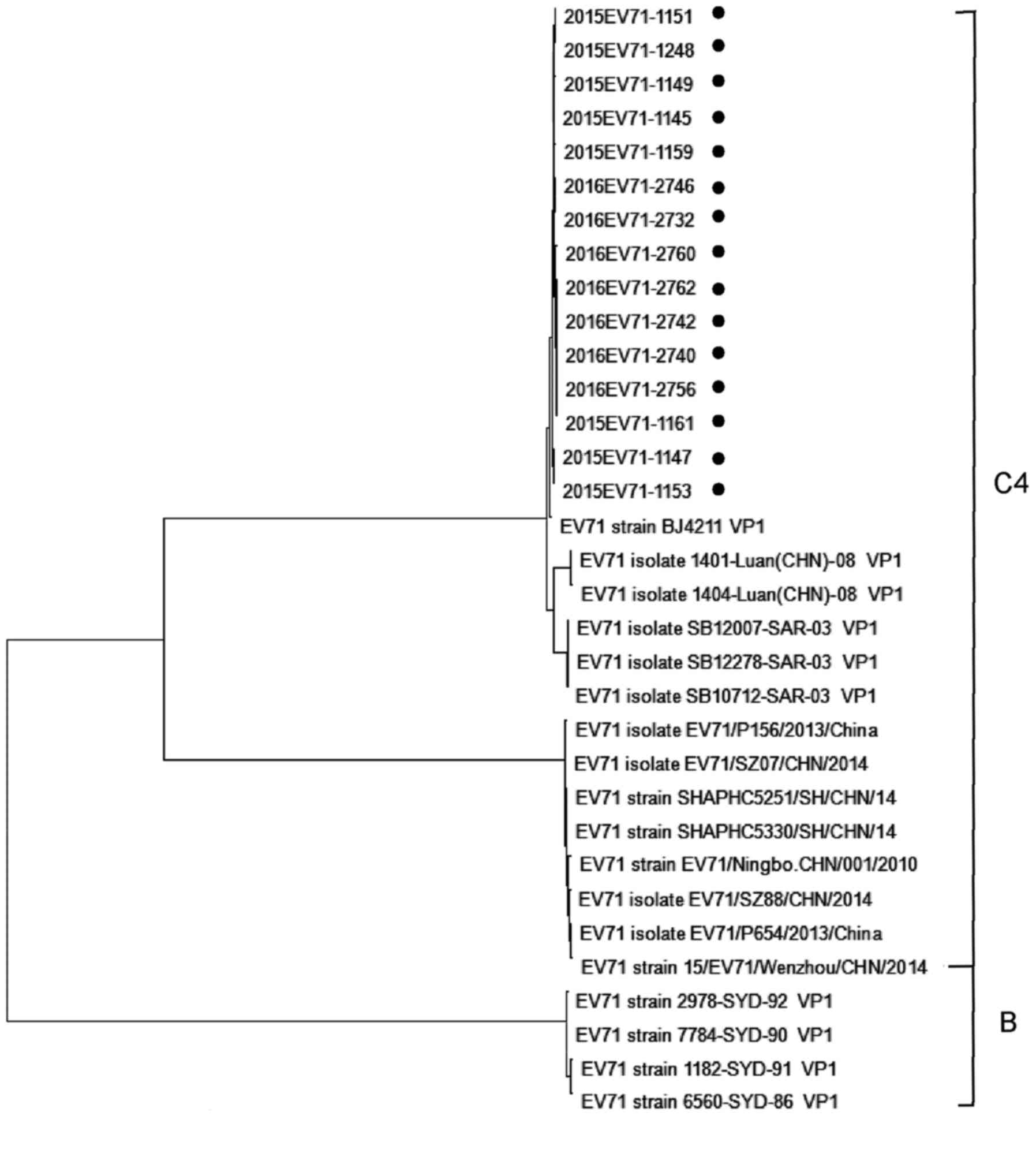
Phylogenetic tree analysis of the EV71
VP1 gene
After sequence alignment, a total of 15 strains with
amino acid sequence variation and homology <99.0% were selected
from the 50 EV71 strains. The phylogenetic tree was constructed
using the adjacency method and Kimura 2-parameter model with MEGA
software (version 4.0; Fig. 3).
Three-dimensional conformation of EV71
virus VP1
EV71 VP1 contains 297 amino acids and is 32 kD in
size. The crystal structure, according to the VP1 type with ‘jelly
roll’ β barrel structure, includes eight β folds arranged in two β
patches, each layer containing four β folds (9). VP1 is the major protective
neutralization site region of the EV71 virus. Compared with other
EV71-associated reference sequences that have been registered in
GenBank, an amino acid substitution occurred in the 144th amino
acid (histidine → tyrosine) encoded by the VP1 gene of the EV71
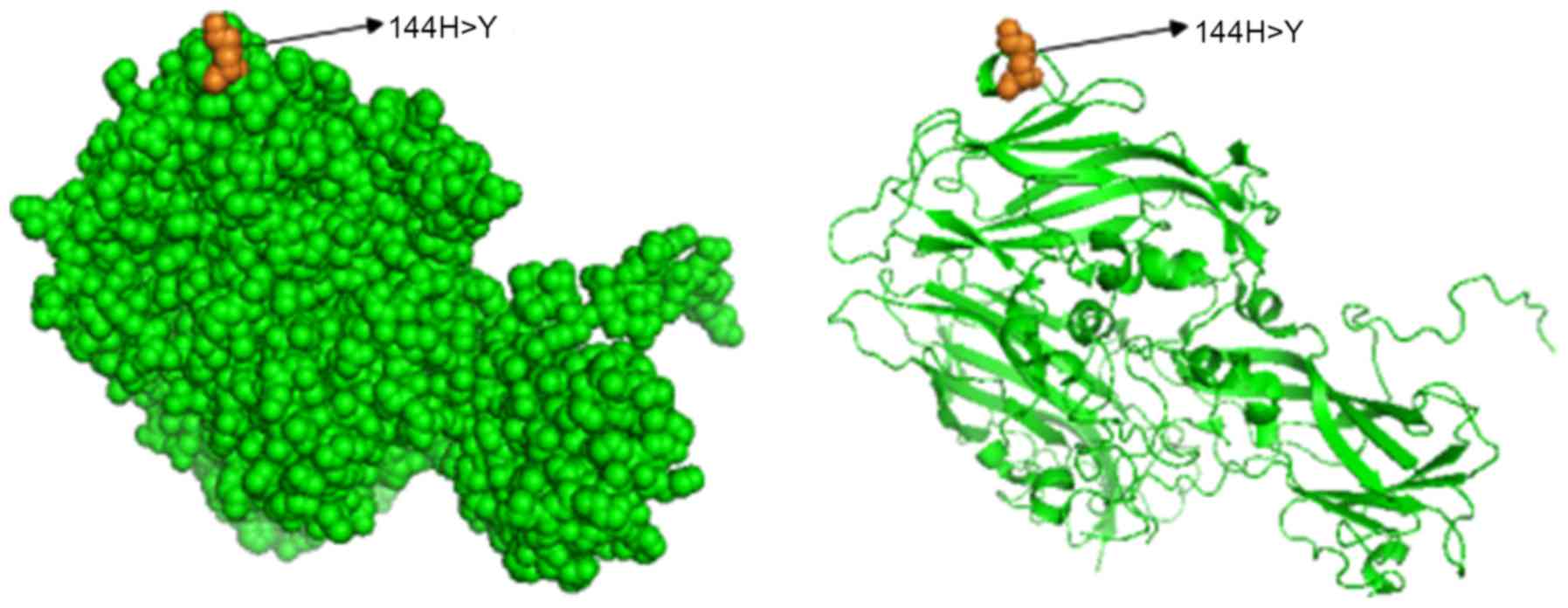
virus strain in the present study. As indicated in the
three-dimensional conformation presented in Fig. 4, the amino acid mutation site H144Y
was located on the surface of the VP1 protein (PDB ID: 3J23
template used for modeling).
Discussion
HFMD is a common infectious disease in children
resulting in skin rashes and lesions on the palms, soles of the
feet, oral mucosa, tongue and buttocks, and may be accompanied by
fever and diarrhea. The disease may be caused by various pathogens,
including EV, eccovirus and Coxsackievirus, the most common of
which are EV71 and CA16. However, other subtypes may also cause
HFMD (10), including CA4, CA6,
CA10, CA12, CB3, CB5 and eccovirus types 4, 19 and 30(11). While HFMD frequently occurs in
preschool children, particularly those aged <5 years, adults may
also be infected and the prevalence of HFMD in adults has been
exhibiting an increasing trend (12,13).
HFMD is prevalent in Southeast Asia, including Taiwan and mainland
China, Singapore, Malaysia and Japan. HFMD was first reported in
mainland China in 1981, in Shanghai, and associated cases have been
since reported in other regions (14,15).
In the present study, a total of 590 fecal samples
were obtained from patients with suspected HFMD and a total of 50
EV71-positive patients were identified by fluorescence PCR, with a
detection rate of 8.47%. Among the 50 EV71-positive patients, 32
were male and 18 were female. Furthermore, 2 female patients were
adults, which is consistent with the reported low incidence of HFMD
in adults and the susceptibility of adult females to viral
infection (16). High psychological
pressure, physical fatigue, malnutrition, low immunity and close
contact with patients with HFMD are important risk factors for HFMD
in adults (17). Adults infected
with HFMD are prone to cluster infection and infecting family
members. Furthermore, adults may be asymptomatic but still highly
infectious. In recent years, there have been reports on severe HFMD
and serious complications in adults (18,19).
Therefore, it is important to prevent and control the prevalence of
HFMD in adults.
EV71 is divided into three genotypes, A, B and C, of
which B and C may be subdivided into B1-B5 subtypes and C1-C5
subtypes. Previous studies have indicated that the D, E and F
subtypes of EV71 have been isolated in India, Hong Kong and Africa
(20,21). The VP1 gene sequences of 50 EV71
strains detected in the present study were compared with the EV71
reference strains in GenBank for sequence comparison and homology
analysis, and it was confirmed that all of the 50 strains were of
the EV71 C4 subtype, which was the same as the EV71 subtype
isolated by Fu et al (22) in
Kunming, Yunnan province in 2011.
In the present study, through sequence comparison
analysis, a total of 15 strains with different amino acid sequences
were screened out from the 50 EV71 strains, including
EV71-VP1-1145-1147, -1149, -1151, -1153, -1159, -1161 and -1248 in
2015 and EV71-VP1-2732, -2740, -2742, -2746, -2756, -2760 and -2762
in 2016. Phylogenetic trees were constructed with MEGA software
(version 4.0) and the adjacency method and a Kimura 2-parameter
model were used to analyze the genetic origin, variation and
association of the virus with other strains in China and in other
countries. The viral strains detected in the present study were
compared with those in Beijing (EV71 strain BJ4211 VP1), Hefei
[EV71 isolate 1401-Luan (CHN)-08 VP1, Anhui, 2008; EV71 isolate
1404-Luan (CHN)-08 VP1] and Sarawak Prefecture, Malaysia [no. EV71
e SB12007-SAR-03 VP1; EV71 isolate 1401-Luan (CHN)-08 VP1].
SB12278-SAR-03 VP1 and EV71 isolate SB10712-SAR-03 VP1 are all on
the same branch and the genetic distance is close (0.01-0.03),
which indicates that the origin of the virus is similar. The
strains isolated from Zhejiang, Ningbo province, in 2010 (EV71
strain EV71/Ningbo. CHN/001/2010), Shanghai in 2014 (EV71 strain
SHAPHC5251/SH/CHN/14), Shenzhen in 2014 (EV71 isolate
EV71/SZ07/CHN/2014; EV71 isolate EV71/SZ88/CHN/2014), Wenzhou in
2013 and 2014 (EV71 isolate EV71/P156/2013/China; EV71 isolate
EV71/P654/2013/China; EV71 strain 15/EV71/Wenzhou/CHN/2014) were in
the same evolutionary lineage, and there were certain differences
in amino acid sequence, with a genetic distance of 1.52. The
genetic distance of strains isolated from Australia in 2006 (EV71
strain 2978-SYD-92 VP1; EV71 strain 7784-SYD-90 VP1; EV71 strain
1182-SYD-91 VP1; EV71 strain 6560-SYD-86 VP1) was 2.11 and the
amino acid sequence was quite different (23).
In the present study, 15 strains of variable EV7l
strains over two years were selected and subjected to genetic
analysis. No major changes in genetic variation were observed over
the 2-year period; however, from an evolutionary perspective, there
were still certain differences. The strains detected in the same
year had relatively closer genetic distances. This may be due to
the small variation of genes and virulence in the region. EV7l
strains from the Sarawak region in Malaysia had high homology with
strains from the southwest border of China. In recent years, trade
and tourism have led to the spread of pathogens, particularly HFMD,
and safety monitoring, prevention and control should be implemented
to avoid more widespread and serious infection.
The present study revealed that an amino acid
substitution occurred in the 144th amino acid of the VP1 protein in
the EV71 virus strain in 2015 and 2016. The three-dimensional
conformation suggested that the amino acid mutation site H144Y was
located on the surface of the VP1 protein. Researchers have used
biological software analysis to predict that EV71 epitopes may be
distributed between 90-120, 150-170, 200-240 and 230-250 amino
acids (24). Through sequence
comparison, Li et al (25)
screened the highly conserved epitopes of EV71 among different gene
subtypes as SP55 (amino acid position 163-177) and SP70 (amino acid
position 208-222) in the VP1 protein and demonstrated that these
epitopes may induce the production of specific neutralizing
antibodies against the EV71 virus in mice, which are ideal targets
for multi-epitope vaccines. Other studies suggested that EV71 is
able to infect white blood cells by binding to specific receptor
molecules called P-selectin ligand-1 (PSGL-1) (26). The residue 145 in the capsid protein
VP1 of the PSGL-1-binding virus was G or Q (VP1-145G or Q), while
that of PSGL-1-unbinding virus was VP1-145e (26). VP1-145 acts as a switch to control
the binding of the virus to PSGL-1 by adjusting the exposure of
VP1-244K (26). Whether the VP1-144
mutation identified in the present study affects the binding of
virus to target cells remains to be further explored.
Acknowledgements
The authors wish to thank Dr Xianghong Cao
(Department of Laboratory, Kunming Yanan Hospital) for assistance
with the design of the present study.
Funding
The present study was funded by the Yunnan Applied
Basic Research Project (grant no. 2014FB080).
Availability of data and materials
The datasets used and/or analyzed during the present
study are available from the corresponding author on reasonable
request.
Authors' contributions
YW, XG and HG conceived and designed the study. YW,
XYF, YL, YY and CP performed the experiments. YW, YL and YY
analyzed the data. YW and YL wrote the manuscript. XG and HG
revised the manuscript. All authors read and approved the final
manuscript.
Ethics approval and consent to
participate
The study was approved by Kunming Yan'an
Hospital/Kunming Medical University Affiliated Yan'an Hospital
Medical Ethics Committee (document no. 2017-035-01). The patients
provided oral informed consent and the requirement for written
informed consent was waived by the ethics committee.
Patient consent for publication
Not applicable
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Chen Z, Sun H, Yan Y, Wang Y, Zhu C, Zhou
W, Huang L, Wang M, Mize M, Tian J and Ji W: Epidemiological
profiles of hand, foot, and mouth disease, including meteorological
factors, in Suzhou, China. Arch Virol. 160:315–321. 2015.PubMed/NCBI View Article : Google Scholar
|
|
2
|
Li L, Yin H, An Z and Feng Z:
Considerations for developing an immunization strategy with
enterovirus 71 vaccine. Vaccine. 33:1107–1112. 2015.PubMed/NCBI View Article : Google Scholar
|
|
3
|
Li H, Li S, Zheng J, Cai C, Ye B, Yang J
and Chen Z: Cerebrospinal fluid Th1/Th2 cytokine profiles in
children with enterovirus 71-associated meningoencephalitis.
Microbiol Immunol. 59:152–159. 2015.PubMed/NCBI View Article : Google Scholar
|
|
4
|
Ang LW, Tay J, Phoon MC, Hsu JP, Cutter J,
James L, Goh KT and Chow VT: Seroepidemiology of coxsackievirus a6,
coxsackievirus a16, and enterovirus 71 infections among children
and adolescents in Singapore, 2008-2010. PLoS One.
10(e0127999)2015.PubMed/NCBI View Article : Google Scholar
|
|
5
|
Huang X, Wei H, Wu S, Du Y, Liu L, Su J,
Xu Y, Wang H, Li X, Wang Y, et al: Epidemiological and etiological
characteristics of hand, foot, and mouth disease in Henan, China,
2008-2013. Sci Rep. 5(8904)2015.PubMed/NCBI View Article : Google Scholar
|
|
6
|
Zhou SL, Ying XL, Han X, Sun XX, Jin Q and
Yang F: Characterization of the enterovirus 71 VP1 protein as a
vaccine candidate. J Med Virol. 87:256–262. 2015.PubMed/NCBI View Article : Google Scholar
|
|
7
|
Zhang S, Wang Y and Gao H: Research
progress in laboratory detection of common pathogens of hand foot
and mouth disease. J Kunming Med University. 37:1–4. 2016.
|
|
8
|
Ministry of health of the People's
Republic of China. Hand-foot-mouth disease prevention and control
guidelines (2008 edition). Capital Public Health. 2:146–148.
2008.
|
|
9
|
Zhiqin Zhang and Guoshi Xiang: Research
progress of enterovirus 71 VP1 protein. Chin J Zoonoses.
31:377–379, 390. 2015.
|
|
10
|
Mao Q, Wang Y, Yao X, Bian L, Wu X, Xu M
and Liang Z: Coxsackievirus A16: Epidemiology, diagnosis, and
vaccine. Hum Vaccin Immunother. 10:360–367. 2014.PubMed/NCBI View
Article : Google Scholar
|
|
11
|
Yao X, Bian LL, Mao QY, Zhu FC, Ye Q and
Liang ZL: Echovirus 7 associated with hand, foot, and mouth disease
in mainland China has undergone a recombination event. Arch Virol.
160:1291–1295. 2015.PubMed/NCBI View Article : Google Scholar
|
|
12
|
Guoquan Zhang: Observation on the curative
effect of treating HFMD in adults by integrated Chinese and western
medicine. Zhejiang J Integrated Chin Western Med 2014: 825-826.
|
|
13
|
Linghua Yu, Xinguang Yin, Xingju Wang, et
al: Analysis of correlation characteristics of 41 adult hand, foot
and mouth disease patients. Chin J Infectious Dis. 31:560–563.
2013.
|
|
14
|
Chen GP, Wu JB, Wang JJ, Pan HF, Zhang J,
Shi YL, Cao C, Li FR, Fan YG, Meng FY and Ye DQ: Epidemiological
characteristics and influential factors of hand, foot and mouth
disease (HFMD) reinfection in children in Anhui province. Epidemiol
Infect. 144:153–160. 2016.PubMed/NCBI View Article : Google Scholar
|
|
15
|
Zhou X, Zhu Q, Xia W, He F, Hu M, Ni X,
Gao M, Chen H and Chen S: Molecular epidemiology of an outbreak of
hand, foot, and mouth disease associated with subgenotype C4a of
enterovirus A71 in Nanchang, China in 2014. J Med Virol.
87:2154–2158. 2015.PubMed/NCBI View Article : Google Scholar
|
|
16
|
Eshima N, Tokumaru O, Hara S, Bacal K,
Korematsu S, Karukaya S, Uruma K, Okabe N and Matsuishi T:
Age-specific sex-related differences in infections: A statistical
analysis of national surveillance data in Japan. PLoS One.
7(e42261)2012.PubMed/NCBI View Article : Google Scholar
|
|
17
|
Jianbo Wang, Yan Li, Xiaohong Li, et al:
Four cases of hand foot and mouth disease in adults and virological
observation. Chin J Dermatol. 47:659–660. 2014.
|
|
18
|
Yubao Lian and Jinqiu Yu: A case of adult
severe hand, foot and mouth disease. Chin J Clin Infect Dis. 3:376.
2010.
|
|
19
|
Sheng Zhang, Lingjia Li, Hui Gao, et al:
Clinical report of 2 adult cases of hand foot and mouth disease and
its family pathogen aggregation. Lab Med. 31:733–734. 2016.
|
|
20
|
Mingyu Xie, Qi Peng, Xiaomei Lu, et al:
Research progress on human enterovirus 71 vaccine. Chin J Clin
Pediatrics. 31:1508–1512. 2016.
|
|
21
|
Kataoka C, Suzuki T, Kotani O,
Iwata-Yoshikawa N, Nagata N, Ami Y, Wakita T, Nishimura Y and
Shimizu H: The role of VP1 amino acid residue 145 of enterovirus 71
in viral fitness and pathogenesis in a cynomolgus monkey model.
PLoS Pathog. 11(e1005033)2015.PubMed/NCBI View Article : Google Scholar
|
|
22
|
Zongmin Fu, Xiaoling Xia, Lin Zhao, Ping
Zhao and Ling Zhao: Pathogen detection and clinical characteristics
analysis of hand, foot and mouth disease in kunming in 2011. J Clin
Pediatrics. 31:48–51. 2013.
|
|
23
|
Sanders SA, Herrero LJ, McPhie K, Chow SS,
Craig ME, Dwyer DE, Rawlinson W and McMinn PC: Molecular
epidemiology of enterovirus 71 over two decades in an Australian
urban community. Arch Virol. 151:1003–1013. 2006.PubMed/NCBI View Article : Google Scholar
|
|
24
|
Wenxiang He, Yongjun Zhang, Guangmin Chen,
Ying Zhu, Wei Chen and Yuwei Weng: Analysis of prevalence and
genetic variation of EV71 in fujian province. Chin J Zoonoses.
33:136–142. 2017.
|
|
25
|
Li YX, Zhao H, Cao RY, Deng YQ, Han JF,
Zhu SY, Ma J, Liu L, Qin ED and Qin CF: Recombinant tandem
multi-linear neutralizing epitopes of human enterovirus 71 elicited
protective immunity in mice. Virol J. 11(79)2014.PubMed/NCBI View Article : Google Scholar
|
|
26
|
Nishimura Y, Lee H, Hafenstein S, Kataoka
C, Wakita T, Bergelson JM and Shimizu H: Enterovirus 71 Binding to
PSGL-1 on Leukocytes: VP1-145 Acts as a Molecular Switch to Control
Receptor Interaction. PLoS Pathog. 9(e1003511)2013.PubMed/NCBI View Article : Google Scholar
|