Introduction
Breast cancer is the most commonly diagnosed type of
cancer and the leading cause of cancer-related mortality in women
worldwide (1). Breast cancer has
been estimated to account for 24.2% of all new cancer diagnoses and
15.0% of all cancer-related deaths among women (1). Although the overall survival and
prognosis of patients with breast cancer has significantly improved
in recent years, metastasis remains the leading cause of mortality
in patients with breast cancer (2).
For example, patients with metastases have a 5-year survival rate
of only 26% compared with a 5-year survival rate of 90% across all
patients with breast cancer (3).
Hence, it is important to further understand the mechanisms
underlying the development and progression of breast cancer to
identify novel targets for the treatment of this disease.
Baculoviral IAP repeat containing 8 (ILP2) is an
inhibitor belonging to the inhibitor of apoptosis protein family
(4), which can prevent
pro-apoptotic stimulation and inhibit apoptosis. A previous study
revealed that the expression levels of inhibitor of apoptosis-like
protein-2 (ILP2) were upregulated in patients with breast cancer
(5). In addition, the viability and
migratory ability of human breast cancer cell lines (HCC-1937, MX-1
and MCF-7) were significantly inhibited following knockdown of ILP2
expression, and apoptosis was increased, compared with those in the
control group (5). Collectively,
these findings indicated that ILP2 may promote the proliferation,
migration and invasion of breast cancer cells.
Transcription factors are proteins that bind to the
DNA helix at specific regulatory sequences to activate or inhibit
transcription through a transactivation or transrepression domain
(6). Determining the activity of
transcription factors is crucial for understanding the regulation
of gene expression (7). A number of
studies have reported that transcription factors serve an important
role in the occurrence and development of numerous types of tumors.
Therefore, the targeting of transcription factors may represent a
novel method for tumor treatment (6,8).
Homeobox (HOX) genes function as master regulatory transcription
factors, and their expression levels have been found to be
regularly altered in cancer (9).
The expression levels of HOX genes were reported to be upregulated
or downregulated in different tumor types, depending on the
specific HOX gene involved and the type of cancer being
investigated. HOXD8 is an important member of the HOX gene family,
and was found to serve an important role in colorectal cancer,
non-small cell lung cancer, advanced epithelial ovarian cancer,
lung cancer and hepatocellular carcinoma (10-14).
Therefore, the present study was undertaken to investigate the role
of HOXD8 in the occurrence and development of breast cancer.
Bioinformatics software was used to examine the
expression of HOXD8 in breast cancer tissues and predict its
downstream genes. The aim of the present study was to examine the
association between HOXD8 and its downstream genes, and to explore
their roles and mechanism of action in breast cancer cells, in the
hope that the results may uncover novel targets for the treatment
of breast cancer.
Materials and methods
Cell lines and culture
Breast cancer cell lines (MCF-7, MDA-MB-231,
SUM190PT and SK-BR-3) and a normal human breast epithelial cell
line (MCF-10A) were purchased from Procell Life Science &
Technology Co., Ltd. MCF-7 and MDA-MB-231 cells were cultured in
DMEM (Gibco; Thermo Fisher Scientific, Inc.); SUM190PT cells were
cultured in Ham's F-12 medium (Sigma-Aldrich; Merck KGaA); SK-BR-3
cells were cultured in McCoy's 5A medium (Sigma-Aldrich; Merck
KGaA); and MCF-10A cells were cultured in RPMI-1640 medium
(Sigma-Aldrich; Merck KGaA) (15).
All media aforementioned were all supplemented with 10% FBS (Gibco;
Thermo Fisher Scientific, Inc.) and 100 U/ml
penicillin/streptomycin (Thermo Fisher Scientific, Inc.). All cells
were maintained in an incubator containing 5% CO2 at
37˚C.
Cell transfection
Gene overexpression was performed with a pcDNA3.1
vector and gene knockdown with a pLVX-shRNA2 lentiviral vector. The
full length coding sequence of HOXD8 (accession no. AH010089.2;
https://www.ncbi.nlm.nih.gov/nuccore/AH010089.2/)
was found from National Center for Biotechnology Information (NCBI;
https://www.ncbi.nlm.nih.gov/). Fragment
with restriction sites was synthesized and inserted into the
pcDNA3.1 vector (pcDNA-HOXD8) by Hunan Fenghui Biotechnology Co.,
Ltd., whereas the empty vector was the negative control (pcDNA-NC).
Two types of short hairpin (sh)RNA-HOXD8 (shRNA-HOXD8-1 and
shRNA-HOXD8-2), two types of shRNA-ILP2 (shRNA-ILP2-1 and
shRNA-ILP2-2) and non-targeted shRNA as negative control (shRNA-NC)
were purchased from Guangzhou RiboBio Co., Ltd. and transfected
into MCF-7 cells. shRNA-1 and shRNA-2 correspond to different
sequences being incorporated into the same vector. Briefly, cells
(5x105 cells/well) were plated into six-well plates and
cultured until they reached 50-70% confluence. Cells were
transfected with 10 nM pcDNAs and shRNAs using
Lipofectamine® 2000 reagent (Invitrogen; Thermo Fisher
Scientific, Inc.) at 37˚C, according to the manufacturer's
protocol. Following 48 h of transfection, the expression levels of
HOXD8 and ILP2 were analyzed, and the cells were used for
subsequent experiments. The sequences of shRNA are as follows:
shRNA-HOXD8-1
5'-CCGGAGCCGAAGGCCTGACAAATTACTCGAGTAATTTGTCAGGCCTTCGGCTTTTTTG-3';
shRNA-HOXD8-2
5'-CCGGGCCGAAGGCCTGACAAATTAACTCGAGTTAATTTGTCAGGCCTTCGGCTTTTTG-3';
shRNA-ILP2-1
5'-CCGGACGGTGGACAAGTCCTATATTCTCGAGAATATAGGACTTGTCCACCGTTTTTTG-3';
shRNA-ILP2-2
5'-CCGGTTTGGGCCACAACGTTAATATCTCGAGATATTAACGTTGTGGCCCAAATTTTTG-3';
shRNA-NC
5'-CCGGCAACAAGATGAAGAGCACCAACTCGAGTTGGTGCTCTTCATCTTGTTGTTTTTG-3'.
Western blotting
All cells (5x105 cells/well) were seeded
into a six-well plate separately and total protein was extracted
from cells using RIPA lysis buffer (Beyotime Institute of
Biotechnology). Total protein concentration was quantified using
the BCA method and 20 µg protein/lane was separated by 10%
SDS-PAGE. The separated proteins were subsequently transferred onto
PVDF membranes and blocked with 5% skimmed milk diluted in 5% BSA
(Beijing Solarbio Science & Technology Co., Ltd.) for 2 h at
room temperature. The membranes were then incubated with the
following primary antibodies at a dilution of 1:1,000 overnight at
4˚C: Anti-HOXD8 (cat. no. sc-515357; Santa Cruz Biotechnology,
Inc.), anti-MMP2 (cat. no. 40994; Cell Signaling Technology, Inc.),
anti-MMP9 (cat. no. 13667; Cell Signaling Technology, Inc.),
anti-ILP2 (cat. no. ab9664; Abcam) and anti-GAPDH (cat. no. 5174,
Cell Signaling Technology, Inc.). Following primary antibody
incubation, the membranes were incubated with an HRP-conjugated
goat anti-rabbit or goat anti-mouse IgG secondary antibody
(1:100,000; cat. nos. G-21234 and G-21040, respectively;
Invitrogen; Thermo Fisher Scientific, Inc.) for 1.5 h at room
temperature. GAPDH served as the internal reference control.
Protein bands were visualized using an ECL kit (cat. no. 21342;
Beyotime Institute of Biotechnology) and densitometric analysis was
performed using ImageJ v1.51 software (National Institutes of
Health).
Cell Counting Kit-8 (CCK-8) assay
MCF-7 cells were seeded into 96-well plates at the
density of 5x103 cells/well and cultured in an incubator
containing 5% CO2 at 37˚C for 24, 48 or 72 h. In total,
10 µl CCK-8 solution (cat. no. ab228554; Abcam) was added to each
well at each time point. Following incubation for another 2 h at
37˚C, the optical density was measured at a wavelength of 450 nm
using a microplate reader (Thermo Fisher Scientific, Inc.).
Colony formation assay
MCF-7 cells (3x103 cells/well) were
seeded into six-well plates and incubated with DMEM supplemented
with 10% FBS at 37˚C. Following 14 days of incubation, the cells
were fixed with 4% methanol for 15 min at room temperature and
stained with 0.1% crystal violet dye solution for another 10 min at
room temperature. The number of cell clone clusters containing
>50 cells was counted under a light microscope (magnification,
x100; Olympus Corporation). The assay was performed in triplicate
to determine the number of colonies.
Wound healing assay
MCF-7 cells (5x105 cells/well) were
plated into six-well plates and incubated at 37˚C until they
reached 90% confluence. A scratch was subsequently created in the
cell monolayer using a 200-µl pipette tip to generate an artificial
wound. At 0 and 48 h of culture in serum-free medium at 37˚C,
images of the wound area were captured in the same field using an
inverted light microscope (Olympus Corporation; magnification,
x100) and analyzed by the ImageJ v1.51 software (National
Institutes of Health). Cell migration rate = wound area difference
between 0 and 48 h/wound area at 0 h x 100%.
Transwell assay
Matrigel (cat. no. 356234; Corning, Inc.) was thawed
overnight at 4˚C and diluted with serum-free medium (1:8) before 50
µl of this Matrigel was inoculated in the upper chamber of
Transwell plates (cat. no. CLS3422; 8.0 µm; Corning, Inc.) at 37˚C.
A total of 5x104 MCF-7 cells in 100 µl serum-free DMEM
were seeded into the upper chamber of the Transwell plates and the
lower chamber was filled with 500 µl DMEM supplemented with 10%
FBS. Following 48 h of incubation at 37˚C, the cells in the lower
chamber were fixed with 10% formaldehyde for 15 min at room
temperature and stained with 0.1% crystal violet solution for 15
min at room temperature. The number of invasive cells was counted
using a light microscope (magnification, x200) and analyzed by
ImageJ v1.51 software (National Institutes of Health). Cell
invasion rate=the number of invasive cells/number of inoculated
cells x100%.
Chromatin immunoprecipitation (ChIP)
assay
To determine whether HOXD8 bound to the promoter of
the ILP2 gene, a ChIP assay was performed in shRNA-NC- and
shRNA-HOXD8-transfected cells using a SimpleChIP®
enzymatic chromatin IP kit (Cell Signaling Technology, Inc.)
according to the manufacturer's protocol. After MCF-7 cells were
lysed, 50 µg protein G agarose beads was added to 1 ml supernatant
and incubated at 4˚C for 1 h. The supernatant was then taken after
centrifugation at 1,000 x g at 4˚C for 3 min before being divided
into two groups. Afterwards, 3 µg IgG or HOXD8 antibodies were
added to 500 µg protein samples and incubated overnight at 4˚C. The
next day, 50 µg protein G agarose beads was added and the
precipitate was collected after incubating at 4˚C for 6 h and
centrifugated at 1,000 x g at 4˚C for 3 min. The precipitate was
washed with 5X lysis buffer and resuspended in 150 µl 1X ChIP
Elution Buffer. Chromatin from beads were eluted with gentle
vortexing (1,200 rpm) at 65˚C for 30 min. DNA was purified using
the DNA Purification kit (cat. no. D0033; Beyotime Institute of
Biotechnology). Relative enrichment was performed by quantitative
PCR (qPCR) analysis. The antibody against IgG (cat. no. 5415S; Cell
Signaling Technology, Inc.) was diluted to the same concentration
(2 µg/ml) as the HOXD8 antibody (cat. no. sc-515357; Santa Cruz
Biotechnology, Inc.).
Dual-luciferase reporter assay
The full length (FL) was the fragment located at
position 100-2,000 of ILP2 (Sequence ID, NC_000019.10,
53293434-53291335). Three deletion mutants were the fragments
containing the predicted binding sites located at positions 420-427
(element 3, E3), 1601-1608 (E2), 1747-1754 (E1) separately mutated
(Hunan Fenghui Biotechnology Co., Ltd.) and cloned into the pGL3
basic vector (BioVector NTCC Inc.). MCF-7 cells (1x105
cells/well) were seeded into 24-well plates and transfected with
100 ng luciferase vectors (FL; E3 Deletion, E3 Del; E2 Del or E1
Del) and 10 nM expression vectors (shRNA-NC or shRNA-HOXD8) using
Lipofectamine® 2000 reagent (Thermo Fisher Scientific,
Inc.) at room temperature. Following 5 h of incubation, the
transfection solution was replaced by 500 µl DMEM and cells were
incubated for another 24 h at 37˚C. The relative luciferase
activity of cells was measured using a Dual-Luciferase Reporter
Assay System (Promega Corporation) according to the manufacturer's
protocol. Firefly luciferase activity was normalized to
Renilla luciferase activity.
Database analysis
The Gene Expression Profiling Interactive Analysis
(GEPIA, http://gepia.cancer-pku.cn) database
is a web server for cancer and normal gene expression profiling and
interactive analyses (16). GEPIA
analysis contains the expression analysis of RNA sequencing data
from 9,736 tumors and 8,587 normal samples in The Cancer Genome
Atlas (http://cancergenome.nih.gov) and
Genotype-Tissue Expression (GTEx, http://commonfund.nih.gov/GTEx) projects. A total of
1,085 tumor samples and 291 normal samples were obtained from the
breast invasive carcinoma (BRCA) data set of GEPIA, with
|Log2FC|>1 and P<0.01 as the cutoff; where FC is
fold-change.
JASPAR (http://jaspar.genereg.net) is a database of
transcription factor binding profiles. There were a total of three
profiles on HOXD8 found in the JASPAR database; the FASTA-formatted
sequence of ILP2 was input to scan with the selected profile
(Species: Homo sapiens; ID: MA0910.2), and three predicted
sequences with high scores were selected as promising binding
sites.
Statistical analysis
Statistical analysis was performed using GraphPad
Prism 8.0 software (GraphPad Software, Inc.) and the data are
presented as the mean ± SD of triplicate experiments. The
statistical significance of the differences between two groups were
determined using an unpaired Student's t-test and among multiple
groups using one-way ANOVA followed by a Dunnett's post hoc test.
P<0.05 was considered to indicate a statistically significant
difference.
Results
HOXD8 expression is downregulated in
breast cancer tissues and cell lines
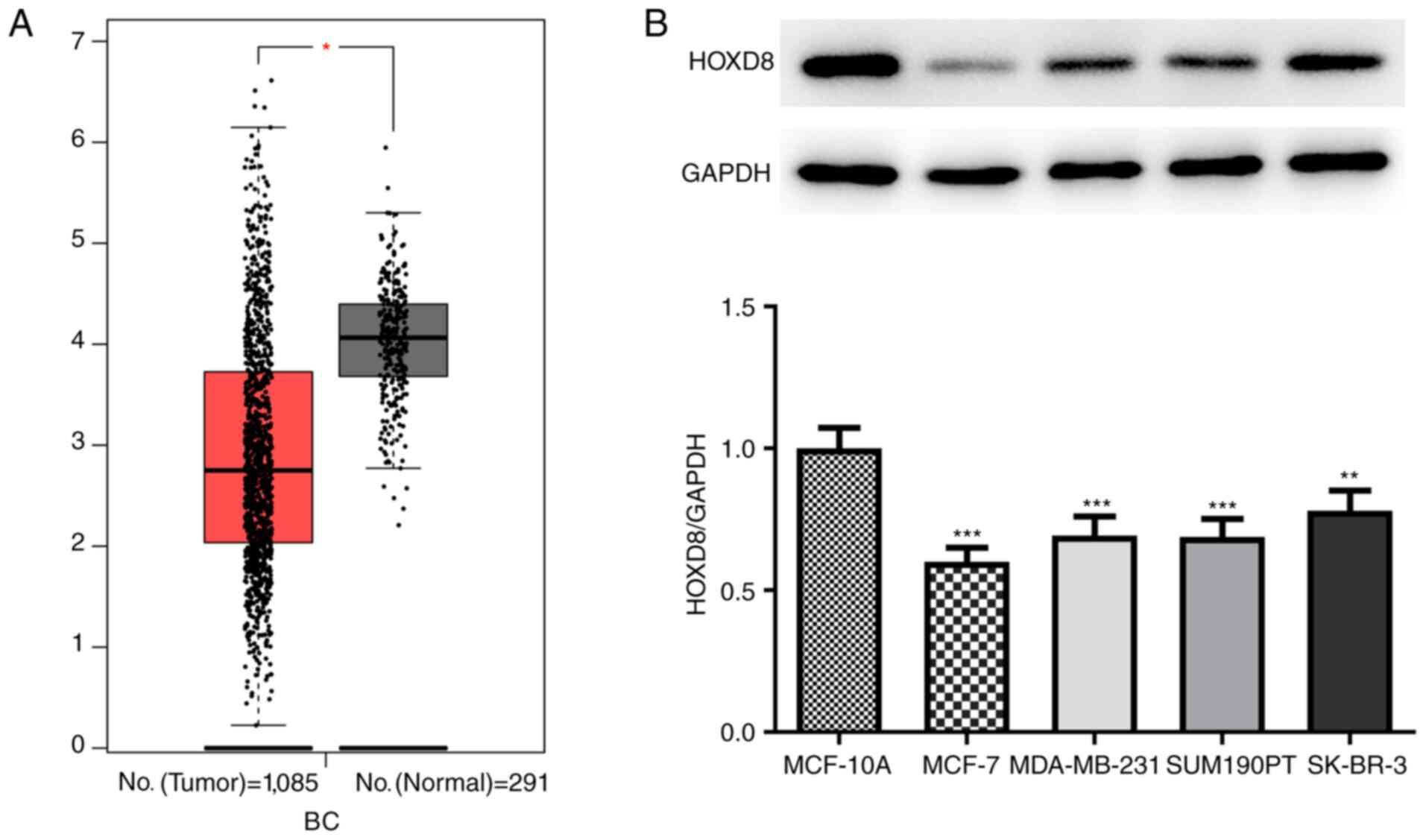
Data from 1,085 tumor samples and 291 normal samples
were obtained from the BRCA dataset from the GEPIA database; it was
revealed that the expression levels of HOXD8 were significantly
downregulated in breast cancer tissues (Fig. 1A). In addition, the expression
levels of HOXD8 were analyzed in four breast cancer cell lines
(MCF-7, MDA-MB-231, SUM190PT and SK-BR-3), and the results revealed
that HOXD8 protein expression was significantly lower in breast
cancer cell lines compared with expression in the MCF-10A normal
human breast epithelial cell line (Fig.
1B). HOXD8 expression was the lowest in MCF-7 cells; therefore,
this cell line was selected for use in subsequent experiments.
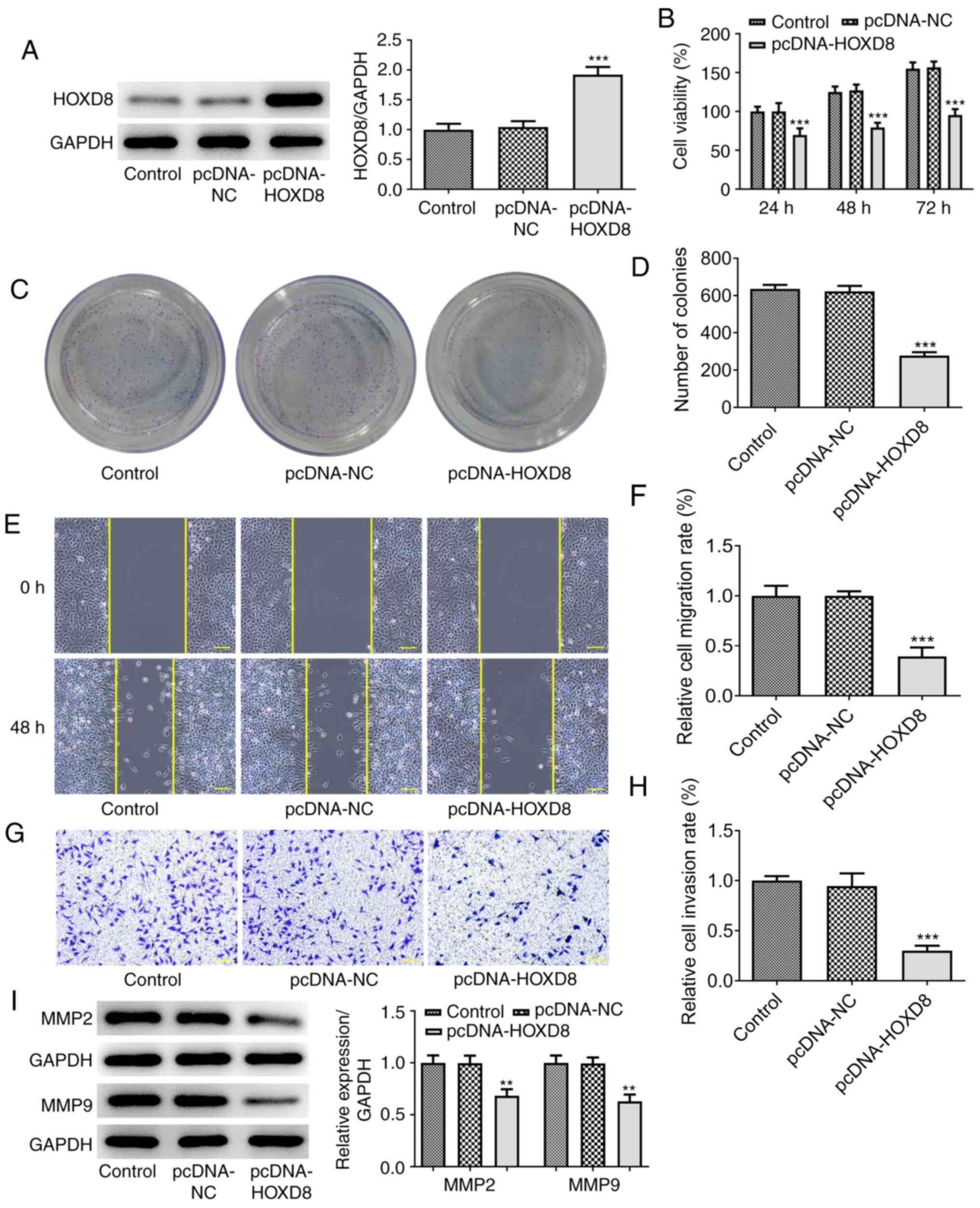
HOXD8 overexpression inhibits breast
cancer cell proliferation, migration and invasion
To further investigate the biological functions of
HOXD8 in breast cancer, MCF-7 cells were transfected with
pcDNA-HOXD8 or pcDNA-NC empty vector. Western blotting revealed
that HOXD8 expression levels were significantly upregulated
following transfection with pcDNA-HOXD8 compared with the
untransfected MCF-7 cells (the control) and pcDNA-NC groups
(Fig. 2A), indicating the
successful transfection of pcDNA-HOXD8 into MCF-7 cells. The
results of the CCK-8 assay demonstrated that the overexpression of
HOXD8 significantly inhibited MCF-7 cell proliferation (Fig. 2B), and the number of cell clone
clusters containing >50 cells was counted, the results of the
colony formation assays were consistent with these findings
(Fig. 2C-D). Moreover, the results
obtained from the Matrigel and wound healing assays revealed that
the overexpression of HOXD8 inhibited the invasive and migratory
ability, respectively, of MCF-7 cells compared with the control and
pcDNA-NC groups (Fig. 2E-H). In
addition, western blotting demonstrated that the overexpression of
HOXD8 significantly downregulated the expression levels of
migration-related proteins, MMP2 and MMP9, compared with the
control and pcDNA-NC groups (Fig.
2I). These results suggested that HOXD8 may act as a tumor
suppressor in breast cancer cells.
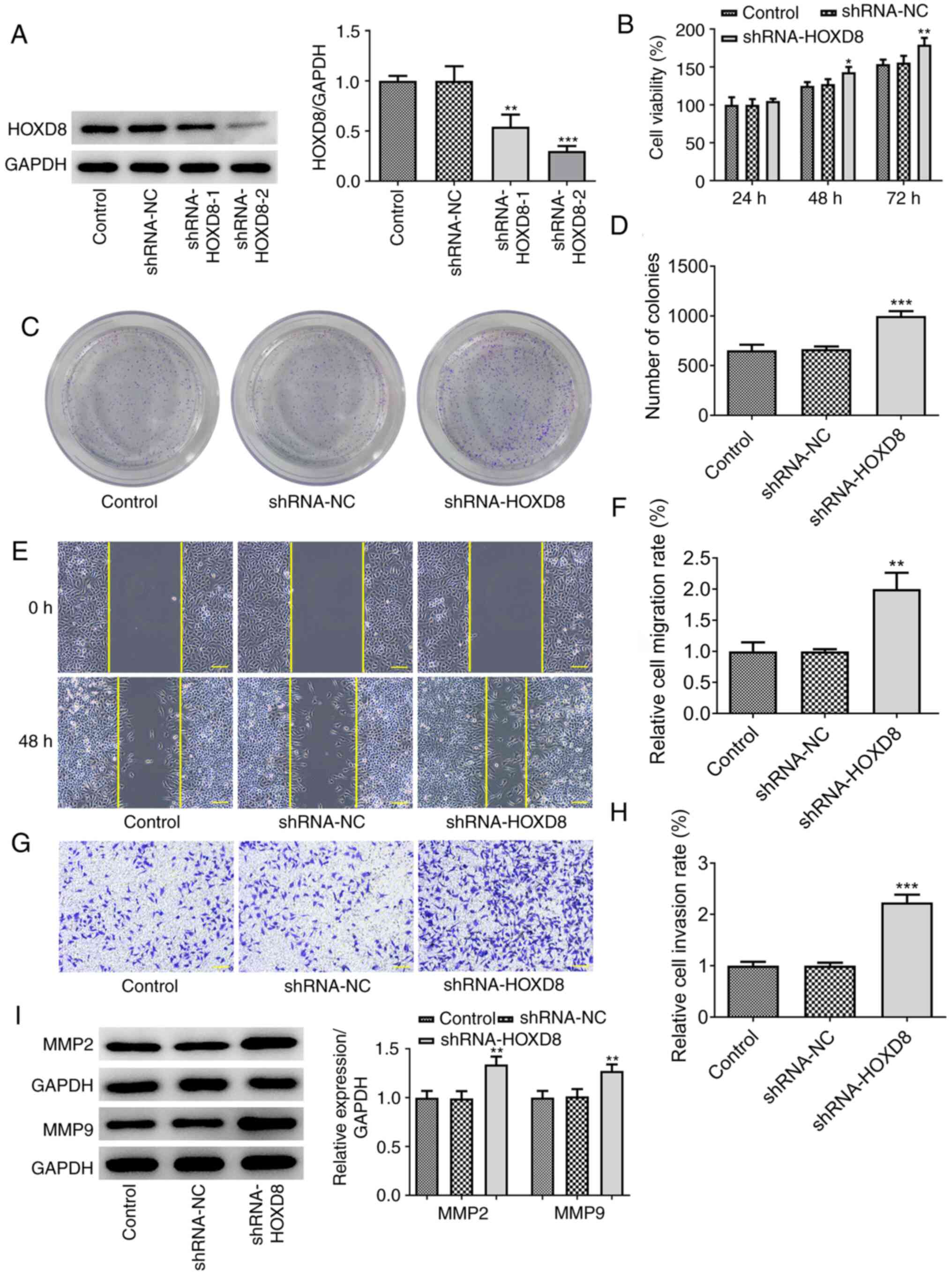
HOXD8 knockdown promotes breast cancer
cell proliferation, migration and invasion
It was next investigated whether the knockdown of
HOXD8 could promote the proliferation and colony forming ability of
MCF-7 cells. Two types of shRNA-HOXD8 were separately transfected
into MCF-7 cells to silence HOXD8 expression and shRNA-HOXD8-2 was
selected for subsequent assays because it reduced the expression
levels of HOXD8 to a greater extent compared with that of
shRNA-HOXD8-1 (Fig. 3A). The
results revealed that the proliferation (Fig. 3B), colony forming ability (Fig. 3C-D), invasion and migration
(Fig. 3E-H) of MCF-7 cells were
increased following HOXD8 knockdown. The protein expression levels
of MMP2 and MMP9 were also upregulated following knockdown of HOXD8
compared with the control and shRNA-NC groups (Fig. 3I). These findings further suggested
that HOXD8 may play a tumor-suppressive role in breast cancer.
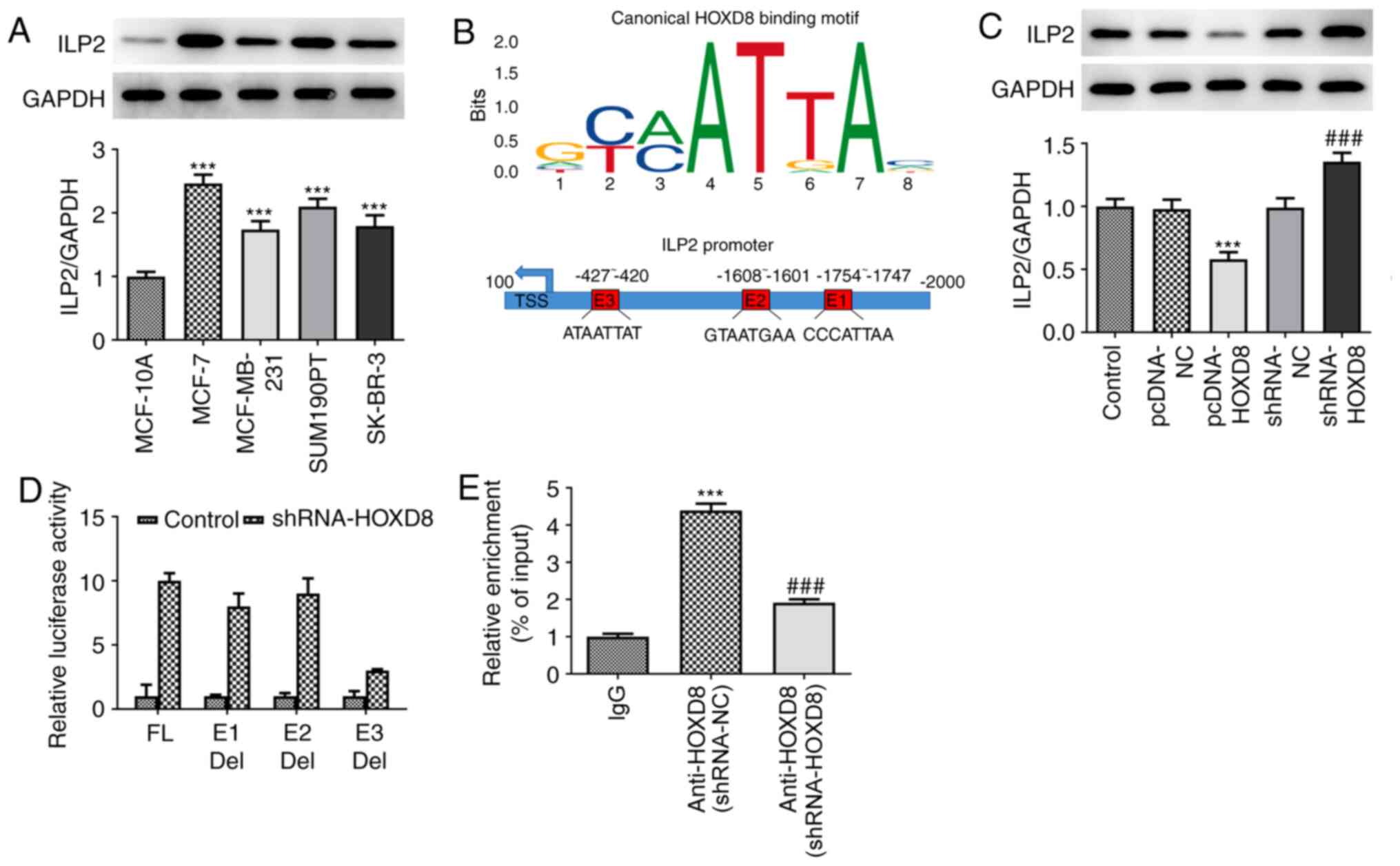
ILP2 is a direct target of HOXD8 in
MCF-7 breast cancer cells
To determine the mechanism underlying the regulatory
role of HOXD8 in breast cancer cells, the present study
investigated potential targets using the JASPAR database; ILP2 was
identified as a potential target and was investigated in further
experiments. As shown in Fig. 4A,
ILP2 expression levels were found to be significantly upregulated
in breast cancer cell lines compared with MCF-10A cells, which
indicated a potential role for ILP2 in breast cancer tumorigenesis
and progression. Notably, in Fig.
4B, each column of HOXD8 sequence motif corresponds to a base
position, and each base position is accumulated by the bases that
will appear at that position. The larger the letter, the greater
the total amount of information, and the greater the probability of
the base appearing. Through combining the motif with the input
sequence of ILP2, three putative HOXD8-binding elements (E1:
CCCATTAA; E2: GTAATGAA; E3: ATAATTAT) were identified within the
ILP2 promoter region using the JASPAR database. Subsequently,
whether HOXD8 could regulate ILP2 expression was further
determined. Western blotting demonstrated that HOXD8 overexpression
downregulated ILP2 protein expression levels, whereas HOXD8
knockdown markedly upregulated ILP2 expression levels (Fig. 4C). To determine which binding site
was required for HOXD8-mediated ILP2 expression, the three
predicted HOXD8-binding sites were individually deleted and used in
separate luciferase assays. The results revealed that HOXD8 almost
failed to promote ILP2 transcriptional activity without the E3
element (Fig. 4D), which indicated
that the E3 element may be essential for HOXD8 to activate ILP2
transcription. Notably, the enrichment of ILP2 from the ChIP assay
and qPCR analysis demonstrated that HOXD8 bound to the promoter of
the ILP2 gene (Fig. 4E).
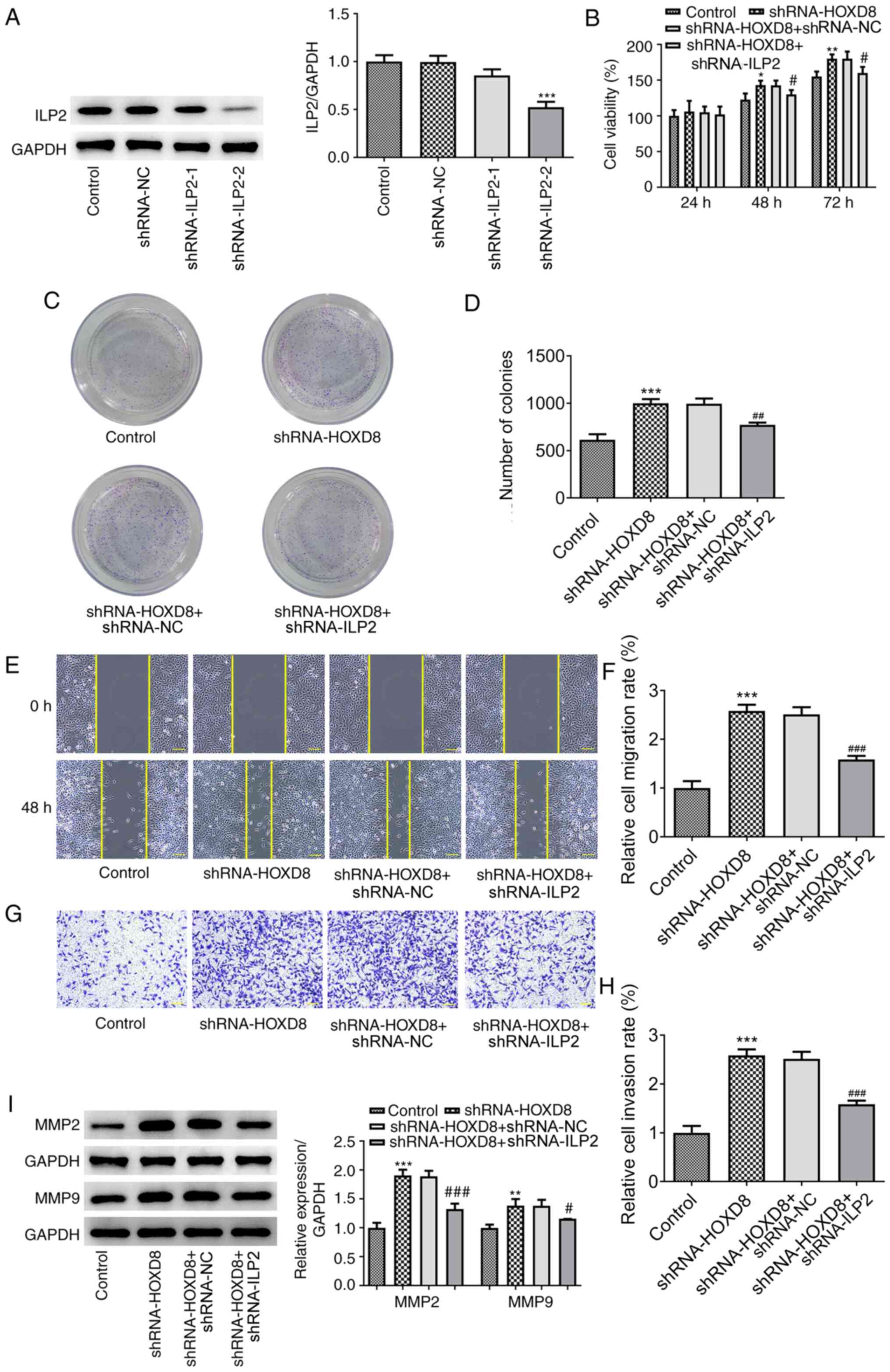
ILP2 knockdown reverses the promoting
effects of HOXD8 knockdown on breast cancer cell proliferation,
migration and invasion
To determine whether ILP2 influenced the effects of
HOXD8 on the proliferation, migration and invasion, two types of
shRNA-ILP2 were transfected into MCF-7 breast cancer cells to
downregulate ILP2 expression, and shRNA-ILP2-2 was selected for
subsequent experiments due to the higher potency of ILP2 knockdown
compared with shRNA-ILP2-1 (Fig.
5A). As previously discussed, HOXD8 knockdown notably increased
the proliferation (Fig. 5B),
colony-forming ability (Fig. 5C-D),
invasion and migration (Fig. 5E-H)
of breast cancer cells, and these effects could be partially
reversed by ILP2 knockdown. In addition, western blotting results
demonstrated that transfection with shRNA-ILP2 downregulated the
HOXD8 knockdown-induced upregulated expression levels of MMP2 and
MMP9 (Fig. 5I). These results
suggested that HOXD8 may regulate breast cancer cell progression by
binding to the ILP2 promoter.
Discussion
Numerous studies have demonstrated the important
tumor-suppressive role of HOXD8. For example, HOXD8 expression
levels were found to be downregulated in colorectal cancer tissues,
which inhibited the proliferation, colony forming ability and
invasion of colorectal cancer cells and upregulated the expression
levels of apoptosis-associated proteins (10). A similar effect was also observed in
hepatocellular carcinoma (14).
However, to the best of our knowledge, the effects of HOXD8 in
breast cancer have yet to be determined. Therefore, the present
study aimed to investigate the role of HOXD8 in breast cancer.
First, the expression levels of HOXD8 in multiple breast cancer
cell lines were analyzed, and based on the results, the expression
level of HOXD8 in the MCF-7 cell line was the lowest. In addition,
since MCF-7 cell line is a representative in vitro model of
HER2-negative luminal breast cancer (17), the MCF-7 cell line was selected for
use in subsequent experiments. By constructing overexpression and
knockdown vectors, the effects of HOXD8 overexpression or knockdown
on cell proliferation, migration and invasion were determined. The
results revealed that the overexpression of HOXD8 inhibited the
proliferation, invasion and migration of breast cancer cells. The
expression levels of MMP2 and MMP9 were also investigated to
further determine the effects of HOXD8 on cell migration, and it
was demonstrated that HOXD8 overexpression inhibited, and HOXD8
knockdown increased, the expression of MMPs. MMPs are a family of
zinc and calcium ion-dependent proteases that can target and
degrade numerous proteins in the extracellular matrix (ECM)
(18), which is key to tumor
invasion and metastasis, as the ECM constitutes the first barrier
for tumor cells to overcome to effectively metastasize. The main
component of the ECM is type IV collagen, which can be degraded by
MMP2 and MMP9. Notably, the expression of MMP2 and MMP9 has been
found to be implicated in the progression of several types of
cancer (19-22).
These findings suggested that HOXD8 may serve a role in breast
cancer cell migration and invasion.
Apoptosis inhibitor proteins can inhibit cell
apoptosis and promote cell proliferation, and have been shown to
serve an important role in the occurrence and development of
several types of cancer (23-25).
In normal tissues, ILP2 is only expressed in the testis and
lymphoblasts, where it inhibits cell apoptosis (4). Previous research on the role of ILP2
in breast cancer demonstrated that the expression levels of ILP2
were significantly upregulated in breast cancer tissues and cell
lines, whereas knockdown of ILP2 expression significantly inhibited
the proliferation, migration and invasion of breast cancer cells,
and increased apoptosis (26). The
results of our previous study revealed that ILP2 may serve as a
serum biomarker for breast cancer, which is important for the
diagnosis and treatment of this disease (5). In addition, a previous study reported
that ILP2 overexpression exerted no inhibitory effects on
TNF-mediated apoptosis, but effectively inhibited apoptosis
mediated by Bax and apoptotic peptidase activating factor
1/caspase-9. In addition, an interaction was identified between
ILP2 and caspase-9(4). Therefore,
it was suggested that devising an inhibitor for the ILP2/caspase
interaction may be an effective anticancer measure (27). The results of the present study
revealed that HOXD8 exerted a tumor-suppressive effect on breast
cancer by targeting the ILP2 promoter. Therefore, regulation of the
HOXD8/ILP2 interaction may also represent a potential effective
anticancer treatment.
In conclusion, the results of the present study
demonstrated that the expression levels of HOXD8 were downregulated
in breast cancer tissues and cell lines. In addition, HOXD8
overexpression was found to inhibit the proliferation, migration
and invasion of breast cancer cells by targeting ILP2. These
findings provide a novel insight into potential therapeutic target
for breast cancer; however, the exact mechanism underlying the role
of ILP2 in breast cancer requires further investigation.
Acknowledgements
Not applicable.
Funding
Funding: No funding was received.
Availability of data and materials
The datasets used and/or analysed during the present
study are available from the corresponding author on reasonable
request.
Authors' contributions
XW designed the study and wrote the manuscript; YC
performed the experiments and analyzed the data; XF participated in
the experiments, examined the data and critically revised the
manuscript for important intellectual content. All authors confirm
the authenticity of all the raw data. All authors have read and
approved the final manuscript.
Ethics approval and consent to
participate
Not applicable.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Bray F, Ferlay J, Soerjomataram I, Siegel
RL, Torre LA and Jemal A: Global cancer statistics 2018: GLOBOCAN
estimates of incidence and mortality worldwide for 36 cancers in
185 countries. CA Cancer J Clin. 68:394–424. 2018.PubMed/NCBI View Article : Google Scholar
|
|
2
|
Miller KD, Nogueira L, Mariotto AB,
Rowland JH, Yabroff KR, Alfano CM, Jemal A, Kramer JL and Siegel
RL: Cancer treatment and survivorship statistics, 2019. CA Cancer J
Clin. 69:363–385. 2019.PubMed/NCBI View Article : Google Scholar
|
|
3
|
Peart O: Metastatic breast cancer. Radiol
Technol. 88:519M–539M. 2017.PubMed/NCBI
|
|
4
|
Richter BW, Mir SS, Eiben LJ, Lewis J,
Reffey SB, Frattini A, Tian L, Frank S, Youle RJ, Nelson DL, et al:
Molecular cloning of ILP-2, a novel member of the inhibitor of
apoptosis protein family. Mol Cell Biol. 21:4292–4301.
2001.PubMed/NCBI View Article : Google Scholar
|
|
5
|
Xiang M, Zhou W, Gao D, Fang X and Liu Q:
Inhibitor of apoptosis protein-like protein-2 as a novel
serological biomarker for breast cancer. Int J Mol Sci.
13:16737–16750. 2012.PubMed/NCBI View Article : Google Scholar
|
|
6
|
Lambert M, Jambon S, Depauw S and
David-Cordonnier MH: Targeting transcription factors for cancer
treatment. Molecules. 23(1479)2018.PubMed/NCBI View Article : Google Scholar
|
|
7
|
Bemer M, van Dijk ADJ, Immink RGH and
Angenent GC: Cross-family transcription factor interactions: An
additional layer of gene regulation. Trends Plant Sci. 22:66–80.
2017.PubMed/NCBI View Article : Google Scholar
|
|
8
|
Bushweller JH: Targeting transcription
factors in cancer-from undruggable to reality. Nat Rev Cancer.
19:611–624. 2019.PubMed/NCBI View Article : Google Scholar
|
|
9
|
Brotto DB, Siena ADD, de B II, Carvalho
SDCES, Muys BR, Goedert L, Cardoso C, Plaça JR, Ramão A, Squire JA,
et al: Contributions of HOX genes to cancer hallmarks: Enrichment
pathway analysis and review. Tumour Biol.
42(1010428320918050)2020.PubMed/NCBI View Article : Google Scholar
|
|
10
|
Mansour MA and Senga T: HOXD8 exerts a
tumor-suppressing role in colorectal cancer as an apoptotic
inducer. Int J Biochem Cell Biol. 88:1–13. 2017.PubMed/NCBI View Article : Google Scholar
|
|
11
|
Sun P, Song Y, Liu D, Liu G, Mao X, Dong
B, Braicu EI and Sehouli J: Potential role of the HOXD8
transcription factor in cisplatin resistance and tumour metastasis
in advanced epithelial ovarian cancer. Sci Rep.
8(13483)2018.PubMed/NCBI View Article : Google Scholar
|
|
12
|
Liu Y, Miao L, Ni R, Zhang H, Li L, Wang
X, Li X and Wang J: microRNA-520a-3p inhibits proliferation and
cancer stem cell phenotype by targeting HOXD8 in non-small cell
lung cancer. Oncol Rep. 36:3529–3535. 2016.PubMed/NCBI View Article : Google Scholar
|
|
13
|
Zhu W, Wang JP, Meng QZ, Zhu F and Hao XF:
MiR-142-5p reverses the resistance to gefitinib through targeting
HOXD8 in lung cancer cells. Eur Rev Med Pharmacol Sci.
24:4306–4313. 2020.PubMed/NCBI View Article : Google Scholar
|
|
14
|
Sun S, Wang N, Sun Z, Wang X and Cui H:
MiR-5692a promotes proliferation and inhibits apoptosis by
targeting HOXD8 in hepatocellular carcinoma. J BUON. 24:178–186.
2019.PubMed/NCBI
|
|
15
|
Zhao W, Geng D, Li S, Chen Z and Sun M:
LncRNA HOTAIR influences cell growth, migration, invasion, and
apoptosis via the miR-20a-5p/HMGA2 axis in breast cancer. Cancer
Med. 7:842–855. 2018.PubMed/NCBI View Article : Google Scholar
|
|
16
|
Tang Z, Li C, Kang B, Gao G, Li C and
Zhang Z: GEPIA: A web server for cancer and normal gene expression
profiling and interactive analyses. Nucleic acids research.
45:W98–W102. 2017.PubMed/NCBI View Article : Google Scholar
|
|
17
|
Wei HC: Mathematical modeling of tumor
growth: The MCF-7 breast cancer cell line. Math Biosci Eng.
16:6512–6535. 2019.PubMed/NCBI View Article : Google Scholar
|
|
18
|
Jabłońska-Trypuć A, Matejczyk M and
Rosochacki S: Matrix metalloproteinases (MMPs), the main
extracellular matrix (ECM) enzymes in collagen degradation, as a
target for anticancer drugs. J Enzyme Inhib Med Chem. 31 (Suppl
1):177–183. 2016.PubMed/NCBI View Article : Google Scholar
|
|
19
|
Avădanei R, Căruntu ID, Amălinei C,
Lozneanu L, Balan R, Grigoraş A, Ciobanu Apostol D and Giuşcă SE:
High variability in MMP2/TIMP2 and MMP9/TIMP1 expression in
secondary liver tumors. Rom J Morphol Embryol. 54:479–485.
2013.PubMed/NCBI
|
|
20
|
Kalhori V and Törnquist K: MMP2 and MMP9
participate in S1P-induced invasion of follicular ML-1 thyroid
cancer cells. Mol Cell Endocrinol. 404:113–122. 2015.PubMed/NCBI View Article : Google Scholar
|
|
21
|
Sokołowska J and Urbańska K:
Immunohistochemical assessment of metalloproteinases MMP2 and MMP9
expression in canine various subtypes of lymphomas in relation with
proliferative and apoptotic markers. Pol J Vet Sci. 22:203–211.
2019.PubMed/NCBI View Article : Google Scholar
|
|
22
|
Wang X, Yang B, She Y and Ye Y: The lncRNA
TP73-AS1 promotes ovarian cancer cell proliferation and metastasis
via modulation of MMP2 and MMP9. J Cell Biochem. 119:7790–7799.
2018.PubMed/NCBI View Article : Google Scholar
|
|
23
|
Varfolomeev E and Vucic D: Inhibitor of
apoptosis proteins: Fascinating biology leads to attractive tumor
therapeutic targets. Future Oncol. 7:633–648. 2011.PubMed/NCBI View Article : Google Scholar
|
|
24
|
Fulda S: Exploiting inhibitor of apoptosis
proteins as therapeutic targets in hematological malignancies.
Leukemia. 26:1155–1165. 2012.PubMed/NCBI View Article : Google Scholar
|
|
25
|
Fulda S: Inhibitor of apoptosis proteins
as targets for anticancer therapy. Expert Rev Anticancer Ther.
7:1255–1264. 2007.PubMed/NCBI View Article : Google Scholar
|
|
26
|
Zhu L, Zhou W, Zhu X, Xiang S, Wang S,
Peng Y, Lu B, Tang P, Chen Q, Wu M, et al: Inhibitor of apoptosis
protein-like protein-2: A novel growth accelerator for breast
cancer cells. Oncol Rep. 40:2047–2055. 2018.PubMed/NCBI View Article : Google Scholar
|
|
27
|
Khalili S, Mohammadpour H, Shokrollahi
Barough M and Kokhaei P: ILP-2 modeling and virtual screening of an
FDA-approved library: A possible anticancer therapy. Turk J Med
Sci. 46:1135–1143. 2016.PubMed/NCBI View Article : Google Scholar
|