Introduction
Hepatocellular carcinoma (HCC) is reported as the
sixth most common type of cancer worldwide (1). Patients with HCC generally have a
poor prognosis due to the highly invasive nature of the disease
(2). A large number of studies
have reported that the inactivation of tumor suppressor genes and
the activation of oncogenes is responsible for the metastasis of
HCC (3-5). However, the mechanisms underlying
recurrence and metastasis in HCC have not yet been fully
determined.
Epithelial-mesenchymal transition (EMT),
characterized by the alteration in the cancer cell phenotype from
an epithelial one to a motile mesenchymal one, is a well-known
initiating and essential event in the progression and metastasis of
various types of cancer (6-8).
MicroRNAs (miRNAs or miRs) are a class of long non-coding RNAs
which are 18-24 nucleotides in length. These RNAs may serve as
vital modulators which regulate a number of biological processes in
cancer, including invasion and metastasis (9-11). A
number of miRNAs have been shown to be involved in the metastasis
of HCC, including miRNA-92a, miRNA-7, miRNA-3910, miRNA-1301,
miRNA-203a-3p and miRNA-93 (11-17).
Researchers have focused on the function of miR-103 in
tumorigenesis. miR-103 belongs to the miR-103/107 family, and is
capable of inducing EMT of mammary epithelial cells. It is of
interest that miR-103 seems to play differential roles in various
types of cancer. For example, miR-103 functions as a tumor
suppressor gene in non-small cell lung cancer (18), while in gastric cancer, colorectal
cancer and prostate cancer, miR-103 functions as an oncogene
(19-21). A recent study indicated that
miR-103 promoted HCC cell growth by targeting A kinase anchor
protein 12 (AKAP12) (22).
However, to date, at least to the best of our knowledge, no causal
association has been established between miR-103 and metastasis and
EMT in HCC. Thus, studies focusing on the exact mechanisms of
action of miR-103 in HCC progression are necessary.
Therefore, in the current study, we aimed to
determine the role and mechanism of action of miR-103 in metastasis
and EMT in HCC. For this purpose, assays were performed using human
specimens, HCC cell lines and animals. The findings of this study
may aid in the development of novel targeted therapies for HCC.
Materials and methods
Patients and tissue samples
Tissue samples for this study were collected from
120 patients with HCC (including patients with or without
metastasis) who underwent surgical resection between March, 2010
and March, 2011 at the Second Affiliated Hospital of Xi'an Jiaotong
University (Xi'an, China). The Ethics Committee of the Medical
School of Xi'an Jiaotong University reviewed and approved the
study, and written informed consent was obtained from each
participant at each examination phase. The study complied with the
principles of the Helsinki Declaration. The clinicopathological
characteristics of all the patients are presented in Table I. None of the patients underwent
prior chemotherapy or radiotherapy. Fresh HCC tissues paired with
distant non-cancerous tissues samples (≥2 cm distance from the
tumor margin) were immediately frozen in liquid nitrogen during the
surgical resection. They were stored at −80°C [for use in western
blot analysis or reverse transcription-quantitative PCR (RT-qPCR)
assay] or in paraffin (for use in immunohistochemistry).
 | Table IAssociation between miR-103 and LATS2
expression levels with the clinicopathological characteristic of
patients with hepatocellular carcinoma. |
Table I
Association between miR-103 and LATS2
expression levels with the clinicopathological characteristic of
patients with hepatocellular carcinoma.
| Variable | Total no. of
patients, n=120 (%) | miR-103
| P-value | LATS2
| P-value |
|---|
| Low expression | High
expression | Low expression | High
expression |
|---|
| Age (years) | | | | 0.057 | | | 0.120 |
| <50 | 49 (40.8) | 10 | 39 | | 37 | 12 | |
| ≥50 | 71 (59.2) | 26 | 45 | | 44 | 27 | |
| Sex | | | | 0.496 | | | 0.349 |
| Female | 21 (17.5) | 5 | 16 | | 16 | 5 | |
| Male | 99 (82.5) | 31 | 68 | | 65 | 34 | |
| HBsAg | | | | 0.955 | | | 0.903 |
| Positive | 76 (63.3) | 15 | 61 | | 51 | 25 | |
| Negative | 44 (36.7) | 21 | 23 | | 30 | 14 | |
| AFP (ng/ml) | | | |
0.01a | | | 0.422 |
| <400 | 97 (80.8) | 24 | 73 | | 50 | 27 | |
| ≥400 | 23 (19.2) | 12 | 11 | | 31 | 12 | |
| Cirrhosis | | | | 0.741 | | | 0. 903 |
| Yes | 76 (63.3) | 22 | 54 | | 51 | 25 | |
| No | 44 (36.7) | 14 | 30 | | 30 | 14 | |
| Tumor size
(cm) | | | |
0.007a | | |
0.032a |
| <5 | 48 (40) | 21 | 27 | | 27 | 21 | |
| ≥5 | 72 (60) | 15 | 57 | | 54 | 18 | |
| Tumor
multiplicity | | | | 0.228 | | | 0.992 |
| Single | 77 (64.2) | 26 | 51 | | 52 | 25 | |
| Multiple | 43 (35.8) | 10 | 33 | | 29 | 14 | |
|
Differentiation | | | | 0.318 | | | 0.383 |
| Well-moderate | 53 (44.2) | 17 | 36 | | 38 | 15 | |
|
Poor-undifferentiated | 67 (55.8) | 19 | 48 | | 43 | 24 | |
| Microscopic
vascular invasion | | | |
<0.001b | | |
0.001a |
| Yes | 39 (32.5) | 28 | 11 | | 18 | 21 | |
| No | 81 (67.5) | 8 | 73 | | 63 | 18 | |
| Stage | | | |
<0.001b | | |
0.021a |
| I–II | 78 (65) | 32 | 46 | | 47 | 31 | |
| III–IV (with
metastasis) | 42 (35) | 4 | 38 | | 34 | 8 | |
Immunohistochemistry (IHC)
The specimens were cut into 3-μm-thick
sections and placed on histological slides. The sections were then
deparaffinized, rehydrated and immersed into 3% hydrogen peroxide.
The steps followed for immunohistochemistry and the staining
intensity determined were performed following procedures reported
previously (23). The primary
antibodies used in this study were all specific for
immunohistochemistry, including anti-large tumor suppressor kinase
2 (LATS2) antibody (bs-4081R; at a dilution of 1:50),
anti-E-cadherin antibody (bs-1016R; at a dilution of 1:100) and
anti-vimentin antibody (bs-8533R; at a dilution of 1:100) (all from
Beijing Bioss Biotechnology, Beijing, China). Horseradish
peroxidase (HRP)-labelled anti-rabbit second antibodies were used
in this study (bse-0302R; Beijing Bioss Biotechnology; at a
dilution of 1:6,000). Images were acquired using a Leica TCS SP2
confocal laser-scanning microscope (Leica Microsystems).
RT-qPCR
miR-103 and LATS2 expression in both samples and HCC
cells, and CYR61, AREG, CTGF and CXCL5 expression in the HCC
samples was assessed by RT-qPCR. RT-qPCR was performed using
SYBR-Green (Shanghai GeneCore Biotechnologies Co., Ltd., Shanghai,
China). Firstly, RNeasy reagent (Qiagen, Shanghai, China) was used
to extract total RNA from samples or HCC cells. Subsequently, a
NanoDrop 2000 spectrophotometer (Thermo Fisher Scientific, Waltham,
MA, USA) was used to measure the total RNA concentration. Total RNA
was then reverse transcribed to obtain cDNA for RT-qPCR using a
reverse transcription kit (ABI, Life Technologies, Carlsbad, CA,
USA). The RT-qPCR conditions were as follows: Pre-heating for 10
min at 95°C; repeating 40 cycles at 95°C for 15 sec and 60 sec at
60°C. The specific primers were designed and synthesized by Takara
(Dalian, China), and the sequences for PCR are shown as follows:
miR-103, 5′-CCCGCCAAGCCCTTACC-3′ (forward) and
5′-GCCGTCGGTGATGCTTTTTTGG-3′ (reverse); LATS2,
5′-ACCCCAAAGTTCGGACCTTAT-3′ (forward) and 5′-CAT
TTGCCGGTTCACTTCTGC-3′ (reverse); cysteine-rich angiogenic inducer
(61CYR61), 5′-CCCTGAACTTGTGGATGTCATTG-3′ (forward) and
5′-GTCATGATGATCCAGTCCTGC AAA-3′ (reverse); amphiregulin (AREG),
5′-TGCTGGATTGG ACCTCAATG-3′ (forward) and 5′-TCCCGAGGACGGTTCAC
TAC-3′ (reverse); connective tissue growth factor (CTGF),
5′-GAAAAGAUUCCCACCCAU-3′ (forward) and 5′-AUU GGGUGGGAAUCUUUUC-3′
(reverse); C-X-C motif chemo-kine 5 (CXCL5),
5′-GTTCCATCTCGCCATTCATGC-3′ (forward) and
5′-GCGGCTATGACTGAGGAAGG-3′ (reverse); GAPDH,
5′-AATGGACAACTGGTCGTGGAC-3′ (forward) and
5′-CCCTCCAGGGGATCTGTTTG-3′ (reverse). RT-qPCR was performed in
triplicate. The level of U6 RNA was used as internal control for
miR-103. The level of GAPDH was used as the internal control for
LATS2, Cyr61, AREG, CTGF and CXCL5. The relative mRNA expression
against the GAPDH levels was assessed using 2−ΔΔCq
method (24).
Cell culture and transfection
HCC cell lines (MHCC-97H, MHCC-97L and Hep3B) and
L02 normal liver cells were purchased from ATCC (Manassas, VA, USA)
and cultured in DMEM containing 10% FBS with 100 μg/ml
streptomycin and 100 U/ml penicillin (Sigma-Aldrich, St. Louis, MO,
USA) in an incubator at 37°C with 5% CO2. miR-103 mimic
was purchased from GeneCopoeia (Guangzhou, China). The LATS2 vector
was purchased from Addgene (Cambridge, MA, USA). These vectors or
mimics were transfected into the HCC cells using Lipofectamine
2000. The sequences of the vectors and the mimics were as follows:
LATS2 target vector, TTC ACCTTCCGAAGGTTCT; negative control (NC)
vector, TCG TACTCTCGTCTTCGAT; miR-103 mimic sense, AGCAGCA
UUGUAAGGGCUAUGA and antisense, AUAGCCCUGUAC AAUGCUGCUUU; miR-103 NC
mimic sense, AGCAGCAG UUUAGGGCACUAUGA and antisense, AUAGUGCCCUAA
ACUGCUGCUUU. The transfection concentrations used in this study
were as follows: miR-103 NC mimic and miR-103 mimic, 50 nM; LATS2
NC vector and LATS2 vector, 100 μg/ml. Total RNA and protein
were subsequently collected at 48 h following transfection.
Antibodies and western blot analysis
The cells were lysed in RIPA buffer and the protein
concentrations were determined by BCA assays (Cell Signaling
Technology, Inc., Danvers, MA, USA). The total protein for each
sample was quantified by the Bradford method. Total proteins (25
μg) were resolved by 12% sodium dodecyl
sulfate-polyacrylamide gel electrophoresis and subsequently
transferred onto fluoride membranes. The membranes were then
treated with the primary antibodies followed by blocking in 5%
non-fat milk for 1 h. The primary antibodies used were as follows:
anti-LATS2 antibody (bs-4081R; Beijing Bioss Biotechnology; at a
dilution of 1:500), anti-yes-associated protein (YAP) antibody
(#14074; at a dilution of 1:600), anti-phospho-YAP (ser127)
antibody (#13008; at a dilution of 1:500), anti-vimentin antibody
(#5741; at a dilution of 1:500), anti-E-cadherin antibody (#3195;
at a dilution of 1:300), CYR61 (#14479; at a dilution of 1:300)
(all from Cell Signaling Technology, Inc., Danvers, MA, USA), AREG
(ab99975; at a dilution of 1:300), CTGF (ab6922; at a dilution of
1:300), CXCL5 (ab9802; at a dilution of 1:500) (all from Abcam,
Cambridge, MA, USA) and GAPDH (G5262; Sigma-Aldrich; at a dilution
of 1:3,000). The membranes were then carefully kept in a
refrigerator overnight at 4°C. The membranes were washed and
subsequently incubated with goat anti-rabbit secondary antibodies
for LATS2, YAP, p-YAP (ser127), E-cadherin, vimentin and goat
anti-mouse secondary antibody for GAPDH (anti-rabbit:
bs-40295G-IRDye8; anti-mouse: bs-40296G-IRDye8; both at a dilution
of 1:10,000) (both from Beijing Bioss Biotechnology) at 37°C for 2
h. Finally, the membranes were tested using a bio-imaging system
(DNR Bio-Imaging Systems, Jerusalem, Israel). ImageJ version 1.6.0
software (National Institutes of Health, Bethesda, MD, USA) was
used to quantify the intensity of protein bands and normalized by
GAPDH.
Animal model assay
The animal experiments carried out in this study
were approved by the Experimental Animal Ethical Committee of Xi'an
Jiaotong University. A total of 24 male C57BL/6 mice (weight, 13-15
g; age, 6 weeks; 4 groups, 6 in each group) used in our study were
obtained from SLAC Laboratory Animal Co. (SCXK-2007-004; Shanghai,
China), and raised at 22±2°C with a 12-h light/dark cycle under
pathogen-free environment. All mice were freely accessed autoclaved
standard food and water. One week later, the HCC cells transfected
with LATS2 target vector/miR-103 mimic (1×106) or HCC
cells transfected with negative control (NC) vector/NC mimic
(1×106) were injected into the lateral tail vein of the
nude mice. Another 4 weeks later, the mice were sacrificed. The
number of nodules in the lungs was counted and the tumor tissues
were cleaned with PBS, fixed with 4% paraformaldehyde for 2 h, and
subsequently placed in 20% sucrose solution for 24 h.
Cell migration and invasion assay
(Transwell assay)
For the cell migration assays, 2×104 HCC
cells in 200 μl FBS-free medium were placed in the upper
Transwell chamber of uniformly, while 600 μl of normal
culture medium (with 10% FBS) was placed into the bottom chambers.
Following 48 h of incubation at 37°C in a 5% CO2
incubator, the cells remaining on the membrane in the top chamber
side were carefully wiped off using a cotton swab, while the cells
that had migrated through the membrane into the surface of the
bottom chambers were softly cleaned and fixed with methanol. The
migrated cells were stained with 0.1% crystal violet solution
(Amresco, Solon, OH, USA) for 15 min at room temperature. The
number of migrated cells was carefully counted in 10 random fields
under a microscope (Olympus Corporation, Tokyo, Japan). For the
invasion assay, the same procedures were carried out as described
above, with the exception that the membrane in the top chamber was
coated with 200 mg/ml Matrigel (BD Biosciences, San Jose, CA,
USA).
MTT assay
The HCC cells (500 per well in 100 μl medium)
were added to 96-well plate and incubated at 37°C for 1, 2, 3, 4,
5, 6 and 7 days, respectively. The medium was replaced with new
fresh medium every 48 h in the 2, 3, 4, 5, 6 and 7 days group.
Following incubation for each period, 10 μl MTT solution (5
mg/ml in PBS) were added to each well and maintained at 37°C. After
4 h, the medium was removed, and 200 μl of dimethyl
sulfoxide were then added. Cell viability were read at a 492 nm
wavelength using a plate reader (Bio-Rad Laboratories, Hercules,
CA, USA). The number of repeated tests for each group was
tripled
Target gene prediction for miRNA
miRNA targets were identified using the
bioinformatics software TargetScan Human version 7.1 (www.targetscan.org/vert_71/).
Reporter luciferase assay
HCC control cells and HCC cells were transfected
with miR-103 inhibitor, mimics, GL3-LATS2-3′-UTR WT or
pGL3-LATS2-3′-UTR MUT luciferase reporter vectors (GenePharma Co.,
Ltd., Shanghai, China) using Lipofectamine 3000, according to the
manufacturer's instructions. The cells were seeded into a 24-well
plate. Following transfection for 48 h, all the cells were
collected, and measured using the Dual-Luciferase Assay System
(Promega, Madison, WI, USA). The relative Firefly luciferase
activity was measured by normalizing to Renilla luciferase
activity.
Statistical analysis
All the experiments were in triplicate, and the
results are presented either as the means ± SD or as one
representative experiment. All the statistical analyses were
performed using SPSS 20.0 software for Windows. Chi-square tests
(χ2) were used to assess the association between miR-103
expression and the patient clinicopathological parameters. The
Student's t-test was used to compare the means between 2 groups,
one-way analysis of ANOVA with Dunnett's multiple comparisons test
were used to compare the means among multiple groups. Spearman's
correlation analysis was used to analyze the correlation between
miR-103, and E-cadherin, vimentin and LATS2 expression. The
Kaplan-Meier method (the log-rank test) was used to plot the
survival curves. A two-sided P-value <0.05 was considered to
indicate a statistically significant difference.
Results
Upregulation of miR-103 is associated
with the progression of human HCC
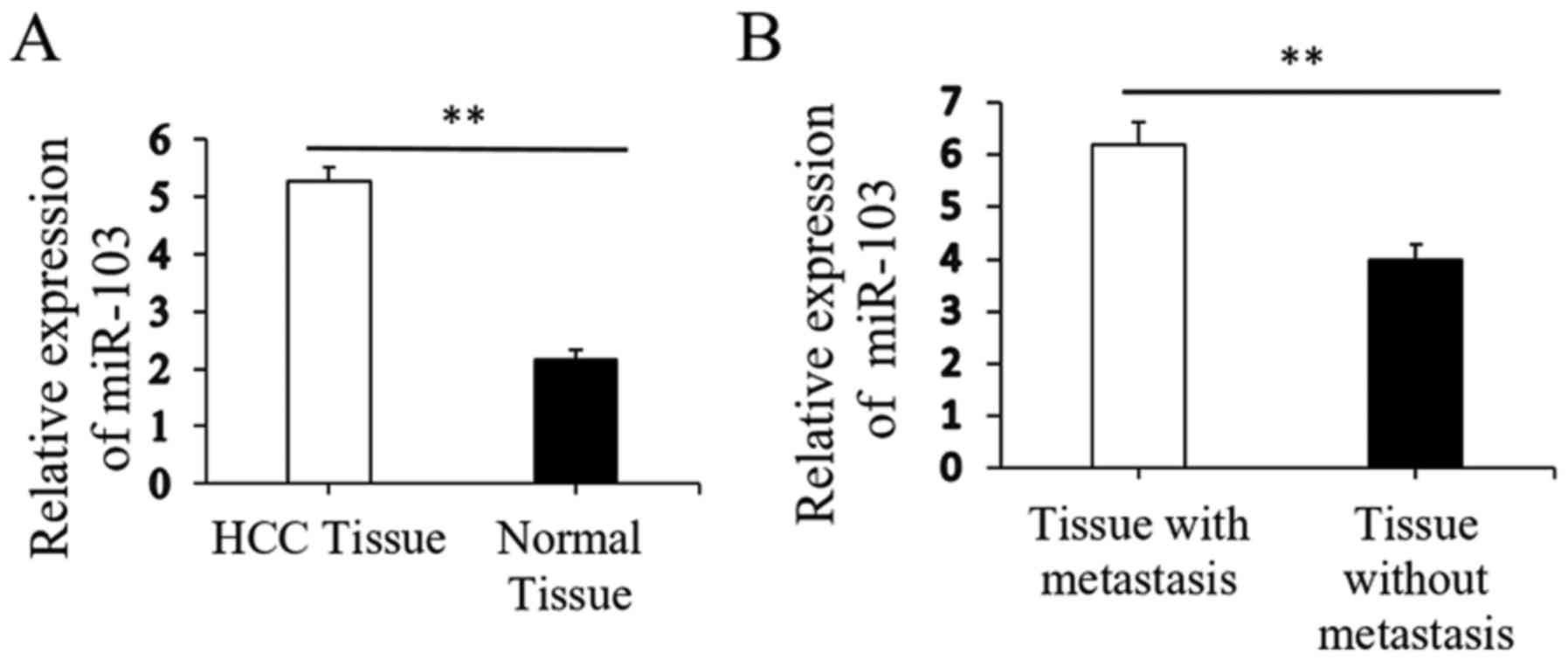
Initial RT-qPCR analysis indicated that the
expression of miR-103 was markedly higher in the HCC tissues than
in the paired distant non-cancerous tissues (5.28±0.23 vs.
2.15±0.19, P<0.01; Fig. 1A).
Further analysis revealed that the miR-103 level in the HCC tissues
from patients with metastasis was significantly increased compared
to that in patients without metastasis (6.19±0.43 vs. 3.98±0.29,
P<0.01; Fig. 1B). The clinical
significance of miR-103 in patients with HCC was then further
evaluated. The miR-103 expression level exhibited a direct
association with microscopic vascular invasion (P<0.001), a high
level of AFP (P=0.01), a larger tumor size (P=0.007) and an
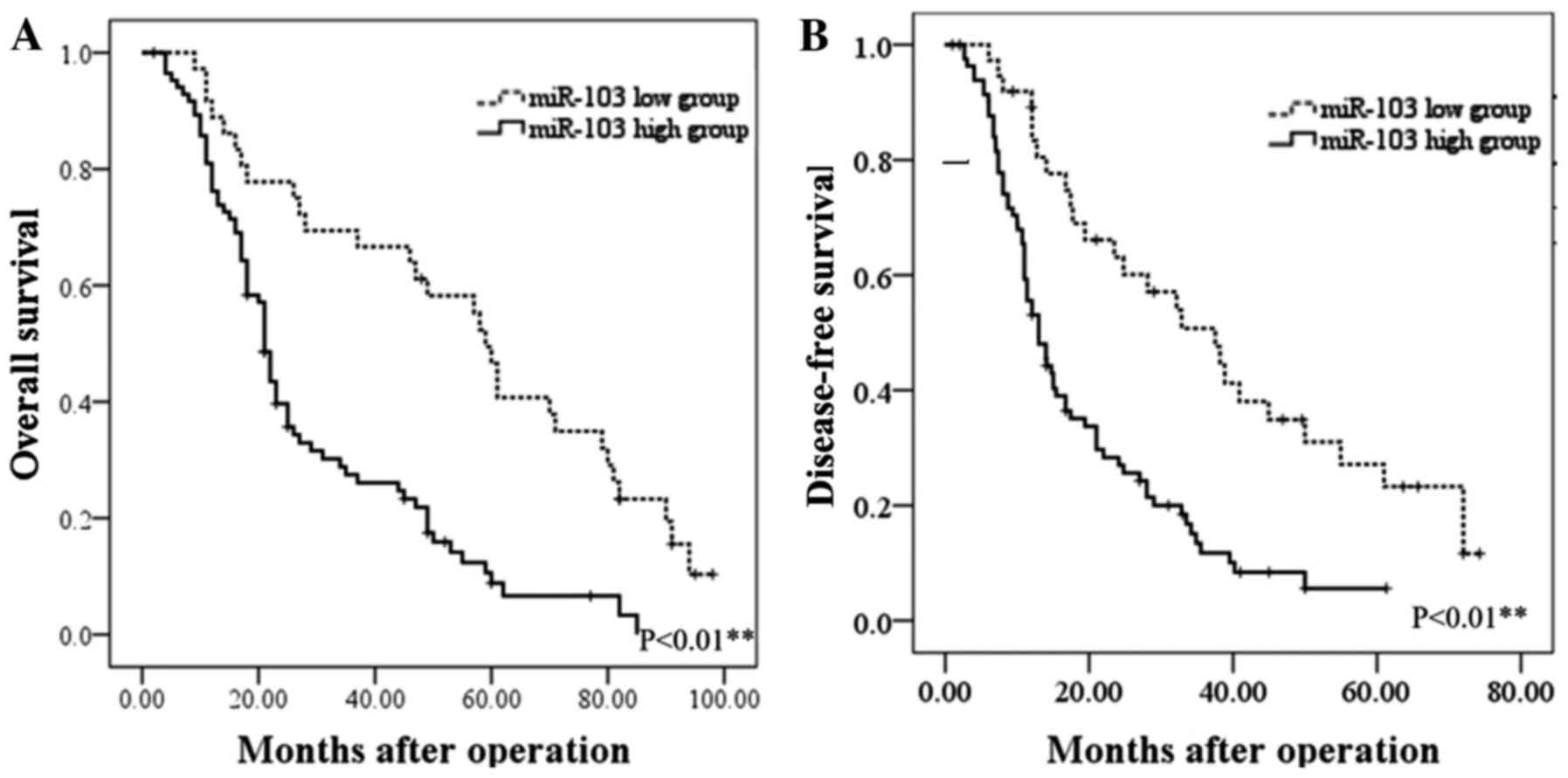
advanced TNM stage in HCC (P<0.001; Table I). Furthermore, the prognostic
impact of the miR-103 level on the survival rate of patients with
HCC was determined. The results revealed that patients with HCC
with a higher level of miR-103 had a shorter overall survival (OS)
and disease-free survival (DFS) than those with a low level
(P<0.01; Fig. 2). These
findings indicate increased miR-103 level is associated with the
metastasis, progression and poor prognosis of HCC.
miR-103 promotes the growth, migratory
and invasive capacity of HCC cells
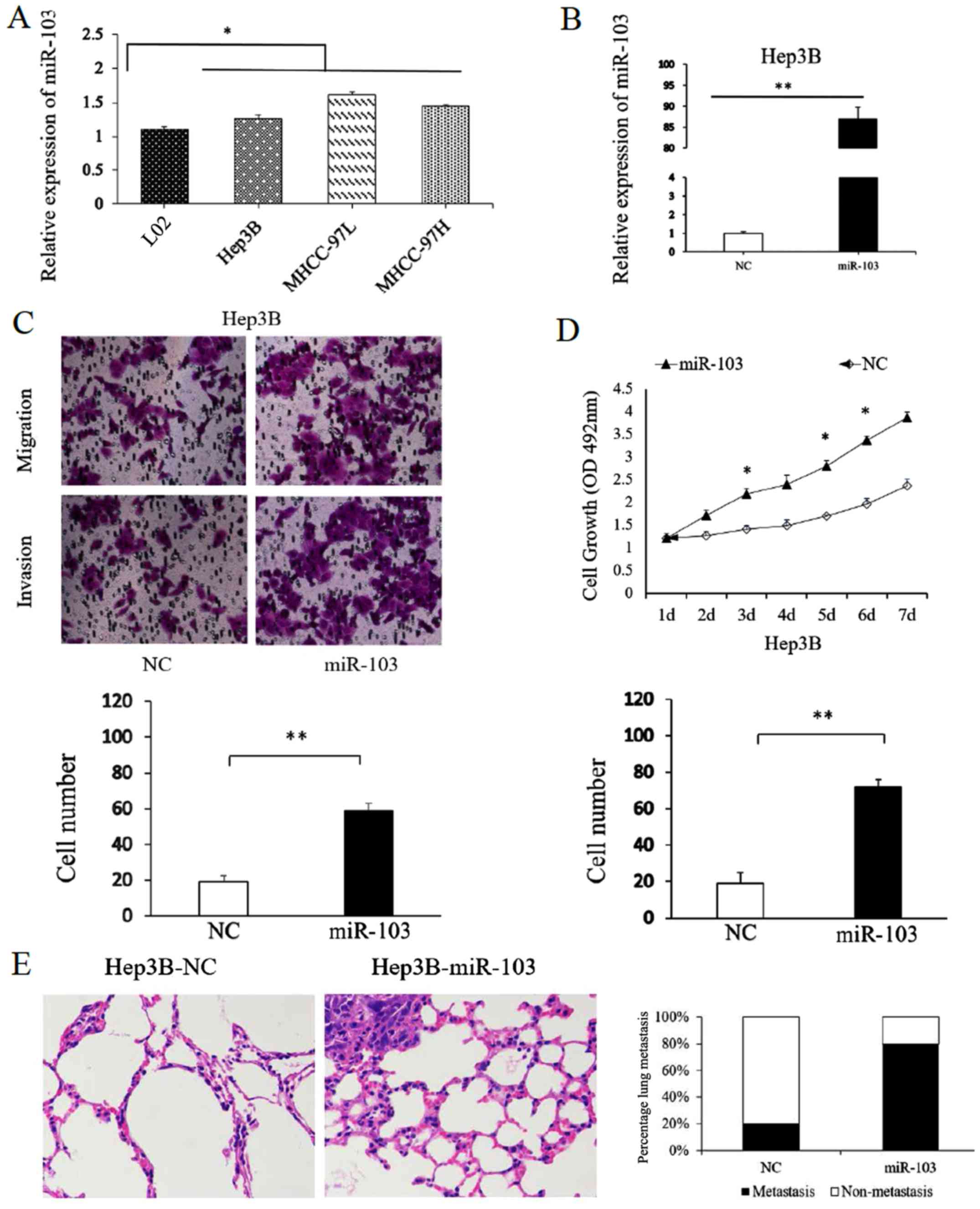
To assess the function of miR-103 in HCC cells, the
miR-103 levels in HCC cell lines and in the normal human liver cell
line, L02, were compared. Consistent with our findings obtained
with the HCC tissues, the results confirmed that miR-103 level was
markedly increased in the HCC cell lines compared with the L02
cells. Among the MHCC-97H, MHCC-97L and Hep3B HCC cell lines, the
miR-103 level was the lowest in the Hep3B cells (Fig. 3A). Thus, the Hep3B cells were
transfected with the miR-103 mimic. Transfection with the miR-103
mimic markedly increased miR-103 expression in the Hep3B cells
(Fig. 3B). The results of
Transwell assay suggested that the migratory and invasive abilities
of the Hep3B cells significantly increased with the upregulation of
miR-103 expression (P<0.01; Fig.
3C). Similarly, the results of MTT assay revealed that the
overexpression of miR-103 significantly increased the proliferative
ability of the Hep3B cells (P<0.05; Fig. 3D).
To further confirm the function of miR-103 in
metastasis of HCC cells in vitro, we also performed tumor
xenograft metastasis assays. The results revealed that the
overexpression of miR-103 significantly increased the lung
metastasis of Hep3B cells (P<0.05; Fig. 3E). These data demonstrate that
miR-103 promotes the metastatic behavior of HCC cells in
vitro and in vivo.
miR-103 promotes the EMT of HCC
cells
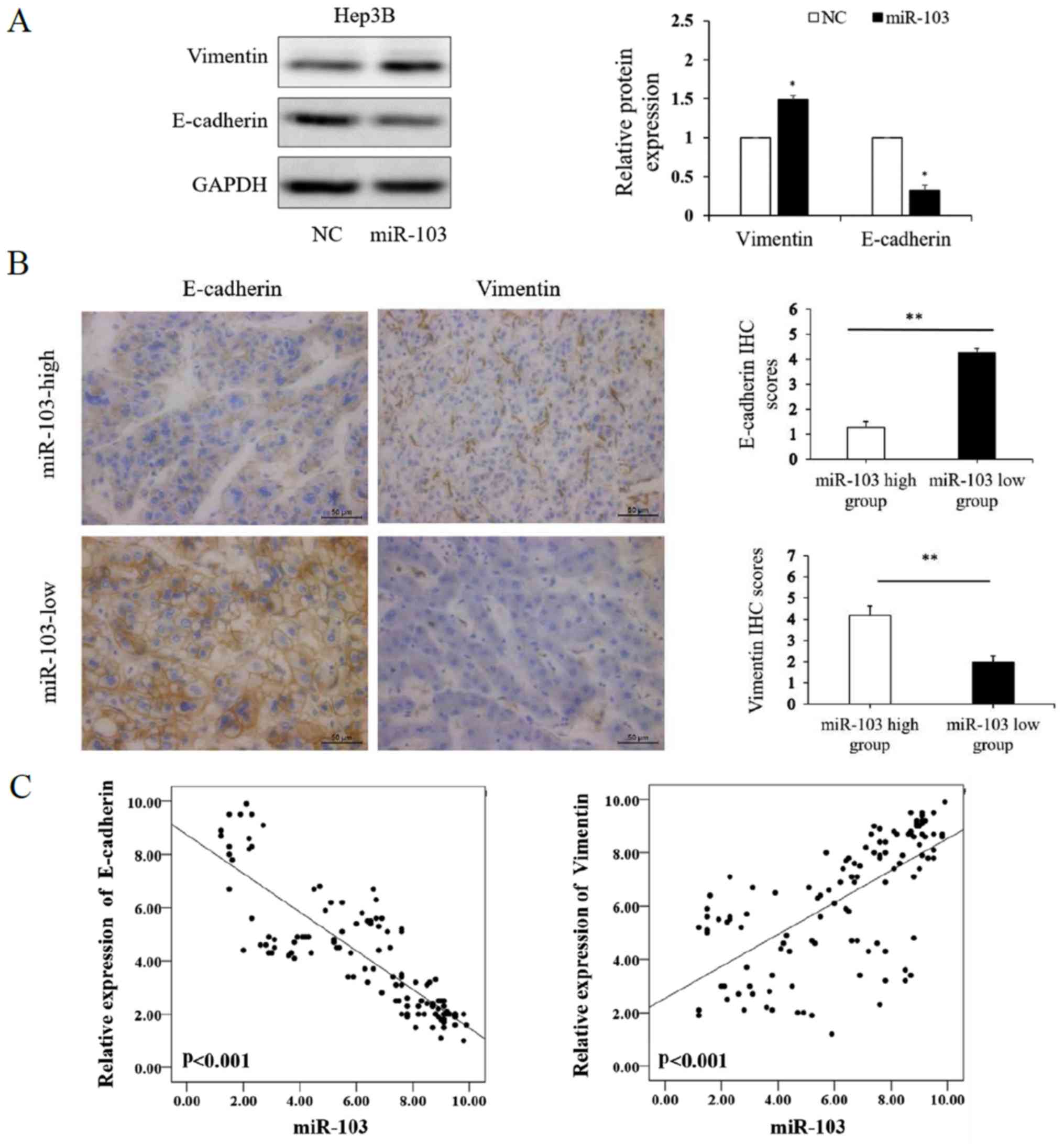
We further investigated whether miR-103 modulates
the EMT of HCC cells. The results of western blot analysis revealed
that the over-expression of miR-103 markedly increased the
expression of vimentin and decreased the expression of E-cadherin
in the Hep3B cells (P<0.01; Fig.
4A). We further compared the expression level of E-cadherin and
vimentin in the HCC specimens with a high miR-103 expression to
those in HCC tissues with a low miR-103 expression by IHC, which
yielded similar results (P<0.01; Fig. 4B). Spearman's correlation analysis
was performed to determine the correlation between the expression
of miR-103, and that of E-cadherin and vimentin derived from
RT-qPCR analysis. The results revealed that a high miR-103
expression was strongly associated with a low E-cadherin expression
(P<0.001, r=−0.864) and high vimentin expression (P<0.001,
r=0.712; Table II and Fig. 4C). These data indicate that miR-103
promotes the invasion, metastasis and EMT of HCC.
 | Table IIAssociation between miR-103 and
EMT-related protein expression levels in hepatocellular carcinoma
tissue specimens. |
Table II
Association between miR-103 and
EMT-related protein expression levels in hepatocellular carcinoma
tissue specimens.
| Variable | miR-103
| r value | P-value |
|---|
| High
expression | Low expression |
|---|
| E-cadherin | | | -0.864 |
P<0.001a |
| High
expression | 16 | 22 | | |
| Low
expression | 68 | 14 | | |
| Vimentin | | | | |
| High
expression | 68 | 15 | 0.712 |
P<0.001a |
| Low
expression | 16 | 21 | | |
LATS2 may serve as a direct target of
miR-103 in HCC
We further accessed the public database, TargetScan
to search for miR-103 target genes. The data indicated that the
complementary sequence of miR-103 was contained in the 3′-UTR of
LATS2 mRNA (Fig. 5A). LATS2 is a
well known pivotal tumor suppressor and a critical regulatory
factor of cancer metastasis (25,26).
Combined with evidence from bioinformatics analysis, we
hypothesized miR-103 may promote EMT by regulating LATS2 in HCC. To
confirm this hypothesis, the mRNA expression levels of LATS2 were
assessed in miR-103- overexpressing HCC cells. The results of
RT-qPCR revealed that the overexpression of miR-103 significantly
decreased the mRNA level of LATS2 in the Hep3B cells (P<0.01;
Fig. 5D, left panel). To confirm
the regulation of LATS2 by miR-103 in HCC, we examined the
expression of LATS2 in HCC tissues. In addition, the results from
IHC revealed that compared with the HCC tissue samples with a low
expression level of miR-103, LATS2 expression was markedly
decreased in the HCC tissues with a high expression of miR-103
(P<0.05; Fig. 5B). The results
of RT-qPCR also revealed that the expression of LATS2 was lower in
the HCC tissues than in the distant non-cancerous tissues
(2.14±0.12 vs. 4.83±0.24, P<0.01; Fig. 5C). In addition, the level of LATS2
in the HCC samples from patients with metastasis was markedly lower
than that in those without metastasis (1.51±0.09 vs. 3.19±0.21,
P<0.01; Fig. 5C). Spearman's
correlation analysis was also performed to examine the correlation
between the expression of miR-103, and that of E-cadherin and
vimentin derived from RT-qPCR analysis. A statistically significant
inverse correlation was observed between the expression of miR-103
and that of LATS2 in the HCC tissues (r=−0.791, P<0.001;
Table III and Fig. 5E). However such a correlation was
not observed between miR-103 and LATS2 expression in the distant
non-cancerous tissues (r=−0.055, P>0.05; Table III and Fig. 5E). On the other hand, we performed
luciferase reporter gene assays to confirm the direct binding
between miR-103 and LATS2. As shown in Fig. 5D (right panel), the overexpression
of miR-103 in the Hep3B cells inhibited the luciferase activity of
LATS2 with a wt 3′-UTR. However, the overexpression of miR-103 did
not regulate the luciferase activity of the mutant (mt) 3′-UTR
LATS2 significantly. These findings obtained from both the clinical
samples and HCC cells demonstrate that LATS2 may act as a direct
target of miR-103 in HCC.
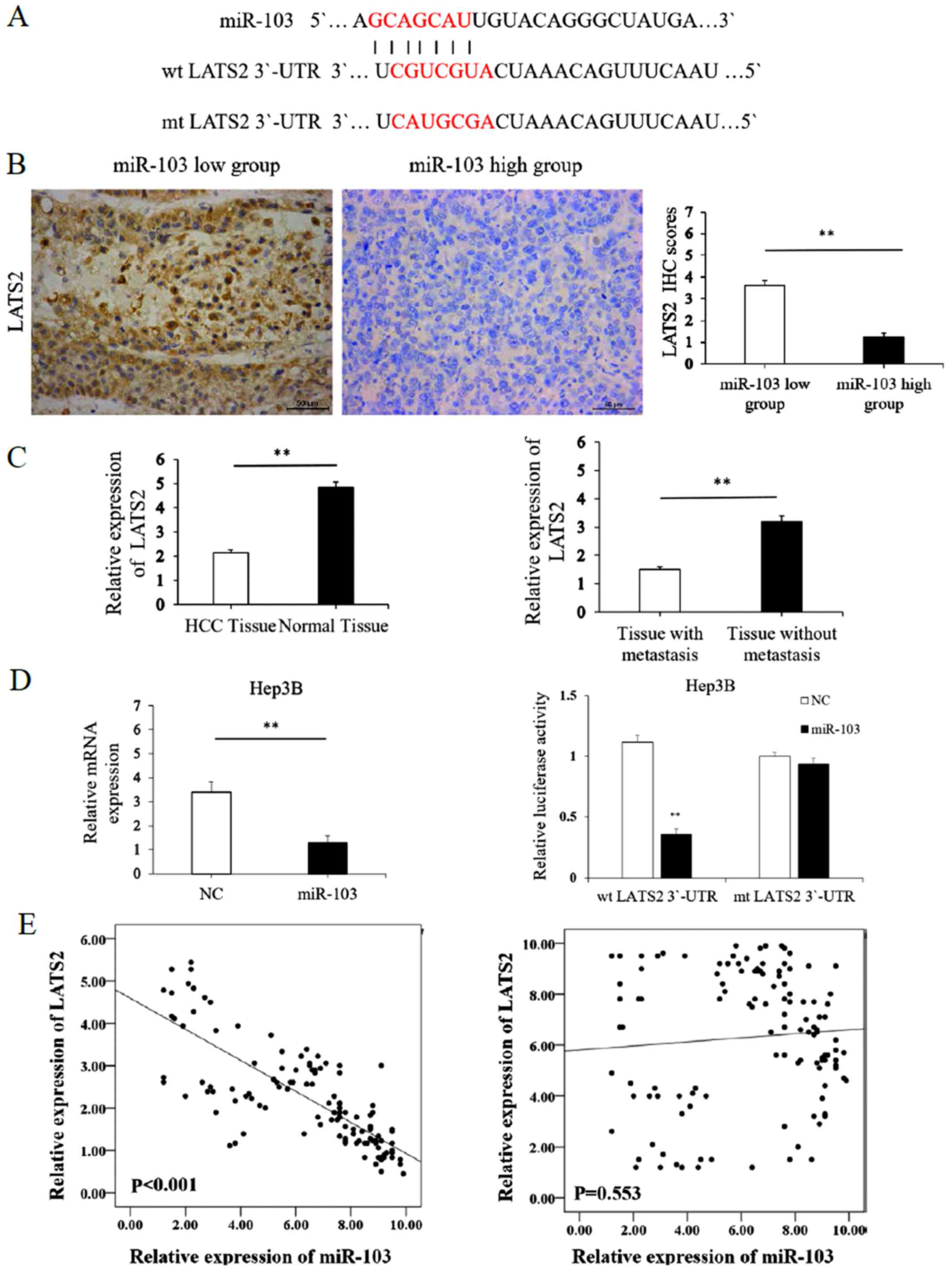 | Figure 5LATS2 acts as a direct downstream
target of miR-103. (A) miR-103 and its putative binding sequence in
the 3′-UTR of LATS2. (B) Immunohistochemistry of LATS2 comparing
tissues with a high miR-103 level and those with a low miR-103
level (**P<0.01; magnification, ×400). (C) Left
panel, relative mRNA expression of LATS2 in HCC compared to distant
non-cancerous HCC tissues; right panel, relative mRNA expression of
LATS2 in HCC tissues from patients with metastasis cases and those
without metastasis cases (**P<0.01). (D) Left panel,
the mRNA expression level of LATS2 was significantly decreased
following transfection of miR-103 mimics into HCC cells
(**P<0.01); right panel, the overexpression of
miR-103 significantly regulated the luciferase activity of cells
that carried the wild-type, but not the mutant type 3′-UTR of LATS2
in HCC cells (**P<0.01). (E) Left panel, miR-103
expression negatively correlates with LATS2 expression in HCC
tissues (Spearman's r=−0.791; see Table III); right panel, no significant
correlation was observed between miR-103 and LATS2 in distant
non-cancerous tissues. HCC, hepatocellular carcinoma; NC, negative
control; LATS2, large tumor suppressor kinase 2. |
 | Table IIIAssociation between miR-103 and LATS2
expression levels in hepatocellular carcinoma tissue specimens. |
Table III
Association between miR-103 and LATS2
expression levels in hepatocellular carcinoma tissue specimens.
| Variable | miR-103
| r value | P-value |
|---|
| High
expression | Low expression |
|---|
| LATS2 | | | | |
| Tumor | | | | |
| High
expression | 15 | 19 | −0.791 |
<0.001a |
| Low
expression | 69 | 17 | | |
| Normal | | | −0.055 | 0.553 |
| High
expression | | | | |
| Low
expression | | | | |
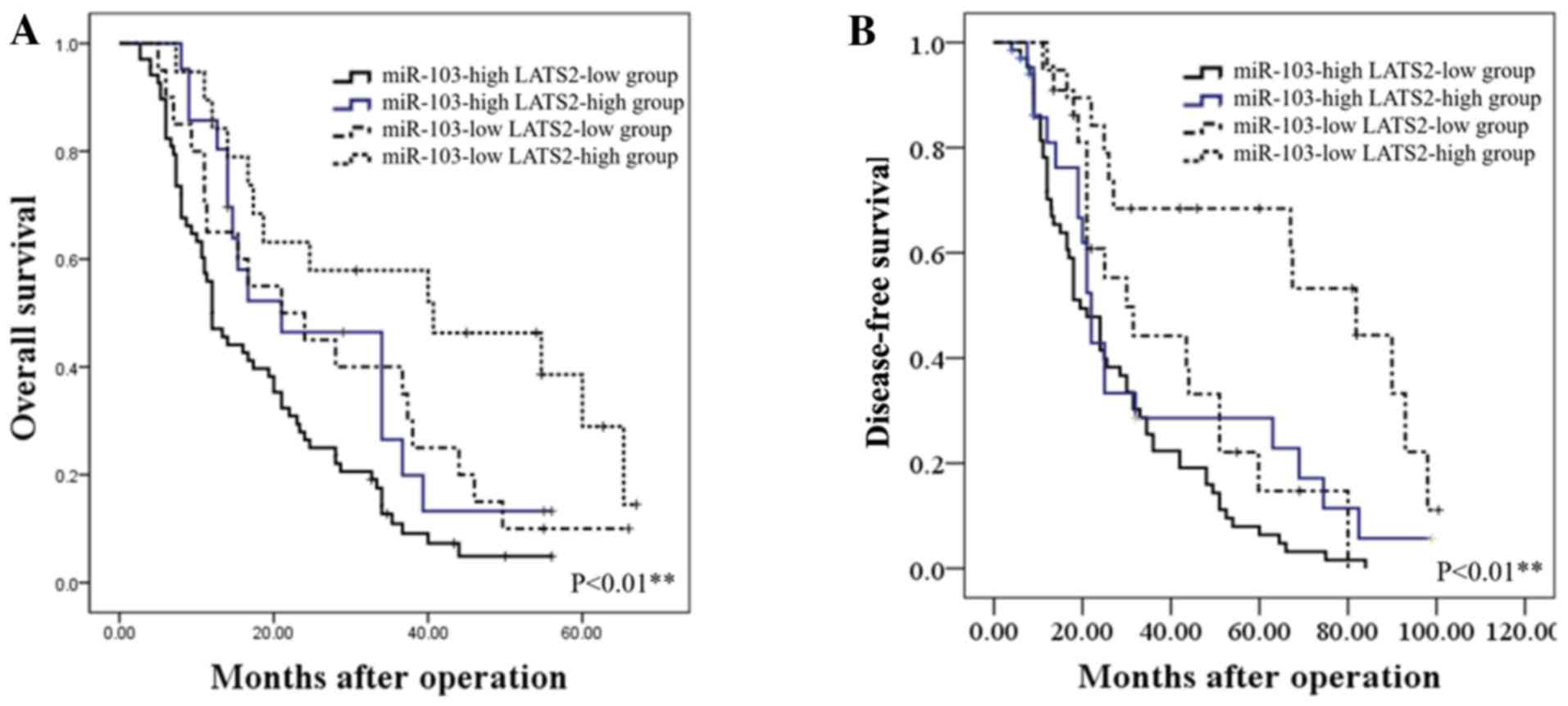
Prognostic significance of miR-103 and
LATS2 in patients with HCC
We further assessed whether the combination of
miR-103 and LATS2 may serve as a prognostic predictor for OS and
DFS in HCC. The combination analysis revealed that patients with a
high miR-103 expression/low LATS2 expression had the lowest OS and
DFS. By contrast, patients with HCC with a low miR-103
expression/high LATS2 expression had the most favorable OS and DFS
(P<0.01; Fig. 6).
miR-103 regulates Hippo-YAP signaling via
LATS2
Previous studies have demonstrated that LATS2 is
related to the Hippo downstream effector, YAP phosphorylation
(27,28). Thus, in this study, to determine
whether miR-103 functions in HCC in association with the Hippo
signaling pathway via LATS2, we assessed the protein level of
LATS2, phosphorylated YAP (p-S127) and YAP. As shown in Fig. 7A, the LATS2 expression level
significantly decreased with the increased miR-103 level in HCC
(P<0.05). Our study here showed increased YAP and decreased
p-YAP (p-S127) because of increased miR-103 (P<0.05; Fig. 7A). The results of western blot
analysis and RT-qPCR further revealed that the downstream target
genes of YAP, including CYR61, AREG, CTGF and CXCL5, were regulated
by miR-103 (Fig. 7). Herein, we
demonstrated that miR-103 promoted the activation of YAP and may
thus serve as an upstream regulator of the Hippo signaling
pathway.
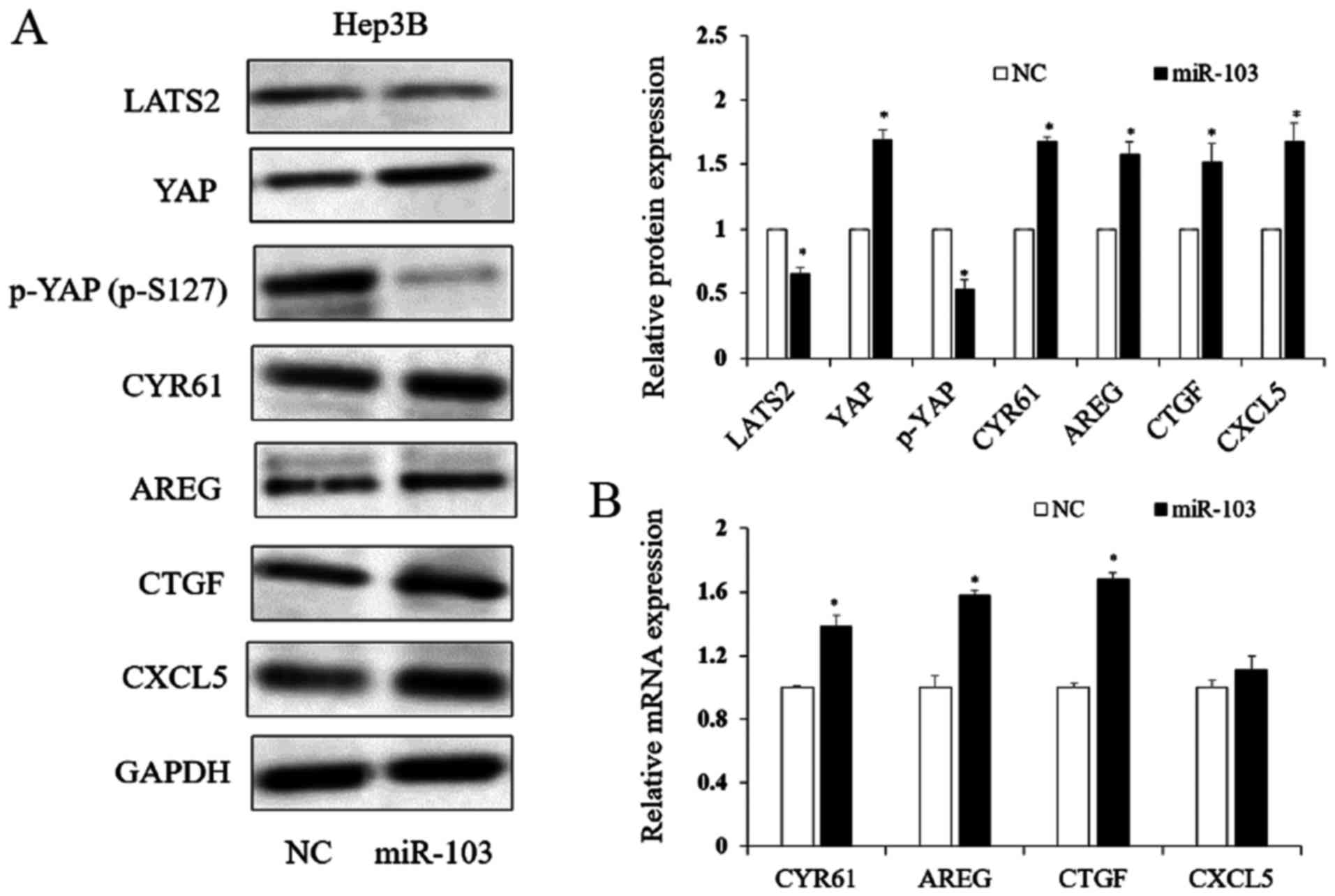 | Figure 7The expression of miR-103 regulates
the LATS2/YAP signaling pathway. (A) miR-103 regulates the protein
level of LATS2, YAP, p-YAP (p-S127), CYR61, AREG, CTGF and CXCL5
(*P<0.05). (B) RT-qPCR analysis suggested that an
increased miR-103 expression regulated the expression level of
downstream of YAP target genes (*P<0.05). LATS2,
large tumor suppressor kinase 2; YAP, yes-associated protein;
CYR61, cysteine-rich angiogenic inducer; AREG, amphiregulin; CTGF,
connective tissue growth factor; CXCL5, C-X-C motif chemokine
5. |
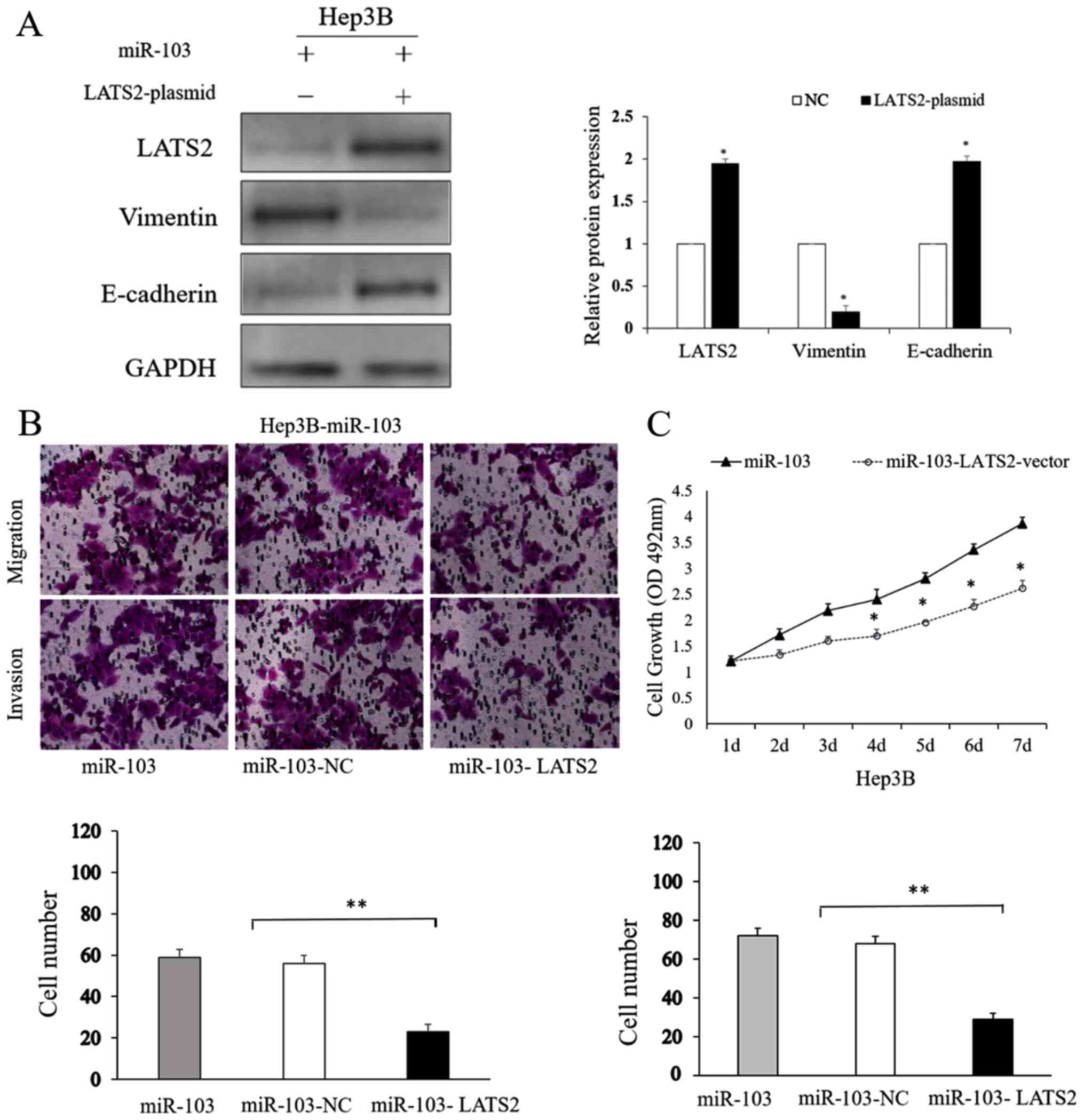
LATS2 counteracts the functional effects
of miR-103 on HCC cells
To further explore the effects of LATS2 on the
process of miR-103-induced metastasis, and on the growth and EMT of
HCC cells, we treated transfected the miR-103-overexpressing Hep3B
cells with a LATS2 expression plasmid. The LATS2 plasmid
upregulated LATS2 expression in the Hep3B cells (P<0.05;
Fig. 8A). The results of western
blot analysis shown in Fig. 8A
indicated that the overexpression of LATS2 reversed the
EMT-promoting effects induced by the overexpression of miR-103 in
HCC cells (P<0.05). The results of Transwell assay also
indicated that LATS2 overexpression attenuated the migratory and
invasive ability of the Hep3B cells which had been promoted by the
overexpression of miR-103 (P<0.01; Fig. 8B). MTT assay showed us altering
LATS2 level could counteract the proliferation capability induced
by miR-103 (P<0.01; Fig. 8C).
On the whole, these results indicate that LATS2 is not only a
direct target of miR-103, but is also an important determinant of
miR-103.
Discussion
In this study, at least to the best of our
knowledge, for the first time, we demonstrate that miR-103 promotes
HCC cell metastasis and EMT by directly targeting LATS2. A recent
study indicated that miR-103 promoted HCC cell growth and that
miR-103 functioned as a novel oncogene in HCC (22); however, the mechanisms involved are
not yet fully clear and further studies to determined the clinical
significance of miR-103 in HCC are warranted. Results from analyses
using clinical tissue samples, in vitro and in vivo,
may provide valuable information regarding the role of miR-103 in
tumorigenesis.
Numerous miRNAs have been confirmed to serve as
critical prognostic biomarkers in human cancers. It has been
demonstrated miR-103 overexpression in gastric cancer tissues is
associated with a prognosis of patients with gastric cancer
(19). In addition, in a previous
study, Kaplan-Meier analysis of colorectal cancer patients
confirmed that an increased miR-103 expression was associated with
a shorter OS (20). The findings
of this study demonstrated that miR-103 expression was
significantly increased in HCC tissues, and was associated with
poor OS of patients with HCC. These data indicate that miR-103 may
function as a promising prognostic marker in HCC.
A number of miRNAs have been confirmed to act as
crucial modulators of not only metastasis (29,30),
but also EMT in HCC (31-33). The findings of this study support
those of previous findings in that miR-103 promotes HCC cell
growth. We further confirmed that an increased miR-103 expression
promoted the migration and EMT of HCC cells. The discovery of the
possible mechanisms of EMT has been recognized as the key for
exploror HCC; thus, the findings of this study may provide
potential targets for clinical interventions.
As an important suppressor of the Hippo signaling
pathway, LATS2 encodes a putative Ser/Thr protein kinase and plays
a key role in oncogenesis by regulating cell proliferation,
apoptosis and EMT. LATS2/YAP has been found to regulate the growth,
proliferation and invasion of tumor cells (34). This study demonstrated that miR-103
directly targeted the 3′-UTR of LATS2, resulting in the
downregulation of LATS2, YAP activation and the regulation of YAP
downstream signaling. These data indicate that miR-103 promotes the
EMT of HCC cells by the direct suppression of LATS2, which is
supported by the fact that the LATS2/YAP signaling pathway is
involved in EMT in human cancers, including HCC (35). Further analyses in this study
demonstrated that the overexpression of LATS2 indeed counteracted
the effects of miR-103 in HCC cells, including the promoting
effects of this miRNA on proliferation, invasion, migration and
EMT.
However, it should be mentioned that LATS2 is not
the sole target of miR103, as several other targets have been
identified in other cancer cells, including A-kinase anchoring
protein 12 (AKAP12), vascular endothelial growth factor (VEGF),
Wnt3a and p57 (36-38). Thus, it is possible that miR-103
can promote the proliferation, migration, invasion and EMT of HCC
cells through other targets. Among these targets of miR-103, the
one that is the most important determinant of the properties of
miR-103 remains unclear. The exact mechanisms underlying the
effects of miR-103 on the progression of HCC warrant further
investigation.
Taken together, the findings of the current study
uncover the biological and clinical significance of miR-103 in HCC.
We identified that miR-103 promotes HCC metastasis and EMT, relying
on the direct suppression of LATS2. These data suggest that the
targeting of the miR-103/LATS2/YAP axis may prove to be a novel
approach for the suppression of the development and metastasis of
HCC.
Acknowledgments
The authors would like to thank Professor Chen Huang
of Xi'an Jiaotong University (Xi'an, China) for providing the
experimental platform and expert opinions.
Funding
This study was supported by grants from the Natural
Science Basic Research Plan in Shaanxi Province of China (no.
1191329734) and China Postdoctoral Science General Financial Grant
(no. 2017M623193).
Availability of data and materials
The datasets used and/or analyzed during the current
study are available from the corresponding authors on request.
Authors' contributions
LLH collected the clinical samples and performed
most of the experiments. XRY collected data from public datasets
and analyzed the data, performed RT-qPCR assay and the statistical
analysis. SQZ conceived and designed the study and assisted in the
drafting of the manuscript. All authors have read and approved the
final manuscript.
Ethics approval and consent to
participate
The Ethics Committee of the Medical School of Xi'an
Jiaotong University reviewed and approved the study, and written
informed consent was obtained from each participant at each
examination phase. The study complied with the principles of the
Helsinki Declaration. The animal experiments carried out in this
study were approved by the Experimental Animal Ethical Committee of
Xi'an Jiaotong University.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Finn RS: Current and future treatment
strategies for patients with advanced hepatocellular carcinoma:
Role of mTOR inhibition. Liver Cancer. 1:247–256. 2012. View Article : Google Scholar
|
|
2
|
Zhang M, Wu R, Jiang J, Minuk GY and Niu
J: The presence of hepatitis B core antibody is associated with
more advanced liver disease in alcoholic patients with cirrhosis.
Alcohol. 47:553–558. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Jin H, Ko YS and Kim HJ: P2Y2R-mediated
inflammasome activation is involved in tumor progression in breast
cancer cells and in radiotherapy-resistant breast cancer. Int J
Oncol. 53:1953–1966. 2018.PubMed/NCBI
|
|
4
|
Sun G, Ding X, Bi N, Wu L, Wang J, Zhang
W, Dong X, Lv N, Song Y, Zhan Q and Wang L: MiR-423-5pin brain
metastasis: Potential role in diagnostics and molecular biology.
Cell Death Dis. 9:9362018. View Article : Google Scholar
|
|
5
|
Zhang J, Mo HQ, Tian FJ, Zeng WH, Liu XR,
Ma XL, Li X, Qin S, Fan CF and Lin Y: EIF5A1 promotes trophoblast
migration and invasion via ARAF-mediated activation of the
integrin/ERK signaling pathway. Cell Death Dis. 9:9262018.
View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Mathias RA, Gopal SK and Simpson RJ:
Contribution of cells undergoing epithelial-mesenchymal transition
to the tumour microenvironment. J Proteomics. 78:545–557. 2013.
View Article : Google Scholar
|
|
7
|
Neelakantan D, Zhou H, Oliphant MUJ, Zhang
X, Simon LM, Henke DM, Shaw CA, Wu MF, Hilsenbeck SG, White LD, et
al: EMT cells increase breast cancer metastasis via paracrine GLI
activation in neighbouring tumour cells. Nat Commun. 8:157732017.
View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Kalluri R and Weinberg RA: The basics of
epithelial-mesenchymal transition. J Clin Invest. 119:1420–1428.
2009. View
Article : Google Scholar : PubMed/NCBI
|
|
9
|
Bartel DP: MicroRNAs: Genomics,
biogenesis, mechanism, and function. Cell. 116:281–297. 2004.
View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Lujambio A and Lowe SW: The microcosmos of
cancer. Nature. 482:347–355. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Zimmerman AL and Wu S: MicroRNAs, cancer
and cancer stem cells. Cancer Lett. 300:10–19. 2011. View Article : Google Scholar
|
|
12
|
Wang L, Wu J and Xie C: miR-92a promotes
hepatocellular carcinoma cells proliferation and invasion by FOXA2
targeting. Iran J Basic Med Sci. 20:783–790. 2017.PubMed/NCBI
|
|
13
|
Kabir TD, Ganda C, Brown R, Beveridge D,
Richardson K, Chaturvedi V, Candy P, Epis M, Wintle L, George J, et
al: A miR-7/GAS6/TYRO3 axis regulates the growth and invasiveness
of sorafenib-resistant cells in human hepatocellular carcinoma.
Hepatology. 67:216–231. 2018. View Article : Google Scholar
|
|
14
|
Cheng L, Wang H and Han S: MiR-3910
promotes the growth and migration of cancer cells in the
progression of hepatocellular carcinoma. Dig Dis Sci. 62:2812–2820.
2017. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Yang C, Xu Y, Cheng F, Hu Y, Yang S, Rao J
and Wang X: miR-1301 inhibits hepatocellular carcinoma cell
migration, invasion, and angiogenesis by decreasing Wnt/β-catenin
signaling through targeting BCL9. Cell Death Dis. 8:e29992017.
View Article : Google Scholar
|
|
16
|
Huo W, Du M, Pan X, Zhu X, Gao Y and Li Z:
miR-203a3p1 targets IL-24 to modulate hepatocellular carcinoma cell
growth and metastasis. FEBS Open Bio. 7:1085–1091. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Ji C, Liu H, Yin Q, Li H and Gao H: miR-93
enhances hepatocellular carcinoma invasion and metastasis by EMT
via targeting PDCD4. Biotechnol Lett. 39:1621–1629. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Yang D, Wang JJ, Li JS and Xu QY: miR-103
functions as a tumor suppressor by directly targeting programmed
cell death 10 in NSCLC. Oncol Res. Jul 21–2017.Epub ahead of print.
View Article : Google Scholar
|
|
19
|
Zheng J, Liu Y, Qiao Y, Zhang L and Lu S:
miR-103 promotes Proliferation and metastasis by targeting KLF4 in
gastric cancer. Int J Mol Sci. 18:E9102017. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Chen HY, Lin YM, Chung HC, Lang YD, Lin
CJ, Huang J, Wang WC, Lin FM, Chen Z, Huang HD, et al: miR-103/107
promote metastasis of colorectal cancer by targeting the metastasis
suppressors DAPK and KLF4. Cancer Res. 72:3631–3641. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Xue D, Zhou C, Lu H, Xu R, Xu X and He X:
LncRNA GAS5 inhibits proliferation and progression of prostate
cancer by targeting miR-103 through AKT/mTOR signaling pathway.
Tumour Biol. 37:142016. View Article : Google Scholar
|
|
22
|
Xia W, Ni J, Zhuang J, Qian L, Wang P and
Wang J: MiR-103 regulates hepatocellular carcinoma growth by
targeting AKAP12. Int J Biochem Cell Biol. 71:1–11. 2016.
View Article : Google Scholar
|
|
23
|
Han LL, Nan HC, Tian T, Guo H, Hu TH, Wang
WJ, Ma JQ, Jiang LL, Guo QQ, Yang CC, et al: Expression and
significance of the novel tumor-suppressor gene SMG-1 in
hepatocellular carcinoma. Oncol Rep. 31:2569–2578. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar
|
|
25
|
Hao Y, Chun A, Cheung K, Rashidi B and
Yang X: Tumor suppressor LATS1 is a negative regulator of oncogene
YAP. J Biol Chem. 283:5496–5509. 2008. View Article : Google Scholar
|
|
26
|
Zhang J, Smolen GA and Haber DA: Negative
regulation of YAP by LATS1 underscores evolutionary conservation of
the Drosophila Hippo pathway. Cancer Res. 68:2789–2794. 2008.
View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Ling HH, Kuo CC, Lin BX, Huang YH and Lin
CW: Elevation of YAP promotes the epithelial-mesenchymal transition
and tumor aggressiveness in colorectal cancer. Exp Cell Res.
350:218–225. 2017. View Article : Google Scholar
|
|
28
|
Yuan Y, Li D, Li H, Wang L, Tian G and
Dong Y: YAP overexpression promotes the epithelial-mesenchymal
transition and chemoresistance in pancreatic cancer cells. Mol Med
Rep. 13:237–242. 2016. View Article : Google Scholar
|
|
29
|
Gong F, Ren P, Zhang Y, Jiang J and Zhang
H: MicroRNAs-491-5psuppresses cell proliferation and invasion by
inhibiting IGF2BP1 in non-small cell lung cancer. Am J Transl Res.
8:485–495. 2016.
|
|
30
|
Fite K and Gomez-Cambronero J:
Down-regulation of MicroRNAs (MiRs) 203, 887, 3619 and 182 prevents
vimentin-triggered, Phospholipase D (PLD)-mediated Cancer Cell
Invasion. J Biol Chem. 291:719–730. 2016. View Article : Google Scholar :
|
|
31
|
Jaca A, Govender P, Locketz M and Naidoo
R: The role of miRNA-21 and epithelial mesenchymal transition (EMT)
process in colorectal cancer. J Clin Pathol. 70:331–356. 2017.
View Article : Google Scholar
|
|
32
|
Tang O, Chen XM, Shen S, Hahn M and
Pollock CA: MiRNA-200b represses transforming growth
factor-β1-induced EMT and fibronectin expression in kidney proximal
tubular cells. Am J Physiol Renal Physiol. 304:F1266–F1273. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Hu J, Shan Z, Hu K, Ren F, Zhang W, Han M,
Li Y, Feng K, Lei L and Feng Y: miRNA-223 inhibits
epithelial-mesenchymal transition in gastric carcinoma cells via
Sp1. Int J Oncol. 49:325–335. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Zhang Y, Hu CF, Chen J, Yan LX, Zeng YX
and Shao JY: LATS2 is de-methylated and overexpressed in
nasopharyngeal carcinoma and predicts poor prognosis. BMC Cancer.
10:5382010. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Wang S, Li H, Wang G, Zhang T, Fu B, Ma M,
Quan Z and Chen G: Yes-associated protein (YAP) expression is
involved in epithelial-mesenchymal transition in hepatocellular
carcinoma. Clin Transl Oncol. 18:172–177. 2016. View Article : Google Scholar
|
|
36
|
Shi FP, Wang XH, Zhang HX, Shang MM, Liu
XX, Sun HM and Song YP: MiR-103 regulates the angiogenesis of
ischemic stroke rats by targeting vascular endothelial growth
factor (VEGF). Iran J Basic Med Sci. 21:318–324. 2018.PubMed/NCBI
|
|
37
|
Zhang Z, Wu S, Muhammad S, Ren Q and Sun
C: miR-103/107 promote ER stress-mediated apoptosis via targeting
the Wnt3a/β-catenin/ATF6 pathway in preadipocytes. J Lipid Res.
59:843–853. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Wang X, Lin Y, Peng L, Sun R, Gong X, Du J
and Zhang X: MicroRNA-103 promotes proliferation and inhibits
apoptosis in spinal osteosarcoma cells by targeting p57. Oncol Res.
26:22018. View Article : Google Scholar
|