Introduction
Non-small-cell lung cancer (NSCLC) may establish an
immunosuppressive tumor microenvironment that is conducive to tumor
growth (1). Natural killer (NK)
cells play a pivotal role in innate immunity and immunological
surveillance (2). Activation of NK
cells partly depends on the interactions between killer-cell
immunoglobulin-like receptors (KIRs) and human leukocyte antigen
(HLA) class I ligands (3). KIRs are
a family of cell surface receptors expressed by NK cells and
certain subpopulations of T lymphocytes. The KIR family consists of
inhibitory as well as activating receptors characterized by both
allelic (high numbers of variants) and haplotypic (different
numbers of genes for inhibitory and activating receptors on
individual chromosomes) polymorphisms (4). The ligands of KIRs are specific
epitopes on HLA class I molecules (5). HLA-C allotypes fall into the C1 or C2
groups (asparagine and lysine, respectively, at position 80) that
are recognized by KIR2DL2/KIR2DL3 and KIR2DL1/KIR2DS1,
respectively. Two previous studies have implicated HLA-C1 and
HLA-C2 as potential ligands for KIR2DS4 (6,7).
KIR3DL1/KIR3DS1 recognizes HLA-Bw4 epitopes of HLA-A and HLA-B
alleles, which are classified according to whether isoleucine or
threonine is present at position 80 (4).
An increased understanding of the complexities of
the biology of the KIR/HLA system has provided opportunities to
leverage NK cell function as a novel avenue of immunotherapy for
cancer (8). Epidemiological studies
have associated particular KIR and HLA genotypes with the
susceptibility to and clinical outcome of leukemia and certain
types of solid tumors (9–11). There is a lack of clinical
information on the role of KIR/HLA interactions in metastatic NSCLC
(mNSCLC). Therefore, the present study was performed to investigate
the effect of KIR/HLA ligand on the prognosis of Chinese Han mNSCLC
patients treated with chemotherapy.
Patients and methods
Patients and follow-up
A total of 70 eligible Chinese Han patients who
attended the Fudan University Shanghai Cancer Center (Shanghai,
China) were enrolled in the present study. Eligible patients were
aged ≥18 years and had pathologically confirmed metastatic or
recurrent NSCLC. Patients were considered ineligible if they had
received previous systemic anticancer therapy <1 year prior, had
severe drug allergies, had an active infection or other serious
diseases or conditions, or were unable to provide consent. Blood
samples and laboratory blood test results were collected prior to
chemotherapy. The performance status of the cancer patients was
defined according to the Eastern Cooperative Oncology Group (ECOG).
Eligible patients received platinum-based first-line chemotherapy.
All the patients were followed up for 2 years after enrollment.
Follow-up data for all patients were obtained from their most
recent medical review, which consisted of a clinical examination
and an assessment of chest X-rays or computed tomography scans.
Progression-free survival (PFS) was defined as the time from the
date of inclusion to the date of disease progression or death,
whichever came first. Overall survival (OS) was defined as the time
from the date of inclusion to the date of death. Data were censored
if patients were non-progressing or alive at the time point of
evaluation. If a patient missed a follow-up or had an unknown event
date, the patient was censored at the last radiological assessment
for PFS and the last contact date for OS. The Ethics Committee of
Fudan University Shanghai Cancer Center approved this study and all
the included patients provided written informed consent.
Sample preparation
Genomic DNA was extracted from peripheral blood
mononuclear cells using the QIAamp DNA Mini kit (Qiagen GmbH,
Hilden, Germany) according to the manufacturer's instructions.
Total RNA was extracted from whole-blood samples with the PAXgeneÔ
Blood RNA system (PreAnalytiX GmbH, Hombrechtikon, Switzerland).
cDNA was then synthesized with the QuantiTect® reverse
transcription kit (Qiagen).
KIR and HLA ligand typing
The presence or absence of 15 human KIR genes plus
two pseudogenes at the level of genomic DNA and mRNA was analyzed
by sequence-specific primer polymerase chain reaction (SSP-PCR)
using the Miltenyi Biotec KIR typing kit (Miltenyi Biotec GmbH,
Bergisch Gladbach, Germany) according to the manufacturer's
protocol. Differentiation between KIR2DS4del (the 22 bp-deleted
form of KIR2DS4) vs. KIR2DS4full (the full-length form of KIR2DS4)
was also possible. The presence or absence of KIR-HLA ligands,
including HLA-CAsn80 (HLA-C1), HLA-CLys80 (HLA-C2),
HLA-BBw4+Thr80 (HLA-Bw4T80), HLA-BBw4+Ile80
(HLA-Bw4I80), HLA-ABw4+ and
HLA-BBw4+Asp77,Thr80 was determined by SSP-PCR using the
Olerup SSP KIR HLA Ligand kit (Olerup SSP, Stockholm, Sweden)
according to the manufacturer's protocol.
Statistical analysis
Differences in categorical variables were analyzed
with the Fisher's exact test. Frequencies of >20% and <80%
for KIRs or HLA ligands were included in the univariate analysis.
Survival curves were estimated using the Kaplan-Meier method and
compared with the log-rank test. Only variables with P<0.1 in
the univariate analysis (Tables II
and III), apart from the epidermal
growth factor receptor gene (EGFR) mutation status (17.1% of
patients not tested), were entered in the multivariate model. The
Cox proportional hazard regression model was applied for
multivariate survival analysis, with the stepwise selection, to
determine independent predictors of survival. The
Benjamini-Hochberg method (12) was
used to perform multitest correction. The contingency analysis was
computed and the Fisher's exact test was used to investigate the
independence between the significant KIR/HLA ligand and other
variables. P<0.05 was considered to indicate statistically
significant differences. All the analyses were conducted using
SAS® 9.4 software (SAS Institute Inc., Cary, NC,
USA).
 | Table II.Association of KIR and HLA class I
ligands with survival prognosis of patients with mNSCLC by
univariate analysis. |
Table II.
Association of KIR and HLA class I
ligands with survival prognosis of patients with mNSCLC by
univariate analysis.
|
| PFS (negative vs.
positive) | OS (negative vs.
positive) |
|---|
|
|
|
|
|---|
| KIR and HLA typing
(n=70) | HR (95% CI) | P-value | HR (95% CI) | P-value |
|---|
| KIR genotype |
|
|
|
|
|
KIR2DL5all | 0.58
(0.32–1.05) | 0.072 | 1.05
(0.54–2.05) | 0.888 |
|
KIR2DL5A | 0.59
(0.32–1.10) | 0.095 | 0.96
(0.49–1.90) | 0.912 |
|
KIR2DL5B | 0.64
(0.35–1.17) | 0.148 | 1.06
(0.54–2.08) | 0.870 |
|
KIR2DS4del | 1.07
(0.62–1.85) | 0.797 | 0.60
(0.31–1.16) | 0.126 |
|
KIR2DS4full | 1.29
(0.69–2.41) | 0.430 | 1.39
(0.66–2.91) | 0.387 |
|
KIR2DS4del+/KIR2DS4full− | 0.97
(0.50–1.91) | 0.935 | 0.60
(0.26–1.37) | 0.227 |
|
KIR2DS4full+/KIR2DS4del− | 1.11
(0.64–1.92) | 0.716 | 1.57
(0.81–3.06) | 0.182 |
|
KIR2DS4ful+/KIR2DS4del+ | 1.20
(0.67–2.16) | 0.540 | 0.85
(0.40–1.82) | 0.673 |
|
KIR3DS1 | 0.70
(0.38–1.28) | 0.243 | 0.99
(0.50–1.95) | 0.979 |
|
KIR2DS1 | 0.56
(0.30–1.05) | 0.071 | 0.93
(0.47–1.85) | 0.843 |
|
KIR2DS2 | 0.80
(0.38–1.69) | 0.557 | 0.89
(0.38–2.11) | 0.795 |
|
KIR2DS3 | 0.58
(0.34–2.14) | 0.736 | 1.48
(0.55–3.97) | 0.436 |
|
KIR2DS5 | 0.60
(0.32–1.14) | 0.118 | 1.12
(0.56–2.26) | 0.744 |
| KIR cDNA type |
|
|
|
|
|
KIR2DL5all | 0.70
(0.39–1.27) | 0.244 | 0.90
(0.45–1.80) | 0.760 |
|
KIR2DS1 | 0.72
(0.39–1.34) | 0.300 | 1.43
(0.62–3.27) | 0.399 |
|
KIR2DS4del | 1.18
(0.64–2.18) | 0.588 | 0.51
(0.26–1.01) | 0.049 |
|
KIR2DS4full | 1.22
(0.67–2.24) | 0.516 | 1.55
(0.77–3.11) | 0.219 |
|
KIR3DL1 | 1.40
(0.80–2.46) | 0.231 | 1.16
(0.58–2.31) | 0.669 |
|
KIR3DS1 | 0.59
(0.33–1.06) | 0.075 | 0.96
(0.48–1.91) | 0.908 |
|
KIR2DS4full+/KIR2DS4del− | 1.19
(0.68–2.09) | 0.540 | 1.84
(0.93–3.66) | 0.082 |
| KIR HLA ligand |
|
|
|
|
|
HLA-C2 | 1.21
(0.67–2.18) | 0.523 | 1.23
(0.59–2.57) | 0.574 |
|
HLA-Bw4T80 | 0.69
(0.39–1.22) | 0.199 | 0.54
(0.28–1.05) | 0.066 |
|
HLA-Bw4I80 | 0.70
(0.39–1.25) | 0.225 | 0.93
(0.43–1.99) | 0.851 |
|
HLA-ABw4 | 1.31
(0.73–2.35) | 0.364 | 1.72
(0.81–3.67) | 0.156 |
| HLA-C
genotype |
|
|
|
|
|
C1C1 | 1.31
(0.69–2.47) | 0.614 | 1.33
(0.60–2.95) | 0.746 |
|
C1C2 | 0.78
(0.24–2.57) |
| 0.86
(0.20–3.62) |
|
|
C2C2 | 0.60
(0.17–2.12) |
| 0.64
(0.14–3.04) |
|
 | Table III.Patient characteristics and their
value in predicting survival by univariate analysis. |
Table III.
Patient characteristics and their
value in predicting survival by univariate analysis.
|
|
|
| OS | PFS |
|---|
|
|
|
|
|
|
|---|
| Variables | Subgroups | No. (%) | Median | P-value | Median | P-value |
|---|
| Age, years | ≤60 | 41 (58.6) | 19.3 | 0.583 | 10.5 | 0.178 |
|
| >60 | 29 (41.4) | 17.5 |
| 4.4 |
|
| Gender | Female | 29 (41.4) | NR | <0.001 | 12.1 | 0.006 |
|
| Male | 41 (58.6) | 11.9 |
| 4.5 |
|
| Smoking status | Never smoker | 32 (45.7) | NR | 0.001 | 10.5 | 0.152 |
|
| Ever smoker | 38 (54.3) | 11.9 |
| 4.4 |
|
| ECOG PS | 0 | 13 (18.8) | 23.2 | 0.073 | 6.5 | <0.001 |
|
| 1 | 49 (71.0) | 19.3 |
| 10.5 |
|
|
| 2/3 | 8 (11.4) | 4.5 |
| 2.2 |
|
| EGFRa | Mutation | 21 (36.2) | NR | <0.001 | 11.6 | 0.082 |
|
| Wildtype | 37 (63.8) | 15.6 |
| 6.3 |
|
|
Histologyb | AD | 58 (86.6) | NR | <0.001 | 10.3 | 0.002 |
|
| SCC | 9 (13.4) | 10.0 |
| 2.8 |
|
| Metastatic
site | Brain/liver | 13 (18.6) | 14.4 | 0.266 | 7.3 | 0.622 |
|
| Others | 57 (81.4) | 21.4 |
| 8.1 |
|
| CRP
(mg/l)d | ≤10 | 45 (67.2) | NR | 0.001 | 10.3 | 0.140 |
|
| >10 | 22 (32.8) | 11.6 |
| 3.5 |
|
Results
Concordance between KIR typing using
DNA and mRNA in NSCLC patients
Raw frequency data of all tested KIRs at the DNA and
mRNA level, as well as HLA ligands, were calculated (data not
shown). The framework genes 3DL3, 3DP1, 2DL4 and 3DL2 were positive
in all samples at the DNA level, while the framework genes 3DL3
(0%) and 3DP1 (4.3%) were absent in nearly all individuals at the
mRNA level. The frequencies of 2DL1, 2DL2, 2DL3, 3DS1, 2DS4full and
2DP1 at the DNA level were similar to those at the mRNA level,
whereas the frequencies of other genes at the DNA level were
significantly higher compared with those at the mRNA level.
Furthermore, the frequencies of the full-length and deleted
versions of KIR2DS4 were analyzed and the difference between the
DNA and mRNA levels was compared (Table
I). The majority of the 70 samples [62 at the mRNA (88.6%) and
65 (92.9%) at the DNA level] from the Chinese Han population were
positive for KIR2DS4. The KIR2DS4del variant was found in 47.1% of
the 70 individuals at the DNA level, similar to what was previously
reported (13). However, at the mRNA
level, the presence of the KIR2DS4del variant decreased to 27.1% in
this population. The ratio of deleted:full-length form of KIR2DS4
was 1:2.3 at the DNA level and 1:3.9 at the mRNA level (Table I).
 | Table I.Frequency of KIR2DS4 alleles at the
mRNA and DNA level. |
Table I.
Frequency of KIR2DS4 alleles at the
mRNA and DNA level.
| Type | 2DS4del | 2DS4full |
2DS4full-positive/2DS4del-positive |
2DS4full-positive/2DS4del-negative |
2DS4full-positive/2DS4del-positive | 2DS4negative | 2DS4dela/2DS4fullb |
|---|
| mRNA (n=70) | 27.1% (19) | 72.9% (51) | 15.7% (11) | 61.4% (43) | 11.4% (8) | 11.4% (8) | 1:3.9 |
| DNA (n=70) | 47.1% (33) | 72.9% (51) | 20.0% (14) | 45.7% (32) | 25.7%
(18) | 7.1%
(5) | 1:2.3 |
Effect of individual KIR and HLA class
I ligands on the prognosis of mNSCLC
The vast majority of the patients (95.7%) were
positive for HLA-C1, a frequency which is similar to that of the
Japanese population (14). Among the
15 types of KIRs and the 6 types of HLA ligands, with the exception
that positive HLA-Bw4I80 expression was associated with a poor
performance status, there were no other significant associations
between KIRs/HLA ligands and patient characteristics (data not
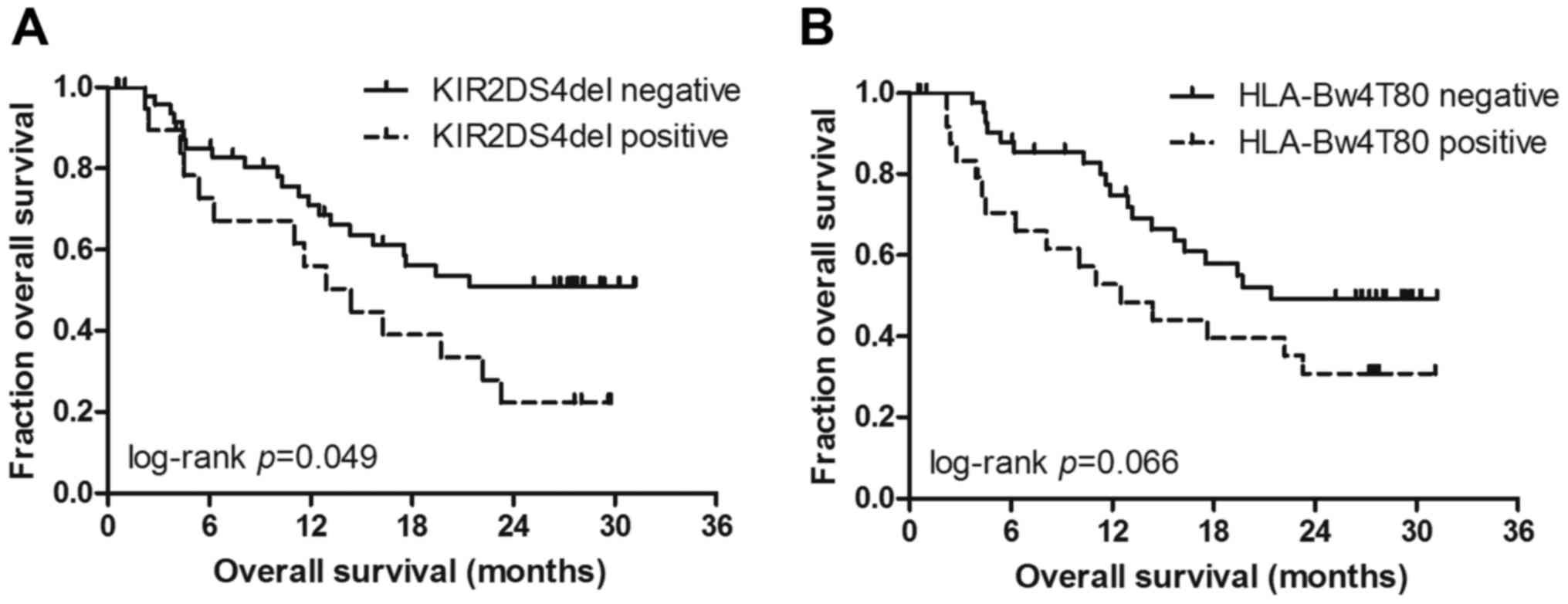
shown). Patients without KIR2DS4del gene expression (Fig. 1A) exhibited a significantly better OS
[hazard ratio (HR)=0.51, P=0.049]. The median OS was not reached in
patients without KIR2DS4del gene expression vs. 14.5 months in
patients with KIR2DS4del gene expression. As regards the HLA
genotype, only HLA-Bw4T80 was found to be associated with decreased
OS at the 10% level (HR=0.51, P=0.066) (Fig. 1B). The median OS was 12.5 months in
the HLA-Bw4T80-positive group vs. 21.4 months in the
HLA-Bw4T80-positive group. No other KIRs or HLA ligands were
significantly associated with clinical outcome according to the
univariate analysis (Table II).
Multivariate analysis considering
patient characteristics
The effect of known clinicopathological confounding
factors [age, gender, smoking status, performance status,
EGFR mutation status, histology, metastatic site and
C-reactive protein (CRP) level] on OS was evaluated by univariate
analysis (Table III). The results
revealed that male gender, adenocarcinoma and a performance status
of 2 were significantly associated with disease progression. Male
gender, adenocarcinoma, smoking, wild-type EGFR status and
CRP level were factors that significantly affected OS. In the
stepwise multivariate Cox model, which includes significant
clinical factors and KIR/HLA ligands at the 10% level, positive
KIR2DS4del gene expression (KIR2DS4del mRNA) (adjusted P=0.014,
HR=3.33), positive HLA-Bw4T80 (adjusted P=0.012, HR=3.58), smoking
(adjusted P=0.02, HR=2.87) and a CRP level >10 mg/l (adjusted
P=0.004, HR=6.48) remained independent predictors of decreased OS;
no KIR/HLA type or clinicopathological variables were selected as
independent predictors of PFS following multitest correction
(Table IV).
 | Table IV.Multivariate Cox analysis of PFS and
OS. |
Table IV.
Multivariate Cox analysis of PFS and
OS.
|
|
| Multivariate
analysis |
|---|
|
|
|
|
|---|
| Variables | Groups | HR (95% CI) | P-value | Adjusted
P-value |
|---|
| OS (n=64) |
|
|
|
|
| CRP,
mg/l | >10 vs. ≤10 | 6.48
(2.11–19.91) | 0.001 | 0.004 |
|
Smoking | Yes vs. no | 2.87
(1.18–6.96) | 0.020 | 0.020 |
|
HLA-Bw4T8 | Positive vs.
negative | 3.58
(1.51–8.47) | 0.004 | 0.012 |
|
KIR2DS4del mRNA | Positive vs.
negative | 3.33
(1.39–7.94) | 0.007 | 0.014 |
| PFS (n=64) |
|
|
|
|
| CRP,
mg/l | >10 vs. ≤10 | 2.94
(1.36–6.37) | 0.006 | 0.054 |
|
Histology | SCC vs. AD | 2.38
(0.95–5.95) | 0.065 | 0.195 |
|
Gender | Male vs.
female | 1.71
(0.84–3.49) | 0.138 | 0.204 |
| ECOG
PS | 2 vs. 0 | 4.20
(1.10–16.09) | 0.036 | 0.160 |
|
HLA-Bw4T8 | Positive vs.
negative | 2.41
(1.21–4.83) | 0.013 | 0.091 |
| KIR2DS1
DNA | Positive vs.
negative | 17.08
(1.28–227.54) | 0.032 | 0.160 |
|
KIR2DL5all DNA | Positive vs.
negative | 34.14
(1.52–765.94) | 0.026 | 0.156 |
|
KIR2DL5A DNA | Positive vs.
negative | 0.047
(0.005–0.461) | 0.047 | 0.072 |
|
KIR2DL5B DNA | Positive vs.
negative | 0.063
(0.002–1.737) | 0.063 | 0.204 |
Discussion
Several studies have investigated the association
between KIR/HLA genotypes and autoimmune disease, infectious
disease, transplantation, stem cell disease and malignancy
(15–17). Regarding malignant diseases, certain
KIR/HLA types have been associated with the susceptibility or worse
clinical outcome in several types of cancer, such as leukemia,
colorectal cancer and lung cancer (18–20).
In lung cancer, Wisniewski et al reported a
possible protective effect of the HLA-C C1C2 genotype on
susceptibility to NSCLC and its association with the
KIR2DL2/KIR2DS2/C1C1 genotype, which was correlated with better
response to therapy and longer survival in the Caucasian population
(20). Due to the ethnic differences
and small sample size, we did not observe these consistent findings
in our samples. In the present study, both cDNA and genomic DNA
were typed, in contrast to other reports where genomic DNA was
used, considering that only expressed genes may play a biological
role and non-expressed KIR gene(s) may exert an effect on a
different, causative gene, due to linkage disequilibrium. We
observed that the majority of KIR genes had a different typing
profile at the DNA and mRNA levels. Furthermore, only KIR cDNA
exerted a significant effect on the outcome of mNSCLC in this study
population following multitest correction. We observed that
KIR2DS4del expression was associated with a significantly decreased
survival time, and KIR2DS4full-positive/KIR2DS4del-negative
genotype was significantly associated with long-term survival
according to the univariate analysis. However, following
multivariate Cox regression analysis and multitest correction, only
KIR2DS4del mRNA expression was considered as an independent factor
associated with OS.
The activating KIR2DS4 receptor has been reported to
bind to group 1 and 2 HLA-C, HLA-A11 and non-HLA class I ligands
(21,22). KIR2DS4del is the mutant form of
KIR2DS4 with a 22-bp deletion in exon 5, resulting in a frameshift
mutation. This truncated KIR2DS4 protein would be secreted due to
the loss of the transmembrane/cytoplasmic domains. Individuals who
are homozygous for haplotype A and with a deletion in both alleles
are likely to have a dysfunctional KIR receptor activator,
considering that a proper allele of KIR2DL4 may function as an
activating receptor. KIR2DS4del frequencies vary among different
populations. In the Caucasian population, the ratio of
full-length:deleted KIR2DS4 is ~1:2. However, in the Chinese,
Japanese and Korean populations, the ratio of KIR2DS4full to
KIR2DS4del is reversed (13), which
is similar to what was observed in the present study.
The associations and functions of KIR2DS4 in
infectious diseases have been extensively investigated. KIR2DS4full
is associated with increased viral loads and transmission rates of
HIV-1 (23,24). Additionally, KIR2DS4full promotes
HIV-1 pathogenesis during chronic infection, probably through the
maintenance of an excessive pro-inflammatory state (25). The combination of KIR2DS4 and
KIR2DS4del was associated with disease progression from hepatitis
to hepatocellular carcinoma (HCC) development via cirrhosis
(26). In terms of malignant
diseases, Giebel et al (27)
reported that the absence of KIR2DS4full was associated with
susceptibility to chronic myeloid leukemia, but the effect of
KIR2DS4full appeared to be the opposite in endometrioid ovarian
cancer (28). Although the
associations of all known 15 human KIRs and their known HLA ligands
with the prognosis of mNSCLC were evaluated in the present study,
we only observed that the KIR2DS4full-positive/KIR2DS4del-negative
type combined with its potential ligand HLA-C1 was significantly
associated with improved OS in the univariate analysis (data not
shown). Almost all patients expressed HLA-C1 (95.7% positive) in
our samples; therefore, it was hypothesized that the improving
effect of KIR2DS4full on OS may be associated with potentiating NK
cell-mediated cytotoxicity and tumor surveillance through the
interaction between HLA-C1 and activating KIR2DS4full.
KIR3DL1 will only inhibit NK cytotoxicity against
target cells expressing a discrete subset of HLA-B and HLA-A
allotypes. In consideration of the clinicopathological variables,
we observed that HLA-Bw4T80 was the only HLA type significantly
affecting OS in multivariate analyses following multitest
correction. This observation is consistent with a previous report
(29), which suggested an
association of HLA-Bw4T80 with decreased OS in patients with HCC.
HLA-Bw4T80 is an allotype of the HLA-Bw4 motif with a threonine at
position 80, which exhibits lower affinity for KIR3DL1 compared
with the HLA-Bw4 motif with an isoleucine at the same position
(30). It is possible that
HLA-Bw4T80 binds with KIR3DL1 to deliver inhibitory signals to NK
cells, which would then enhance the ability of tumor cells to
escape immune detection (31).
However, a positive association of the combination of
KIR3DL1/HLA-Bw4T80 with survival was not observed (data not
shown).
Notably, the KIR/KLA system exhibits extensive
genetic diversity. It would be interesting to perform a similar
analysis of an association of KIR2DS4del mRNA and HLA-Bw4T80 with
clinical outcome in other ethnic populations. To the best of our
knowledge, this is the first study addressing the relevance of KIRs
and HLA ligands regarding the clinical outcome of mNSCLC in a
Chinese population. If confirmed in a larger cohort of patients in
independent studies, KIR2DS4del mRNA and HLA-Bw4T80 may be
prognostic indicators for treatment stratification and patient
management. Furthermore, functional studies will be required to
determine the role of KIR2DS4del mRNA and HLA-Bw4T80 in the immune
response against mNSCLC.
Acknowledgements
The present study was supported by Transgene SA,
France and by grants from the Wu Jieping Medical Foundation (no.
320675014278) and the Shanghai Municipal Commission of Health and
Family Planning (no. 201440423).
Glossary
Abbreviations
Abbreviations:
|
KIR
|
killer-cell immunoglobulin-like
receptor
|
|
HLA
|
human leukocyte antigen
|
|
NK
|
natural killer
|
|
NSCLC
|
non-small-cell lung cancer
|
|
mNSCLC
|
metastatic non-small-cell lung
cancer
|
|
SSP-PCR
|
sequence-specific primer polymerase
chain reaction
|
|
OS
|
overall survival
|
|
PFS
|
progression-free survival
|
|
CRP
|
C-reactive protein
|
References
|
1
|
Carbone DP, Gandara DR, Antonia SJ,
Zielinski C and Paz-Ares L: Non-small-cell lung cancer: Role of the
immune system and potential for immunotherapy. J Thorac Oncol.
10:974–984. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Guillerey C and Smyth MJ: NK cells and
cancer immunoediting. Curr Top Microbiol Immunol. 395:115–145.
2016.PubMed/NCBI
|
|
3
|
Höglund P and Brodin P: Current
perspectives of natural killer cell education by MHC class I
molecules. Nat Rev Immunol. 10:724–734. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Parham P, Norman PJ, Abi-Rached L and
Guethlein LA: Human-specific evolution of killer cell
immunoglobulin-like receptor recognition of major
histocompatibility complex class I molecules. Philos Trans R Soc
Lond B Biol Sci. 367:800–811. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Parham P: MHC class I molecules and KIRs
in human history, health and survival. Nat Rev Immunol. 5:201–214.
2005. View
Article : Google Scholar : PubMed/NCBI
|
|
6
|
Campbell KS, Cella M, Carretero M,
Lopez-Botet M and Colonna M: Signaling through human killer cell
activating receptors triggers tyrosine phosphorylation of an
associated protein complex. Eur J Immunol. 28:599–609. 1998.
View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Katz G, Markel G, Mizrahi S, Arnon TI and
Mandelboim O: Recognition of HLA-Cw4 but not HLA-Cw6 by the NK cell
receptor killer cell Ig-like receptor two-domain short tail number
4. J Immunol. 166:7260–7267. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Benson DM Jr and Caligiuri MA: Killer
immunoglobulin-like receptors and tumor immunity. Cancer Immunol
Res. 2:99–104. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Al Omar S, Middleton D, Marshall E, Porter
D, Xinarianos G, Raji O, Field JK and Christmas SE: Associations
between genes for killer immunoglobulin-like receptors and their
ligands in patients with solid tumors. Hum Immunol. 71:976–981.
2010. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Boyton RJ and Altmann DM: Natural killer
cells, killer immunoglobulin-like receptors and human leucocyte
antigen class I in disease. Clin Exp Immunol. 149:1–8. 2007.
View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Martin MP, Borecki IB, Zhang Z, Nguyen L,
Ma D, Gao X, Qi Y, Carrington M and Rader JS: HLA-Cw group 1
ligands for KIR increase susceptibility to invasive cervical
cancer. Immunogenetics. 62:761–765. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Ferreiraa JA and Nyangomab SO: A
multivariate version of the Benjamini-Hochberg method. Journal of
Multivariate Analysis. 99:2108–2124. 2008. View Article : Google Scholar
|
|
13
|
Bao X, Hou L, Sun A, Qiu Q, Yuan X, Chen
M, Chen Z and He J: Distribution of killer cell immunoglobulin-like
receptor genes and 2DS4 alleles in the Chinese Han population. Hum
Immunol. 71:289–292. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Yawata M, Yawata N, Draghi M, Little AM,
Partheniou F and Parham P: Roles for HLA and KIR polymorphisms in
natural killer cell repertoire selection and modulation of effector
function. J Exp Med. 203:633–645. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Díaz-Peña R, Vidal-Castiñeira JR, Mulero
J, Sánchez A, Queiro R and López-Larrea C: Activating killer
immunoglobulin-like receptors genes are associated with increased
susceptibility to ankylosing spondylitis. Clin Exp Immunol.
180:201–206. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
La Manna G, Corsini S, Iannelli S,
Cappuccilli ML, Comai G, Iorio M, Todeschini P, Carretta E, Scolari
MP, Bontadini A and Stefoni S: Influence of the immunogenetic KIR
and HLA systems on long-term renal transplant outcome. Ann
Transplant. 18:611–621. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Olvera A, Pérez-Álvarez S, Ibarrondo J,
Ganoza C, Lama JR, Lucchetti A, Cate S, Hildebrand W, Bernard N,
Gomez L, et al: The HLA-C*04: 01/KIR2DS4 gene combination and human
leukocyte antigen alleles with high population frequency drive rate
of HIV disease progression. AIDS. 29:507–517. 2015.PubMed/NCBI
|
|
18
|
Babor F, Manser AR, Fischer JC,
Scherenschlich N, Enczmann J, Chazara O, Moffett A, Borkhardt A,
Meisel R and Uhrberg M: KIR ligand C2 is associated with increased
susceptibility to childhood ALL and confers an elevated risk for
late relapse. Blood. 124:2248–2251. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
De Re V, Caggiari L, De Zorzi M, Talamini
R, Racanelli V, Da M, Buonadonna A, Zagonel V, Cecchin E, Innocenti
F and Toffoli G: Genetic diversity of the KIR/HLA system and
outcome of patients with metastatic colorectal cancer treated with
chemotherapy. PLoS One. 9:e849402014. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Wisniewski A, Jankowska R,
Passowicz-Muszyńska E, Wiśniewska E, Majorczyk E, Nowak I, Frydecka
I and Kuśnierczyk P: KIR2DL2/S2 and HLA-C C1C1 genotype is
associated with better response to treatment and prolonged survival
of patients with non-small cell lung cancer in a Polish Caucasian
population. Hum Immunol. 73:927–931. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Graef T, Moesta AK, Norman PJ, Abi-Rached
L, Vago L, Aguilar AM Older, Gleimer M, Hammond JA, Guethlein LA,
Bushnell DA, et al: KIR2DS4 is a product of gene conversion with
KIR3DL2 that introduced specificity for HLA-A*11 while diminishing
avidity for HLA-C. J Exp Med. 206:2557–2572. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Katz G, Gazit R, Arnon TI, Gonen-Gross T,
Tarcic G, Markel G, Gruda R, Achdout H, Drize O, Merims S and
Mandelboim O: MHC class I-independent recognition of NK-activating
receptor KIR2DS4. J Immunol. 173:1819–1825. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Hong HA, Paximadis M, Gray GE, Kuhn L and
Tiemessen CT: KIR2DS4 allelic variants: Differential effects on in
utero and intrapartum HIV-1 mother-to-child transmission. Clin
Immunol. 149:498–508. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Merino A, Malhotra R, Morton M, Mulenga J,
Allen S, Hunter E, Tang J and Kaslow RA: Impact of a functional
KIR2DS4 allele on heterosexual HIV-1 transmission among discordant
Zambian couples. J Infect Dis. 203:487–495. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Merino AM, Dugast AS, Wilson CM, Goepfert
PA, Alter G, Kaslow RA and Tang J: KIR2DS4 promotes HIV-1
pathogenesis: New evidence from analyses of immunogenetic data and
natural killer cell function. PLoS One. 9:e993532014. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Pan N, Jiang W, Sun H, Miao F, Qiu J, Jin
H, Xu J, Shi Q, Xie W and Zhang J: KIR and HLA loci are associated
with hepatocellular carcinoma development in patients with
hepatitis B virus infection: A case-control study. PLoS One.
6:e256822011. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Giebel S, Nowak I, Wojnar J,
Krawczyk-Kulis M, Holowiecki J, Kyrcz-Krzemien S and Kusnierczyk P:
Association of KIR2DS4 and its variant KIR1D with leukemia.
Leukemia. 22:2129–2131. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Giebel S, Boratyn-Nowicka A, Karabon L,
Jedynak A, Pamula-Pilat J, Tecza K, Kula D, Kowal M, Frydecka I and
Grzybowska E: Associations between genes for killer
immunoglobulin-like receptors and their ligands in patients with
epithelial ovarian cancer. Hum Immunol. 75:508–513. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Cariani E, Pilli M, Zerbini A, Rota C,
Olivani A, Zanelli P, Zanetti A, Trenti T, Ferrari C and Missale G:
HLA and killer immunoglobulin-like receptor genes as outcome
predictors of hepatitis C virus-related hepatocellular carcinoma.
Clin Cancer Res. 19:5465–5473. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Purdy AK and Campbell KS: Natural killer
cells and cancer: Regulation by the killer cell Ig-like receptors
(KIR). Cancer Biol Ther. 8:2211–2220. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
de Smith AJ, Walsh KM, Ladner MB, Zhang S,
Xiao C, Cohen F, Moore TB, Chokkalingam AP, Metayer C, Buffler PA,
et al: The role of KIR genes and their cognate HLA class I ligands
in childhood acute lymphoblastic leukemia. Blood. 123:2497–2503.
2014. View Article : Google Scholar : PubMed/NCBI
|