Introduction
Trisomy 21 (Down syndrome) is the most common cause
of prenatal chromosomal abnormalities (1), with an incidence of 1/800-1/600
pregnancies (2). Thus, prenatal
screening for trisomy 21 is extremely important in order to
decrease the prevalence of Down syndrome births. Over that past few
years, second-trimester maternal serum screening for trisomy 21 has
further developed, and a number of serum biochemical markers have
been reported in the screening for Down syndrome, such as
α-fetoprotein (AFP), unconjugated oestriol (uE3), maternal serum
human chorionic gonadotropin (HCG), free β-HCG and inhibin-A
(3). Common dual serological
indicators include AFP combined with total or free β-HCG (4–5), and the
detection rate of AFP combined with total HCG is ~56%, with a 4.9%
false-positive rate (6). The
combination of AFP, uE3 and HCG is the common triple test (7). A recent report revealed that, with a 5%
false-positive rate, the detection rate of the triple test was
66.7%, whereas it was only 50% with AFP and free β-HCG (8), demonstrating that the triple test
achieved a higher detection rate compared with dual serum marker
testing. Quadruple testing markers include AFP, β-HCG (total or
free), uE3 and inhibin-A. The combined detection of different serum
markers or different detection methods may yield different results.
It has been demonstrated that second-trimester screening quadruple
testing with AFP, HCG, uE3 and inhibin-A had a higher detection
rate for trisomy 21 compared with a dual or triple test. The
authors also reported that the detection of two or three indicators
(AFP, uE3, total or free β-HCG) was better compared with that of
any single indicator, and the detection rate was 60–70%, with a
false-positive rate of 5%. In addition, it was demonstrated that
the quadruple testing with AFP, HCG, uE3 and inhibin-A had the
highest diagnostic yield, but there was no significant advantage
when compared with the traditional triple test (AFP, uE3 and HCG)
(3). It has been widely recognized
that different serum markers, cut-off values of risks, or the
determination of multiples of the median, may all affect the
detection rate. For second-trimester screening, the detection of
AFP and free β-HCG in maternal serum has been selected for several
years in the Center of Prenatal Diagnosis, Obstetrics and
Gynecology hospital Affiliated to Nanjing Medical University, after
balancing the benefit against the significantly higher cost of the
triple screening test. The aim of the present study was to
summarize our results on the in-depth screening for trisomy 21.
Through extensive research, it has been demonstrated
that cell-free fetal DNA is present in the maternal serum (9); based on that finding, non-invasive
prenatal testing (NIPT) for aneuploidy of chromosomes 21, 13, 18, X
and Y rapidly developed (10) and
has attracted significant attention. Regarding the difference
between fetal and maternal DNA fragments, NIPT may detect the
number of fetal DNA fragments; when trisomy 21 occurs, the
difference between the normal and abnormal number of fetal DNA
fragments may be significant and trisomy 21 may be identified based
on that difference (11). NIPT has a
high sensitivity (100%) for Down syndrome, and a 99.7% specificity
by multiplexed massively parallel shotgun sequencing (12). NIPT may help avoid the risks
associated with chorionic villus sampling and amniocentesis, such
as abortion and premature delivery (13,14).
Furthermore, NIPT may offer an opportunity for the prenatal
treatment of Down syndrome (15).
Although non-invasive prenatal diagnosis may have certain
advantages, it cannot confirm the presence of chromosomal
abnormalities, as possible chromosomal mosaicism may cause
false-positive results (16); thus,
for patients with positive results, amniocentesis is required.
Materials and methods
Population selection and gestational
age calculation
A total of 221,288 normal singleton pregnancies who
underwent second-trimester screening at the Center of Prenatal
Diagnosis (Obstetrics and Gynecology Hospital Affiliated to Nanjing
Medical University, Nanjing, China) between October 2004 and
October 2013 were included in the present study, and blood samples
were collected from each patient at 15+0 and
20+6 weeks of gestation. If the menstrual cycle was
regular, gestational age was estimated from the date of the last
menstruation; if not, gestational age was calculated by type B
ultrasonic testing.
Quality control
Maternal venous blood samples were collected and
centrifuged at 4,000 × g for 3 min, then stored at −20°C until
detection. AFP and free β-HCG in the serum were detected using a
Wallac AutoDELFIA® hAFP/Free hCGβ Dual kit (PerkinElmer, Turku,
Finland). In order to guarantee the reliability of the experiment,
control serum samples (Hangzhou Biosan Biological Technology Co.,
Ltd., Hangzhou, China), including low, median and high
concentrations, were processed along with the serum specimens.
Assessment of median and risk
value
The calculation method of gestational age was as
mentioned above. First, the original medians of each gestational
week were obtained. Then, the optimal curve regression of this set
of data was selected, and the best fitted curve was obtained.
Finally, the regressive median for each week after regression was
calculated. Comparisons between our median and the corresponding
medians provided by 2T software of the same gestational week were
performed.
For risk calculation, medians of AFP or free β-HCG
were converted into multiple of the median (MoM) for gestational
age, and adjusted by maternal weight. The 2T risk analysis software
was applied (the medians were the original presented), and a risk
value >1/300 was considered to be positive.
Confirmation of Down syndrome
For pregnant women with high risk (≥1/300), advanced
age (≥35 years), an extremely high free β-HCG value (MoM ≥10), or
abnormal ultrasound findings, the results were confirmed by
chromosomal analysis via amniocentesis or umbilical cord blood
sampling (cordocentesis). All Down syndrome cases were confirmed by
amniocentesis or cordocentesis, and all screened subjects received
telephone follow-ups; however, some of the subjects could not be
contacted. The number of residual cases (low- and intermediate-risk
cases that resulted in Down syndrome births) in 1/1,000-1/300 and
<1/1,000 were compared, in order to demonstrate the significance
of recommending intermediate-risk cases for further
examination.
Statistical analysis
DataFit software (http://www.oakdaleengr.com/index.html) was used for
curve regression analysis, and SPSS 17.0 software (SPSS, Inc.,
Chicago, IL, USA) was also applied to construct or compare charts.
Mann-Whitney U tests were applied to compare the median difference
of each gestational week (from week 15 to 20); Chi-square tests
were also used and P<0.05 was considered to indicate
statistically significant differences.
Results
The medians of AFP and free β-HCG for
each region were statistically significant when compared with
medians embedded in the 2T software
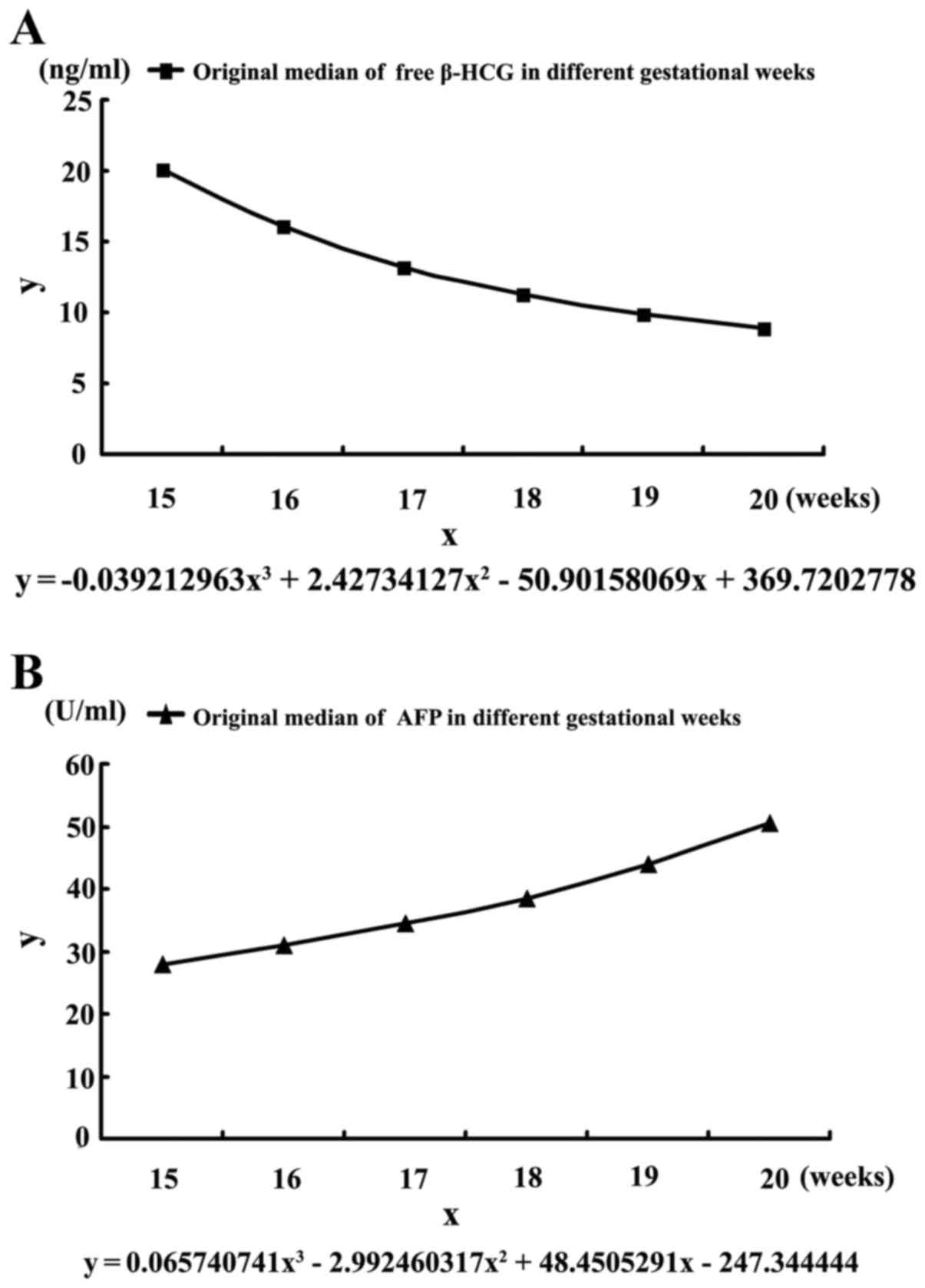
In order to obtain the regressive medians of free
β-HCG and AFP in Nanjing, curve regression analysis was performed,
and the best curve regression equations among these were obtained.
The equation for free β-HCG was y=−0.039212963x3 +
2.42734127x2 − 50.90158069x + 369.7202778 (Fig. 1A, R2=0.999); and the curve
equation of AFP was y=0.065740741x3 −
2.992460317x2 + 48.4505291x − 247.344444 (Fig. 1B, R2=0.999). Then, the
regressive medians of 15–20 weeks were obtained. The Mann-Whitney U
test demonstrated that the medians of free β-HCG each week in our
region were higher compared with the data provided by the 2T
software (P<0.01). In terms of AFP, it was observed that from
15–17 weeks, the medians of AFP in our data were higher compared
with those provided by the 2T software. However, after 18–20 weeks,
an opposite trend was observed (P<0.01, Table I). All the abovementioned results
revealed that the differences in the medians of free β-HCG and AFP
between our region and the corresponding medians in the 2T
software, which were obtained from Caucasians, were statistically
significant, indicating that our own medians of free β-HCG and AFP
must be set up.
 | Table I.Median comparisons of free β-HCG and
AFP between data of our hospital and data embedded in the 2T
software. |
Table I.
Median comparisons of free β-HCG and
AFP between data of our hospital and data embedded in the 2T
software.
|
|
| Free β-HCG
(ng/ml) | AFP (U/ml) |
|---|
|
|
|
|
|
|---|
| Gestational week | Cases (n) | Original median | Median of the
original median predicted | Median embedded in
the 2T software | Original median | Median of the
original median predicted | Median embedded in
the 2T software |
|---|
| 15 | 5,955 | 20.0 | 20.0 | 19.0 | 28.0 | 28.0 | 27.6 |
| 16 | 42,410 | 16.1 | 16.1 | 15.4 | 31.0 | 31.0 | 30.6 |
| 17 | 80,045 | 13.2 | 13.2 | 13.0 | 34.6 | 34.5 | 34.2 |
| 18 | 58,758 | 11.3 | 11.2 | 10.8 | 38.5 | 38.6 | 39.3 |
| 19 | 23,966 | 9.9 | 9.9 | 8.9 | 43.9 | 43.9 | 45.8 |
| 20 | 10,036 | 8.9 | 8.9 | 8.0 | 50.6 | 50.6 | 50.8 |
Screening analysis of the combined
detection of AFP and free β-HCG
In order to determine the detection rate, the cases
with confirmed Down syndrome were analyzed. Among the 221,288
screened pregnancies, 118 had Down syndrome. The detection rate and
false-positive rate varied with different cut-off values (Table II). When the cut-off value was set
at 1/270, the detection rate was 59.3% and the false-positive rate
was 4.43%. However, if the cut-off value was set at 1/300, the
detection rate was 66.1% and the false-positive rate was 5.22%.
Furthermore, when the cut-off value was set at 1/1,000, the
detection rate and false-positive rate was 90.6 and 19.21%,
respectively.
 | Table II.DR and FPR for DS at different cut-off
values. |
Table II.
DR and FPR for DS at different cut-off
values.
| Cut-off value | DR for DS (%) | FPR for DS (%) |
|---|
| 1/200 | 51.7 | 2.72 |
| 1/250 | 55.9 | 3.94 |
| 1/270 | 59.3 | 4.43 |
| 1/280 | 63.6 | 5.05 |
| 1/300 | 66.1 | 5.22 |
| 1/350 | 73.7 | 6.36 |
| 1/500 | 78.8 | 9.61 |
| 1/1,000 | 90.6 | 19.21 |
| 1/1,500 | 91.5 | 26.74 |
Analysis of the impact of different
medians on the detection rate and false-positive rate of Down
syndrome
Our previous conclusion revealed a significant
difference in medians between those embedded in the 2T software and
those of our laboratory. However, it remains unknown whether the
difference may affect the detection rate or false-positive rate. In
order to resolve these issues, 2,575 specimens within different
batches were analyzed. First, the original Down syndrome risk
values of each specimen were recorded. Then, the new medians were
applied and a new risk value was obtained. Finally, the cut-off
values with a 5% false-positive rate were obtained, and the
original and new cut-off value was applied to analyze the detection
rate for Down syndrome. Furthermore, the false-positive rate using
the 2T medians and the medians from our laboratory was also
analyzed at a cut-off value of 1/280.
Our results revealed that, if a 5% false-positive
rate was used to select a cut-off value, the original cut-off value
would be 1/280, and the new cut-off value would be 1/310. For the
118 Down syndrome cases, if the cut-off value was 1/280, a total of
75 cases would be detected (63.6%). Furthermore, if a cut-off value
of 1/310 was applied, 80 of 118 cases would be detected (67.8%).
However, there was no significant difference on the detection rate
(P>0.05, Table III). If 1/280
was used as the cut-off value, the original false-positive rate
would be 5.05% (130/2,575), and taking into consideration the new
risk values calculated by our own medians, the false-positive rate
was 4.38% (113/2,575). Although the latter false-positive rate was
lower compared with the former, the difference was not
statistically significant (P>0.05, Table IV). Our results suggested that the
difference in medians between our laboratory and those embedded in
2T, did not statistically significantly affect either the detection
rate or false-positive rate. However, considering the increase in
detection rate and decrease in false-positive rate, we recommend
the use of our own medians of AFP and free β-HCG.
 | Table III.Comparison of different medians for
detection rate for DS. |
Table III.
Comparison of different medians for
detection rate for DS.
| Median | Cut-off value of 5%
FPR | DS cases
detected | P-value |
|---|
| Original | 1/280 | 75/118 | >0.05 |
| New | 1/310 | 80/118 |
|
 | Table IV.Effect of different medians on
false-positive rate for DS. |
Table IV.
Effect of different medians on
false-positive rate for DS.
| Median | Positive cases
ratio | FPR for DS at 1/280
(%) | P-value |
|---|
| Original | 130/2575 | 5.05 | >0.05 |
| New | 113/2575 | 4.38 |
|
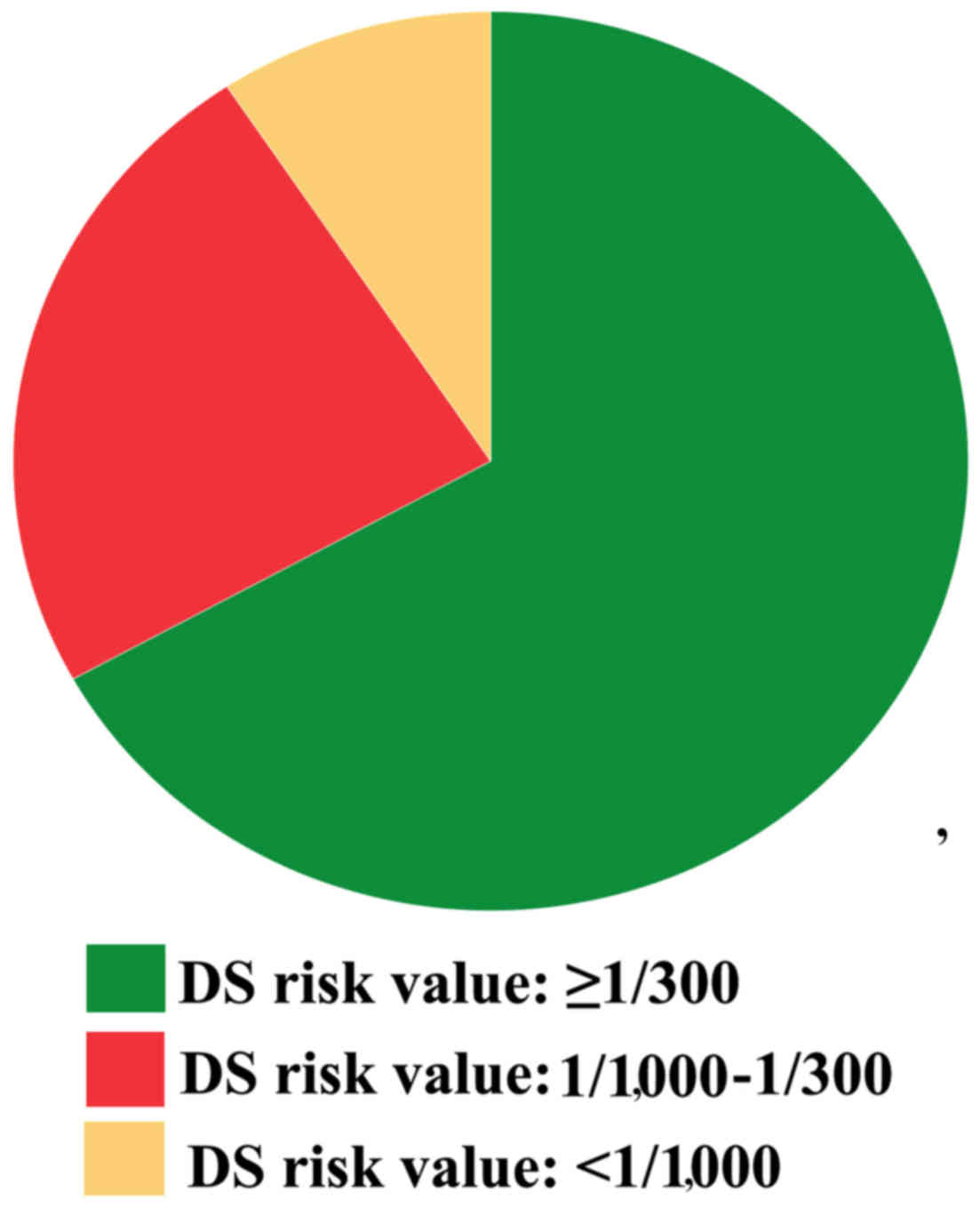
Residual Down syndrome cases are
mainly concentrated in a certain risk range
After a long period of research, it was found that
residual Down syndrome cases were primarily concentrated at risk
values between 1/1,000 and 1/300 (Table
V). A total of 221,288 screened cases were counted in our
hospital, and the value at risk within 1/300 were 11,636 cases,
accounting for 5.3% of the total number of screened cases; the
number of cases with risk values within 1/1,000-1/300 was 30,997,
accounting for 14.0%; and the number of cases with risk values
<1/1,000 was 178,655 (80.7%). It may be concluded that Down
syndrome cases were mainly concentrated at risk values ≥1/300
(66.95%), suggesting that our screening was efficient. However,
there were certain residual cases: For example, there were 39
residual cases and 28 had a risk value within 1/1000-1/300
(23.73%); in addition, only 9.32% (11 residual cases) had a risk
value of <1/1,000, indicating that residual cases were mainly
concentrated at risk values between 1/1,000 and 1/300 (P<0.01,
Fig. 2). In summary, close attention
should be paid to cases with a risk value 1/1,000-1/300 to
significantly reduce the number of Down syndrome births.
 | Table V.Composition of DS pregnancy in
different ranges of risk value. |
Table V.
Composition of DS pregnancy in
different ranges of risk value.
| Risk value range of
DS | Number of
screenings | Proportion of the
total sample (%) | Number of pregnancies
with DS | Proportion of
pregnancies with DS (%) |
|---|
| ≥1/300 | 11,636 | 5.3 | 79 | 66.95 |
| 1/1,000-1/300 | 30,997 | 14.0 | 28 | 23.73 |
| <1/1,000 | 178,655 | 80.7 | 11 | 9.32 |
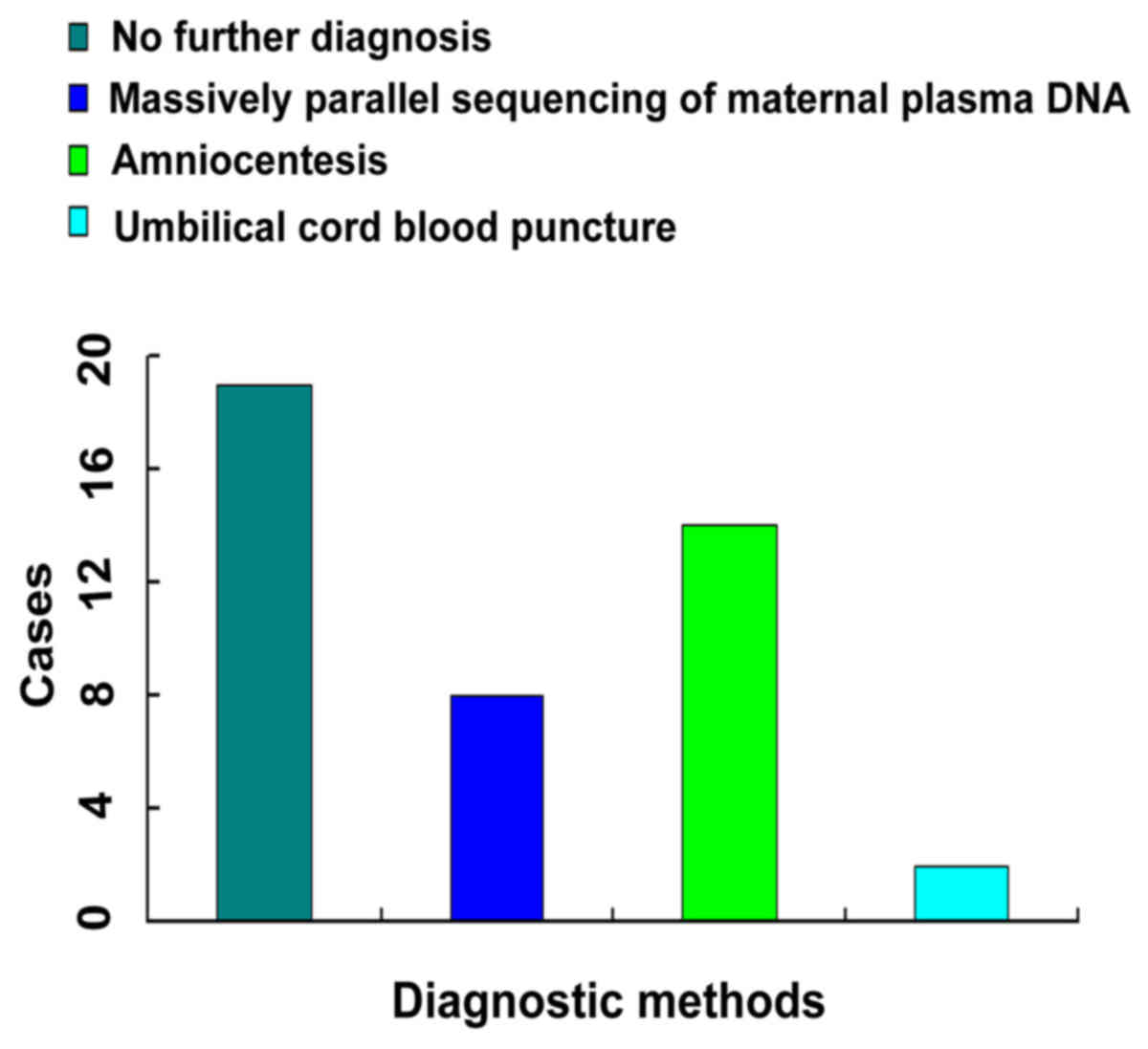
Further diagnosis for the different
ranges of risk value
According to the abovementioned findings, the
results were divided into three categories. First, a risk value
≥1/300 was considered as high-risk. For such cases, amniocentesis
or cordocentesis would be advisable. Second, a risk value of
1/1,000-1/300 was considered to be in the extenuation range, and it
would require a rational suggestion. If the risk value was within
1/1,000-1/300, but the value of free β-HCG was extremely high
(MoM>10), or the ultrasound revealed abnormal results that may
be associated with chromosomal abnormalities, amniocentesis or
cordocentesis is recommended. If the correction of MoM of free
β-HCG was in the normal range, massive parallel sequencing of
maternal plasma DNA may be applied. Finally, if the risk value was
<1/1,000 and there were abnormal ultrasound results possibly
associated with chromosomal abnormalities, amniocentesis or
cordocentesis would be recommended. Through this approach, leak
cases may significantly decrease (Fig.
3). For risk values <1/300, there were 18 Down syndrome
births, as these subjects did not undergo further examinations; 7
Down syndrome cases were detected by massive parallel sequencing;
amniocentesis detected the majority of the cases (12 cases); only 2
cases underwent cordocentesis and terminated their pregnancies. It
may be concluded that amniocentesis remains the primary method for
confirming Down syndrome, and NIPT is a new method developing
rapidly, as it may be used to avoid the risk of intrauterine
infection or abortion caused by amniocentesis or cordocentesis.
Appropriate advice must be offered to subjects with extenuation
risk values, in order to reduce the number of Down syndrome
births.
Discussion
Trisomy 21 is associated with relatively common
chromosomal abnormalities, which affect human health and the
quality of life. Hence, scholars have been searching for more
accurate screening methods to prevent Down syndrome births.
Serological screening is a simple and relatively risk-free method,
and has been attracting increasing attention. A good screening
index may increase positive detection rate and reduce missed Down
syndrome pregnancies; therefore, close attention should be given to
these serological markers.
There were differences in the medians of AFP and
free β-HCG. Data provided by Wang et al (8) revealed that the medians of AFP were
higher compared with those in Caucasian women for 15–20 weeks,
which was different from our results; however, the medians of free
β-HCG were similar, which was higher compared with the medians of
the 2T software. They also reported that, using the total β-HCG
(HCG) and AFP, the detection rate for Down syndrome was 50%, and
that of triple screening was 66.7%, with a 5% false-positive rate.
It was reported that, when using AFP and free β-HCG, the detection
rate for Down syndrome was 56.25% by Lifecycle software (17). In our data, the detection rate is
63.6%, with a false-positive rate of 5.05%. It may be concluded
that our detection rate was higher compared with the previously
reported detection rate of double screenings. Although our
laboratory medians differed from the medians embedded in the 2T
software, there was no significant statistical impact on the
detection rate or the false-positive rate; however, there was an
increase in the detection rate and a decrease in the false-positive
rate. Taking this finding into consideration, we recommend the use
of our own medians of AFP and free β-HCG.
If risk values are >1/1,000, further diagnostic
investigation is crucial. It is recommended that high-risk cases
undergo amniocentesis or cordocentesis. Our results revealed that a
risk value between 1/1,000-1/300 was the main range of residual
cases. Thus, providing more rational suggestions may lead to fewer
residual cases. Amniocentesis or cordocentesis are effective
diagnostic methods, but are associated with a risk of miscarriage
(~1%); however, NIPT is considered as a safe method (18). Subjects with results raising clinical
suspicion who decline further diagnostic examinations constitute a
major cause of Down syndrome births. Thus, if the results are
abnormal, a doctor should be consulted as soon as possible.
Thorough screening and prenatal diagnosis of Down
syndrome is crucial, as it may help reduce the financial burden on
families, reduce stress, and exert an overall beneficial effect on
society. Further in-depth studies are required on this subject to
design screening or diagnostic methods that are more accurate and
cost-effective.
References
|
1
|
Smith M and Visootsak J: Noninvasive
screening tools for Down syndrome: A review. Int J Womens Health.
5:125–131. 2013.PubMed/NCBI
|
|
2
|
Avent ND: Maternal plasma biomarkers for
down syndrome: Present and future. Drugs Today (Barc). 49:145–152.
2013. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Alldred SK, Deeks JJ, Guo B, Neilson JP
and Alfirevic Z: Second trimester serum tests for Down's Syndrome
screening. Cochrane Database Syst Rev. 13:CD0099252012.
|
|
4
|
Androutsopoulos G, Gkogkos P and Decavalas
G: Mid-trimester maternal serum HCG and alpha fetal protein levels:
Clinical significance and prediction of adverse pregnancy outcome.
Int J Endocrinol Metab. 11:102–106. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Extermann P, Bischof P, Marguerat P and
Mermillod B: Second-trimester maternal serum screening for Down's
syndrome: Free beta-human chorionic gonadotrophin (HCG) and
alpha-fetoprotein, with or without unconjugated oestriol, compared
with total HCG, alpha-fetoprotein and unconjugated oestriol. Hum
Reprod. 13:220–223. 1998. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Shaw SW, Hsu JJ, Lee CN, Hsiao CH, Chen
CP, Hsieh TT and Cheng PJ: First- and second-trimester Down
syndrome screening: Current strategies and clinical guidelines.
Taiwan J Obstet Gynecol. 47:157–162. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Sayin NC, Canda MT, Ahmet N, Arda S, Süt N
and Varol FG: The association of triple-marker test results with
adverse pregnancy outcomes in low-risk pregnancies with healthy
newborns. Arch Gynecol Obstet. 277:47–53. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Wang YY, Luo J, Zhu MW, Liu LN and Ma X:
Second-trimester double or triple screening for Down syndrome: A
comparison of Chinese and Caucasian populations. Int J Gynaecol
Obstet. 94:67–72. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Sifakis S, Papantoniou N, Kappou D and
Antsaklis A: Noninvasive prenatal diagnosis of Down syndrome:
Current knowledge and novel insights. J Perinat Med. 40:319–327.
2012. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Nicolaides KH, Syngelaki A, Gil M,
Atanasova V and Markova D: Validation of targeted sequencing of
single-nucleotide polymorphisms for non-invasive prenatal detection
of aneuploidy of chromosomes 13, 18, 21, X and Y. Prenat Diagn.
33:575–579. 2013. View
Article : Google Scholar : PubMed/NCBI
|
|
11
|
Shaw SW, Chen CP and Cheng PJ: From Down
syndrome screening to noninvasive prenatal testing: 20 years'
experience in Taiwan. Taiwan J Obstet Gynecol. 52:470–474. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Ehrich M, Deciu C, Zwiefelhofer T, Tynan
JA, Cagasan L, Tim R, Lu V, McCullough R, McCarthy E, Nygren AO, et
al: Noninvasive detection of fetal trisomy 21 by sequencing of DNA
in maternal blood: A study in a clinical setting. Am J Obstet
Gynecol. 204:205.e1–11. 2011. View Article : Google Scholar
|
|
13
|
Baffero GM, Somigliana E, Crovetto F,
Paffoni A, Persico N, Guerneri S, Lalatta F, Fogliani R and Fedele
L: Confined placental mosaicism at chorionic villous sampling: Risk
factors and pregnancy outcome. Prenat Diagn. 32:1102–1108. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Bamberg C, Fotopoulou C, Thiem D, Roehr
CC, Dudenhausen JW and Kalache KD: Correlation of midtrimester
amniotic fluid cytokine concentrations with adverse pregnancy
outcome in terms of spontaneous abortion, preterm birth and
preeclampsia. J Matern Fetal Neonatal Med. 25:812–817. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Guedj F and Bianchi DW: Noninvasive
prenatal testing creates an opportunity for antenatal treatment of
Down syndrome. Prenat Diagn. 33:614–618. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Van Opstal D and Srebniak MI: Cytogenetic
confirmation of a positive NIPT result: Evidence-based choice
between chorionic villus sampling and amniocentesis depending on
chromosome aberration. Expert Rev Mol Diagn. 16:513–520. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Yu DY, Fu P, Zhang ZH, Wang F, Han MY, Ren
HY, Zhao W, Zhang K, Li S and Jiang N: Evaluation of Down's
syndrome screening methods using maternal serum biochemistry in the
second trimester pregnancy. Zhonghua Yi Xue Yi Chuan Xue Za Zhi.
28:332–335. 2011.(In Chinese). PubMed/NCBI
|
|
18
|
Lim JH, Park SY and Ryu HM: Non-invasive
prenatal diagnosis of fetal trisomy 21 using cell-free fetal DNA in
maternal blood. Obstet Gynecol Sci. 56:58–66. 2013. View Article : Google Scholar : PubMed/NCBI
|