Introduction
Esophageal carcinoma is a common malignancy
worldwide, with a 5-year survival rate of ~30%, even with
chemotherapy, surgery and radiation therapy, due to tumor
heterogeneity (1–9). It is well-known that early diagnosis
and timely treatment may improve the survival rate in early-stage
cancer patients, with a 5-year survival rate as high as 90%
(10,11). Cancer Research UK (http://www.cancerresearchuk.org/about-cancer/cancer-symptoms/why-is-early-diagnosis-important)
clearly reported the association of survival rate with stage at
diagnosis for several cancers, such as breast, ovarian, lung and
bowel cancer. However, the majority of esophageal cancer patients
are often diagnosed at an advanced stage, as there are no obvious
symptoms in the early stages of the disease. Therefore, it is
crucial to identify an effective biomarker for early diagnosis.
Recently, the matrix-assisted laser desorption/ionization
time-of-flight mass spectrometry (MOLDI-TOF-MS) and
surface-enhanced laser desorption/ionization time-of-flight mass
spectrometry (SELDI-TOF-MS) techniques have been widely employed in
the search for cancer biomarkers in brain cancer (12), oral squamous cell carcinoma (13), pancreatic cancer (14), lung cancer (15), esophageal squamous cell carcinoma
(16) and breast cancer (17,18),
among others. There have been several attempts to identify
esophageal carcinoma biomarkers using these techniques (16,19–22).
However, those studies failed to identify any single biomarker for
the diagnosis of esophageal cancer, but rather indicated several
proteins. In addition, the identified proteins differed among
different research groups. This makes it difficult to establish a
unified standard for rapid and accurate diagnosis of this type of
cancer. In the present study, a single protein, anaphylatoxin C3a,
which is one of the complement proteins, was investigated as a
diagnostic biomarker to distinguish between esophageal cancer
patients and healthy individuals.
Materials and methods
Research subjects
A total of 40 serum samples were included in this
study, 20 of which were collected from esophageal carcinoma
patients from Yancheng First People's Hospital (group A), whereas
the others were collected from non-esophageal carcinoma
participants recruited from the First Hospital of Nanjing Medical
University (group B) between November 2014 and May 2015. The
participants were aged 25–76 years, with a mean age of 55.7 years.
In group A, there were 8 cases of patients who were undergoing
therapy. All the blood collections were performed an overnight
fast. Blood was collected in 5-ml blood collection tubes without
anticoagulant and the serum samples were stored at −80°C for
further use.
Ethics statement
All patients signed an informed consent form and the
study protocol was approved by the Institutional Review Board of
the First People's Hospital of Yancheng (Yancheng, China). The
study was reviewed and approved by Yancheng Medical Ethics
Committee.
SELDI-TOF-MS assay
SELDI-TOF-MS (Ciphergen Biosystems, Fremont, CA,
USA) was applied to collect the raw mass spectrometry data.
Biomarker Wizard software (Ciphergen Biosystems, Fremont, CA, USA)
was used to export the raw data into a digital format, according to
the standard Excel format. A t-test was conducted with the Excel
data using SPSS 17.0 software (SPSS Inc., Chicago, IL, USA).
Enzyme-linked immunoabsorbent assay
(ELISA)
The concentrations of C3a in the serum samples were
quantified by ELISA according to the manufacturer's protocol (cat.
no. ab133037; Abcam, Cambridge, MA, USA). Briefly, 50 µl standard
samples and 50 µl serum samples were diluted 5 times with
phosphate-buffered saline, placed into a 96-well plate and cultured
at 37°C for 30 min. The plate was washed 5 times, after which time
50 µl reagent A was added into each well, followed by 50 µl reagent
B. The samples were mixed well and cultured for a further 15 min at
37°C in the dark; stop solution was then added into each well. An
ELISA plate reader (Synergy HXT; BioTek Instruments Inc., Winooski,
VT, USA) was used at a wavelength of 450 nm; the inter-assay and
intra-assay coefficients of variation of the ELISA kits for C3a
were <10%.
Statistical analysis
The SPSS 19.0 software package (SPSS Inc., Chicago,
IL, USA) was used for data analysis, and the data are expressed as
means ± standard deviation. The comparison between the groups was
performed using one-way analysis of variance. P<0.05 indicates
that the difference was statistically significant.
Results
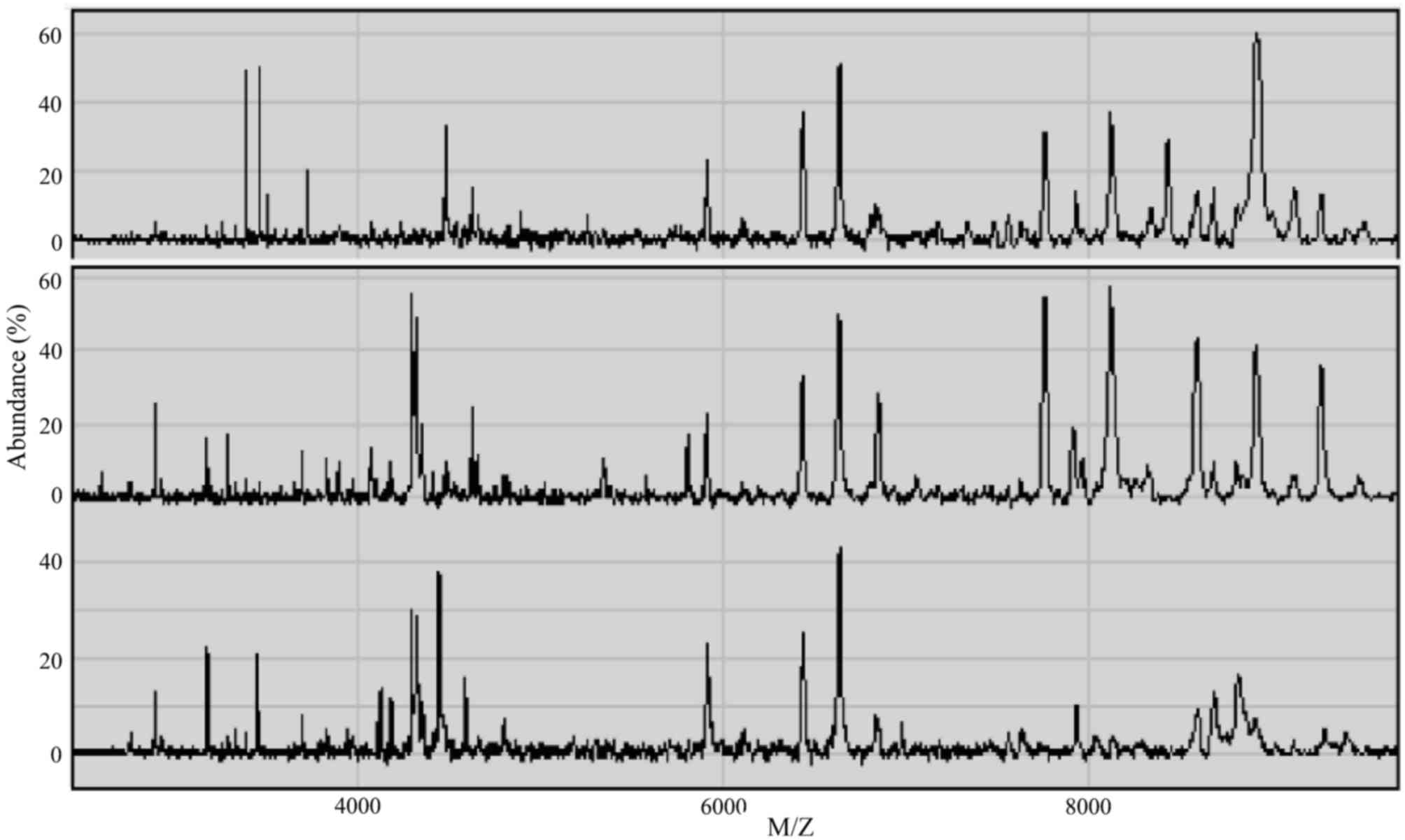
Serum proteomic profiles
SELDI-TOF-MS was applied to examine the serum
samples within a window of 1–10 kDa, and 253 protein peaks were
detected in the sera of esophageal cancer patients. There were 144
statistically significant difference peaks (P<0.05): 56 peaks
had higher protein expression and 88 had lower protein expression
in esophageal cancer patients. The mass spectra (MS) of the serum
proteins of esophageal cancer patients and non-esophageal cancer
participants are shown in Fig.
1.
Establishment of diagnostic
biomarker
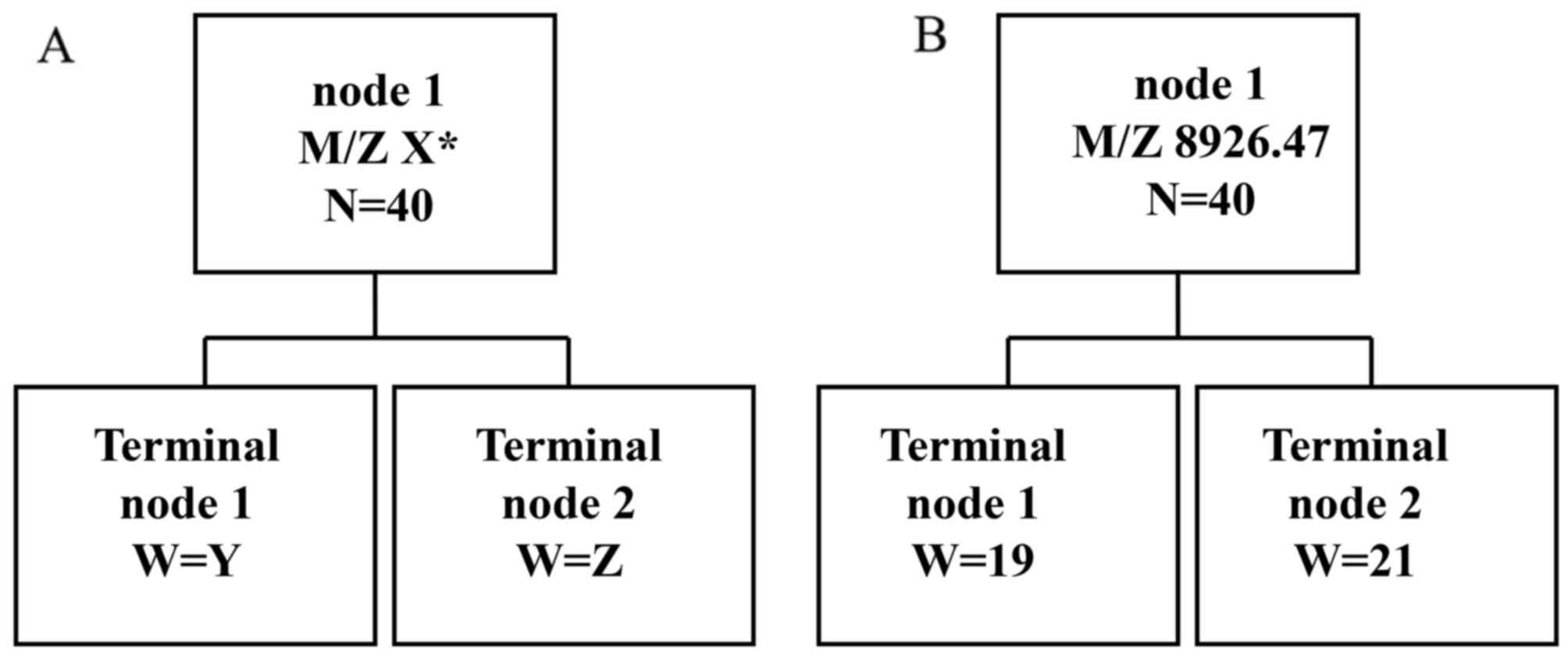
In order to establish the diagnostic biomarker, 6 MS
peaks, i.e., M/Z 2,748.87 (P=2.38×10−7), M/Z 4,119.31
(P=3.18×10−7), M/Z 4,425.94 (P=2.38×10−7),
M/Z 4,798.62 (P=3.67×10−7), M/Z 9,136.76
(P=6.45×10−7) and M/Z 8,926.47 (P=7.33×10−8),
exhibiting statistically significant differences, were further
analyzed. The classical tree model was employed to examine the
predictor variables (Fig. 2A). The
protein M/Z 8,926.47 had the best prediction result (Fig. 2B). The sensitivity and specificity of
the biomarker were 95% (19/20) and 97.5% (39/40), respectively. The
positive predictive accuracy was 100% (19/19) and the negative
predictive accuracy was 95.2% (20/21). Thus, the protein at M/Z
8,926.47 may be an optimal biomarker for distinguishing the
esophageal cancer and non-esophageal cancer sera with high accuracy
and high specificity.
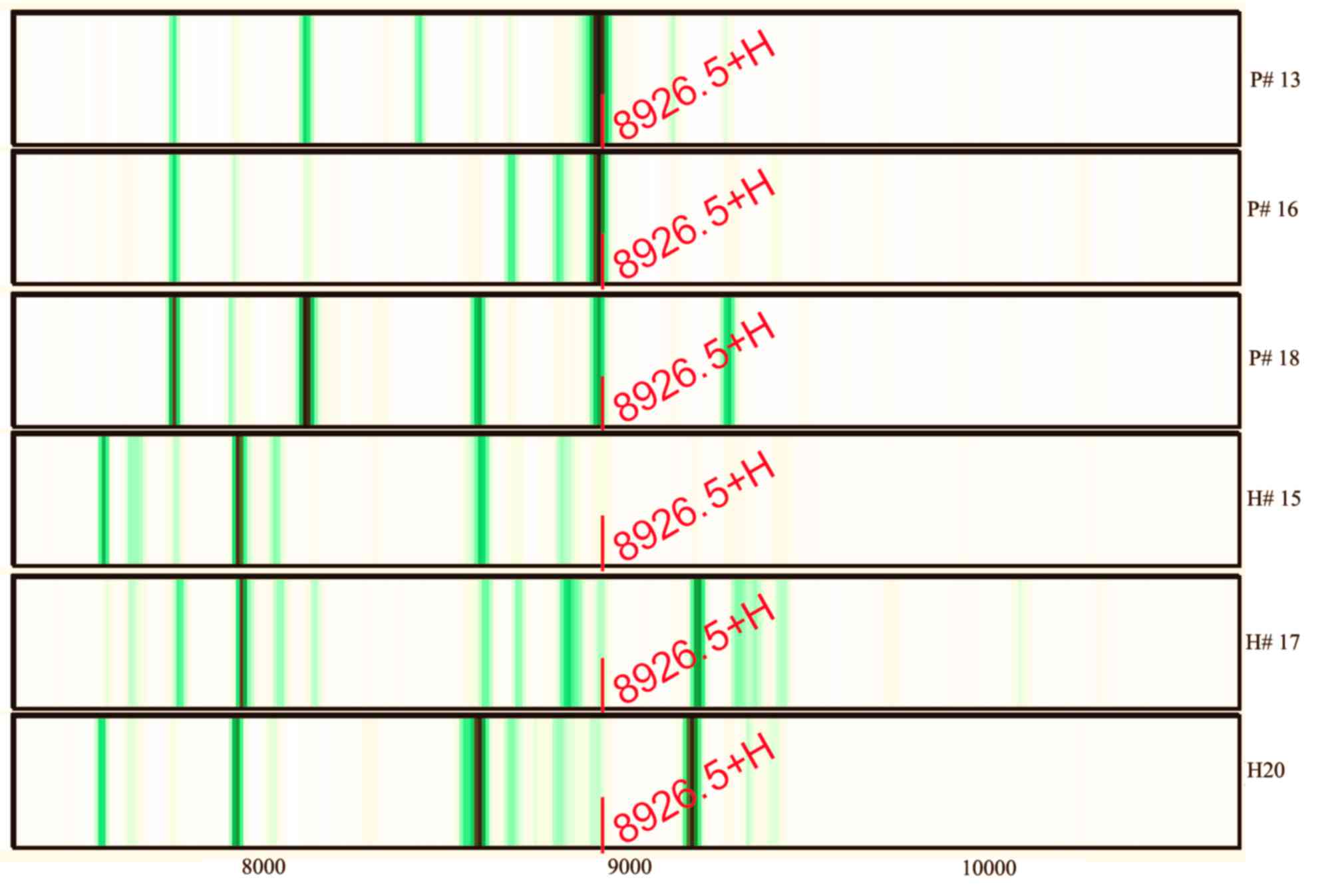
Further study indicated that the abundance of M/Z
8,926.47 was reduced from 60.69 prior to therapy to 43.39 after
therapy (the mean abundance of this peak was 6.81 for
non-esophageal carcinoma participants). Three of the serum protein
MS of esophageal cancer patients and non-esophageal carcinoma
participants are shown in Fig. 3.
The intensities of M/Z 8,926.47 in the non-esophageal carcinoma
participants were distinctly lower compared with those in
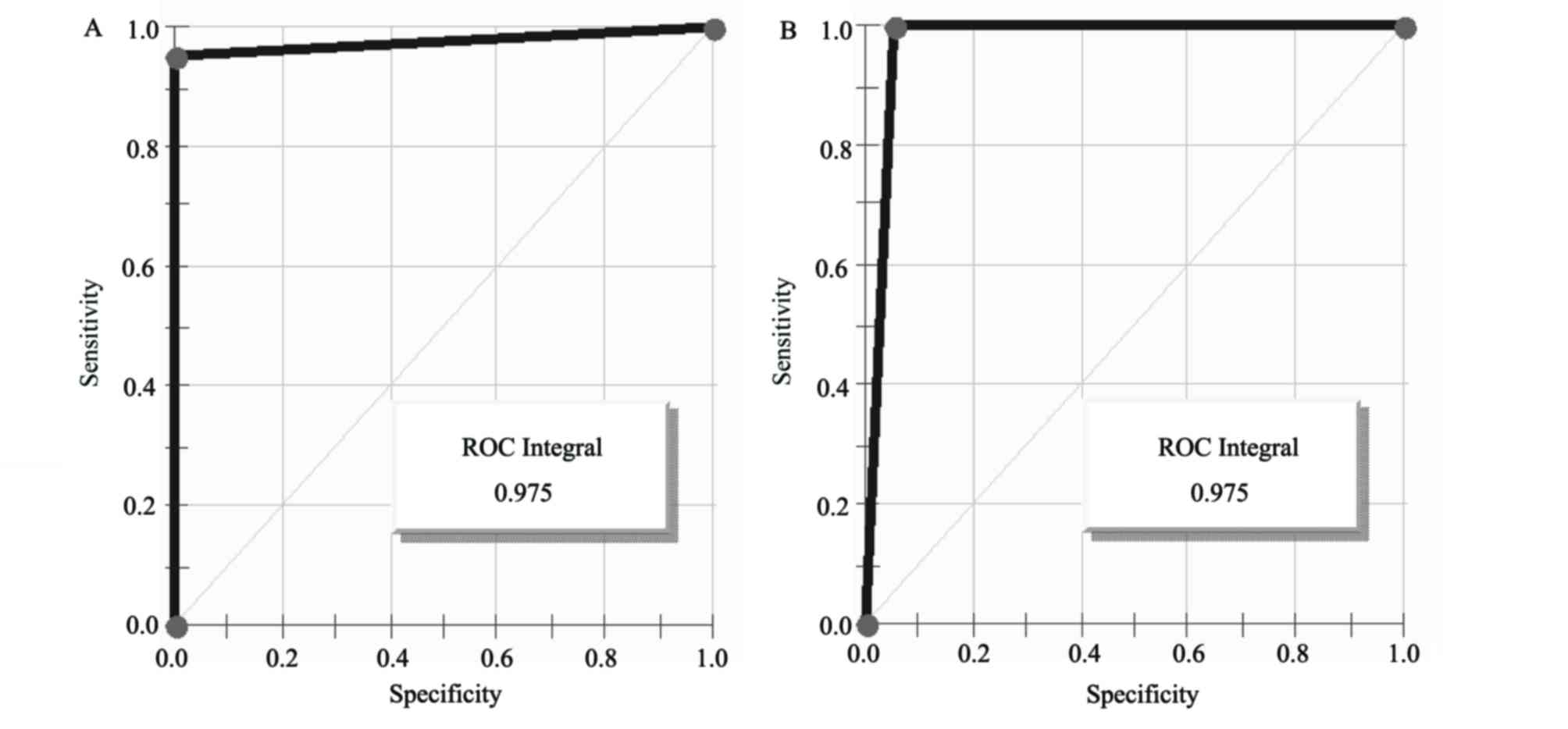
esophageal carcinoma patients. The areas under the receiver
operating characteristic curves of the diagnostic biomarker were
97.5% (Fig. 4).
Determination of the concentration of
C3a via ELISA
The human C3a ELISA kit was used to determine the
concentration of anaphylatoxin C3a using the obtained serum
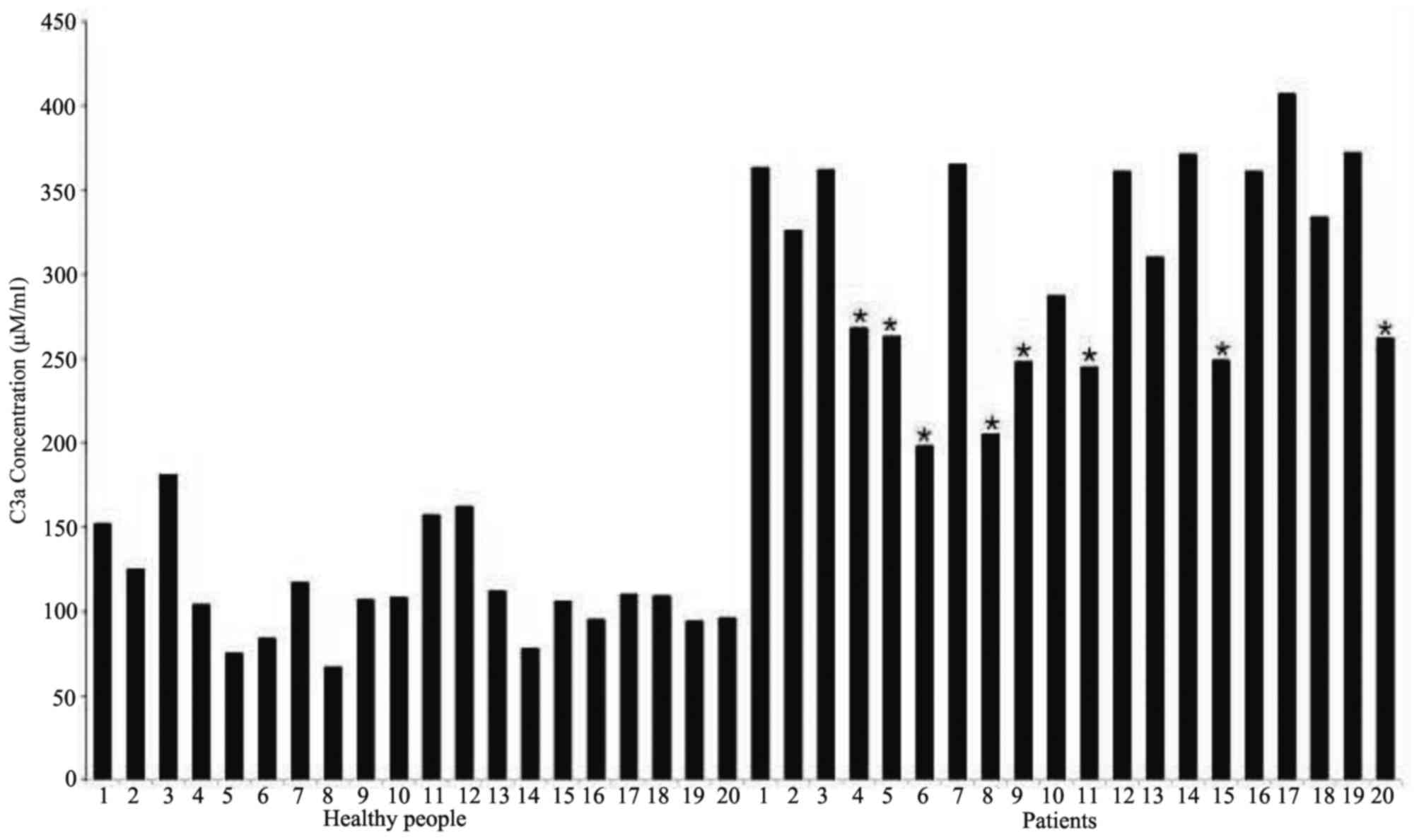
samples. The ELISA experimental results (Fig. 5) revealed that the concentration of
anaphylatoxin C3a in the sera of esophageal cancer patients (mean,
308±60 ng/ml) were significantly higher compared with those in
healthy participants (mean, 112±30 ng/ml). Among esophageal
carcinoma patients, the mean concentration of C3a in the sera of
those without therapy and those undergoing therapy was 352±31 and
242±25 ng/ml, respectively.
Discussion
Regarding the SELDI-TOF MS results, significant
differences were observed in the serum samples between the
esophageal carcinoma patients and healthy participants for 6
protein peaks (P<10−6). Among those, the abundance of
the peak M/Z 8,926.47 (P=7.33×10−8) markedly increased
from 6.81 (non-esophageal carcinoma) to 60.69 (esophageal
carcinoma), suggesting that the expression of M/Z 8,926.47 was
significantly increased in the plasma of esophageal carcinoma
patients. This particular peak distinguished between the sera of
esophageal carcinoma patients and non-esophageal carcinoma
participants with high accuracy (100%) and high efficiency (97.5%),
indicating that M/Z 8,926.47 may be a promising biomarker for early
diagnosis. Based on our experience, this peak is likely to be
complementary to the C3a protein.
The ELISA results demonstrated that the
concentrations were statistically significantly different
(P<0.01) and the mean concentration of anaphylatoxin C3a in
esophageal carcinoma and non-esophageal carcinoma samples was
308±60 and 112±30 ng/ml, respectively, which was in agreement with
the MS peak abundance of 53.98 and 6.81. Furthermore, among
esophageal cancer samples, the C3a concentration was 352±31 and
242±25 ng/ml for samples before and after treatment, respectively,
which was also in agreement with the MS peak abundances of 60.69
and 43.39. This finding indicates that, after treatment, the
concentration of C3a in the plasma was markedly decreased.
According to the ELISA results, it may be concluded that, when the
serum C3a concentration is >120 ng/ml, the risk of esophageal
cancer is increased.
C3a is a serum protein first discovered in 1896,
which plays a key role in either antitumor immune response
(23–27), or in promoting tumor growth and
progression (28,29). However, the role of C3a in esophageal
cancer has not been determined to date. According to the MS and
ELISA results, the anaphylatoxin C3a concentrations in the sera of
treated patients are significantly lower compared with those
without treatment. It may be hypothesized that C3a plays a key role
in promoting esophageal tumorigenesis. As previously reported,
anaphylatoxin C3a may contribute to cancer cell immune escape via
promoting local immunosuppression (30,31).
However, more studies should be conducted to elucidate the
mechanisms through which C3a promotes esophageal tumorigenesis.
Acknowledgements
The authors would like to acknowledge the Department
of Human Resources and Social Security of Jiangsu Province
(swyy-030) and Foundation of Jiangsu Educational Committee
(16KJB180031) for financially supporting the present study.
Esophageal carcinoma serum samples were provided by Dr Zhou
Xiaoning (Oncology Department, Yancheng First People's Hospital).
Healthy participants' serum samples were provided by Dr Shao
Qianwen (Oncology Department, the First Hospital of Nanjing Medical
University).
References
|
1
|
Fisher R, Pusztai L and Swanton C: Cancer
heterogeneity: Implications for targeted therapeutics. Br J Cancer.
108:479–485. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Graff BA, Kvinnsland Y, Skretting A and
Rofstad EK: Intratumour heterogeneity in the uptake of
macromolecular therapeutic agents in human melanoma xenografts. Br
J Cancer. 88:291–297. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Kim JH, Ko ES, Lim Y, Lee KS, Han BK, Ko
EY, Hahn SY and Nam SJ: Breast cancer heterogeneity: MR imaging
texture analysis and survival outcomes. Radiology. 282:665–675.
2017. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Beca F and Polyak K: Intratumor
heterogeneity in breast cancer. Adv Exp Med Biol. 882:169–189.
2016. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Aleskandarany MA, Green AR, Ashankyty I,
Elmouna A, Diez-Rodriguez M, Nolan CC, Ellis IO and Rakha EA:
Impact of intratumoural heterogeneity on the assessment of Ki67
expression in breast cancer. Breast Cancer Res Treat. 158:287–295.
2016. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Suda K, Murakami I, Sakai K, Tomizawa K,
Mizuuchi H, Sato K, Nishio K and Mitsudomi T: Heterogeneity in
resistance mechanisms causes shorter duration of epidermal growth
factor receptor kinase inhibitor treatment in lung cancer. Lung
Cancer. 91:36–40. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Bashir U, Siddique MM, Mclean E, Goh V and
Cook GJ: Imaging heterogeneity in lung cancer: Techniques,
applications, and challenges. AJR Am J Roentgenol. 207:534–543.
2016. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Obulkasim A, Ylstra B, van Essen HF,
Benner C, Stenning S, Langley R, Allum W, Cunningham D, Inam I,
Hewitt LC, et al: Reduced genomic tumor heterogeneity after
neoadjuvant chemotherapy is related to favorable outcome in
patients with esophageal adenocarcinoma. Oncotarget. 7:44084–44095.
2016. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Cao W, Wu W, Yan M, Tian F, Liu HS, Wang
JW, Zhang QW, Li YJ and Li M: Multiregion sequencing reveals
intratumor heterogeneity in esophageal squamous cell carcinoma.
Zhonghua Zhong Liu Za Zhi. 38:660–666. 2016.(In Chinese).
PubMed/NCBI
|
|
10
|
Chen LQ, Hu CY, Ghadirian P and Duranceau
A: Early detection of esophageal squamous cell carcinoma and its
effects on therapy: An overview. Dis Esophagus. 12:161–167. 1999.
View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Schweigert M, Dubecz A and Stein HJ:
Oesophageal cancer-an overview. Nat Rev Gastroenterol Hepatol.
10:230–244. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Spalding K, Board R, Dawson T, Jenkinson
MD and Baker MJ: A review of novel analytical diagnostics for
liquid biopsies: Spectroscopic and spectrometric serum profiling of
primary and secondary brain tumors. Brain Behav. 6:e005022016.
View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Xie X, Jiang Y, Yuan Y, Wang P, Li X, Chen
F, Sun C, Zhao H, Zeng X, Jiang L, et al: MALDI imaging reveals
NCOA7 as a potential biomarker in oral squamous cell carcinoma
arising from oral submucous fibrosis. Oncotarget. 7:59987–60004.
2016. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Potjer TP, Mertens BJ, Nicolardi S, van
der Burgt YE, Bonsing BA, Mesker WE, Tollenaar RA and Vasen HF:
Application of a serum protein signature for pancreatic cancer to
separate cases from controls in a pancreatic surveillance cohort.
Transl Oncol. 9:242–247. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Ren J, Zhang D, Liu Y, Zhang R, Fang H,
Guo S, Zhou D, Zhang M, Xu Y, Qiu L and Li Z: Simultaneous
quantification of serum nonesterified and esterified fatty acids as
potential biomarkers to differentiate benign lung diseases from
lung cancer. Sci Rep. 6:342012016. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Jia K, Li W, Wang F, Qu H, Qiao Y, Zhou L,
Sun Y, Ma Q and Zhao X: Novel circulating peptide biomarkers for
esophageal squamous cell carcinoma revealed by a magnetic
bead-based MALDI-TOFMS assay. Oncotarget. 7:23569–23580. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Wang S, Chen X, Luan H, Gao D, Lin S, Cai
Z, Liu J, Liu H and Jiang Y: Matrix-assisted laser
desorption/ionization mass spectrometry imaging of cell cultures
for the lipidomic analysis of potential lipid markers in human
breast cancer invasion. Rapid Commun Mass Spectrom. 30:533–542.
2016. View
Article : Google Scholar : PubMed/NCBI
|
|
18
|
Kawaguchi-Sakita N, Kaneshiro-Nakagawa K,
Kawashima M, Sugimoto M, Tokiwa M, Suzuki E, Kajihara S, Fujita Y,
Iwamoto S, Tanaka K and Toi M: Serum immunoglobulin G Fc region
N-glycosylation profiling by matrix-assisted laser
desorption/ionization mass spectrometry can distinguish breast
cancer patients from cancer-free controls. Biochem Biophys Res
Commun. 469:1140–1145. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Zhao J, Fan YX, Yang Y, Liu DL, Wu K, Wen
FB, Zhang CY, Zhu DY and Zhao S: Identification of potential plasma
biomarkers for esophageal squamous cell carcinoma by a proteomic
method. Int J Clin Exp Pathol. 8:1535–1544. 2015.PubMed/NCBI
|
|
20
|
Schwacke J, Millar TP, Hammond CE, Saha A,
Hoffman BJ, Romagnuolo J, Hill EG and Smolka AJ: Discrimination of
normal and esophageal cancer plasma proteomes by MALDI-TOF mass
spectrometry. Dig Dis Sci. 60:1645–1654. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Fan NJ, Gao CF and Wang XL: Tubulin beta
chain, filamin A alpha isoform 1, and cytochrome b-c1 complex
subunit 1 as serological diagnostic biomarkers of esophageal
squamous cell carcinoma: A proteomics study. OMICS. 17:215–223.
2013. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Zhai XH, Yu JK, Lin C, Wang LD and Zheng
S: Combining proteomics, serum biomarkers and bioinformatics to
discriminate between esophageal squamous cell carcinoma and
pre-cancerous lesion. J Zhejiang Univ Sci B. 13:964–971. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Weiner LM, Surana R and Wang S: Monoclonal
antibodies: Versatile platforms for cancer immunotherapy. Nat Rev
Immunol. 10:317–327. 2010. View
Article : Google Scholar : PubMed/NCBI
|
|
24
|
Rutkowski MJ, Sughrue ME, Kane AJ, Mills
SA and Parsa AT: Cancer and the complement cascade. Mol Cancer Res.
8:1453–1465. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Sliwkowski MX and Mellman I: Antibody
therapeutics in cancer. Science. 341:1192–1198. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Taylor RP and Lindorfer MA: The role of
complement in mAb-based therapies of cancer. Methods. 65:18–27.
2014. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Baig NA, Taylor RP, Lindorfer MA, Church
AK, LaPlant BR, Pettinger AM, Shanafelt TD, Nowakowski GS and Zent
CS: Induced resistance to ofatumumab-mediated cell clearance
mechanisms, including complement-dependent cytotoxicity, in chronic
lymphocytic leukemia. J Immunol. 192:1620–1629. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Markiewski MM, DeAngelis RA, Benencia F,
Ricklin-Lichtsteiner SK, Koutoulaki A, Gerard C, Coukos G and
Lambris JD: Modulation of the antitumor immune response by
complement. Nat Immunol. 9:1225–1235. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Khan MA, Assiri AM and Broering DC:
Complement and macrophage crosstalk during process of angiogenesis
in tumor progression. J Biomed Sci. 22:582015. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Sayegh ET, Bloch O and Parsa AT:
Complement anaphylatoxins as immune regulators in cancer. Cancer
Med. 3:747–758. 2014. View
Article : Google Scholar : PubMed/NCBI
|
|
31
|
Denko NC, Fontana LA, Hudson KM, Sutphin
PD, Raychaudhuri S, Altman R and Giaccia AJ: Investigating hypoxic
tumor physiology through gene expression patterns. Oncogene.
22:5907–5914. 2003. View Article : Google Scholar : PubMed/NCBI
|