Introduction
Breast cancer, the most common malignancy in women
(1), is treated by surgical removal
of the cancer. In addition, adjuvant systemic therapy is given
depending on the perceived aggressiveness of the removed cancer.
Currently the established prognostic parameter include histological
grade, tumor size, presence of lymph node metastasis, tumor cell
proliferation (Ki67 labeling index; Ki67LI) as well as hormonal
receptor and HER2 status (2-4)
(Ki67) (5). In many patients,
supplementary molecular parameters are analyzed (6-8).
These molecular classifiers are built on multiplexed analysis of
the mRNAs of 21-70 genes (9-11).
The rising interest in mitochondrial function and
dysfunction on cancer development has been reviewed by Davis and
Williams and Hsu et al (12,13).
The loss of proliferation control in cancer cells may result in
cellular masses that extend beyond the capacity of the supporting
vasculature, leading to oxygen and nutrient deprivation. Hence,
tumor cells must adapt to overcome these restrictions. Mitochondria
are key organelles for energy production in normal and neoplastic
cells. Quantity and activity of mitochondria are essential for
tumor growth (reviewed in refs. 12-16). Mutations in mitochondrial
genes or aberrant mitochondrial content have been described to
occur in various cancer types (17-20).
An increased mitochondria quantity has earlier been linked to
aggressive tumor phenotype and poor prognosis in lung (21), colorectal (22,23),
prostate (24), gastric (25), cervical (18), and ovarian cancer (26). In glioma, however, high mitochondria
content was linked to favorable prognosis (27). In one study, on 76 breast
carcinomas, a prognostic impact of the mitochondria count was also
suggested (28). Focused on these
reports, we assumed that the cellular mitochondria content of
breast cancer cells might potentially be clinically relevant in
breast cancer.
The mitochondrially encoded cytochrome c
oxidase II (MTCO2) monoclonal antibody recognizes a 60 kDa
non-glycosylated protein subunit of cytochrome c oxidase in
mitochondria found in human cells and has been used to reveal the
mitochondrial content of tumor cells in previous studies (24,28,29).
We tested the clinical relevance of the cellular mitochondria
content in breast cancer on a pre-existing breast cancer tissue
microarray (TMA) containing more than 2,000 cancers. The data show
that a ʻmitochondrion-rich phenotype’ represents a strong and
independent predictor of patient prognosis in breast cancer.
Materials and methods
Patients
A total of 2,197 human breast cancer samples from
paraffin-embedded tissue specimens fixed in 4% neutral buffered
formalin were used (30). The
breast cancer samples were consecutively collected between 1984 and
2000 and follow-up data were retrospectively collected. The median
patient's age was 63 (range, 25-101) years. Overall survival data
were available from 1,982 patients (713 patients with and 1,508
without event). The mean follow-up time was 63 months (range, 1-176
months). The TMA was produced as detailed earlier in (31). In short, from each patient one 0.6
mm core was taken from a representative cancer tissue block. All
tissues were distributed among 6 TMA blocks, each containing
263-522 tumor samples. Four-micrometre sections of the TMA blocks
were transferred to an adhesive coated slide system (Instrumedics
Inc.) for immunohistochemistry (IHC) analysis. Molecular data used
in this study were available from previously published studies.
These included amplification/deletion data obtained by fluorescence
in situ hybridization for HER2, MYC, 8p21,
9p21, and PTEN, as well as Ki67LI (30,32-34).
IHC
Freshly cut TMA sections were processed the same
day. Slides were deparaffinized and exposed to heat-induced antigen
retrieval for 5 min at 121˚C in pH 7.8 Tris-EDTA-Citrate buffer
prior to incubation with the mouse monoclonal antibody MTCO2
(Abcam; #ab3298; 1/450 dilution). Bound antibody was visualized
using the EnVision kit (Dako). MTCO2 staining was homogenous in the
analyzed tissue samples and staining intensity of all cases was
semiquantitatively assessed in four categories: Negative (no
visible staining), weak (1+ staining intensity), moderate (2+
stainong intensity) and strong (3+ staining intensity).
Statistical analysis
Contingency tables were calculated to study
associations between MTCO2 expression and clinicopathological
variables, and the chi-square (likelihood) test was used to find
significant relationships. Analysis of variance and F-test was
applied to find associations between MTCO2 staining levels and
tumor cell proliferation as measured by the Ki67LI. Kaplan-Meier
curves were generated using overall survival as the clinical
endpoint. The log-rank test was applied to test the significance of
differences between stratified survival functions. Cox proportional
hazards regression analysis was performed to test the statistical
independence and significance between pathological and molecular
variables. JMP 12.0 software (SAS Institute Inc.) was used.
Results
Technical issues
A total of 1,797 (81.8%) of the 2,197 arrayed tumor
samples were interpretable in our TMA analysis. Non-informative
cases (400 spots; 18.2%) were due to missing tissue samples or the
absence of unequivocal cancer tissue in the TMA spot.
MTCO2 immunostaining in normal breast
tissue and breast cancer
There were 20 normal breast tissue samples included
in our TMA. Normal breast tissues showed negative to moderate MTCO2
staining in luminal cells under the chosen experimental conditions.
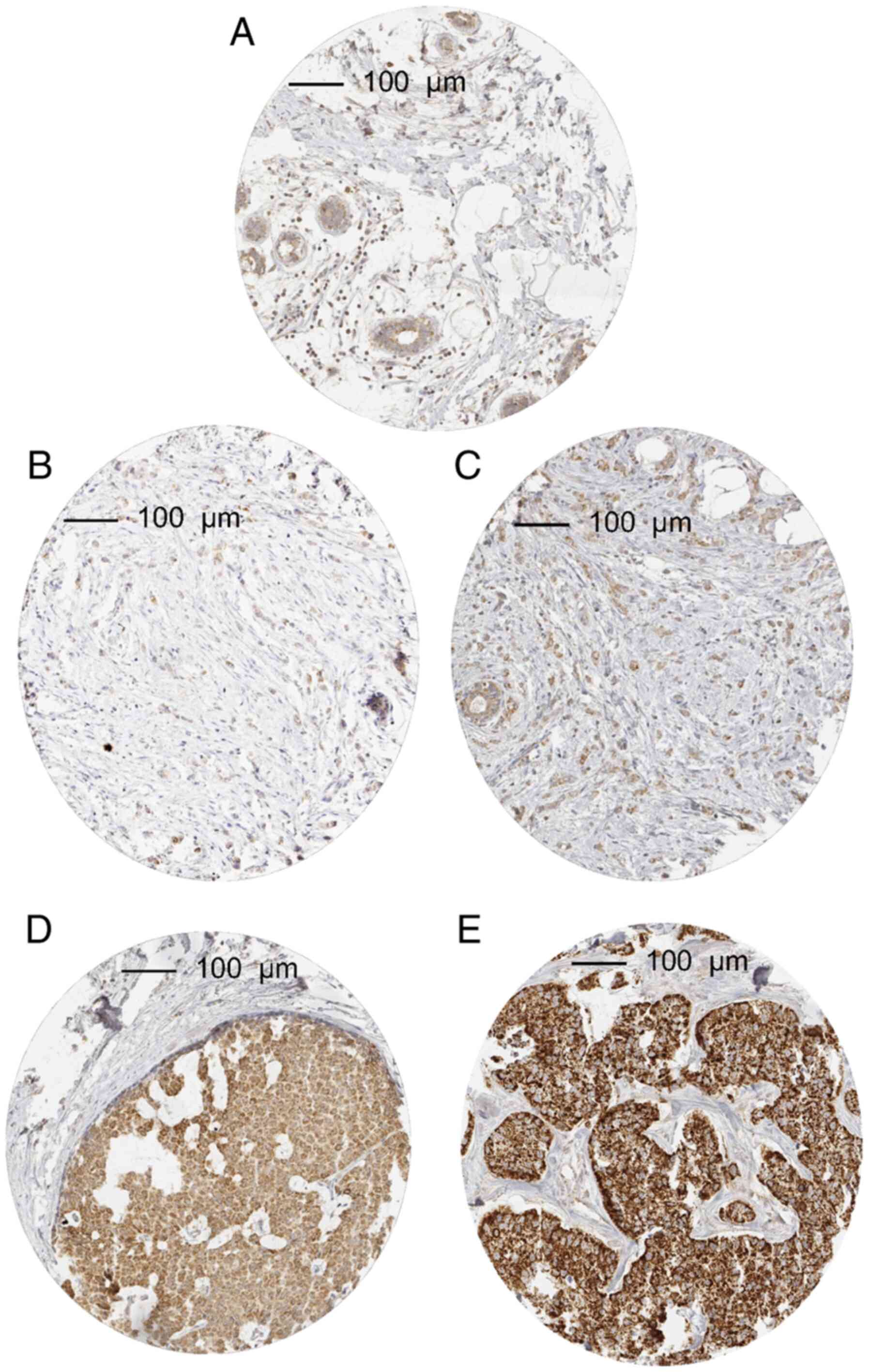
In cancer, MTCO2 immunostaining was considered weak in 34.6%,
moderate in 22.3% and strong in 15.0% of tumors. A total of 506
(28.2%) showed no detectable MTCO2 staining and were categorized as
negative. Characteristic images of MTCO2 immunostainings are shown
in Fig. 1. The intensity of MTCO2
immunostaining varied between histological breast cancer subtypes
(Table I). Strong MTCO2 staining
was significantly more common in medullary (27.9%), papillary
(16.0%) and cancers of no special type (NST; 16.6%) than in lobular
(6.9%) or tubular carcinomas (4.9%). Strong MTCO2 staining was also
commonly seen in some of the rare breast cancer subtypes such as in
3 of 13 carcinomas with apocrine differentiation, 17 of 61
carcinomas with medullary features and 2 of 12 glycogen-rich clear
cell type carcinomas (Table
SI).
 | Table IAssociation between MTCO2 staining
and breast cancer phenotype. |
Table I
Association between MTCO2 staining
and breast cancer phenotype.
| | MTCO2 staining | |
|---|
|
Characteristics | N | Negative, % | Weak, % | Moderate, % | Strong, % | P-value |
|---|
| All cases | 1,797 | 28.2 | 34.6 | 22.3 | 15.0 | |
| Histology | | | | | | |
|
NST | 1,281 | 24.2 | 36.0 | 23.2 | 16.6 | |
|
Lobular
carcinoma | 233 | 46.4 | 33.1 | 13.7 | 6.9 |
<0.0001a |
|
Medullary
carcinoma | 61 | 18.0 | 26.2 | 27.9 | 27.9 | 0.0761a |
|
Cribriform
carcinoma | 55 | 34.6 | 27.3 | 21.8 | 16.4 | 0.3461a |
|
Mucinous
carcinoma | 51 | 45.1 | 33.3 | 19.6 | 2.0 | 0.0005a |
|
Tubular
carcinoma | 41 | 48.8 | 39.0 | 7.3 | 4.9 | 0.0004a |
|
Papillary
carcinoma | 25 | 20.0 | 28.0 | 36.0 | 16.0 | 0.5425a |
|
Apocrine
carcinoma | 13 | 23.1 | 15.4 | 38.5 | 23.1 | 0.3460a |
|
Other rare
typesb | 22 | 9.1 | 27.3 | 50.0 | 13.6 | 0.0399a |
| pT stage | | | | | | |
|
pT1 | 631 | 36.3 | 40.3 | 17.6 | 5.9 | <0.0001
(<0.0001c) |
|
pT2 | 851 | 24.2 | 32.1 | 24.7 | 19.0 | |
|
pT3 | 98 | 25.5 | 30.6 | 24.5 | 19.4 | |
|
pT4 | 209 | 21.1 | 30.1 | 24.9 | 23.9 | |
| BRE grade | | | | | | |
|
G1 | 423 | 41.6 | 37.8 | 13.5 | 7.1 | <0.0001
(<0.0001c) |
|
G2 | 673 | 29.9 | 35.5 | 23.5 | 11.1 | |
|
G3 | 564 | 15.4 | 26.6 | 29.6 | 28.4 | |
| Nodal stage | | | | | | |
|
pN0 | 761 | 33.5 | 34.2 | 22.7 | 9.6 | <0.0001
(0.0063c) |
|
pN1 | 644 | 25.6 | 36.2 | 20.3 | 17.9 | |
|
pN2 | 103 | 18.5 | 38.8 | 24.3 | 18.5 | |
|
pN3 | 0 | 0.0 | 0.0 | 0.0 | 0.0 | |
| Estrogen
receptor | | | | | | |
|
Negative | 406 | 16.01 | 29.31 | 28.82 | 25.86 | <0.0001
(<0.0001c) |
|
Positive | 1,296 | 31.33 | 36.5 | 20.45 | 11.73 | |
| Progesterone
receptor | | | | | | |
|
Negative | 1,059 | 29.27 | 32.01 | 22.95 | 15.77 | <0.0001
(0.0795c) |
|
Positive | 569 | 26.19 | 40.07 | 19.68 | 14.06 | |
Association with tumor phenotype and
molecular features
High levels of MTCO2 immunostaining were
significantly associated with high pT stage, high BRE grade,
estrogen and progesterone receptor negativity as well as HER2
overexpression or amplification (P<0.0001 each, Tables I and II). This was also seen for NST carcinomas
(P≤0.01, Table I). Further analyses
with previously described frequent and prognostic relevant
molecular features of breast cancers such as HER2(35), and c-MYC- amplification (32) as well as deletions of 8p21(34), 9p21(33), and 10q23(36) showed significant associations with
high MTCO2 staining intensity (Table
II).
 | Table IIAssociation between MTCO2 staining
and molecular alterations. |
Table II
Association between MTCO2 staining
and molecular alterations.
| | MTCO2 staining | |
|---|
| Molecular
alterations | N | Negative, % | Weak, % | Moderate, % | Strong, % | P-value |
|---|
| HER2 normal | 1,141 | 29.2 | 36.3 | 21.2 | 13.3 | <0.0001 |
| HER2 amplified | 239 | 15.5 | 32.6 | 30.1 | 21.8 | |
| MYC normal | 1,232 | 26.9 | 34.9 | 23.1 | 15.1 | <0.0001 |
| MYC amplified | 64 | 7.8 | 29.7 | 28.1 | 34.4 | |
| 8p21 normal | 578 | 27.7 | 39.1 | 20.9 | 12.3 | <0.0001 |
| 8p21 deletion | 553 | 17.0 | 27.7 | 29.5 | 25.9 | |
| 9p21 normal | 835 | 25.0 | 33.1 | 24.4 | 17.5 | 0.0182 |
| 9p21 deletion | 150 | 16.7 | 28.0 | 32.0 | 23.3 | |
| 10q23 normal | 904 | 25.0 | 35.0 | 22.9 | 17.1 | <0.0001 |
| 10q23 deletion | 216 | 11.6 | 26.4 | 36.1 | 25.9 | |
Association with tumor cell
proliferation
Data on tumor cell proliferation as evaluated by the
Ki67LI were available from a previous study with the same TMA
(30). The mean Ki67LI increased
from 19.62±0.66 in MTCO2 negative cancers to 37.75±0.93 in cancers
with strong MTCO2 staining (P<0.0001). This statistically
significant relationship was also seen in tumor subsets with
identical pT or pN stage, lobular and medullary carcinoma, BRE
grade and HER2 status as well as 8p and PTEN
deletion. All data are summarized in Table III.
 | Table IIIAssociation between MTCO2 staining
and Ki67LI. |
Table III
Association between MTCO2 staining
and Ki67LI.
| Cases | MTCO2 staining | N | Ki67LI | P-value |
|---|
| All cases | Negative | 428 | 19.6±0.7 | <0.0001 |
| | Weak | 523 | 27.0±0.6 | |
| | Moderate | 338 | 33.0±0.8 | |
| | Strong | 216 | 37.8±0.9 | |
| No special
type | Negative | 264 | 20.7±0.8 | <0.0001 |
| | Weak | 383 | 27.8±0.7 | |
| | Moderate | 253 | 33.3±0.9 | |
| | Strong | 168 | 38.0±1.1 | |
| Lobular cancer | Negative | 92 | 16.2±1.2 | <0.0001 |
| | Weak | 66 | 20.3±1.4 | |
| | Moderate | 24 | 28.4±2.3 | |
| | Strong | 15 | 26.9±2.9 | |
| Medullary
cancer | Negative | 9 | 29.9±5.2 | 0.0109 |
| | Weak | 15 | 43.7±4.1 | |
| | Moderate | 16 | 50.2±3.9 | |
| | Strong | 15 | 50.9±4.0 | |
| HER2 amplified | Negative | 32 | 26.7±2.3 | <0.0001 |
| | Weak | 67 | 34.2±1.6 | |
| | Moderate | 64 | 40.3±1.6 | |
| | Strong | 43 | 41.3±1.9 | |
| MYC amplified | Negative | 4 | 28.5±7.4 | 0.3927 |
| | Weak | 19 | 38.3±3.4 | |
| | Moderate | 17 | 41.6±3.6 | |
| | Strong | 21 | 41.6±3.2 | |
| 8p deletion | Negative | 86 | 24.8±1.5 | <0.0001 |
| | Weak | 135 | 30.2±1.2 | |
| | Moderate | 145 | 35.3±1.2 | |
| | Strong | 116 | 40.3±1.3 | |
| PTEN deletion | Negative | 24 | 30.6±3.2 | 0.0118 |
| | Weak | 55 | 37.7±2.1 | |
| | Moderate | 75 | 41.7±1.8 | |
| | Strong | 44 | 42.2±2.3 | |
| pT1 | Negative | 192 | 19.0±0.9 | <0.0001 |
| | Weak | 200 | 23.8±0.9 | |
| | Moderate | 90 | 29.9±1.3 | |
| | Strong | 31 | 37.8±2.3 | |
| pT2 | Negative | 170 | 19.9±1.1 | <0.0001 |
| | Weak | 238 | 29.6±0.9 | |
| | Moderate | 179 | 35.3±1.1 | |
| | Strong | 127 | 37.9±1.3 | |
| pT3 | Negative | 23 | 18.2±3.1 | <0.0001 |
| | Weak | 27 | 31.2±2.9 | |
| | Moderate | 22 | 30.1±3.2 | |
| | Strong | 16 | 43.8±3.7 | |
| pT4 | Negative | 41 | 21.9±2.2 | <0.0001 |
| | Weak | 57 | 25.3±1.7 | |
| | Moderate | 44 | 31.9±1.9 | |
| | Strong | 41 | 34.9±2.1 | |
| BRE G1 | Negative | 150 | 15.5±0.8 | <0.0001 |
| | Weak | 127 | 19.5±0.9 | |
| | Moderate | 45 | 21.4±1.5 | |
| | Strong | 25 | 26.4±1.9 | |
| BRE G2 | Negative | 170 | 18.8±0.9 | <0.0001 |
| | Weak | 208 | 23.7±0.8 | |
| | Moderate | 134 | 28.9±01.0 | |
| | Strong | 63 | 31.4±1.4 | |
| BRE G3 | Negative | 76 | 29.4±1.7 | <0.0001 |
| | Weak | 130 | 37.5±1.3 | |
| | Moderate | 145 | 40.4±1.2 | |
| | Strong | 124 | 43.2±1.3 | |
| pN0 | Negative | 219 | 19.5±0.9 | <0.0001 |
| | Weak | 216 | 26.5±0.9 | |
| | Moderate | 144 | 34.5±1.1 | |
| | Strong | 59 | 38.9±1.8 | |
| pN1 | Negative | 138 | 19.4±1.2 | <0.0001 |
| | Weak | 198 | 27.3±1.0 | |
| | Moderate | 111 | 32.2±1.3 | |
| | Strong | 92 | 39.1±1.4 | |
| pN2 | Negative | 18 | 25.6±3.2 | 0.0064 |
| | Weak | 34 | 29.4±2.3 | |
| | Moderate | 23 | 33.4±2.8 | |
| | Strong | 16 | 41.4±3.4 | |
Prognostic significance of MTCO2
expression
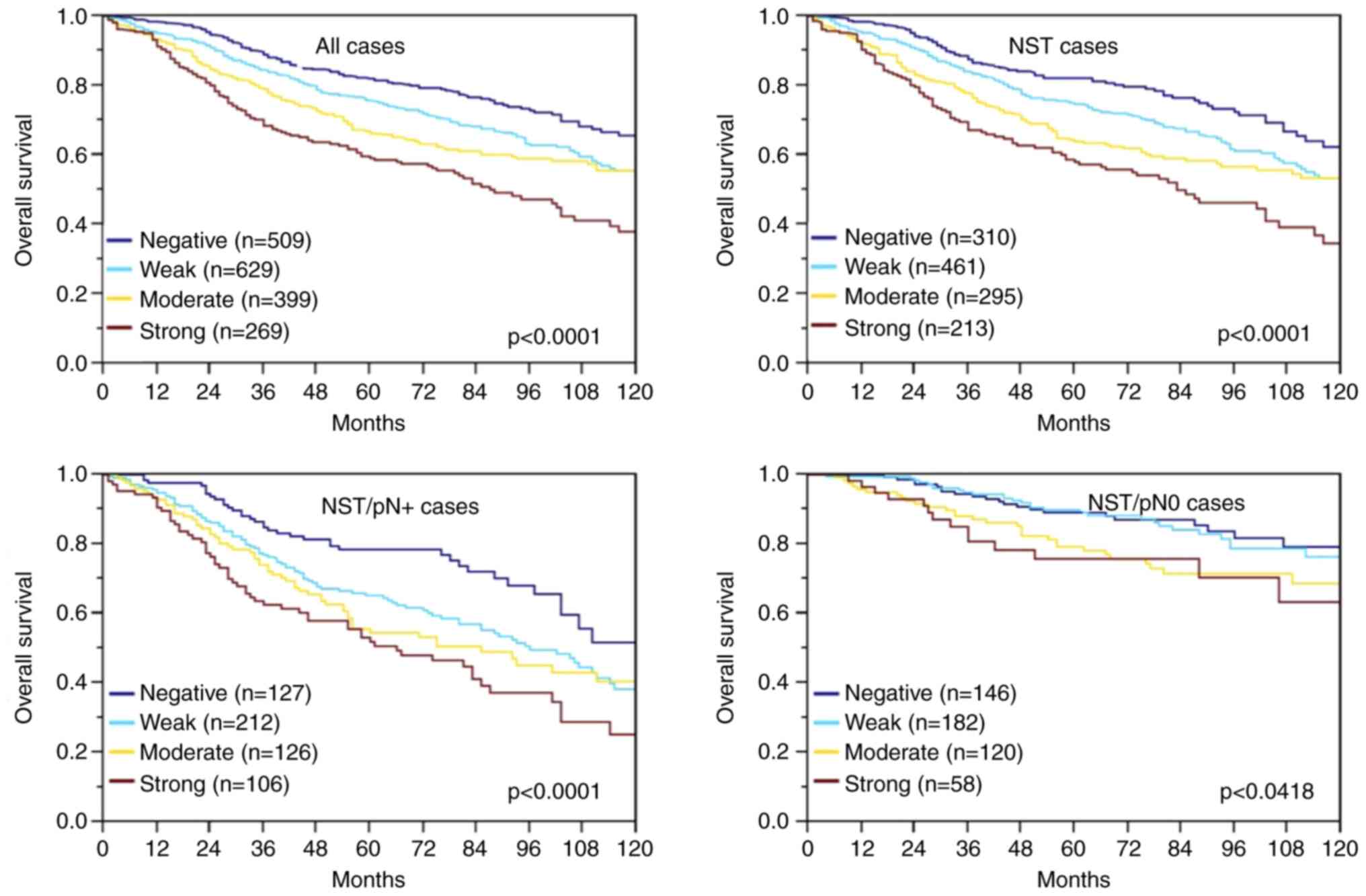
Survival data were available for 1,806 cancers with
interpretable IHC results. The rate of surviving patients
continuously decreased with increasing levels of MTCO2
immunostaining (P=0.0001; Fig. 2).
The association between strong MTCO2 immunostaining and poor
prognosis was also seen in the subgroup of NST cancers
(P<0.0001; Fig. 2) and in the
nodal positive subset (P<0.0001; Fig. 2) and to a much lesser extent also in
nodal negative NST cancers (P=0.0418; Fig. 2). Multivariate analysis for NST
cancers including pT stage, nodal status, and BRE grade did not
identify MTCO2 immunostaining as an independent prognosticator of
survival, however (Table IV).
 | Table IVMultivariate analysis in all breast
cancer cases (n=1,377). |
Table IV
Multivariate analysis in all breast
cancer cases (n=1,377).
|
Characteristics | Hazard ratio | P-value | Overall
P-value |
|---|
| pT stage | | | |
|
2 vs. 1 | 1.52 | 0.0010 | <0.0001 |
|
3 vs. 2 | 1.05 | 0.7700 | |
|
4 vs. 2 | 1.65 | 0.0006 | |
|
4 vs. 3 | 1.56 | 0.2230 | |
| BRE grade | | | <0.0001 |
|
G2 vs.
G1 | 1.35 | 0.0522 | |
|
G3 vs.
G1 | 2.81 | <0.0001 | |
|
G3 vs.
G2 | 2.08 | <0.0001 | |
| pN | | | <0.0001 |
|
1 vs. 0 | 2.26 | <0.0001 | |
|
2 vs. 1 | 2.33 | <0.0001 | |
|
2 vs. 0 | 5.27 | <0.0001 | |
| MTCO2 staining | | | 0.1464 |
|
Weak vs.
negative | 1.33 | 0.0396 | |
|
Moderate vs.
weak | 1.01 | 0.9109 | |
|
Strong vs.
moderate | 0.88 | 0.4133 | |
Discussion
Our study shows that high mitochondria content is
significantly linked to disadvantageous tumor phenotype and bad
prognosis in breast cancer.
MTCO2 immunostaining is highly specific for the
mitochondrial DNA encoded second subunit of cytochrome c
oxidase and can thus be used to quantitate the mitochondria content
by IHC (29). Although mitochondria
are present in every normal and neoplastic human cell, 28.2% of our
tumors had a negative staining result. This was due to our approach
to define experimental conditions, which distinguish cancers with
low and high mitochondria quantities. The higher level of MTCO2
immunostaining in breast cancers as compared to normal breast
tissues fits with the concept that neoplastic transformation goes
along with higher cellular activity requiring more active
mitochondria. That a striking further increase of MTCO2
immunostaining was detected with rising tumor grade and stage,
demonstrates that elevated numbers of mitochondria are also
supporting cancer progression. This is consistent with increasing
energy requirement and a rearranged metabolism during tumor
progression. Our data fit well with findings in multiple other
cancer types, including lung (21),
colorectal (22,23), prostate (24), gastric (25), cervical (18), and ovarian cancer (26), where a similar link between high
levels of MTCO2 with adverse tumor phenotype and bad prognosis was
shown.
In this study, a ubiquitously expressed protein was
quantitated by IHC. The TMA approach is optimal for the
identification of subtle staining differences of proteins that are
abundantly present in cancer, such as mitochondrial components,
because TMAs enable maximal experimental standardization at all
levels. In our study, more than 1,700 breast cancers were analyzed
the same day for maximal standardization. Moreover, all TMA
sections were cut on one day immediately before staining in order
to avoid unequal decay of a tissues reactivity to antibody binding
(37). Finally, one pathologist
interpreted all immunostainings in one continuous session to enable
maximal standardization of staining interpretation. In earlier
studies, this breast cancer TMA enabled us to validate the
prognostic impact of several well-established prognostic
biomarkers, such as HER2 alterations, estrogen and
progesterone receptor expression (30), high Ki67LI, nuclear p53 accumulation
(30), and PTEN deletion
(34). These earlier data
demonstrate the utility of our patient cohort to identify
prognostic biomarkers.
The molecular database that has been collected
during earlier studies for our set of cancers offers the advantage
that biomarkers of interest can always be compared with preexisting
data. For the purpose of this study, we had selected HER2
amplification as well as estrogen and progesterone receptor
expression because of their central role in breast cancer. The
strong link between MTCO2 expression and these important features
further illustrates the importance of the mitochondria quantity in
breast cancer. Our analyses also included Ki67LI as another pivotal
parameter for cellular activity and various further chromosomal
deletions and amplifications because of the role of some of them
for regulating mitochondrion homeostasis.
Mitochondrial homeostasis is critical for cancer. A
sufficiently high production of mitochondria is required to suffice
the needs for energy production and cell metabolism. The prominent
association found between c-Myc amplification and high MTCO2
expression fits well with the key role of c-Myc as an activator of
mitochondrial biogenesis in cancer (38-40).
The transcription factor c-Myc is best known for its critical role
in cell cycle regulation, cell growth, metabolism and apoptosis
(41-43).
However, c-Myc also targets more than 400 different mitochondrial
genes (38-41,44).
Studies have demonstrated that an elevated or reduced c-Myc protein
quantity leads to an increased/diminished mitochondrial mass
(45,46). This couples c-Myc's role of a key
activator of cell cycle activity with mitochondrial biogenesis. As
such, c-Myc increases cellular biosynthetic and respiratory
capacity by upregulating mitochondrial metabolism to complement its
effects on stimulating cell cycle progression to coordinate rapid
cell growth (45,47).
A critical role of high mitochondrion count for cell
proliferation in breast cancer is supported by our data showing a
striking link between MTCO2 expression and a high Ki67LI which was
also visible in the vast majority of groups defined by identical
morphological or molecular features.
The PTEN-induced putative kinase 1 (PINK1)/Parkin
pathway is a major inducer of mitophagy. It is triggered by
mitochondrial membrane depolarization, a signal of mitochondrial
dysfunction that results from lack of reducing equivalents, hypoxia
and impaired electron transport [reviewed in (48)]. The conspicuous relationship between
PTEN deletion and high MTCO2 staining in our study may thus
indicate that high mitochondria quantities may also be caused by
reduced mitophagy. Although clearance of damaged mitochondria via
mitophagy is viewed to be also critical for cellular fitness since
dysfunctional mitochondria can impair the electron transport chain
function, reduced mitophagy can also promote cancer reviewed in
ref. 49). Mitophagy-deficient Parkin null mice develop spontaneous
hepatic tumors (50). Decreased
mitophagy may allow for a permissive threshold of dysfunctional
mitochondria to persist, generating increased tumor-promoting free
oxygen radicals reviewed in ref. 49).
Cytochrome oxidase subunit 2 is a key enzyme of the
respiratory chain, catalyzing electron transfer from NADH and
succinate to molecular oxygen (51). It has no direct tumor related
function but serves as a marker for the cellular mitochondria
content. Increased mitochondria content in cancer cells often
occurs as a result of the elevated metabolism and energy needs of
expanding tumor cell populations (52). Although the mitochondrial content
provided no additional prognostic information in multivariate
analysis, the marked prognostic relevance of MTCO2 immunostaining
found in this study may still suggest ʻmitochondria content’ as a
biomarker with potential clinical utility. Molecular analyses are
frequently done in breast cancer to better assess patient prognosis
and to determine whether adjuvant chemotherapy should be applied
(6-8).
Most currently used tests are analyzing RNAs of multiple genes
forming a prognostic score (9-11,53).
RNA based tests share the disadvantage, however, that the analyzed
RNA always represents a mixture of cancer cells and a variable
fraction of non-neoplastic inflammatory and stromal cells. Now that
multiplex fluorescent-based quantitative IHC becomes increasingly
available, it is well possible that RNA based test will sooner or
later be replaced by IHC based multi-gene tests. MTCO2 might be a
candidate for being part of such a test, also because of the
general biologic importance of mitochondria, which are also the
target of several anti-cancer drugs under development reviewed in
refs. 54-57).
It is a limitation of our study that MTCO2 IHC data
highlight relevant associations between cancer phenotype and
genotype but do not provide mechanistic insights into the putative
cancer biological role of MTCO2. Further studies on the tumor
relevant aspects of mitochondrial density and MTCO2 protein
function are required to better understand the prognostic role of
MTCO2 in breast cancer.
In summary, our findings identify MTCO2
immunostaining as a powerful prognostic biomarker in breast cancer.
MTCO2 measurement, most likely in combination with other antibodies
might be of clinical utility in breast cancer prognosis
assessment.
Supplementary Material
MTC02 staining in rare breast cancer
subtypes.
Acknowledgements
The authors would like to thank Ms. Inge Brandt and
Ms. Sünje Seekamp from the Institute of Pathology of University
Medical Center Hamburg-Eppendorf (Hamburg, Germany) for excellent
technical assistance.
Funding
No funding was received.
Availability of data and materials
The datasets used and/or analyzed during the current
study are available from the corresponding author on reasonable
request.
Authors' contributions
All authors contributed to the conception and design
of the study. PL, KS, MK, IW, LW, PP, LT, CW, UH, VM, BS, IvL, TK,
RHK and FJ prepared the material, and collected and analyzed the
data. PL, EB, RS, MK and GS wrote the first draft of the
manuscript, and all authors commented on previous versions of the
manuscript. RS, MK and GS confirmed the authenticity of all the raw
data. All authors read and approved the final manuscript.
Ethics approval and consent to
participate
The usage of archived diagnostic leftover tissues
for manufacturing the tissue microarrays and their analysis for
research purposes, as well as patient data analysis, has been
approved by local laws (HmbKHG, §12) and by the local ethics
committee (Ethics Commission of the Ärtzekammer Hamburg, Hamburg,
Germany; approval no. WF-049/09). Informed consent was waived by
the ethics committee due to the retrospective nature of the study.
All work has been carried out in compliance with the Declaration of
Helsinki.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Siegel RL, Miller KD and Jemal A: Cancer
statistics, 2020. CA Cancer J Clin. 70:7–30. 2020.PubMed/NCBI View Article : Google Scholar
|
|
2
|
Leong AS and Raymond WA: Prognostic
parameters in breast cancer. Pathology. 21:169–175. 1989.PubMed/NCBI View Article : Google Scholar
|
|
3
|
Taneja P, Maglic D, Kai F, Zhu S, Kendig
RD, Fry EA and Inoue K: Classical and novel prognostic markers for
breast cancer and their clinical significance. Clin Med Insights
Oncol. 4:15–34. 2010.PubMed/NCBI View Article : Google Scholar
|
|
4
|
Soliman NA and Yussif SM: Ki-67 as a
prognostic marker according to breast cancer molecular subtype.
Cancer Biol Med. 13:496–504. 2016.PubMed/NCBI View Article : Google Scholar
|
|
5
|
Cao SS and Lu CT: Recent perspectives of
breast cancer prognosis and predictive factors. Oncol Lett.
12:3674–3678. 2016.PubMed/NCBI View Article : Google Scholar
|
|
6
|
Giuliano AE, Hunt KK, Ballman KV, Beitsch
PD, Whitworth PW, Blumencranz PW, Leitch AM, Saha S, McCall LM and
Morrow M: Axillary dissection vs no axillary dissection in women
with invasive breast cancer and sentinel node metastasis: A
randomized clinical trial. JAMA. 305:569–575. 2011.PubMed/NCBI View Article : Google Scholar
|
|
7
|
McVeigh TP, Hughes LM, Miller N, Sheehan
M, Keane M, Sweeney KJ and Kerin MJ: The impact of Oncotype DX
testing on breast cancer management and chemotherapy prescribing
patterns in a tertiary referral centre. Eur J Cancer. 50:2763–2770.
2014.PubMed/NCBI View Article : Google Scholar
|
|
8
|
Naoi Y and Noguchi S: Multi-gene
classifiers for prediction of recurrence in breast cancer patients.
Breast Cancer. 23:12–18. 2016.PubMed/NCBI View Article : Google Scholar
|
|
9
|
Hornberger J, Cosler LE and Lyman GH:
Economic analysis of targeting chemotherapy using a 21-gene RT-PCR
assay in lymph-node-negative, estrogen-receptor-positive,
early-stage breast cancer. Am J Manag Care. 11:313–324.
2005.PubMed/NCBI
|
|
10
|
Cobleigh MA, Tabesh B, Bitterman P, Baker
J, Cronin M, Liu ML, Borchik R, Mosquera JM, Walker MG and Shak S:
Tumor gene expression and prognosis in breast cancer patients with
10 or more positive lymph nodes. Clin Cancer Res. 11:8623–8631.
2005.PubMed/NCBI View Article : Google Scholar
|
|
11
|
van 't Veer LJ, Dai H, van de Vijver MJ,
He YD, Hart AA, Mao M, Peterse HL, van der Kooy K, Marton MJ,
Witteveen AT, et al: Gene expression profiling predicts clinical
outcome of breast cancer. Nature. 415:530–536. 2002.PubMed/NCBI View
Article : Google Scholar
|
|
12
|
Davis RE and Williams M: Mitochondrial
function and dysfunction: An update. J Pharmacol Exp Ther.
342:598–607. 2012.PubMed/NCBI View Article : Google Scholar
|
|
13
|
Hsu CC, Tseng LM and Lee HC: Role of
mitochondrial dysfunction in cancer progression. Exp Biol Med
(Maywood). 241:1281–1295. 2016.PubMed/NCBI View Article : Google Scholar
|
|
14
|
Boland ML, Chourasia AH and Macleod KF:
Mitochondrial dysfunction in cancer. Front Oncol.
3(292)2013.PubMed/NCBI View Article : Google Scholar
|
|
15
|
Porporato PE, Filigheddu N, Pedro JMB,
Kroemer G and Galluzzi L: Mitochondrial metabolism and cancer. Cell
Res. 28:265–280. 2018.PubMed/NCBI View Article : Google Scholar
|
|
16
|
Grasso D, Zampieri LX, Capeloa T, Van de
Velde JA and Sonveaux P: Mitochondria in cancer. Cell Stress.
4:114–146. 2020.PubMed/NCBI View Article : Google Scholar
|
|
17
|
Abu-Amero KK, Alzahrani AS, Zou M and Shi
Y: High frequency of somatic mitochondrial DNA mutations in human
thyroid carcinomas and complex I respiratory defect in thyroid
cancer cell lines. Oncogene. 24:1455–1460. 2005.PubMed/NCBI View Article : Google Scholar
|
|
18
|
Warowicka A, Kwasniewska A and
Gozdzicka-Jozefiak A: Alterations in mtDNA: A qualitative and
quantitative study associated with cervical cancer development.
Gynecol Oncol. 129:193–198. 2013.PubMed/NCBI View Article : Google Scholar
|
|
19
|
Gao JY, Song BR, Peng JJ and Lu YM:
Correlation between mitochondrial TRAP-1 expression and lymph node
metastasis in colorectal cancer. World J Gastroenterol.
18:5965–5971. 2012.PubMed/NCBI View Article : Google Scholar
|
|
20
|
Qian XL, Li YQ, Gu F, Liu FF, Li WD, Zhang
XM and Fu L: Overexpression of ubiquitous mitochondrial creatine
kinase (uMtCK) accelerates tumor growth by inhibiting apoptosis of
breast cancer cells and is associated with a poor prognosis in
breast cancer patients. Biochem Biophys Res Commun. 427:60–66.
2012.PubMed/NCBI View Article : Google Scholar
|
|
21
|
Sotgia F and Lisanti MP: Mitochondrial
markers predict survival and progression in non-small cell lung
cancer (NSCLC) patients: Use as companion diagnostics. Oncotarget.
8:68095–68107. 2017.PubMed/NCBI View Article : Google Scholar
|
|
22
|
Ambrosini-Spaltro A, Salvi F, Betts CM,
Frezza GP, Piemontese A, Del Prete P, Baldoni C, Foschini MP and
Viale G: Oncocytic modifications in rectal adenocarcinomas after
radio and chemotherapy. Virchows Arch. 448:442–448. 2006.PubMed/NCBI View Article : Google Scholar
|
|
23
|
Wang Y, He S, Zhu X, Qiao W and Zhang J:
High copy number of mitochondrial DNA predicts poor prognosis in
patients with advanced stage colon cancer. Int J Biol Markers.
31:e382–e388. 2016.PubMed/NCBI View Article : Google Scholar
|
|
24
|
Grupp K, Jedrzejewska K, Tsourlakis MC,
Koop C, Wilczak W, Adam M, Quaas A, Sauter G, Simon R, Izbicki JR,
et al: High mitochondria content is associated with prostate cancer
disease progression. Mol Cancer. 12(145)2013.PubMed/NCBI View Article : Google Scholar
|
|
25
|
Sotgia F and Lisanti MP: Mitochondrial
biomarkers predict tumor progression and poor overall survival in
gastric cancers: Companion diagnostics for personalized medicine.
Oncotarget. 8:67117–67128. 2017.PubMed/NCBI View Article : Google Scholar
|
|
26
|
Hu B and Guo Y: Inhibition of
mitochondrial translation as a therapeutic strategy for human
ovarian cancer to overcome chemoresistance. Biochem Biophys Res
Commun. 509:373–378. 2019.PubMed/NCBI View Article : Google Scholar
|
|
27
|
Zhang Y, Qu Y, Gao K, Yang Q, Shi B, Hou P
and Ji M: High copy number of mitochondrial DNA (mtDNA) predicts
good prognosis in glioma patients. Am J Cancer Res. 5:1207–1216.
2015.PubMed/NCBI
|
|
28
|
Ragazzi M, de Biase D, Betts CM, Farnedi
A, Ramadan SS, Tallini G, Reis-Filho JS and Eusebi V: Oncocytic
carcinoma of the breast: Frequency, morphology and follow-up. Hum
Pathol. 42:166–175. 2011.PubMed/NCBI View Article : Google Scholar
|
|
29
|
Williams SL, Valnot I, Rustin P and
Taanman JW: Cytochrome c oxidase subassemblies in fibroblast
cultures from patients carrying mutations in COX10, SCO1, or SURF1.
J Biol Chem. 279:7462–7469. 2004.PubMed/NCBI View Article : Google Scholar
|
|
30
|
Ruiz C, Seibt S, Al Kuraya K, Siraj AK,
Mirlacher M, Schraml P, Maurer R, Spichtin H, Torhorst J, Popovska
S, et al: Tissue microarrays for comparing molecular features with
proliferation activity in breast cancer. Int J Cancer.
118:2190–2194. 2006.PubMed/NCBI View Article : Google Scholar
|
|
31
|
Mirlacher M and Simon R: Recipient block
TMA technique. Methods Mol Biol. 664:37–44. 2010.PubMed/NCBI View Article : Google Scholar
|
|
32
|
Al-Kuraya K, Schraml P, Torhorst J, Tapia
C, Zaharieva B, Novotny H, Spichtin H, Maurer R, Mirlacher M,
Köchli O, et al: Prognostic relevance of gene amplifications and
coamplifications in breast cancer. Cancer Res. 64:8534–8540.
2004.PubMed/NCBI View Article : Google Scholar
|
|
33
|
Lebok P, Roming M, Kluth M, Koop C, Özden
C, Taskin B, Hussein K, Lebeau A, Witzel I, Wölber L, et al: p16
overexpression and 9p21 deletion are linked to unfavorable tumor
phenotype in breast cancer. Oncotarget. 7:81322–81331.
2016.PubMed/NCBI View Article : Google Scholar
|
|
34
|
Lebok P, Mittenzwei A, Kluth M, Özden C,
Taskin B, Hussein K, Möller K, Hartmann A, Lebeau A, Witzel I, et
al: 8p deletion is strongly linked to poor prognosis in breast
cancer. Cancer Biol Ther. 16:1080–1087. 2015.PubMed/NCBI View Article : Google Scholar
|
|
35
|
Yan M, Schwaederle M, Arguello D, Millis
SZ, Gatalica Z and Kurzrock R: HER2 expression status in diverse
cancers: Review of results from 37,992 patients. Cancer Metastasis
Rev. 34:157–164. 2015.PubMed/NCBI View Article : Google Scholar
|
|
36
|
Lebok P, Kopperschmidt V, Kluth M,
Hube-Magg C, Özden C, B T, Hussein K, Mittenzwei A, Lebeau A,
Witzel I, et al: Partial PTEN deletion is linked to poor prognosis
in breast cancer. BMC Cancer. 15:963–910. 2015.PubMed/NCBI View Article : Google Scholar
|
|
37
|
Simon R, Mirlacher M and Sauter G:
Immunohistochemical analysis of tissue microarrays. Methods Mol
Biol. 664:113–126. 2010.PubMed/NCBI View Article : Google Scholar
|
|
38
|
Nieminen AI, Partanen JI, Hau A and
Klefstrom J: c-Myc primed mitochondria determine cellular
sensitivity to TRAIL-induced apoptosis. EMBO J. 26:1055–1067.
2007.PubMed/NCBI View Article : Google Scholar
|
|
39
|
Desbiens KM, Deschesnes RG, Labrie MM,
Desfossés Y, Lambert H, Landry J and Bellmann K: c-Myc potentiates
the mitochondrial pathway of apoptosis by acting upstream of
apoptosis signal-regulating kinase 1 (Ask1) in the p38 signalling
cascade. Biochem J. 372:631–641. 2003.PubMed/NCBI View Article : Google Scholar
|
|
40
|
Klefstrom J, Verschuren E and Evan G:
c-Myc augments the apoptotic activity of cytosolic death receptor
signaling proteins by engaging the mitochondrial apoptotic pathway.
J Biol Chem. 277:43224–43232. 2002.PubMed/NCBI View Article : Google Scholar
|
|
41
|
Dang CV, O'Donnell KA, Zeller KI, Nguyen
T, Osthus RC and Li F: The c-Myc target gene network. Semin Cancer
Biol. 16:253–264. 2006.PubMed/NCBI View Article : Google Scholar
|
|
42
|
Amati B, Frank SR, Donjerkovic D and
Taubert S: Function of the c-Myc oncoprotein in chromatin
remodeling and transcription. Biochim Biophys Acta. 1471:M135–M145.
2001.PubMed/NCBI View Article : Google Scholar
|
|
43
|
Dang CV, Resar LM, Emison E, Kim S, Li Q,
Prescott JE, Wonsey D and Zeller K: Function of the c-Myc oncogenic
transcription factor. Exp Cell Res. 253:63–77. 1999.PubMed/NCBI View Article : Google Scholar
|
|
44
|
Wonsey DR, Zeller KI and Dang CV: The
c-Myc target gene PRDX3 is required for mitochondrial homeostasis
and neoplastic transformation. Proc Natl Acad Sci USA.
99:6649–6654. 2002.PubMed/NCBI View Article : Google Scholar
|
|
45
|
Li F, Wang Y, Zeller KI, Potter JJ, Wonsey
DR, O'Donnell KA, Kim JW, Yustein JT, Lee LA and Dang CV: Myc
stimulates nuclearly encoded mitochondrial genes and mitochondrial
biogenesis. Mol Cell Biol. 25:6225–6234. 2005.PubMed/NCBI View Article : Google Scholar
|
|
46
|
Graves JA, Wang Y, Sims-Lucas S, Cherok E,
Rothermund K, Branca MF, Elster J, Beer-Stolz D, Van Houten B,
Vockley J and Prochownik EV: Mitochondrial structure, function and
dynamics are temporally controlled by c-Myc. PLoS One.
7(e37699)2012.PubMed/NCBI View Article : Google Scholar
|
|
47
|
Miller DM, Thomas SD, Islam A, Muench D
and Sedoris K: c-Myc and cancer metabolism. Clin Cancer Res.
18:5546–5553. 2012.PubMed/NCBI View Article : Google Scholar
|
|
48
|
Leites EP and Morais VA: Mitochondrial
quality control pathways: PINK1 acts as a gatekeeper. Biochem
Biophys Res Commun. 500:45–50. 2018.PubMed/NCBI View Article : Google Scholar
|
|
49
|
Panigrahi DP, Praharaj PP, Bhol CS,
Mahapatra KK, Patra S, Behera BP, Mishra SR and Bhutia SK: The
emerging, multifaceted role of mitophagy in cancer and cancer
therapeutics. Semin Cancer Biol. 66:45–58. 2020.PubMed/NCBI View Article : Google Scholar
|
|
50
|
Wang H, Ni HM, Chao X, Ma X, Rodriguez YA,
Chavan H, Wang S, Krishnamurthy P, Dobrowsky R, Xu DX, et al:
Double deletion of PINK1 and Parkin impairs hepatic mitophagy and
exacerbates acetaminophen-induced liver injury in mice. Redox Biol.
22(101148)2019.PubMed/NCBI View Article : Google Scholar
|
|
51
|
Rak M, Bénit P, Chrétien D, Bouchereau J,
Schiff M, El-Khoury R, Tzagoloff A and Rustin P: Mitochondrial
cytochrome c oxidase deficiency. Clin Sci (Lond).
130:393–407. 2016.PubMed/NCBI View Article : Google Scholar
|
|
52
|
Gundamaraju R, Lu W and Manikam R:
Revisiting Mitochondria Scored Cancer Progression and Metastasis.
Cancers (Basel). 13(432)2021.PubMed/NCBI View Article : Google Scholar
|
|
53
|
Vieira AF and Schmitt F: An update on
breast cancer multigene prognostic tests-emergent clinical
biomarkers. Front Med (Lausanne). 5(248)2018.PubMed/NCBI View Article : Google Scholar
|
|
54
|
Gogvadze V, Orrenius S and Zhivotovsky B:
Mitochondria as targets for cancer chemotherapy. Semin Cancer Biol.
19:57–66. 2009.PubMed/NCBI View Article : Google Scholar
|
|
55
|
Leber B, Geng F, Kale J and Andrews DW:
Drugs targeting Bcl-2 family members as an emerging strategy in
cancer. Expert Rev Mol Med. 12(e28)2010.PubMed/NCBI View Article : Google Scholar
|
|
56
|
Indran IR, Tufo G, Pervaiz S and Brenner
C: Recent advances in apoptosis, mitochondria and drug resistance
in cancer cells. Biochim Biophys Acta. 1807:735–745.
2011.PubMed/NCBI View Article : Google Scholar
|
|
57
|
Dijk SN, Protasoni M, Elpidorou M, Kroon
AM and Taanman JW: Mitochondria as target to inhibit proliferation
and induce apoptosis of cancer cells: The effects of doxycycline
and gemcitabine. Sci Rep. 10(4363)2020.PubMed/NCBI View Article : Google Scholar
|