Introduction
Cervical cancer (CC), is one of the most common
gynecological malignancies in females and has a high death rate
worldwide (1). Each year,
>270,000 females die as a result of cervical cancer, of which
53,000 are located in China (2).
CC originates either from cervical intraepithelial neoplasia (CIN)
or squamous intraepithelial lesions (3). Recently, the occurrence rates of CC
have decreased due to novel screening, diagnostic and treatment
techniques (4). However, a high
number of females still develop and die from this disease every
year (5). Hence, new therapeutic
strategies and molecular diagnostic biomarkers for CC are required.
MicroRNAs (miRNAs/miRs) are a group of non-coding RNAs that are ~22
nucleotides long, and can inhibit mRNA translation or catalyze mRNA
cleavage to block the expression of target mRNAs (6). Therefore, miRNAs have become an area
of interest in oncology research in recent years. Key events in the
development of CC may involve the loss or acquired function of
certain miRNAs (7). miR-145, a
tumor suppressor in various types of cancer, is abnormally
expressed in gastric, ovarian, breast, prostate and lung cancer
(8–11). miR-145 overexpression can suppress
human colon cancer SW620 cells (12). miR-145 regulates the biological
function of tumor cells by regulating the expression levels of
numerous downstream genes (13);
to the best of our knowledge, however, there are no studies about
the function of miR-145 in cervical cancer. Fascin-1 (FSCN1) is an
actin-binding protein which is present in mesenchymal tissues,
however, its expression levels are limited in adult epithelial
cells (14). In the physiological
and tumor environment, fascin proteins stabilize the formation of
parallel actin bundles in cell protrusions, which represent the
initial stage of cell migration (15). FSCN1, a direct target of miR-145 in
esophageal cancer (16), is
involved in a number of cancer processes in vivo (17). For example, FSCN1 suppresses
epithelial mesenchymal transition (EMT) in human ovarian cancer
cells (18). FSCN1 also suppresses
tumors in gastric cancer (19).
Moreover, miR-145 can inactivate CC cells by targeting FSCN1
(20). However, the molecular
mechanisms underpinning the functions of miR-145-5p in CC are not
clear.
The present study aimed to investigate the
functional roles and molecular mechanisms of miR-145-5p in CC and
to reveal the association between miR-145-5p and FSCN1 in CC.
Materials and methods
Tissue collection and cell
culture
In total, 40 paired adjacent tissues (located 0.5 cm
from the cancer lesions) and CC tissues were acquired from the
First Affiliated Hospital of Nanchang University and were stored in
liquid nitrogen prior to use. All patients provided written
informed consent to participate.
ECT1/E6E7, HeLa and C33a cell lines were obtained
from the Cell Bank of Type Culture Collection of the Chinese
Academy of Sciences. These cells were incubated in DMEM (Gibco;
Thermo Fisher Scientific, Inc.) and 10% FBS (Gibco; Thermo Fisher
Scientific, Inc.), penicillin (100 units/ml) and streptomycin (100
µg/ml) at 37°C and 5% CO2. Approval was obtained from
the Ethics Committee of the First Affiliated Hospital of Nanchang
University.
Cell transfection
miR-145-5p mimics or inhibitors (100 nM) and FSCN1
small interfering (si)RNA (50 nM) (GenePharma Co., Ltd.) were
transfected into HeLa cells cultured in 6-well plates (Corning,
Inc.) at 75% confluence using Lipofectamine 3000®
reagent (Invitrogen; Thermo Fisher Scientific, Inc.) at 37°C for 24
h based on the manufacturers' instructions. The sequences of
siRNAs, mimics, inhibitors and controls are as follows: FSCN1 siRNA
sense, 5′-GCUGCUACUUUGACAUCGATT-3′ and antisense,
5′-UCGAUGUCAAAGUAGCAGCTT-3′; scrambled siRNA sense,
5′-AUAUUCCUGCGAUAGCUCGTT-3′ and antisense,
5′-CGAGCUAUCGCAGGAAUAUTT-3′; miR-145-5p mimics sense,
5′-GUCCAGUUUUCCCAGGAAUCCCU-3′ and antisense,
5′-GGAUUCCUGGGAAAACUGGACUU-3′; miR-145-5p mimics control sense
5′-UUCUCCGAACGUGUCACGUTT-3′ and antisense,
5′-ACGUGACACGUUCGGAGAATT-3′; miR-145-5p inhibitors sense,
5′-AGGGAUUCCUGGGAAAACUGGAC-3′ and miR-145-5p inhibitors control
sense, 5′-CAGUACUUUUGUGUAGUACAA-3′.
RNA extraction and RT-qPCR
Total RNA from miR-145-5p mimics or inhibitors and
FSCN1 siRNA-transfected HeLa cells was isolated using
TRIzol® reagent (Invitrogen; Thermo Fisher Scientific,
Inc.). Total RNA content was quantified spectrophotometrically,
then cDNA was prepared with a RT kit (cat. no. RR037A; Takara
Biotechnology Co., Ltd.) at 37°C for 15 min according to the
manufacturer's instructions. miR-145-5p and FSCN1 expression levels
were measured using reverse transcription-quantitative PCR
(RT-qPCR) using an ABI PRISM 7500 sequence detection system (Thermo
Fisher Scientific, Inc.) and SYBR Premix Ex Taq Master Mix (Takara
Bio, Inc.). U6 and GAPDH were used as internal controls. The
following primer pairs (Invitrogen; Thermo Fisher Scientific, Inc.)
were used for the qPCR: miR-145-5p forward,
5′-GUCCAGUUUUCCCAGGAAUCCCU-3′ and reverse,
5′-GGAUUCCUGGGAAAACGUGACUU-3′; U6 forward, 5′-CCCTGGCACCCAGCAC-3′
and reverse, 5′-GCCGATCCACACGGAGTAC-3′; FSCN1 forward,
5′-GGAGACCGACCAGGAGAC-3′ and reverse, 5′-GATTGGACGCCCTCAGTG-3′ and
GAPDH forward, 5′-CAATGACCCCTTCATTGACC-3′ and reverse,
5′-GACAAGCTTCCCGTTCTCAG-3′. A total of 0.5 µl cDNA, 0.5 µl forward
primer, 0.5 µl reverse primer, 2 µl deoxyribonucleotide
triphosphate mixture (2.5 mM), 0.5 µl DNA polymerase (10 U/µl), 6
µl 5X buffer and 15 µl ddH2O were subjected to 38 cycles
at 95°C for 5 min, 95°C for 15 sec and 56°C for 30 sec. miR-145-5p
and FSCN1 expression were analyzed using the 2−ΔΔCq
method (21).
Luciferase reporter assay
The 3′-untranslated region (3′-UTR) of FSCN1 was
located using TargetScan (targetscan.org/vert_71/) was cloned into a pGL4-empty
vector (Promega Corporation). HeLa cells were cultured in a 96-well
plate (2×104 cells/well) and co-transfected with
miR-145-5p mimics, inhibitors or negative control and the 3′-UTR
vector using Lipofectamine® 3000 (Invitrogen; Thermo
Fisher Scientific, Inc.) with Opti-MEM (Thermo Fisher Scientific,
Inc.). Subsequently, a mutant (Mut) 3′-UTR plasmid was generated at
the binding sites for miR-145-5p (Promega Corporation). After 24 h
of culture, the cells were lysed in radioimmunoprecipitation assay
buffer with protease inhibitor cocktail (Roche Diagnostics Shanghai
Co., Ltd.), and luciferase activity was detected using a dual
luciferase reporter assay kit (Promega Corporation). Firefly
luciferase activity was standardized to that of Renilla.
Cell Counting Kit (CCK)-8 assay
The growth of HeLa cells was analyzed using a CCK-8
assay (cat. no. C0038; Beyotime Institute of Biotechnology.). In
brief, cells in 24-well plates were cultured for 24 h and
transfected with miR-145-5p mimics and inhibitors. Subsequently,
transfected cells (100 µl, 1×105 cells/well) and CCK-8
solution (10 µl/well) were added to 96-well plates for 2 h of
culture at 37°C. Viable cells were measured with an Epoch
microplate absorbance tester (BioTek Instruments, Inc.) at a
wavelength of 450 nm.
Migration and invasion assays
The migration and invasion of HeLa and ECT1/E6E7
cells were analyzed using Transwell chambers with 8.0-µm pore size
(Corning, Inc.). In brief, 1×105 transfected cells
without FBS were seeded in the uncoated top chamber, and their
migration or invasion was initiated by adding 20% FBS to the lower
chamber for 24 h. Meanwhile, transfected cells (1×105)
were seeded into a chamber with an 8-µm pore size pre-coated with
BD Matrigel™ Basement Membrane Matrix (BD Biosciences) and placed
in a 24-well plate for 24 h. These cells, along with the coated
membranes, were then placed in the upper chamber and cultured for
migration and invasion assays. These cells were dyed with 0.2%
crystal violet for 15 min at 37°C (Beyotime Institute of
Biotechnology) and washed with PBS. Images were captured and
counted under an upright microscope (magnification, ×40).
Western blotting
Total proteins were obtained using
radioimmunoprecipitation assay buffer (cat. no. 89900; Invitrogen;
Thermo Fisher Scientific, Inc.), and determined using a
bicinchoninic acid kit (cat. no., P0010; Beyotime Institute of
Biotechnology). Proteins (50 µg) were separated using 10% SDS-PAGE.
After membranes were blocked with 5% non-fat milk at 37°C for 1 h,
the membranes were incubated with primary antibodies against FSCN1
(1:1,200; cat. no. ab220195; Abcam) and GAPDH (1:2,000; cat. no:
ab181602; Abcam), followed by HRP-conjugated goat anti-mouse IgG
secondary antibody (1:2,000; cat. no. ab97040; Abcam) overnight at
4°C. Finally, bands were visualized using a FluorChem imaging
device (ProteinSimple). Band intensities were analyzed using ImageJ
(version 1.45; National Institutes of Health).
Statistical analysis
All data were analyzed using SPSS 16.0 (SPSS, Inc.)
or GraphPad Prism 5.0 (GraphPad Software, Inc.). χ2 or
Fisher's exact tests were used to compare categorical variables as
appropriate. Student's t-test, Mann-Whitney U test and one-way
ANOVA with Tukey's post hoc test were used to compare continuous
data. Results were expressed as mean ± SEM. P<0.05 was
considered to indicate a statistically significant difference.
Results
Patient characteristics
The expression of miR-145-5p in CC patients with
different clinicopathological features are summarized in Table I. miR-145-5p expression in both
cancer and adjacent tissues was measured in each patient. The mean
values of miR-145 expression levels in CC tissues were used as the
cutoff values (0.637). The samples were divided into high and low
expression group, and then correlated with the clinicopathological
characteristics of the cases. The cancer histology and
differentiation data are shown in Table I. There are no significant
differences found in clinicopathological factors, including age,
tumor size and differentiation grade, between the low miR-145-5p
expression group and high miR-145-expression group. However, there
was a significant difference in tumor grade and lymph node
metastasis between the miR-145-5p high and low expression
groups.
 | Table I.Association between the expression of
miR-145-5p and clinicopathological factors in patients with
cervical cancer. |
Table I.
Association between the expression of
miR-145-5p and clinicopathological factors in patients with
cervical cancer.
|
|
| miR-145-5p
expression |
|
|---|
|
|
|
|
|
|---|
| Variables | Patients (n=40) | Low (n=25) | High (n=15) | P-value |
|---|
| Age (years) |
|
|
| 0.874 |
| ≤52 | 22 | 14 (56.0) | 8 (53.3) |
|
|
>52 | 18 | 11 (44.0) | 7 (46.7) |
|
| Tumor size (cm) |
|
|
| 0.078 |
| ≤2 | 28 | 15 (60.0) | 13 (86.7) |
|
|
>2 | 12 | 10 (40.0) | 2 (13.3) |
|
| Differentiation |
|
|
| 0.108 |
| Well | 20 | 15 (60.0) | 5 (33.3) |
|
| Moderate,
poor | 20 | 10 (40.0) | 10 (66.7) |
|
| Tumor grade |
|
|
| 0.002 |
| G1 | 17 | 5 (20.0) | 12 (80.0) |
|
| G2 +
G3 | 23 | 20 (80.0) | 3 (20.0) |
|
| Lymph node
metastasis |
|
|
| 0.013 |
| No | 22 | 10 (40.0) | 12 (80.0) |
|
| Yes | 18 | 15 (60.0) | 3 (20.0) |
|
miR-145-5p is downregulated but FSCN1
is overexpressed in CC
The clinicopathological results indicated that
miR-145-5p expression is lower in patients with CC and associated
with lymph node metastasis. Both miR-145-5p and FSCN1 expression in
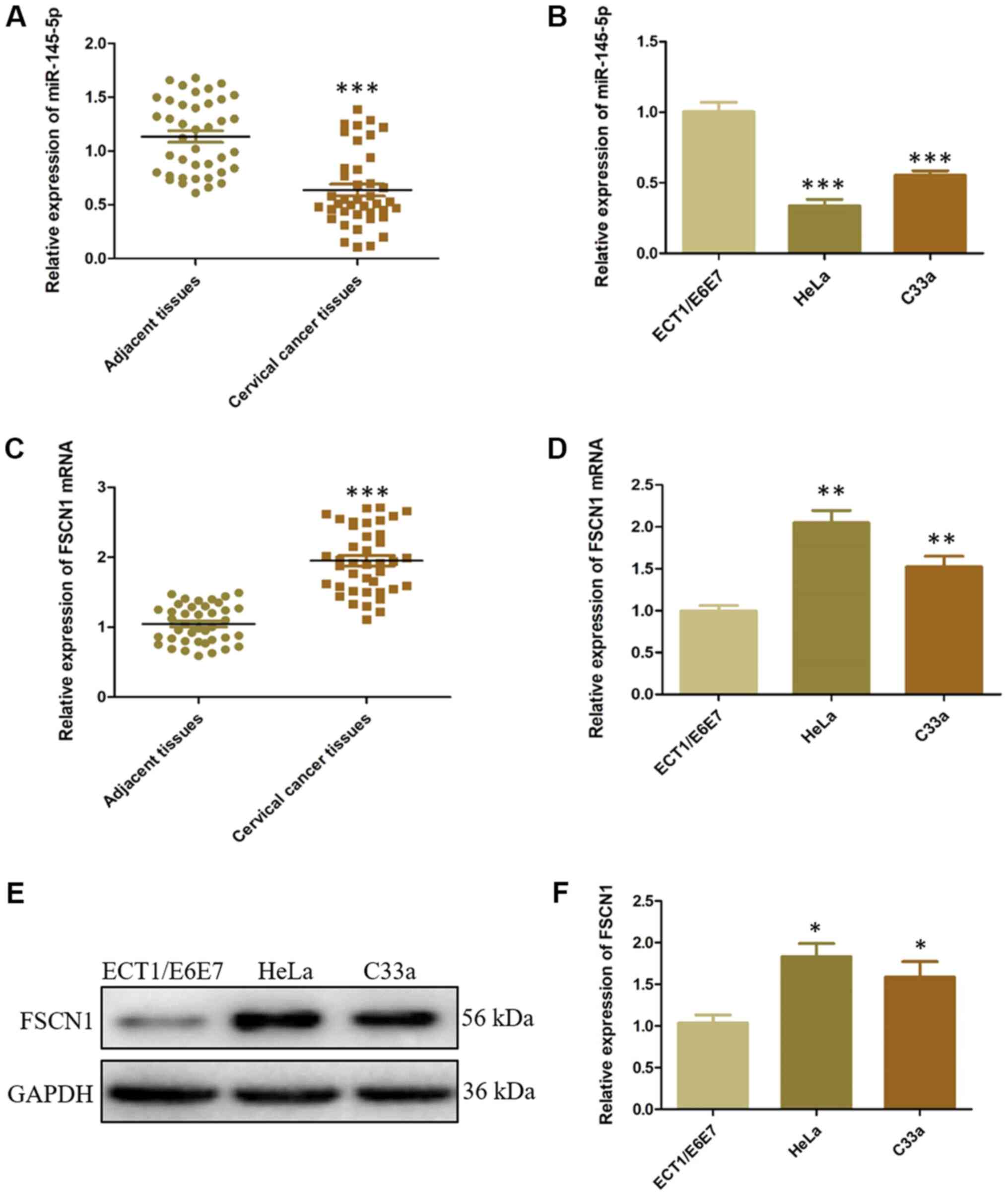
CC tissues and cell lines were analyzed. miR-145-5p expression was
significantly lower in CC tissues and HeLa and C33a cells compared
with the NC controls (Fig. 1A and
B). Simultaneously, FSCN1 expression was significantly higher
in CC tissues and HeLa and C33a cells compared with the NC controls
(Fig. 1C-F). HeLa is a classic
cell line in cervical cancer research and exhibited significant
FSCN1 overexpression in the present study. Thus, HeLa cells were
used for subsequent experiments. These results demonstrated that
both decreased miR-145-5p expression and FSCN1 overexpression
occurs in CC.
miR-145-5p directly targets FSCN1 in
vitro
To investigate the effects of miR-145-5p
downregulation and FSCN1 overexpression in CC, the probable target
genes of miR-145-5p were identified using bioinformatic tests,
which included FSCN1 (Fig. 2A).
Dual-luciferase reporter assays indicated that miR-145-5p mimics
significantly inhibited the luciferase activities of Mut FSCN1, but
did not affect those of wild-type FSCN1 (Fig. 2B). Furthermore, transfection of
miR-145-5p mimics and inhibitors significantly altered miR-145-5p
expression compared with control (Fig.
2C). FSCN1 expression significantly decreased in HeLa cells
treated with miR-145-5p mimics compared with control, while FSCN1
expression significantly increased in cells treated with miR-145-5p
inhibitor compared with NC control (Fig. 2D-F) at both mRNA and protein
levels. These results indicated that miR-145-5p directly targeted
FSCN1.
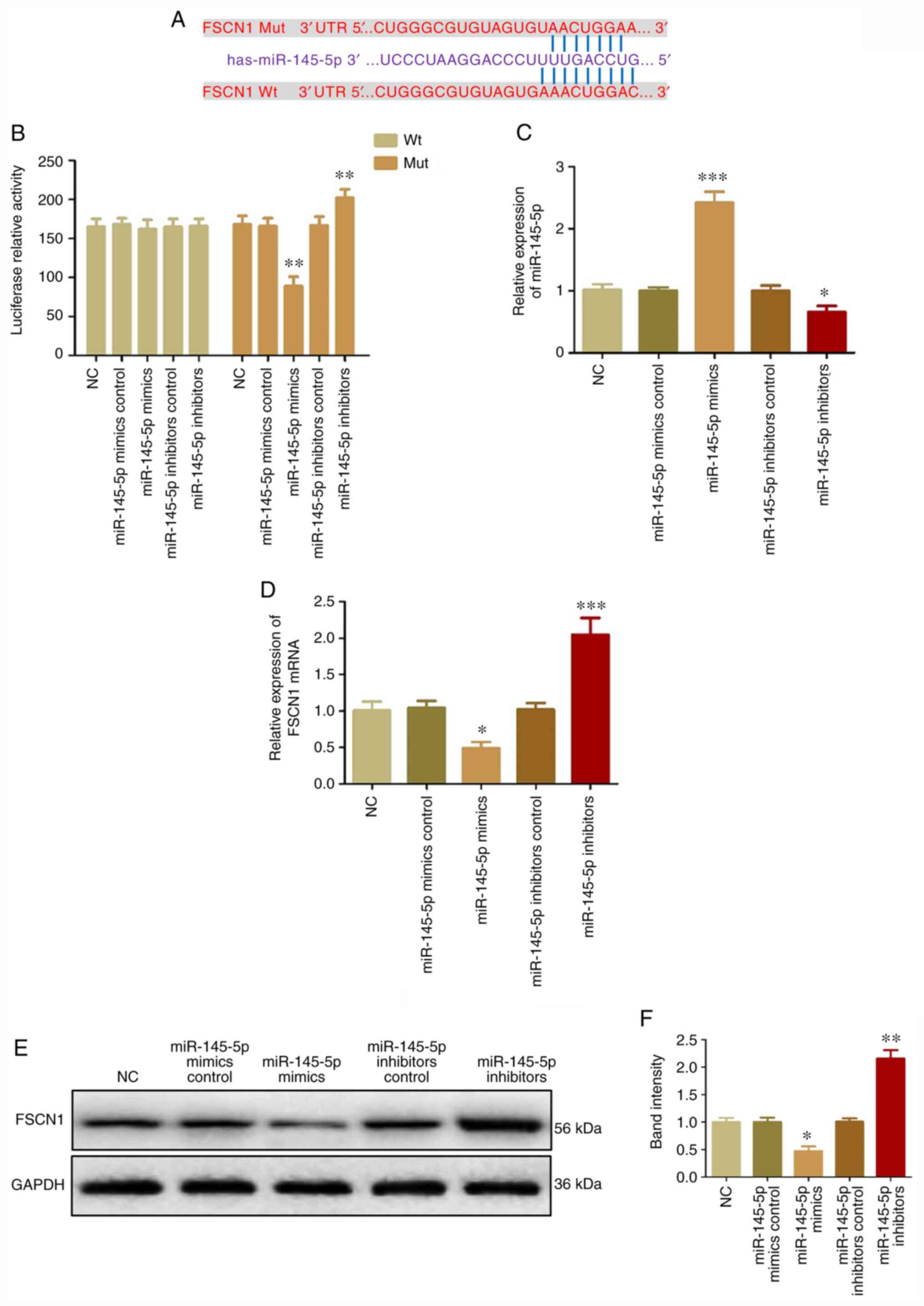 | Figure 2.miR-145-5p directly targets FSCN1 in
cervical cancer cells. (A) Binding sites between miR-145-5p and
FSCN1. (B) Luciferase activities of Wt-FSCN1 mRNA in HeLa cells.
(C) miR-145-5p expression in HeLa cells transfected with miR-145-5p
mimics, inhibitors, mimics control or inhibitors control. FSCN1
expression in HeLa cells transfected with miR-145-5p mimics,
inhibitors, mimics control or inhibitors control was detected at
(D) mRNA and (E and F) protein levels. *P<0.05 vs. NC,
**P<0.01 vs. NC and ***P<0.001 vs. NC. miR, microRNA; FSCN1,
fascin; Wt, wild-type; Mut, mutant; NC, negative control. |
miR-145-5p inhibits the viability,
migration and invasion of HeLa cells
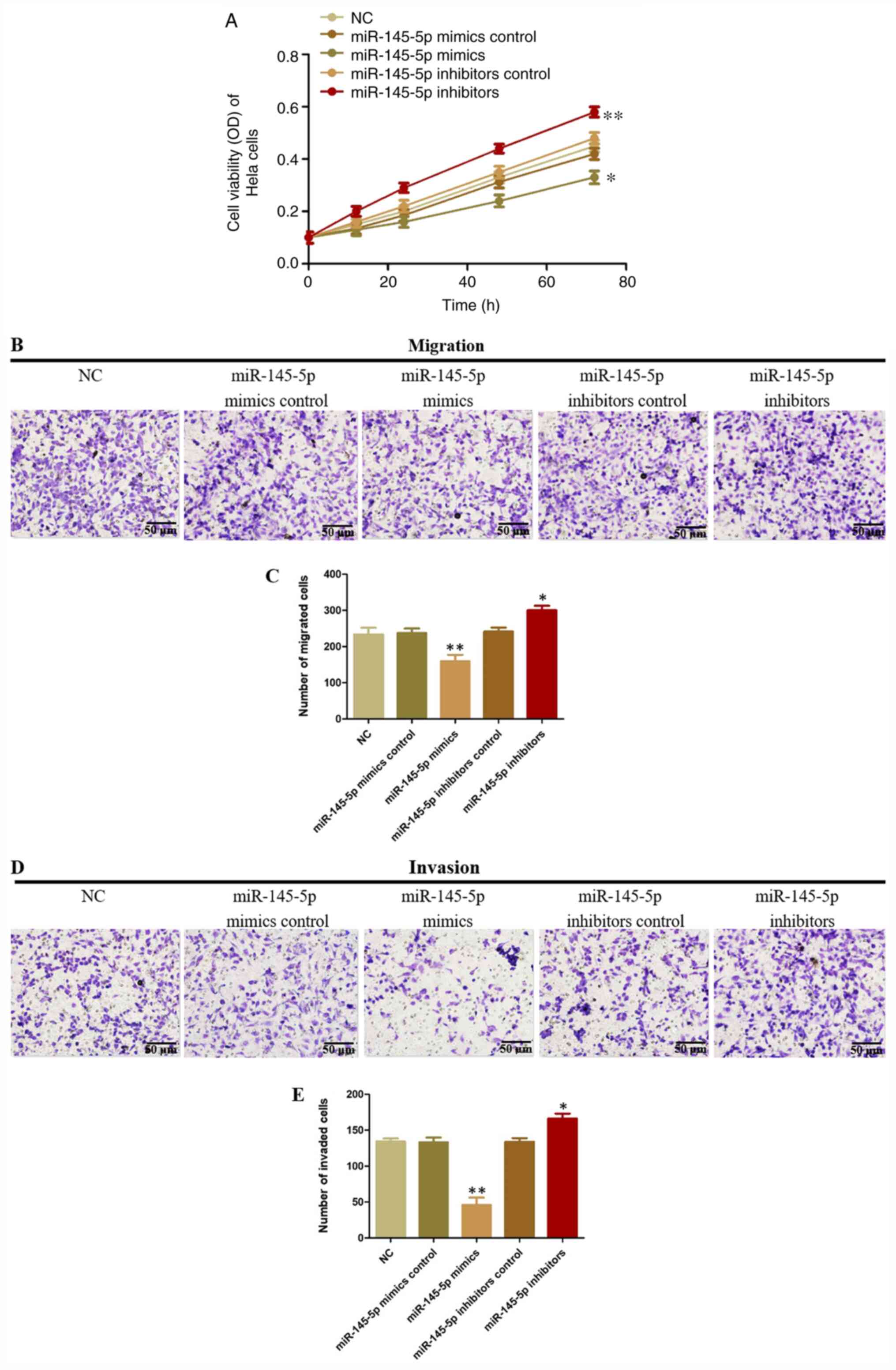
The function of miR-145-5p in CC cells was analyzed
using Transwell and CCK-8 assays. miR-145-5p overexpression
decreased HeLa cell viability, but decreased miR-145-5p expression
significantly promoted HeLa cell viability compared with control
(Fig. 3A). Moreover, miR-145-5p
upregulation significantly suppressed HeLa cell migration compared
with NC control, while miR-145-5p knockdown showed the opposite
effect (Fig. 3B and C). Similar
results were observed in invasion assays (Fig. 3D and E). These results demonstrated
that miR-145-5p can inhibit the migration and invasion of CC
cells.
ECT1/E6E7 viability, migration and
invasion is regulated by miR-145-5p
ECT1/E6E7 is a normal cervical epithelial cell line
and was used to assess the effects of miR-145-5p on tumor
malignancy. The expression levels of miR-145-5p increased in
miR-145-5p mimics-transfected ECT1/E6E7 cells (Fig. 4A) but decreased in miR-145-5p
inhibitors-transfected ECT1/E6E7 cells compared with NC control. It
was found that miR-145-5p overexpression significantly decreased
the viability of ECT1/E6E7 cells (Fig.
4B), but the viability of ECT1/E6E7 cells transfected with
miR-145-5p inhibitors increased compared with NC control.
Meanwhile, miR-145-5p knockdown significantly promoted the
expression of FSCN1 protein in ECT1/E6E7 cells; however, miR-145-5p
overexpression demonstrated the opposite effect (Fig. 4C and D). In addition, the migration
and invasion of ECT1/E6E7 cells significantly decreased following
miR-145-5p overexpression (Fig.
4E-H). However, miR-145-5p knockdown significantly increased
the migration and invasion of these cells.
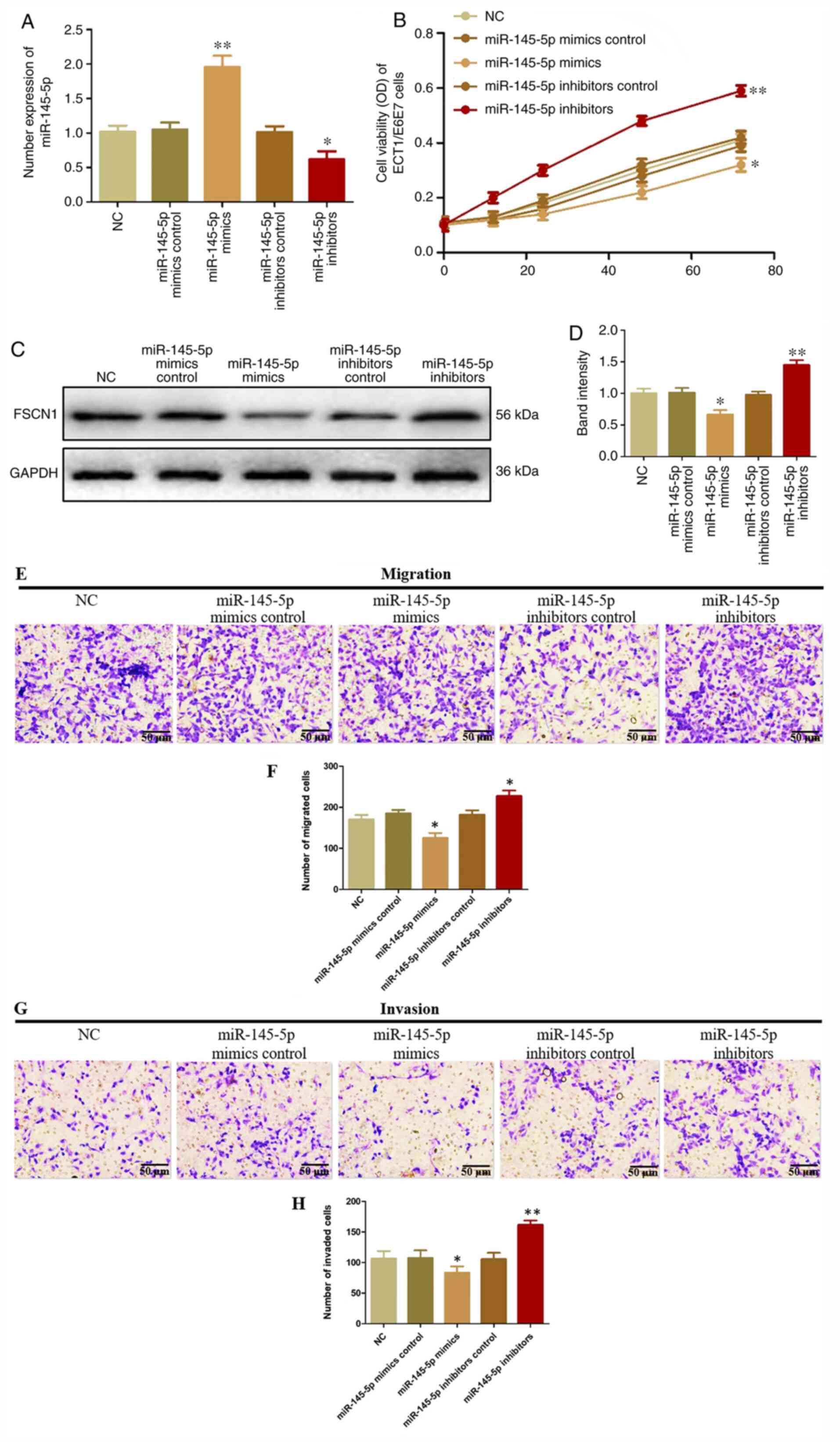 | Figure 4.Suppressive effects of miR-145-5p in
cervical cancer. (A) The expression of miR-145-5p in ETC1/E6E7
cells transfected with miR-145-5p mimics, inhibitors, mimics
control or inhibitors control was evaluated by reverse
transcription-quantitative PCR. (B) Viability of ECT1/E6E7 cells
transfected with miR-145-5p mimics, inhibitors, mimics control or
inhibitors control were detected using a Cell Counting Kit-8 assay.
(C and D) FSCN1 protein expression in ECT1/E6E7 cells transfected
with miR-145-5p mimics, inhibitors, mimics control or inhibitors
control was measured using western blotting. (E and F) Migration of
ECT1/E6E7 cells transfected with miR-145-5p mimics, inhibitors,
mimics control or inhibitors control analyzed using a Transwell
assay. (G and H) Invasion of ECT1/E6E7 cells evaluated following
overexpression and knockdown of miR-145-5p analyzed using a
Transwell assay. Magnification, ×40. *P<0.05 and **P<0.01 vs.
NC. miR, microRNA; FSCN1, fascin; OD, optical density; NC, negative
control. |
ECT1/E6E7 cell metastasis is regulated
by FSCN1 overexpression
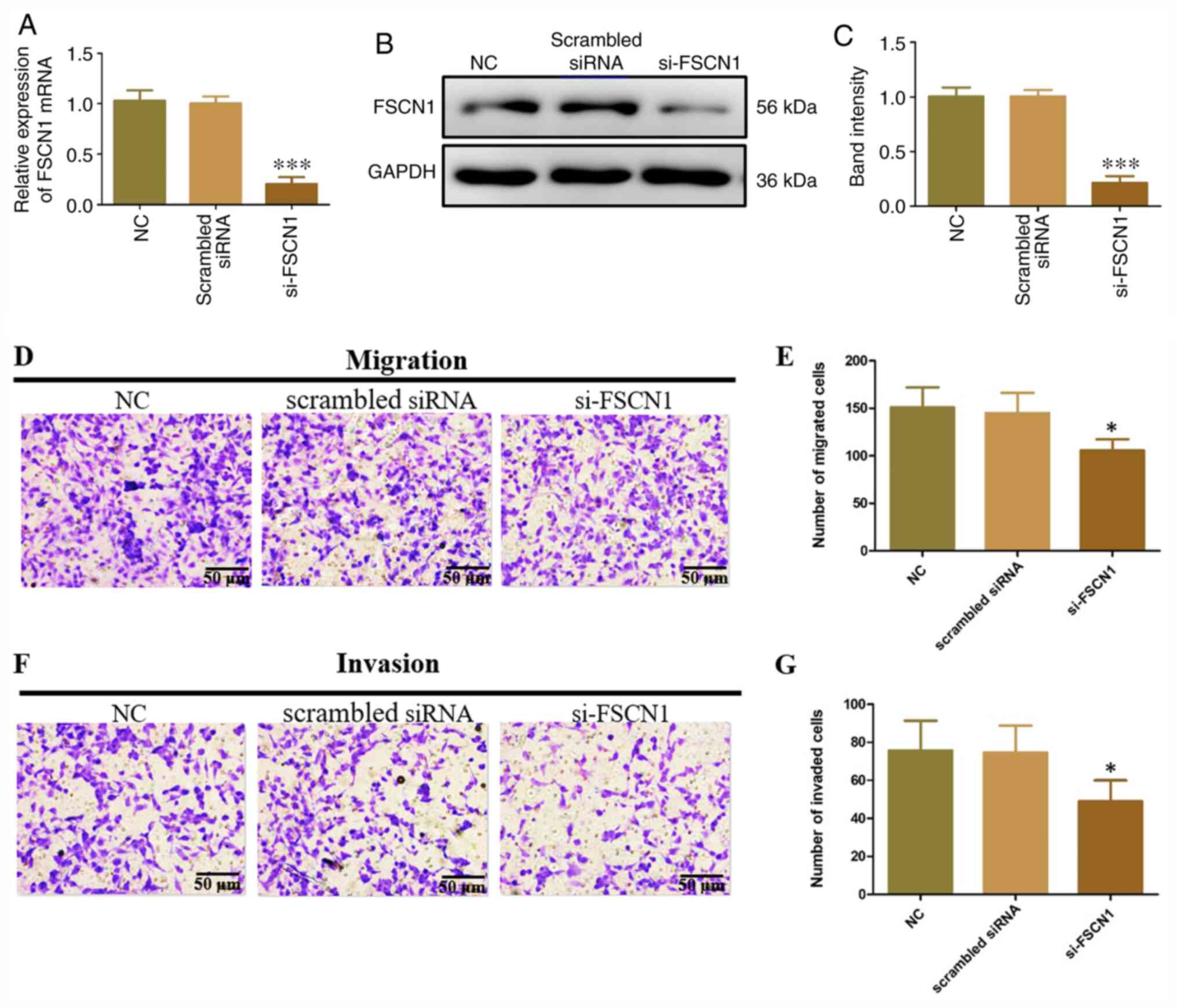
FSCN1 siRNA was transfected into ECT1/E6E7 cells to
investigate its function to the metastasis of ETC1/E6E7 cells.
FSCN1 knockdown significantly decreased the expression of FSCN1
mRNA and protein compared with control and scrambled siRNA
transfection (Fig. 5A-C). In
addition, the migration and invasion of ETC1/E6E7 cells were
significantly inhibited by FSCN1 siRNA (Fig. 5D-G). These findings indicated that
FSCN1 exerted a tumorigenesis function in ECT1/E6E7 cells.
FSCN1 is overexpressed and stimulates
the metastasis of CC
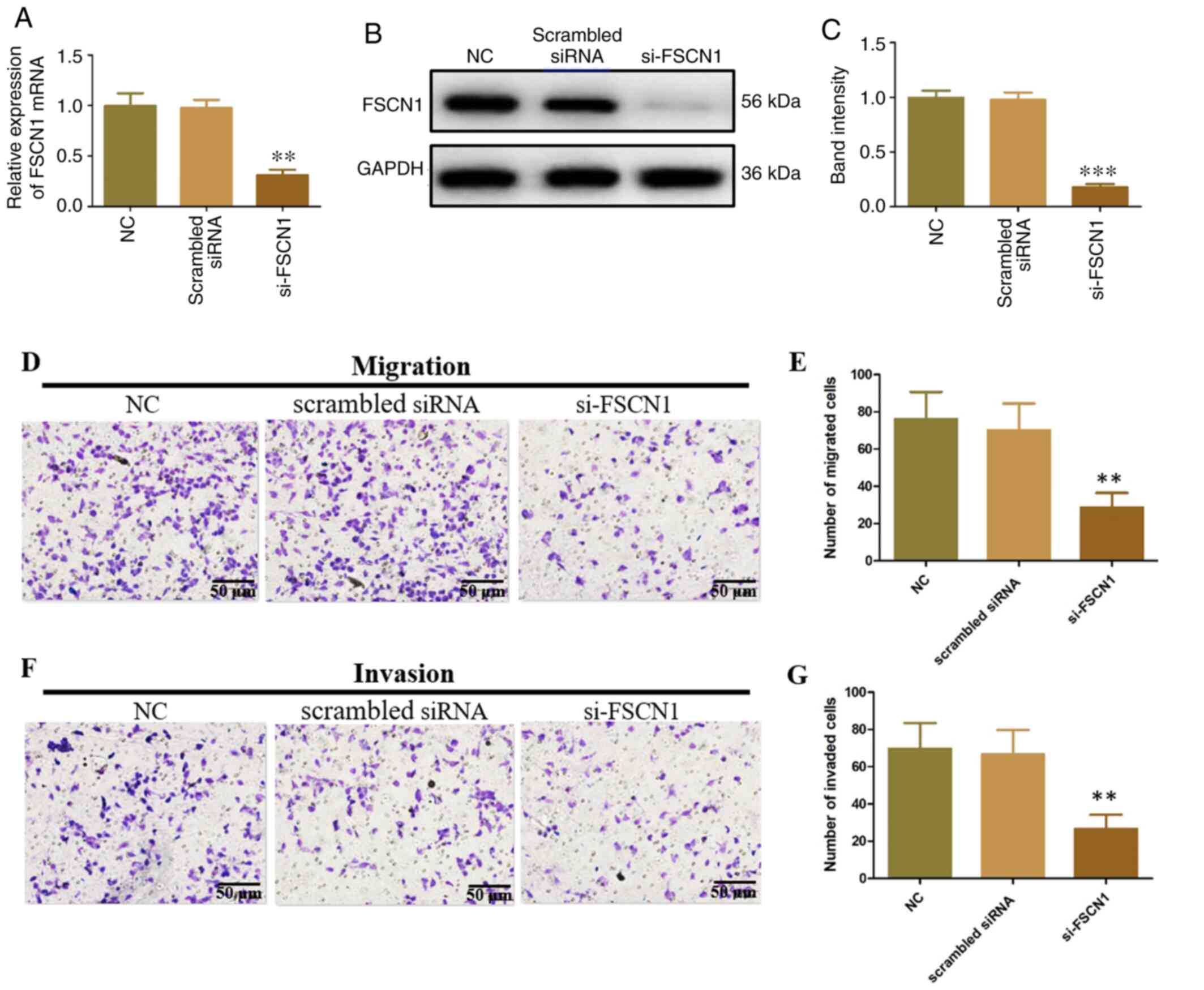
HeLa cells were transfected with FSCN1 siRNA to
investigate the roles of FSCN1 in CC. FSCN1 mRNA and protein
expression were significantly decreased relative to the controls
and scrambled siRNA (Fig. 6A-C).
In addition, FSCN1 siRNA inhibited the migration and invasion of
HeLa cells (Fig. 6D-G), which were
similar to the effects of miR-145-5p overexpression. These findings
revealed that FSCN1 promoted tumorigenesis in CC.
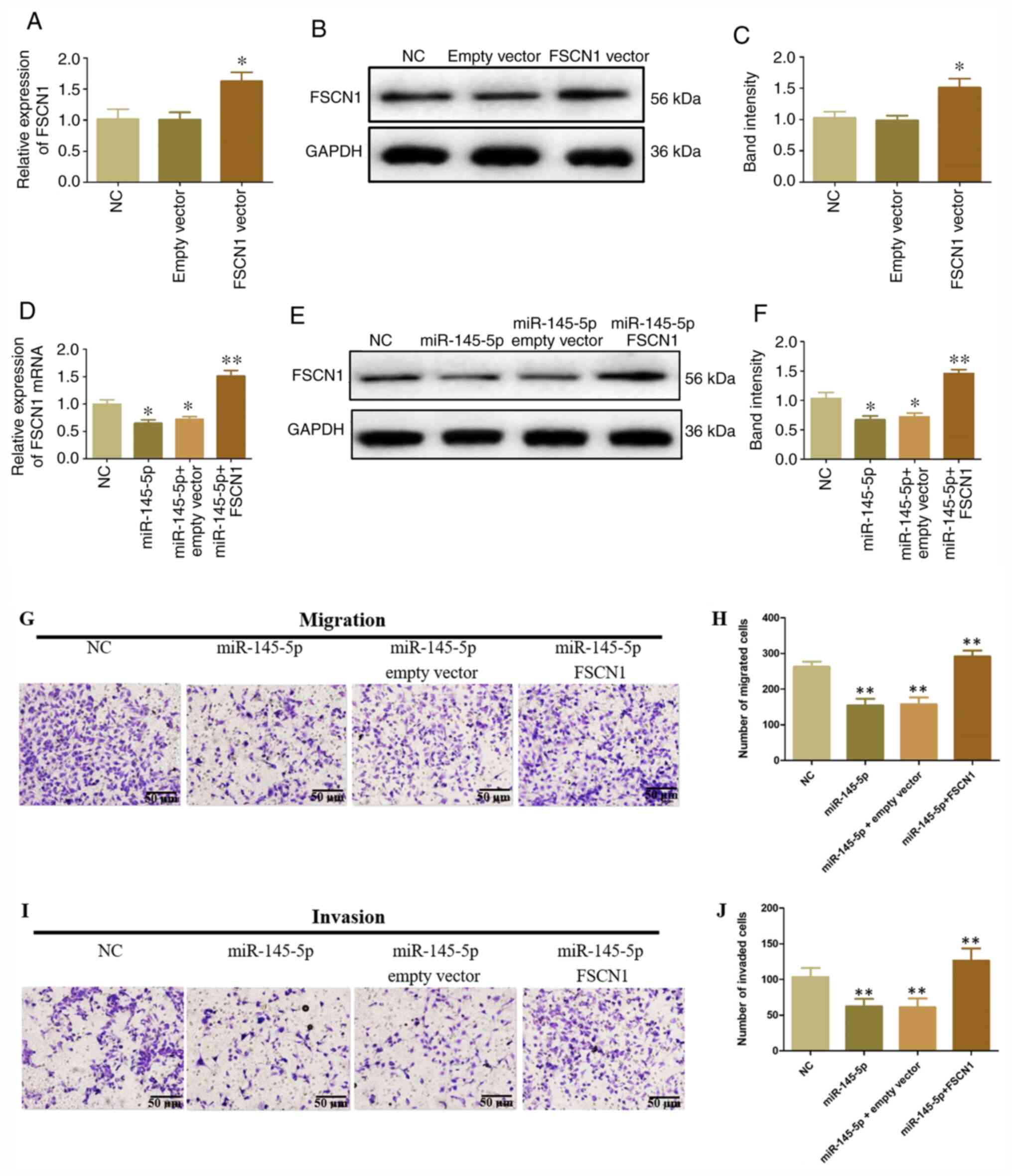
FSCN1 influences the tumor-suppressive
function of miR-145-5p
To confirm whether FSCN1 upregulation affected the
inhibitory role of miR-145-5p on the migration and invasiveness of
HeLa cells, an FSCN1 overexpression vector was transfected into
HeLa cells containing miR-145-5p mimics. FSCN1 expression
significantly increased in HeLa cells transfected with FSCN1 vector
compared with control and empty vectors (Fig. 7A-C). Moreover, FSCN1 mRNA and
protein expression significantly increased following
co-transfection with miR-145-5p and FSCN1 expression vector
(Fig. 7D-F). In addition, FSCN1
overexpression significantly inhibited the tumor-suppressing
function of miR-145-5p on the migration and invasion of HeLa cells
(Fig. 7G-J). These results
indicated that FSCN1 overexpression attenuated the inhibitory
functions of miR-145-5p in CC.
Discussion
miRNAs inhibit mRNA degradation or translation to
regulate posttranscriptional expression of target genes (22). miRNAs are important regulators of
tumor metastasis, including cell differentiation, migration and
invasion (23). miR-145 is
downregulated in multiple types of human cancer and functions as an
important tumor suppressor (24).
In addition, miR-145 is downregulated in laryngeal squamous cell
carcinoma, and its overexpression inhibited the proliferation and
migration of Hep-2 cells by suppressing FSCN1 expression to induce
cell cycle arrest and apoptosis (25). The present study showed that the
expression levels of miR-145-5p were significantly decreased in CC
tissues and cell lines, but FSCN1 levels were significantly
increased in these tissues and cells. Meanwhile, it was found that
miR-145-5p targeted FSCN1 and regulated its expression using
luciferase reporter assays, suggesting that FSCN1 is a target of
miR-145-5p in CC.
FSCN1, an actin-connecting protein, are present in
mesenchymal tissues but is downregulated in adult epithelial cells
(14). FSCN1 stabilizes the
generation of parallel actin bundles in cell-protruding areas
during the early phase of cell movement in physiological and
neoplastic conditions (15). FSCN1
is expressed in various epithelial and non-epithelial neoplasms,
but it is overexpressed in malignant cells, especially at deep
tumor margins (26). FSCN1 is
overexpressed in most cervical intraepithelial neoplasia (CIN)
lesions, which may indicate increased invasion in high grade CIN,
indicating that FSCN1 could have value as a diagnostic biomarker of
superficial stromal invasion (27). The present study reported that
knockdown of miR-145-5p promoted the expression of FSCN1 in HeLa
and ECT1/E6E7 cells; however, overexpression of miR-145-5p
repressed the expression of FSCN1. In addition, the migration and
invasion of HeLa and ECT1/E6E7 cells were enhanced by the
overexpression of miR-145-5p but suppressed by the downregulation
of this miRNA. A previous study reported that the inhibition of
FSCN1 decreased the activity of the MAPK pathway and suppressed the
growth and metastasis of non-small cell lung cancer (NSCLC) cells,
indicating that it may be a potentially effective therapeutic
strategy for treating NSCLC (28).
In addition, FSCN1 is an important mediator of TGF-β1-induced
invasion and migration of kidney cancer cells (KCC) via ERK and JNK
signaling pathways (29). FSCN1
could act as an oncogene and a potential novel prognostic biomarker
for patients with renal cell carcinoma (RCC) after nephrectomy, and
can promote RCC metastasis (30).
However, the signaling pathway of FSCN1 involved in CC is still
unclear and will be investigated in future studies.
In summary, miR-145-5p overexpression significantly
inhibited the proliferation, migration and invasion of HeLa and
ECT1/E6E7 cells, and silencing of FSCN1 expression similarly
restricted the proliferation, migration and invasion of these
cells. The miR-145/FSCN1 axis facilitated the progression of CC by
restricting CC cell proliferation. However, the present study did
not assess the function of miR-145-5p-targeted FSCN1 in animal
experiments. Follow-up studies will focus on their functions in
animal models. Overall, the present findings may provide insight
for further studies investigating the effects of miR-145-5p
functions and underlying mechanisms on the biological behavior of
CC.
Acknowledgements
Not applicable.
Funding
No funding was received.
Availability of data and materials
The datasets used and/or analyzed during the current
study are available from the corresponding author on reasonable
request.
Authors' contributions
SH and SL designed all the experiments, drafted and
revised this manuscript. SH, GY and KP performed the experiments
and analyzed all data. All authors read and approved the final
manuscript.
Ethics approval and consent to
participate
The present study was approved by the Ethics
Committee of the First Affiliated Hospital of Nanchang University
(approval no. 2018023). Written informed consent was obtained from
all patients.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Bray F, Ferlay J, Soerjomataram I, Siegel
RL, Torre LA and Jemal A: Global cancer statistics 2018: GLOBOCAN
estimates of incidence and mortality worldwide for 36 cancers in
185 countries. CA Cancer J Clin. 68:394–424. 2018.PubMed/NCBI
|
|
2
|
Yu Y, Zhang Y and Zhang S: MicroRNA-92
regulates cervical tumorigenesis and its expression is upregulated
by human papillomavirus-16 E6 in cervical cancer cells. Oncol Lett.
6:468–474. 2013.PubMed/NCBI
|
|
3
|
Park TW, Fujiwara H and Wright TC:
Molecular biology of cervical cancer and its precursors. Cancer.
76((10 Suppl)): S1902–S1913. 1995.
|
|
4
|
Peiretti M, Zapardiel I, Zanagnolo V,
Landonia F, Morrowc CP and Maggionia A: Management of recurrent
cervical cancer: A review of the literature. Surg Oncol.
21:e59–e66. 2012.PubMed/NCBI
|
|
5
|
Li N, Cui T, Guo W, Wang D and Mao L:
MiR-155-5p accelerates the metastasis of cervical cancer cell via
targeting TP53INP1. Onco Targets Ther. 12:3181–3196.
2019.PubMed/NCBI
|
|
6
|
Farazi TA, Hoell JI, Morozov P and Tuschl
T: MicroRNAs in human cancer. Adv Exp Med Biol. 774:1–20.
2013.PubMed/NCBI
|
|
7
|
Pardini B, De Maria D, Francavilla A, Di
Gaetano C, Ronco G and Naccarati A: MicroRNAs as markers of
progression in cervical cancer: A systematic review. BMC Cancer.
18:6962018.PubMed/NCBI
|
|
8
|
Iorio MV, Ferracin M, Liu CG, Veronese A,
Spizzo R, Sabbioni S, Magri E, Pedriali M, Fabbri M, Campiglio M,
et al: MicroRNA gene expression deregulation in human breast
cancer. Cancer Res. 65:7065–7070. 2005.PubMed/NCBI
|
|
9
|
Iorio MV, Visone R, Di Leva G, Donati V,
Petrocca F, Casalini P, Taccioli C, Volinia S, Liu CG, Alder H, et
al: MicroRNA signatures in human ovarian cancer. Cancer Res.
67:8699–8707. 2007.PubMed/NCBI
|
|
10
|
Takagi T and Iio AY: Decreased expression
of microRNA-143 and −145 in human gastric cancers. Oncology.
77:12–21. 2009.PubMed/NCBI
|
|
11
|
Liu X, Sempere LF, Galimberti F,
Freemantle SJ, Black C, Dragnev KH, Ma Y, Fiering S, Memoli V, Li
H, et al: Uncovering growth-suppressive MicroRNAs in lung cancer.
Clin Cancer Res. 15:1177–1183. 2009.PubMed/NCBI
|
|
12
|
Li C, Xu N, Li YQ, Wang Y and Zhu ZT:
Inhibition of SW620 human colon cancer cells by upregulating mi
RNA-145. World J Gastroenterol. 22:2771–2778. 2016.PubMed/NCBI
|
|
13
|
Zhang J, Wang L, Li B, Huo M, Mu MU, Liu J
and Han J: miR-145 downregulates the expression of cyclin-dependent
kinase 6 in human cervical carcinoma cells. Ther Med. 8:591–594.
2014.
|
|
14
|
Hashimoto Y, Skacel M and Adams JC: Roles
of fascin in human carcinoma motility and signaling: Prospects for
a novel biomarker? Int J Biochem Cell Biol. 37:1787–804.
2005.PubMed/NCBI
|
|
15
|
Li A, Dawson JC, Forero-Vargas M, Spence
HJ, Yu XZ, König I, Anderson K and Machesky LM: The actin-bundling
protein fascin stabilizes actin in invadopodia and potentiates
protrusive invasion. Curr Biol. 20:339–345. 2010.PubMed/NCBI
|
|
16
|
Liu R, Liao J, Yang M, Sheng J, Yang H,
Wang Y, Pan E, Guo W, Pu Y, Kim SJ and Yin L: The cluster of
miR-143 and miR-145 affects the risk for esophageal squamous cell
carcinoma through co-regulating fascin homolog 1. PLoS One.
7:e339872012.PubMed/NCBI
|
|
17
|
Zhang M, Dong BB, Lu M, Zheng MJ, Chen H,
Ding JZ, Xu AM and Xu YH: miR-429 functions as a tumor suppressor
by targeting FSCN1 in gastric cancer cells. Onco Targets Ther.
9:1123–1133. 2016.PubMed/NCBI
|
|
18
|
Li J, Lu J, Ye Z, Han X, Zheng X, Hou H,
Chen W, Li X and Zhao L: 20(S)-Rg3 blocked epithelial-mesenchymal
transition through DNMT3A/miR-145/FSCN1 in ovarian cancer.
Oncotarget. 8:53375–53386. 2017.PubMed/NCBI
|
|
19
|
Chen JJ, Cai WY, Liu XW, Luo QC, Chen G,
Huang WF, Li N, Cai JC and Yang CF: Reverse correlation between
MicroRNA-145 and FSCN1 affecting gastric cancer migration and
invasion. PLoS One. 10:e0126902015.
|
|
20
|
Ma L and Li LL: miR-145 Contributes to the
progression of cervical carcinoma by directly regulating FSCN1.
Cell Transplant. 28:1299–1305. 2019.PubMed/NCBI
|
|
21
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408.
2001.PubMed/NCBI
|
|
22
|
Bartel DP: MicroRNAs genomics, biogenesis,
mechanism, and function. Cell. 116:281–297. 2004.PubMed/NCBI
|
|
23
|
Iorio MV and Croce CM: MicroRNAs in
cancer: Small molecules with a huge impact. J Clin Oncol.
27:5848–5856. 2009.PubMed/NCBI
|
|
24
|
Slattery ML, Herrick JS, Mullany LE,
Valeri N, Stevens J, Caan BJ, Samowitz W and Wolff RK: An
evaluation and replication of miRNAs with disease stage and
colorectal cancer-specific mortality. Int J Cancer. 137:428–438.
2015.PubMed/NCBI
|
|
25
|
Gao W, Zhang C, Li W, Li H, Sang J, Zhao
Q, Bo Y, Luo H, Zheng X, Lu Y, et al: Promoter
methylation-regulated miR-145-5p inhibits laryngeal squamous cell
carcinoma progression by targeting FSCN1. Mol Ther. 27:365–379.
2019.PubMed/NCBI
|
|
26
|
Stewart CJ, Crook M and Loi S: Fascin
expression in endocervical neoplasia: Correlation with tumour
morphology and growth pattern. J Clin Pathol. 65:213–217.
2012.PubMed/NCBI
|
|
27
|
Koay MH, Crook M and Stewart CJ: Fascin
expression in cervical normal squamous epithelium, cervical
intraepithelial neoplasia, and superficially invasive (stage IA1)
squamous carcinoma of the cervix. Pathology. 46:433–438. 2018.
|
|
28
|
Zhao D, Zhang T, Hou XM and Ling XL:
Knockdown of fascin-1 expression suppresses cell migration and
invasion of non-small cell lung cancer by regulating the MAPK
pathway. Biochem Biophys Res Commun. 497:694–699. 2018.PubMed/NCBI
|
|
29
|
Yang J, Zhang N, Gao R, Zhu Y, Zhang Z, Xu
X, Wang J, Li Z, Liu X, Li Z, et al: TGF-β1 induced fascin1
expression facilitates the migration and invasion of kidney
carcinoma cells through ERK and JNK signaling pathways. Biochem
Biophys Res Commun. 501:913–919. 2018.PubMed/NCBI
|
|
30
|
Zhang M, Zhao Z, Duan X, Chen P, Peng Z
and Qiu H: FSCN1 predicts survival and is regulated by a
PI3K-dependent mechanism in renal cell carcinoma. J Cell Physiol.
233:4748–4758. 2018.PubMed/NCBI
|