Introduction
Triple-negative breast cancer (TNBC) is defined as a
breast cancer subtype that lacks expression of estrogen receptor,
progesterone receptor and human epidermal growth factor receptor 2
(1). Compared with other subtypes,
TNBC typically exhibits higher Ki-67 antigen expression, a greater
mitotic index, more frequent breast cancer susceptibility gene 1
mutations and a more aggressive clinical course (2). In the United States, the mean annual
incidence is ~13.7 per 100,000 women. Higher frequencies are
observed among African American and Hispanic women, as well as in
women with younger age, higher premenopausal body mass index,
earlier age at menarche, higher parity and a history of oral
contraceptive use (2–4). Notably, TNBC is associated with the
lowest 5-year locoregional recurrence-free survival (79.6%) and
local recurrence-free survival (84.6%) among all breast cancer
subtypes (5). The lack of
expression of targetable receptors in TNBC markedly restricts
therapeutic options, with current management primarily involving
surgical resection, radiotherapy and chemotherapeutic agents such
as paclitaxel and doxorubicin (6).
This constraint in treatment options underscores the need to
explore new targets and develop innovative therapies for TNBC.
Cepharanthine (CEP), a bisbenzylisoquinoline
alkaloid derived from Stephania cephalantha, has a wide
range of pharmacological activities, including anti-severe acute
respiratory syndrome coronavirus 2, immunomodulatory,
anti-atherosclerotic and antitumor properties (7). CEP also has notable antiproliferative
and pro-apoptotic activities in cancer cell lines, such as those
derived from hepatocellular carcinoma and colorectal cancer
(8). These effects may be mediated
by the influence of CEP on important signaling pathways, such as
the PI3K/Akt, MAPK and NF-κB pathways, and its modulation of Bcl-2
family protein expression (8).
Despite these well-documented antitumor effects, the direct
molecular targets of CEP remain poorly defined. The aims of the
present study were to evaluate the efficacy of CEP against TNBC
in vitro and elucidate its mechanisms of
action, with a specific focus on identifying the direct molecular
targets.
Materials and methods
Cell culture
All cell lines were maintained at 37°C in 5%
CO2 atmosphere and tested routinely for mycoplasma
contamination. The TNBC cell lines MDA-MB-231 (cat. no. CL-0150;
Procell Life Science & Technology Co., Ltd.) and Hs578T (cat.
no. CL-0114; Procell Life Science & Technology Co., Ltd.) were
cultured in high-glucose DMEM medium (cat. no. C11965500BT; Gibco;
Thermo Fisher Scientific, Inc.), supplemented with 10% fetal bovine
serum (cat. no. 164210; Procell Life Science & Technology Co.,
Ltd.) and 1% penicillin/streptomycin (cat. no. 15140122; Gibco;
Thermo Fisher Scientific, Inc.). The Hs578T cultures were also
supplemented with 10 µg/ml insulin (cat. no. P3376; Beyotime
Institute of Biotechnology). The human normal breast epithelial
cell line MCF10A (cat. no. CL-0525; Procell Life Science &
Technology Co., Ltd.) was cultured in commercially pre-configured
DMEM/F12 medium (cat. no. CM-0525; Procell Life Science &
Technology Co., Ltd.) which was supplemented with 5% donor equine
serum, 20 ng/ml epidermal growth factor, 0.5 µg/ml hydrocortisone,
10 µg/ml insulin, 1% non-essential amino acid solution and 1%
penicillin/streptomycin. Only low-passage cells (<15 passages)
were used in all the experiments.
Cytotoxicity assay
The cytotoxicity of CEP (purity, 99.06%; cat. no.
A0653; Chengdu Must Bio-Technology Co., Ltd.) was evaluated via the
Cell Counting Kit-8 (CCK-8) assay. Briefly, MDA-MB-231, Hs578T and
MCF10A cells were seeded in 96-well plates at a density of 4,000
cells per well and treated with various concentrations (1, 2, 4, 6,
8, 10, 15, 20, 25 and 30 µM) of CEP for 24, 48 or 72 h at 37°C. A
10 µl aliquot of CCK-8 reagent (cat. no. C0005; TargetMol Chemicals
Inc.) was then added to each well and the plates were incubated for
1 h at 37°C. The absorbance at 450 nm was measured using a
Multiskan FC microplate reader (Thermo Fisher Scientific, Inc.).
Dose-response curves and IC50 values were generated
using GraphPad Prism (version 9.0.0; Dotmatics).
Clonogenicity assay
MDA-MB-231 and Hs578T cells were seeded in 6-well
plates at 400 cells per well and incubated for 48 h at 37°C. The
medium was then replaced with fresh medium containing CEP (2, 4, 8,
15 and 30 µM for MDA-MB-231; 1, 2, 4, 8 and 15 µM for Hs578T),
followed by 12 days of incubation at 37°C, with the medium being
refreshed every 3 days. At the end of treatment, colonies were
fixed with 4% paraformaldehyde (cat. no. P0098; Beyotime Institute
of Biotechnology) for 5 min, stained with 1% crystal violet for 30
min room temperature (RT), then air dried at RT. Images of the
colonies were recorded using a ChemiDoc MP imaging system (Bio-Rad
Laboratories, Inc.) and the number of colonies was counted
manually. For counting purposes, a colony was defined as a cell
aggregate having an area >0.05 mm2.
TUNEL assay
MDA-MB-231 and Hs578T cells were seeded in 24-well
plates at a density of 1×104 cells per well and treated
with 10 µM CEP for 24 h at 37°C. Following two PBS washes, cells
were fixed with 4% paraformaldehyde (cat. no. P0098; Beyotime
Institute of Biotechnology) for 15 min at RT, permeabilized with
0.1% Triton X-100 (cat. no. P0097; Beyotime Institute of
Biotechnology) for 10 min at RT and subsequently analyzed using a
One-step TUNEL Apoptosis Assay Kit (cat. no. C1088; Beyotime
Institute of Biotechnology) according to the manufacturer's
instructions. The cell nuclei were stained with 50 µl of DAPI
medium (10 µg/ml) (cat. no. P0131; Beyotime Institute of
Biotechnology) for 10 min at RT, mounted with 50 µl of anti-fade
mounting medium (cat. no. P0126; Beyotime Biotechnology), and then
imaged using a fluorescence microscope (IX73; Olympus Corporation),
with DAPI excitation/emission set at 358/461 nm and FITC
excitation/emission at 495/519 nm. Images from three randomly
selected fields of view per well were captured for the subsequent
analysis of the percentage of TUNEL-positive cells.
Flow cytometry analysis
MDA-MB-231 and Hs578T cells were seeded in 6-well
plates at 1×106 cells per well and exposed to CEP (5, 10
and 20 µM for MDA-MB-231; 2, 4 and 8 µM for Hs578T) for 48 h at
37°C. Following treatment, the cells and supernatant were
collected, centrifuged at 1,000 × g for 5 min at RT and the
supernatant was subsequently discarded. The cell pellet was washed
with PBS and resuspended in 195 µl of annexin V-FITC binding buffer
(cat. no. C1062L; Beyotime Institute of Biotechnology). A total of
5 µl annexin V-FITC was subsequently added to cells with gentle
mixing, followed by 10 µl propidium iodide, also administered with
gentle mixing. The samples were incubated at RT in the dark for 15
min and apoptosis was subsequently assessed using a flow cytometer
(EPICS® XL™; Beckman Coulter, Inc.). The results were
analyzed using FlowJo software (version 10.8.1; FlowJo LLC; BD
Biosciences).
Mitochondrial membrane potential (ΔΨm)
assay
MDA-MB-231 and Hs578T cells were seeded in 35 mm
glass-bottom dishes at a density of 2×105 cells per dish
and treated with CEP (10 µM for MDA-MB-231, and 4 µM for Hs578T)
for 24 h at 37°C. Following two PBS washes, cells were incubated
with 5 µg/ml
5,5′,6,6′-tetrachloro-1,1′,3,3′-tetraethylbenzimidazolylcarbocyanine
iodide (JC-1) (cat. no. T15609; TargetMol Chemicals Inc.) for 20
min at 37°C. The fluorescence of the cells was visualized using a
fluorescence microscope, with JC-1 monomer excitation/emission set
at 510/527 nm and JC-1 aggregate excitation/emission set at 585/590
nm. The shift from red (aggregates) to green (monomers)
fluorescence indicated mitochondrial depolarization.
Western blot analysis
MDA-MB-231 and Hs578T cells were cultured in 10-cm
dishes and treated with different concentrations of CEP (5, 10 and
20 µM for MDA-MB-231; 2, 4 and 8 µM for Hs578T) for 24 h at 37°C.
The cells were lysed in RIPA buffer (cat. no. P0013B; Beyotime
Institute of Biotechnology) at 0°C for 20 min before centrifugation
at 16,000 × g for 10 min at 4°C. Supernatant protein concentrations
were subsequently determined via BCA assay. Equal amounts of
protein (20 µg per lane) were denatured for 7 min at 100°C and then
separated by 12% SDS-PAGE gels at 80 V for 40 min and 120 V for 60
min. Proteins were subsequently transferred to 0.45-µm PVDF
membranes using a Trans-Blot Turbo semi-dry system (1.3 A; 25 V;
10–20 min; Bio-Rad Laboratories, Inc.). The membranes were
subsequently treated with QuickBlock™ western blocking buffer (cat.
no. P0220; Beyotime Biotechnology) for 30 min at RT and incubated
overnight at 4°C with the following primary antibodies (all in a
1:1,000 dilution): Acid ceramidase (ASAH1; cat. no. 11274-1-AP;
Proteintech Group, Inc.), sphingomyelin phosphodiesterase (SMPD1;
cat. no. 14609-1-AP; Proteintech Group, Inc.), cathepsin B (CTSB;
cat. no. 112216-1-AP; Proteintech Group, Inc.), cathepsin D (CTSD;
cat. no. 21327-1-AP; Proteintech Group, Inc.),
phorbol-12-myristate-13-acetate-induced protein 1 (NOXA; cat. no.
14766; Cell Signaling Technology, Inc.), Bax (cat. no. 2772T; Cell
Signaling Technology, Inc.), Bcl-2 (cat. no. 15071T; Cell Signaling
Technology, Inc.) and cleaved caspase-3 (cat. no. 9661T; Cell
Signaling Technology, Inc.). After washing in TBST containing 0.1%
Tween-20 (v/v), the membranes were incubated with HRP-conjugated
secondary antibodies (1:5,000 dilution; cat. no. SA00001-2;
Proteintech Group, Inc.) for 1 h at RT, followed by detection using
the Tanon™ ECL chemiluminescent substrate (cat. no. 180-501;
Shanghai Tanon Life Science Co., Ltd.). The membranes were then
placed in stripping buffer (cat. no. SW3022; Beijing Solarbio
Science & Technology Co., Ltd.) for 3 min and re-probed with
anti-β-actin antibody (cat. no. 66009-1-Ig; Proteintech Group,
Inc.) as the loading control. Semi-quantitative analysis of the
band/intensity was performed using ImageJ software (version 1.53t;
National Institutes of Health).
Proteomics analysis
MDA-MB-231 cells at 80% confluence in 10 cm dishes
were treated with 10 µM CEP or 0.1% DMSO for 24 h at 37°C and then
harvested by scraping on ice. Cells were subject to lysis in urea
buffer (8 M urea; 1% PMSF) using pulsed ultrasonication (200 W; 3
sec on, 10 sec off; 40 cycles) and centrifuged at 12,000 × g at 4°C
for 10 min. The protein content of the supernatants was quantified
using the BCA assay. Following precipitation with 20%
trichloroacetic acid at 4°C for 2 h, the protein isolates were
washed three times with ice-cold acetone and then digested
overnight with trypsin (cat. no. V5280; Promega Corporation) at a
1:50 enzyme-to-protein ratio at 37°C, followed by resuspension in
200 mM tetraethylammonium bromide solution. The isolated peptides
were reduced with 5 mM dithiothreitol at 56°C for 30 min and then
alkylated with 11 mM iodoacetamide at RT for 15 min. The samples
were then dissolved in mobile phase A in 0.1% formic acid and 2%
acetonitrile in water, before samples were separated by
nano-ultra-high performance liquid chromatography (UHPLC) (Bruker
Corporation) using a linear gradient of solvent B (0.1% formic acid
in acetonitrile/water). The gradient was as follows: 6–24% solvent
B (0–14 min), 24–35% solvent B (14–16 min), 35–80% solvent B (16–18
min) and 80% solvent B (18–20 min) at 500 nl/min.
The separated peptides were analyzed using a timsTOF
Pro 2 mass spectrometer (Bruker Corporation) in data-independent
acquisition (DIA) parallel accumulation serial fragmentation
(PASEF) mode. Full MS scans (m/z, 300–1500) were acquired, followed
by 20 PASEF MS/MS scans per cycle for a total of 32 cycles
(isolation window, 7 m/z; scan range, m/z 400–850). The data were
processed using the DIA-NN search engine (version 1.8) (9) and the Homo sapiens UniProt
database (taxon identifier, 9606; release, 2023_12; downloaded from
https://www.uniprot.org/proteomes/UP000005640 in
December 2023) concatenated with a reverse decoy database. Trypsin
was set as the cleavage enzyme with one missed cleavage allowed.
Fixed modifications included N-terminal methionine excision and
cysteine carbamidomethylation. The false discovery rate (FDR) was
controlled at <1%, with a quantitative analysis subsequently
conducted using the robust label-free quantification algorithm of
DIA-NN. Differentially expressed proteins (DEPs) were identified by
a fold change (FC) <0.667 or >1.5 and P<0.05. Functional
annotation of the DEPs was performed using Gene Ontology (GO)
enrichment and the Kyoto Encyclopedia of Genes and Genomes (KEGG)
pathway analyses via the DAVID Bioinformatics Resources (https://david.ncifcrf.gov/) and the KEGG database
(https://www.genome.jp/kegg/),
respectively.
Immunofluorescence analysis
MDA-MB-231 and Hs578T cells were seeded in 35-mm
glass-bottom dishes at a density of 2×105 cells per dish
and treated with CEP (10 µM for MDA-MB-231, and 4 µM for Hs578T)
for 24 h at 37°C. The cells were fixed with 4% paraformaldehyde
(cat. no. P0098; Beyotime Institute of Biotechnology) for 15 min at
RT, permeabilized with 0.1% Triton X-100 (cat. no. P0097; Beyotime
Institute of Biotechnology) for 10 min at RT, and after blocking
with serum-free QuickBlock™ blocking buffer (cat. no. P0260;
Beyotime Biotechnology) for 30 min at RT, incubated with
transcription factor EB (TFEB) primary antibody (1:200; cat. no.
13372-1-AP; Proteintech Group, Inc.) overnight at 4°C. The cells
were then stained with CoraLite594-conjugated secondary antibody
(1:200 dilution; cat. no. SA00013-4; Proteintech Group, Inc.) for 1
h at RT and DAPI (10 µg/ml) for 10 min at RT. Images were recorded
via confocal microscopy (STELLARIS 5; Leica Microsystems, Inc.)
with the following settings: CoraLite594 excitation/emission,
588/604 nm; and DAPI excitation/emission, 358/461 nm. For each
experimental condition, the nuclear-to-cytoplasmic ratio of TFEB
and TFE3 was determined in 5 randomly selected cells using ImageJ
software. The DAPI channel defined the nuclear region, and a 2-µm
expansion from this boundary defined the cytoplasmic region for
intensity measurement.
Lysosomal membrane integrity
assay
MDA-MB-231 and Hs578T cells were seeded in 35-mm
glass-bottom dishes at a density of 2×105 cells per
dish. The lysosomes were pre-loaded with 100 µg/ml Alexa Fluor™
488-dextran (cat. no. D22910; Thermo Fisher Scientific, Inc.) for 6
h at 37°C, followed by a 2-h incubation in fresh DMEM medium at
37°C. After treatment for 24 h with CEP (10 µM for MDA-MB-231, and
4 µM for Hs578T), the retention of dextran within the lysosomes was
assessed by confocal microscopy (STELLARIS 5; Leica Microsystems,
Inc.; Alexa Fluor 488 excitation/emission, 495/519 nm). This
provided a marker of membrane integrity.
Lysosomal pH assay
MDA-MB-231 and Hs578T cells were seeded in 35-mm
glass-bottom dishes at a density of 2×105 cells per dish
and exposed to CEP (10 µM for MDA-MB-231, and 4 µM for Hs578T) for
24 h at 37°C. Lysosomal pH was measured by incubating the cells
with 2 µM LysoSensor™ Green DND-189 (cat. no. 40767ES50; Shanghai
Yeasen Biotechnology Co., Ltd.) for 5 min at 37°C, followed by
washing with PBS and immediate image capture of fluorescence via
confocal microscopy (STELLARIS 5; Leica Microsystems, Inc.);
LysoSensor Green DND-189 excitation/emission, 443/505 nm).
Limited proteolysis-coupled mass
spectrometry (LiP-MS) assay
This assay was conducted using MDA-MB-231 cells.
MDA-MB-231 cells were treated with 10 µM CEP or 0.1% DMSO for 1 h
at 37°C, harvested by ice-cold scraping, lysed in sodium
deoxycholate buffer (5% sodium deoxycholate; 1 mM
KH2PO4; 3 mM Na2HPO4;
155 mM NaCl; pH 7.5) and centrifuged at 16,000 × g for 10 min at
4°C. The cleared lysates were quantified via BCA assay and
processed as followed in parallel: i) The LiP group was subject to
proteinase K (cat. no. P4850; Sigma-Aldrich; Merck KGaA) digestion
(1:100) at 25°C for 1 h, followed by heat inactivation at 100°C for
5 min; and ii) the trypsin (TrP) only control group. All samples
(both LiP and TrP groups) were denatured with 1% sodium
deoxycholate at 95°C for 10 min, reduced with 10 mM
tris(2-carboxyethyl)phosphine for 10 min at RT, and following pH
adjustment to a pH of 9, digested sequentially with C-terminal
lysin (1:100; cat. no. VA1170; Promega Corporation) at 37°C for 4 h
and trypsin (1:100) at 37°C overnight. The isolated peptides were
then acidified to pH≤3 with formic acid, centrifuged at 16,000 × g
for 10 min at 4°C, desalted using Monospin C18 columns (cat. no.
5010-21700; GL Sciences Inc.) and subsequently vacuum-dried.
For the MS analysis, the samples were dissolved in
0.1% formic acid and loaded onto a pulled-tip analytical column
(cat. no. 00G-4053-B0; Phenomenex Inc.) connected to an Easy-nLC
1200 UHPLC system (Thermo Fisher Scientific, Inc.). The mobile
phase consisted of solvent A (0.1% formic acid in water) and
solvent B (0.1% formic acid in 80% acetonitrile). The peptides were
separated using the following gradient: 0–1 min, 8% B; 1–8 min, 12%
B; 8–63 min, 30% B; 63–75 min, 40% B; 75–76 min, 95% B; and 76–90
min, 95% B (flow rate, 300 nl/min). The eluted peptides were then
electrosprayed into a Q Exactive HF-X (Thermo Fisher Scientific,
Inc.), allowing for acquisition of full-scan MS spectra (m/z,
350–1,500) at a resolution of 120,000 (m/z, 200), with the top 40
precursor ions being selected for higher-energy C-trap dissociation
(MS/MS resolution, 15,000; m/z, 200).
The data were processed using Proteome Discoverer
(version 2.4; Thermo Fisher Scientific, Inc.) with the following
parameters: Precursor mass tolerance of ±10 ppm, fragment mass
tolerance of ±0.02 Da and fixed modification that included
carbamidomethylation of cysteine (+57.0214 Da). Oxidation of
methionine (+15.9949 Da) was set as the dynamic modification. The
analysis allowed for up to two missed cleavages and maintained an
FDR threshold of 1%. The abundance of individual peptides was
normalized, imputed, corrected and analyzed for differential
expression using a two-tailed Student's t-test (P<0.01 and FC
>1.5 or <0.667).
Statistical analysis
The data are presented as mean±standard deviation
from three biological replicates. Statistical analyses were
performed using GraphPad Prism (version 9.0.0; Dotmatics). For
comparisons between two groups, unpaired two-tailed Student's
t-tests were used. For comparisons of ≥3 groups, one-way ANOVA was
performed, followed by Dunnett's post hoc test for multiple
comparisons. P<0.05 was considered to indicate a statistically
significant difference.
Results
CEP induces apoptosis in TNBC
cells
CEP (Fig. 1A)
showed dose- and time-dependent cytotoxicity against both TNBC cell
lines (Fig. 1B), with
IC50 values after 48 h treatment of 9.37±0.55 µM for
MDA-MB-231 and 6.07±0.31 µM for Hs578T. The IC50 for the
normal human breast epithelial cell line MCF10A was 13.21±0.54 µM
at 48 h. CEP treatment at higher doses resulted in the clonogenic
formation of TNBC cells being suppressed significantly compared
with untreated cells (Fig. 1C).
Flow cytometric analysis also provided evidence that CEP treatment
significantly induced apoptosis in TNBC cells (Fig. 2), and consistent with this finding,
CEP treatment resulted in a significant increase in TUNEL-positive
cells and caspase-3 activation (Fig.
S1).
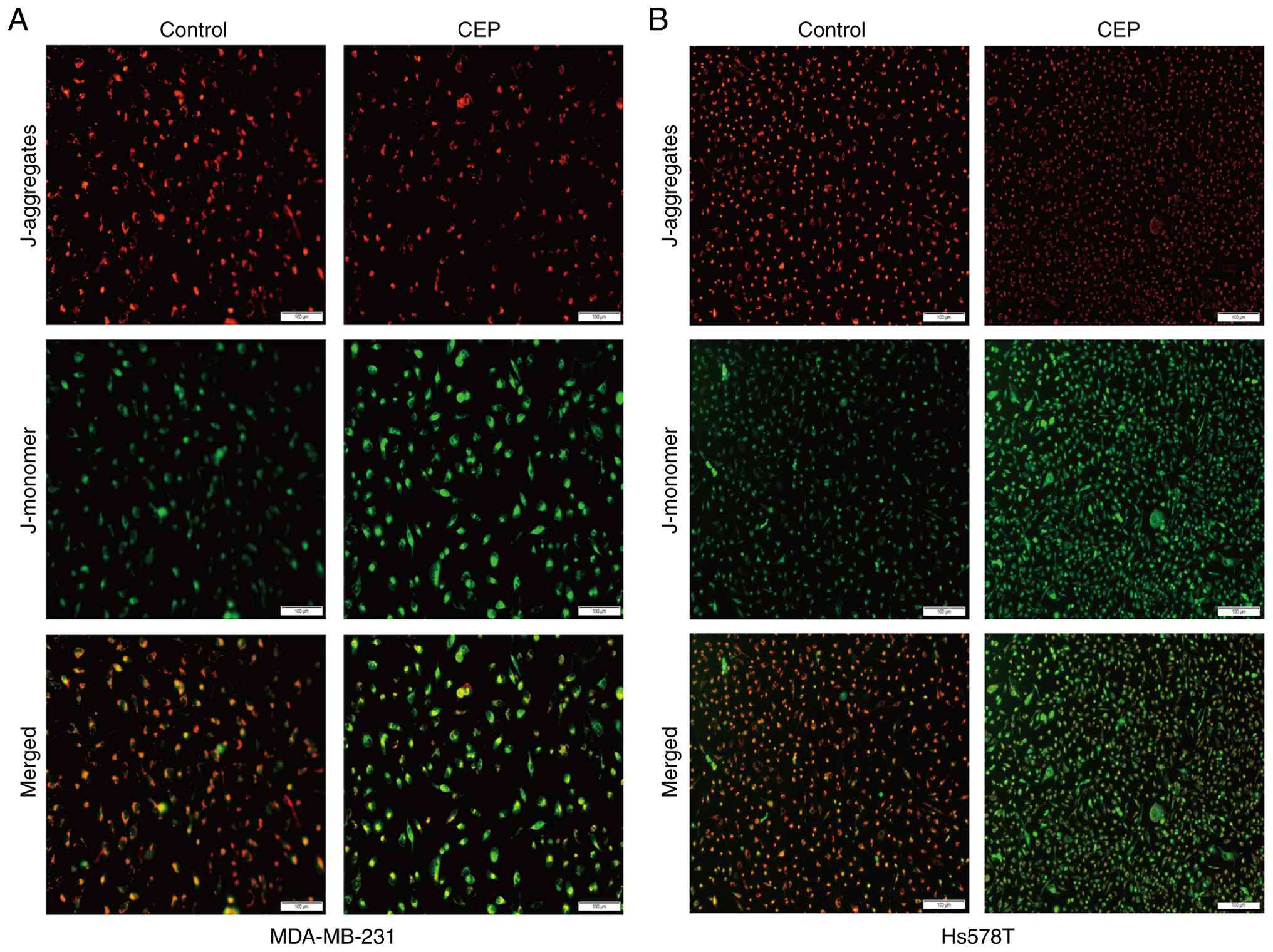
Given that the collapse of the ΔΨm is an important
event in the intrinsic apoptotic pathway (10), the present study subsequently
assessed the effect of CEP on ΔΨm and observed that CEP triggered
notable ΔΨm depolarization, as evidenced by a marked increase in
JC-1 monomer fluorescence and a complementary decrease in aggregate
fluorescence (Fig. 3). Given that
the Bcl-2 protein family is a key regulator of mitochondrial
apoptosis through regulation of ΔΨm (11), the present study examined the
expression of relevant apoptosis-related proteins. Among the Bcl-2
family proteins examined, CEP treatment significantly upregulated
the pro-apoptotic protein NOXA and downregulated the anti-apoptotic
protein Bcl-2, whereas no significant change was observed in the
expression of Bax (Fig. 4). Taken
together, these results demonstrated that CEP induced apoptosis in
TNBC cells, at least in part, by modulating the expression of NOXA
and Bcl-2, leading to mitochondrial dysfunction.
CEP binds to and inhibits lysosomal
enzymes in TNBC cells
The present study used quantitative proteomics to
investigate the mechanism of action of CEP. In MDA-MB-231 cells,
CEP treatment significantly altered the expression of 634 proteins,
of which 521 were upregulated and 113 were downregulated (Table SI). GO and KEGG enrichment
analyses indicated a notable enrichment of lysosomal pathways among
the downregulated proteins (Fig.
5), suggesting that CEP impaired lysosomal integrity or
function. Consistent with this finding, confocal microscopy showed
that CEP treatment triggered a significant increase in the nuclear
accumulation of TFEB, the master regulator of lysosomal biogenesis,
compared with the control group (Fig.
6). This result further supported the interpretation that CEP
compromised lysosomal function and activated the lysosomal stress
response.
Weakly basic compounds can accumulate in lysosomes
via ion trapping, inducing lysosomal membrane permeabilization
(LMP) through detergent-like effects (12–15).
These compounds can also disrupt lysosomal function by either
neutralizing lysosomal pH or destabilizing membrane-associated
enzymes, such as ASAH1, via interactions with anionic lipids
(16–20). Due to the presence of a basic
isoquinoline group in its structure (21,22),
the present study hypothesized that CEP may have induced LMP or
disrupted lysosomal function through similar mechanisms. However,
the Alexa Fluor 488-dextran retention assay showed no cytoplasmic
diffusion after CEP treatment, thereby ruling out induction of LMP
in TNBC cells (Fig. 7A). Lysosomal
pH was assessed by fluorescence imaging using LysoSensor Green
DND-189, a pH-sensitive probe with a fluorescence intensity
inversely related to acidity. This assay showed brighter
fluorescence of LysoSensor™ Green DND-189 in TNBCs cells,
indicating TNBC cells had a reduced lysosomal pH compared with
normal breast epithelial cells (Fig.
S2), and CEP did not increase the level of lysosomal pH in TNBC
cells (Fig. 7B). In addition, CEP
caused no significant alterations in the expression of the
lysosomal membrane-associated enzymes ASAH1 and SMPD1 (Fig. 7C). The present results suggested
that CEP did not promote the degradation of these enzymes in TNBC
cells.
To determine whether CEP directly inhibited
lysosomal enzymes, the present study performed LiP-MS analysis on
TNBC cells. LiP-MS identified 1,163 structurally altered peptides
from 663 proteins, including heat shock protein 90α, a known CEP
target (Table SII), which
confirmed the reliability of the method used in the present study
(23). Among these proteins, two
lysosomal hydrolases, CTSB, comprising 2 peptides, and CTSD,
comprising 4 peptides, exhibited significant conformational changes
induced by CEP (Fig. 8A). Western
blot analysis further showed that CEP significantly impeded the
maturation of CTSB and CTSD, leading to reduced levels of their
active forms (Fig. 8B). Taken
together, these findings demonstrated that CEP interacted directly
with the lysosomal enzymes CTSB and CTSD and hindered their
functional activation.
 | Figure 8.CEP binds to and inhibits lysosomal
enzymes. (A) Structurally altered peptides in the MDA-MB-231 cell
line upon CEP treatment (10 µM, 1 h; FC >1.5 or <0.667, false
discovery rate <1, P<0.01; n=3). (B) CEP treatment suppresses
the maturation of CTSB and CTSD (24 h). *P<0.05, **P<0.01 and
***P<0.001 vs. control. CEP, cepharanthine; FC, fold change;
CTSD, cathepsin D; CTSB, cathepsin B; pro-CTSB pro-cathepsin B;
pro-CTSD, pro-cathepsin D; i-CTSB, inactive cathepsin B; m-CTSB,
mature cathepsin B; m-CTSD, mature cathepsin D. |
Discussion
Our previous study demonstrated that a low dose of
CEP (2 µM) did not induce ΔΨm loss or apoptosis in TNBC cells
(24). However, at this
concentration, CEP was found to enhance the efficacy of epirubicin
against TNBC by blocking autophagic flux, specifically through
inhibiting autophagosome-lysosome fusion (24). This finding prompted further
investigation into the protein targets and pharmacological
mechanisms of CEP in TNBC cells. The present study observed that
relatively higher concentrations of CEP could induce ΔΨm loss (10
µM for MDA-MB-231 and 4 µM for Hs578T cells) and promote apoptosis
in TNBC cells (10–20 µM for MDA-MB-231 and 2–8 µM for Hs578T
cells). This was accompanied by the downregulation of Bcl-2
expression, a result consistent with prior reports in a study
performed by Gao et al (25). However, in contrast to the study by
Gao et al (25), the present study did not observe an
upregulation of Bax in TNBC cells. This discrepancy could be
attributed to the different time points assessed, as evidenced by
the use of 24 h incubations in the present study compared with 48 h
incubations in the reported literature. Additionally, the present
study showed that CEP upregulated the expression of NOXA, another
pro-apoptotic protein in the Bcl-2 family.
Quantitative proteomic analysis revealed significant
alterations in the expression of lysosomal proteins following CEP
treatment, suggesting the potential inhibition of lysosomal
function by CEP. Despite possessing a basic isoquinoline group
(21), CEP did not increase
lysosomal pH or induce LMP at the concentrations tested in the
present study. These observations may have been attributable to the
relatively low doses of CEP used, as previous studies indicate that
low doses of weakly basic drugs such as chloroquine and
clomipramine either fail to elevate or cause only a transient
increase in lysosomal pH (26,27).
Similarly, induction of LMP by these drugs typically requires
higher concentrations, such as 50 µM chloroquine in the 5637 and
T24 cell lines and 100 µM chloroquine in HCT116 cells (28,29).
The present results of the LiP-MS and western blot analyses
provided evidence that CEP bound to the lysosomal hydrolases CTSB
and CTSD and inhibited their maturation. These findings
demonstrated that CEP suppressed lysosomal degradation through
direct enzyme inhibition.
TFEB, a master transcriptional regulator of
lysosomal genes, is activated in response to lysosomal damage or
inhibition (12). Recent studies
have shown that TFEB induces tumor apoptosis by transcriptional
upregulation of cyclic AMP-dependent transcription factor ATF-4
(ATF4) and DNA damage-inducible transcript 3 protein (CHOP)
expression (30–32). As NOXA and Bcl-2 are
transcriptional targets of ATF4 or CHOP (33–35),
the present study proposed that CEP may have activated the
TFEB/ATF4/CHOP axis, thereby upregulating NOXA and downregulating
Bcl-2 expression in TNBC cells. Furthermore, there is evidence that
NOXA undergoes degradation via the autophagy-lysosomal pathway,
which indicates that autophagy inhibition may prevent NOXA
degradation (36–38). Taken together, the findings of the
present study and existing literature indicated that CEP induced
apoptosis in TNBC cells by regulating NOXA and Bcl-2 expression
through lysosomal inhibition of CTSB and CTSD.
Although CEP-mediated lysosomal inhibition remains
poorly elucidated, TNBC cells exhibited heightened susceptibility
to CEP treatment compared with normal epithelial cells of the
MCF10A cell line. This differential sensitivity probably stemmed
from the increased dependence of tumor cells on lysosomal function.
Studies have shown that tumor cells have a higher proliferation
rate and metabolic requirements, which may make them more reliant
on lysosome-mediated degradation and recycling pathways (39,40).
Consistent with this dependency, tumor cells have been shown to
have increased lysosomal abundance and higher lysosomal enzyme
expression and activity than normal cells (41–43).
As a consequence, lysosomal inhibition would be expected to have
more severe effects in malignant cells than in normal cells.
Furthermore, tumor cells maintain a lower lysosomal pH than normal
cells, a property which is beneficial for the maturation and
activity of lysosomal enzymes (44). This acidic environment enhances
proton trapping and subsequent accumulation of CEP within the
lysosomes. Such preferential accumulation of CEP could have
potentially amplified lysosomal inhibition and cytotoxicity toward
TNBC cells. The findings of the present study showed that the CEP
sensitivity gradient of Hs578T>MDA-MB-231>MCF10A aligned with
the gradient of lysosomal acidification across the three cell
lines, which supported this possibility.
Due to the methodological constraints of LiP-MS, the
present study could not exclude additional CEP-binding lysosomal
proteins or non-protein targets. Future investigations are
therefore needed to identify CEP interactors within lysosomes. In
conclusion, the present study established that CEP directly
inhibited lysosomes and triggered mitochondrial apoptosis in TNBC
cells, thereby providing a foundation for further investigation of
the pharmacological mechanisms of CEP activity.
Supplementary Material
Supporting Data
Supporting Data
Supporting Data
Acknowledgements
Not applicable.
Funding
The present work was supported by the National Natural Science
Foundation of China (grant no. 82004009) and the Science and
Technology Plan Program of Guizhou Province (grant no. [2018],
5772-071).
Availability of data and materials
The data generated in the present study are included
in the figures and/or tables of this article. The limited
proteolysis-coupled mass spectrometry and proteomics data generated
in the present study may be found in the iProX database under
accession numbers (project ID: IPX0012108000, proteomeXchange ID:
PXD064430, password: dFLs; proteomeXchange ID: PXD064550, project
ID: IPX0012132000, password: 7eTd) or at the following URLs
(https://www.iprox.cn/page/PSV023.html;?url=1748921339238lMbz;
http://www.iprox.cn/page/PSV023.html;?url=1748921458718JEnw).
Authors' contributions
LS and DY designed experiments and acquired funding.
JL, SZ and YW conducted experiments and collected the data. JL and
YW wrote the manuscript. LS and GO analyzed the results and
reviewed the manuscript. JL, SZ, YW, GO, DY and LS confirm the
authenticity of all the raw data. All authors read and approved the
final version of the manuscript.
Ethics approval and consent to
participate
Not applicable.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
Use of artificial intelligence tools
During the preparation of this work, Deepseek was
used to improve the readability and language of the manuscript, and
subsequently, the authors revised and edited the content produced
by Deepseek as necessary, taking full responsibility for the
ultimate content of the present manuscript.
References
|
1
|
Sung H, Ferlay J, Siegel RL, Laversanne M,
Soerjomataram I, Jemal A and Bray F: Global cancer statistics 2020:
GLOBOCAN estimates of incidence and mortality worldwide for 36
cancers in 185 countries. CA Cancer J Clin. 71:209–249.
2021.PubMed/NCBI
|
|
2
|
Urru SAM, Gallus S, Bosetti C, Moi T,
Medda R, Sollai E, Murgia A, Sanges F, Pira G, Manca A, et al:
Clinical and pathological factors influencing survival in a large
cohort of triple-negative breast cancer patients. BMC Cancer.
18:562018. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Moss JL, Tatalovich Z, Zhu L, Morgan C and
Cronin KA: Triple-negative breast cancer incidence in the United
States: Ecological correlations with area-level sociodemographics,
healthcare, and health behaviors. Breast Cancer. 28:82–91. 2021.
View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Wu Q, Siddharth S and Sharma D: Triple
negative breast cancer: A mountain yet to be scaled despite the
triumphs. Cancers (Basel). 13:36972021. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Baranova A, Krasnoselskyi M, Starikov V,
Kartashov S, Zhulkevych I, Vlasenko V, Oleshko K, Bilodid O,
Sadchikova M and Vinnyk Y: Triple-negative breast cancer: Current
treatment strategies and factors of negative prognosis. J Med Life.
15:153–161. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Bianchini G, De Angelis C, Licata L and
Gianni L: Treatment landscape of triple-negative breast
cancer-expanded options, evolving needs. Nat Rev Clin Oncol.
19:91–113. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Liu K, Hong B, Wang S, Lou F, You Y, Hu R,
Shafqat A, Fan H and Tong Y: Pharmacological activity of
cepharanthine. Molecules. 28:50192023. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Bailly C: Cepharanthine: An update of its
mode of action, pharmacological properties and medical
applications. Phytomedicine. 62:1529562019. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Demichev V, Messner CB, Vernardis SI,
Lilley KS and Ralser M: DIA-NN: Neural networks and interference
correction enable deep proteome coverage in high throughput. Nat
Methods. 17:41–44. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Heerdt BG, Houston MA, Wilson AJ and
Augenlicht LH: The intrinsic mitochondrial membrane potential
(deltapsim) is associated with steady-state mitochondrial activity
and the extent to which colonic epithelial cells undergo
butyrate-mediated growth arrest and apoptosis. Cancer Res.
63:6311–6319. 2003.PubMed/NCBI
|
|
11
|
Czabotar PE and Garcia-Saez AJ: Mechanisms
of BCL-2 family proteins in mitochondrial apoptosis. Nat Rev Mol
Cell Biol. 24:732–748. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Lakpa KL, Khan N, Afghah Z, Chen X and
Geiger JD: Lysosomal stress response (LSR): Physiological
importance and pathological relevance. J Neuroimmune Pharmacol.
16:219–237. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Hu M and Carraway KL III: Repurposing
cationic amphiphilic drugs and derivatives to engage lysosomal cell
death in cancer treatment. Front Oncol. 10:6053612020. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Derendorf H: Excessive lysosomal
ion-trapping of hydroxychloroquine and azithromycin. Int J
Antimicrob Agents. 55:1060072020. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Zhitomirsky B and Assaraf YG: Lysosomal
sequestration of hydrophobic weak base chemotherapeutics triggers
lysosomal biogenesis and lysosome-dependent cancer multidrug
resistance. Oncotarget. 6:1143–1156. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Fu W, Li X, Lu X, Zhang L, Li R, Zhang N,
Liu S, Yang X, Wang Y, Zhao Y, et al: A novel acridine derivative,
LS-1-10 inhibits autophagic degradation and triggers apoptosis in
colon cancer cells. Cell Death Dis. 8:e30862017. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Ellegaard AM, Bach P and Jäättelä M:
Targeting cancer lysosomes with good old cationic amphiphilic
drugs. Rev Physiol Biochem Pharmacol. 185:107–152. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Stark M, Silva TFD, Levin G, Machuqueiro M
and Assaraf YG: The lysosomotropic activity of hydrophobic weak
base drugs is mediated via their intercalation into the lysosomal
membrane. Cells. 9:10822020. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Gebai A, Gorelik A, Li Z, Illes K and
Nagar B: Structural basis for the activation of acid ceramidase.
Nat Commun. 9:16212018. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Scrima S, Lambrughi M, Favaro L, Maeda K,
Jäättelä M and Papaleo E: Acidic sphingomyelinase interactions with
lysosomal membranes and cation amphiphilic drugs: A molecular
dynamics investigation. Comput Struct Biotechnol J. 23:2516–2533.
2024. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Vitello R, Taouba H, Derand M and Liégeois
JF: The bis(1,2,3,4-tetrahydroisoquinoline) alkaloids cepharanthine
and berbamine are ligands of SK channels. ACS Med Chem Lett.
15:215–220. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Jacob J, Varghese N, Rasheed SP, Agnihotri
S, Sharma V and Wakode S: Recent advances in the synthesis of
isoquinoline and its analogue: A review. World J Pharm Pharm Sci.
5:1821–1837. 2016.
|
|
23
|
Haginaka J, Kitabatake T, Hirose I,
Matsunaga H and Moaddel R: Interaction of cepharanthine with
immobilized heat shock protein 90α (Hsp90α) and screening of Hsp90α
inhibitors. Anal Biochem. 434:202–206. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Shen LW, Jiang XX, Li ZQ, Li J, Wang M,
Jia GF, Ding X, Lei L, Gong QH and Gao N: Cepharanthine sensitizes
human triple negative breast cancer cells to chemotherapeutic agent
epirubicin via inducing cofilin oxidation-mediated mitochondrial
fission and apoptosis. Acta Pharmacol Sin. 43:177–193. 2022.
View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Gao S, Li X, Ding X, Qi W and Yang Q:
Cepharanthine induces autophagy, apoptosis and cell cycle arrest in
breast cancer cells. Cell Physiol Biochem. 41:1633–1648. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Mlejnek P, Havlasek J, Pastvova N, Dolezel
P and Dostalova K: Lysosomal sequestration of weak base drugs,
lysosomal biogenesis, and cell cycle alteration. Biomed
Pharmacother. 153:1133282022. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Logan R, Kong AC, Axcell E and Krise JP:
Amine-containing molecules and the induction of an expanded
lysosomal volume phenotype: A structure-activity relationship
study. J Pharm Sci. 103:1572–1580. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Park D and Lee Y: Biphasic activity of
chloroquine in human colorectal cancer cells. Dev Rerprod.
18:225–231. 2014. View Article : Google Scholar
|
|
29
|
Chen HE, Lin JF, Lin YC, Wen SI, Yang SC,
Tsai TF, Chou KY and Hwang IST: Chloroquine induces lysosomal
membrane permeability-mediated cell death in bladder cancer cells.
Formos J Surg. 51:133–141. 2018. View Article : Google Scholar
|
|
30
|
Franco-Juárez B, Coronel-Cruz C,
Hernández-Ochoa B, Gómez-Manzo S, Cárdenas-Rodríguez N,
Arreguin-Espinosa R, Bandala C, Canseco-Ávila LM and Ortega-Cuellar
D: TFEB; beyond its role as an autophagy and lysosomes regulator.
Cells. 11:31532022. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Yang CB, Liu J, Tong BC, Wang ZY, Zhu Z,
Su CF, Sreenivasmurthy SG, Wu JX, Iyaswamy A, Krishnamoorthi S, et
al: TFEB, a master regulator of autophagy and biogenesis,
unexpectedly promotes apoptosis in response to the cyclopentenone
prostaglandin 15d-PGJ2. Acta Pharmacol Sin. 43:1251–1263. 2022.
View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Martina JA, Diab HI, Brady OA and
Puertollano R: TFEB and TFE3 are novel components of the integrated
stress response. EMBO J. 35:479–495. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Zhao X, Kong F, Wang L and Zhang H: c-FLIP
and the NOXA/Mcl-1 axis participate in the synergistic effect of
pemetrexed plus cisplatin in human choroidal melanoma cells. PLoS
One. 1:e01841352017. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Núñez-Vázquez S, Sánchez-Vera I,
Saura-Esteller J, Cosialls AM, Noisier AFM, Albericio F, Lavilla R,
Pons G, Iglesias-Serret D and Gil J: NOXA upregulation by the
prohibitin-binding compound fluorizoline is transcriptionally
regulated by integrated stress response-induced ATF3 and ATF4. FEBS
J. 288:1271–1285. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Armstrong JL, Flockhart R, Veal GJ, Lovat
PE and Redfern CP: Regulation of endoplasmic reticulum
stress-induced cell death by ATF4 in neuroectodermal tumor cells. J
Biol Chem. 285:6091–6100. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Kuroda Y, Koyama D, Kikuchi J, Mori S,
Ichinohe T and Furukawa Y: Autophagic degradation of NOXA underlies
stromal cell-mediated resistance to proteasome inhibitors in mantle
cell lymphoma. Leuk Res. 111:1066722021. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Wang J, Cui D, Gu S, Chen X, Bi Y, Xiong X
and Zhao Y: Autophagy regulates apoptosis by targeting NOXA for
degradation. Biochim Biophys Acta Mol Cell Res. 1865:1105–1113.
2018. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Moriya S, Kazama H, Hino H, Takano N,
Hiramoto M, Aizawa S and Miyazawa K: Clarithromycin overcomes
stromal cell-mediated drug resistance against proteasome inhibitors
in myeloma cells via autophagy flux blockage leading to high NOXA
expression. PLoS One. 18:e02952732023. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Davidson SM and Vander Heiden MG: Critical
functions of the lysosome in cancer biology. Annu Rev Pharmacol
Toxicol. 57:481–507. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Zhang Z, Yue P, Lu T, Wang Y, Wei Y and
Wei X: Role of lysosomes in physiological activities, diseases, and
therapy. J Hematol Oncol. 14:792021. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Chen R, Jäättelä M and Liu B: Lysosome as
a central hub for rewiring pH homeostasis in tumors. Cancers
(Basel). 12:24372020. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Serrano-Puebla A and Boya P: Lysosomal
membrane permeabilization as a cell death mechanism in cancer
cells. Biochem Soc Trans. 46:207–215. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Kirkegaard T and Jäättelä M: Lysosomal
involvement in cell death and cancer. Biochim Biophys Acta.
1793:746–754. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Kroemer G and Jäättelä M: Lysosomes and
autophagy in cell death control. Nat Rev Cancer. 5:886–897. 2005.
View Article : Google Scholar : PubMed/NCBI
|