Introduction
Hepatocellular carcinoma (HCC) is the fifth most
common malignancy worldwide and is the third leading cause of
cancer-related mortality in China (1). HCC is a rapidly growing tumor
associated with a propensity for vascular invasion and metastasis,
which results in poor cancer prognoses (2). Considering that methods for early
detection of HCC are currently unavailable, the majority of
patients present with peritoneal dissemination and distant
metastasis at the time of diagnosis (3). Thus, treatment is often unsuccessful
and overall survival time is short. Consequently, identifying
invasion-related molecules associated with the early and rapid
spread of HCC is a focus of investigation.
Invasion and metastasis are the biological hallmarks
of malignancy (4). The molecular
mechanisms underlying invasion and metastasis are an important area
of investigation. Epithelial-mesenchymal transition (EMT) is a
common event in the plasticity of tumor cells (5,6). As
invasion proceeds, epithelial cell layers lose polarity and
cell-cell interaction, which ultimately results in the complex
remodeling of the cytoskeleton. EMT is an embryonic trait by which
cells adopt a phenotype that is more suitable to migration and
invasion (7). A number of molecules
associated with tumor invasion and EMT in HCC have been reported,
including Twist, Snail and Slug (8–10).
However, the molecular changes associated with the metastatic
ability in HCC progression have not been clearly determined.
Materials and methods
Cell culture and transfection
The HepG2 cell line was obtained from the American
Type Culture Collection (Rockville, MD, USA). The cells were
cultured in Dulbecco’s modified Eagle’s medium (DMEM) supplemented
with 10% fetal bovine serum (FBS; Invitrogen Life Technologies,
Carlsbad, CA, USA). pcDNA3.1/Slug, pcDNA3.1/Snail and pcDNA3.1
empty vector (Clontech, Palo Alto, CA, USA) were transfected into
the cells using polyethylenimine (cat no. 23966; Polysciences,
Inc., Warrington, PA, USA).
Expression plasmids
Full-length Snail and Slug cDNA were generated using
normal human embryo total cDNA, cleaved with
XhoI/EcoRI and subcloned into pcDNA3.1 vectors. The
resulting constructs were confirmed by DNA sequencing. The pcDNA3.1
empty vector was used as a cDNA control. Small interfering
(si)RNA-coding oligonucleotides against human Snail and Slug were
designed and verified for specificity for Snail and Slug. The Snail
siRNA targeting sequence was AAGCTGAGCAAGATTCAGACC and the Slug
siRNA targeting sequence was CAGGACCTCGCCGCTGCAGAC (siB-cell
lymphoma 2 nucleotides 200–221) (11). The U6 promoter with a Snail or Slug
siRNA insert was subcloned into pRNA-U6-Neo. A non-silencing siRNA
sequence (target sequence, AATTCTCCGAACGTGTCACGT) was used as the
negative control.
Invasion and wound healing assays
The cell migration assay was performed using
Transwell cell culture inserts (Invitrogen Life Technologies). The
transfected cells were maintained for 48 h and allowed to migrate
for an additional 24 h. The passaged cells were stained with
crystal violet solution and absorbance was measured at 595 nm. In
the wound healing assays, cell motility was assessed by measuring
the movement of cells towards the scratch. The speed of wound
closure was monitored after 12 and 24 h by measuring the ratio of
the distance of the wound at 0 h. Each experiment was performed in
triplicate.
Colony formation assay
The control and transfected cells were seeded into
six-well plates at a density of 1,000 cells/well. After two weeks,
the clones were fixed in methanol and stained with 2% Giemsa
solution (Merck KGaA, Darmstadt, Germany) for 10 min.
Flow cytometry analysis
Following treatment, the cells were fixed in 75%
ethanol and indirectly labeled by incubation with the primary
anti-SNAI1 rabbit polyclonal (ab82846; Abcam, Cambridge, MA, USA),
anti-SNAI2 mouse monoclonal (ab51772; Abcam), anti-E-cadherin
rabbit polyclonal (sc-7870; Santa Cruz Biotechnology, Inc., Santa
Cruz, CA, USA), anti-vimentin mouse monoclonal (ab20346; Abcam) and
anti-CD133 mouse monoclonal (130080801; Miltenyi Biotech, San
Diego, CA, USA) and the secondary antibodies; goat anti-mouse
polyclonal IgG horse radish peroxidase (HRP) conjugated antibody
(sc-2005, Santa Cruz Biotechnology, Inc.) and goat anti-rabbit
polyclonal IgG-HRP conjugated antibody (sc-2004, Santa Cruz
Biotechnology, Inc.). The percentage of CD133-positive cells was
identified by using flow cytometric analyses with the CD133
monocolonal antibody directly conjugated with phycoerythrin
(Miltenyi Biotec, Auburn, CA, USA). All cells were stained and
incubated with a CD133-phycoerythrin antibody. A C6 flow cytometer
(BD Accuri™, Franklin Lakes, NJ, USA) was used to determine the
percent of CD133-positive cells.
Western blot analysis
Whole cell lysates were resolved by sodium dodecyl
sulfate-polyacrylamide gel electrophoresis (Invitrogen Life
Technologies) and transferred onto polyvinylidene difluoride
membranes (Millipore, Billerica, MA, USA). Blots were blocked and
incubated with the mouse monoclonal antibody (anti-SNAI2; ab51772;
Abcam) followed by incubation with a secondary antibody (1:2,000;
Santa Cruz Biotechnology, Inc.). Blots were developed using an
enhanced chemiluminescence detection kit (Amersham Pharmacia
Biotech, Inc., Piscataway, NJ, USA). Monoclonal β-actin antibody
(1:200; Santa Cruz Biotechnology, Inc.) was used for protein
loading analyses.
Murine xenograft model
Female BALB/c-null mice (age, six weeks) were
obtained from the National Institutes of Health (Bethesda, MD, USA)
and housed in the animal facilities at the Tianjin Medical
University (Tianjin, China), as approved by the Institutional
Animal Care and Use Committee. The HepG2 cells (107
cells/ml) were mixed with Matrigel (BD Biosciences, Franklin Lakes,
NJ, USA) and subcutaneously injected into the backs of the nude
mice (0.1 ml/mouse). The mice were monitored and tumor sizes were
measured daily using a caliper for 25 days. The experiments were
terminated after 25 days due to the tendency of HepG2 cells to
become necrotic and form skin ulcers. The mice were sacrificed by
CO2 asphyxiation following observations. The tumors were
harvested and stored at −80°C for subsequent tests.
Sulfrodamine B (SRB) cell proliferation
assay
Cells were seeded in a 96-well plate (Corning Inc.,
Corning, NY, USA) at a final concentration of 5000 cells per well
follwoing transfection and maintained in DMEM (Gibco-Brl, Grand
Island, NY, USA) containing 10% FBS (Gibco-Brl) at 37°C in a
humidified atmosphere containing 5% CO2. Cell
proliferation was analyzed every 8 hours. Initially the cells were
washed with distilled water after being fixed with trichloroacetic
acid, and were subsequently stained for 10 min with sulforhodamine
B (Sigma-Aldrich, St. Louis, MO, USA) dissolved in 1.0% acetic acid
(Sigma-Aldrich). The plates were washed with 1.0% acetic acid and
allowed to air dry. Finally, 150 μl of 10 mM Tris base
(Sigma-Aldrich) was added to each well in order to solubilize the
dye, and the absorbance at 490 nm was determined with the use of a
microplate reader (Synergy H4, Bio-Tek, Winooski, VT, USA).
Statistical analysis
All the data were evaluated using SPSS software,
version 11.5 (SPSS, Inc., Chicago, IL, USA) and analyzed using the
Student’s t-test and analysis of variance. P<0.05 was considered
to indicate a statistically significant difference. The significant
groups are marked with asterisks as shown in the figures.
Results
Slug upregulation increases HepG2 cell
proliferation in vitro
The effects of Slug expression on cell invasion,
migration and clone formation were investigated. E-cadherin
expression is frequently associated with metastatic ovarian
carcinoma (12). Metastasis is
associated with cell migration and invasion, and the underlying
mechanisms are similar to those of EMT. The invasiveness and
migration capability of HepG2 cells in the Slug and Snail
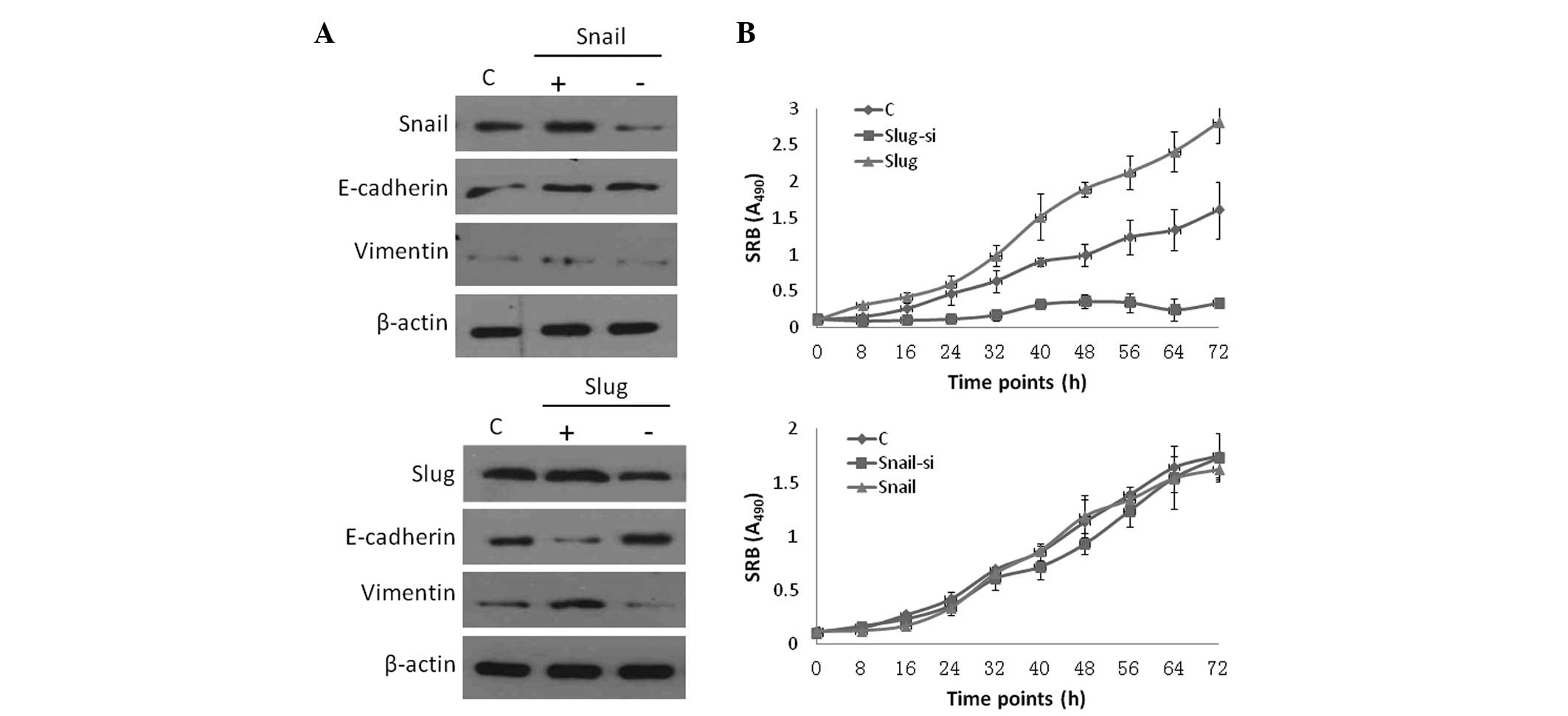
transfection and knockdown groups were examined. Western blot
analysis revealed a significant difference in the ectopic
transfection groups compared with the control group. Slug and Snail
were upregulated in the overexpressed group and were downregulated
in the siRNA group (Fig. 1A). The
sulforhodamine B protein assay was used to measure cell numbers and
a significant difference in cell proliferation was observed between
the Slug overexpression and knockdown groups. The cells exhibited a
significant decrease in cell proliferation in the Slug knockdown
group, whereas cell proliferation was significantly increased in
the overexpressed group compared with the control group
(P<0.01). Snail expression did not significantly affect cell
proliferation (P>0.01).
Slug upregulation increases HCC cell
invasion, migration and clone formation in vitro
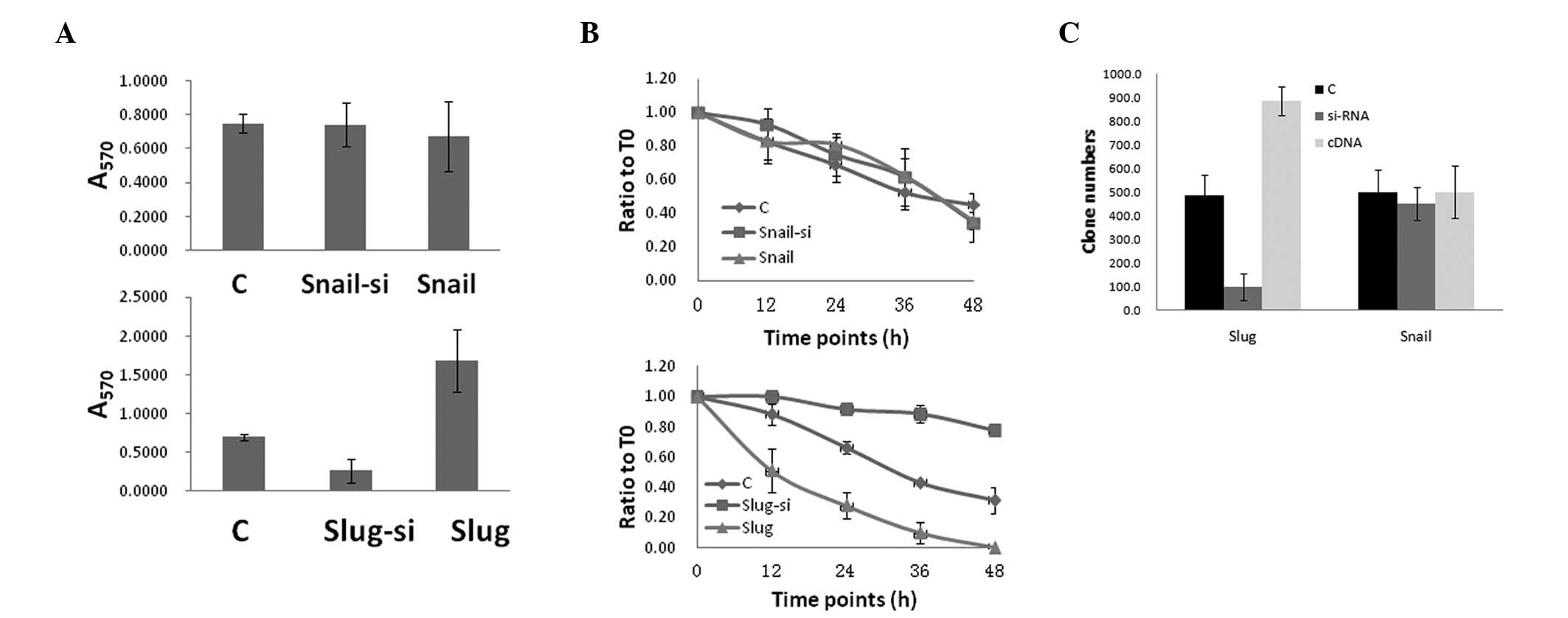
The HepG2 cell cultures were analyzed for functional
changes in migration, invasion and clone formation following
transfection with Slug and Snail, separately. Compared with the
transfection and control groups, the Slug overexpression group
exhibited a significantly increased activity in the migration,
invasion and clonigenicity assays (Fig.
2). The effects of non-uniform transfection efficiency were
minimized by selecting Slug siRNA-transfected cells for slug
knockdown, and the Slug knockdown group revealed decreased activity
in the migration, invasion and clonigenicity assays compared with
the control group. Notably, Snail, which shares structural homology
with Slug, did not promote cell migration.
Slug upregulation increases the
percentage of cluster of differentiation (CD)133+ cells
in HCC cells
Stem cells play an important role in tumor cell
plasticity and proliferation during tumor metastasis, and are
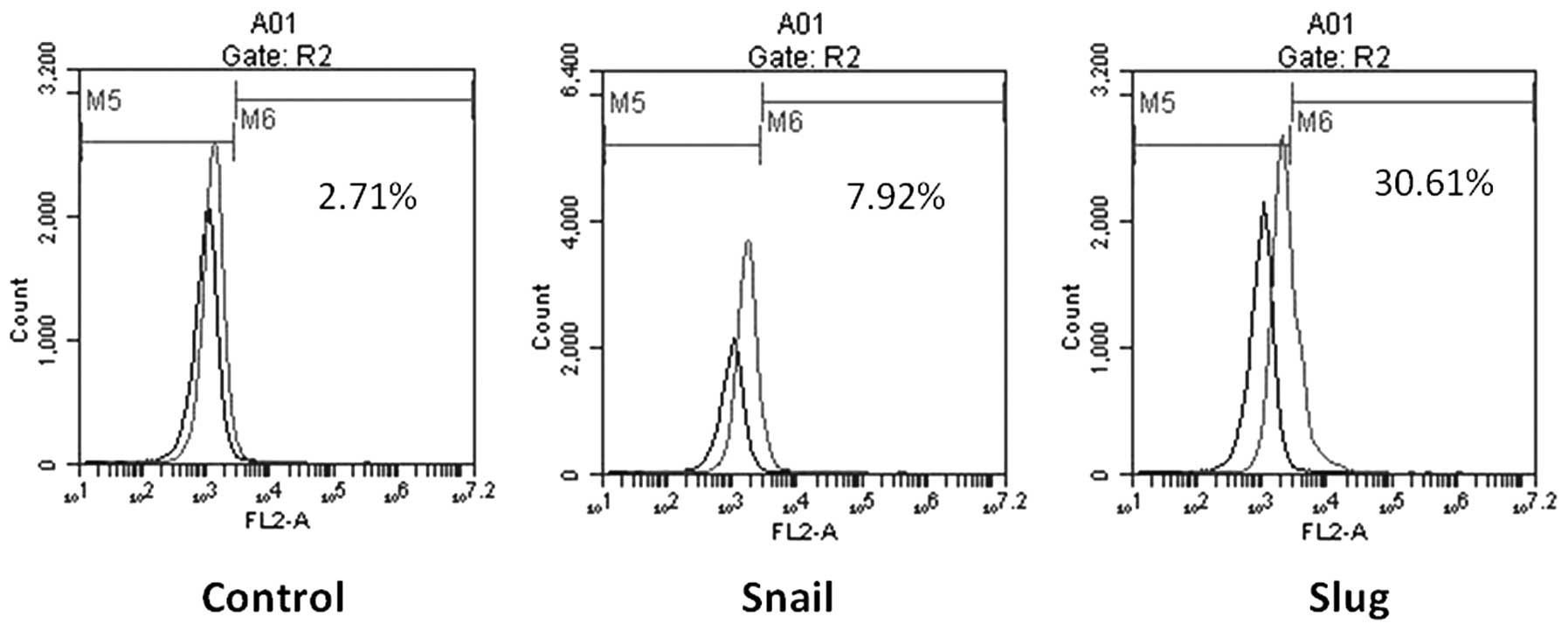
strongly associated with EMT. In this study, flow cytometry
examined the percentage of CD133+ cells in the Slug
overexpression and knockdown groups. The CD133+ cells
accounted for 47.5% of the cells in the Slug overexpression group,
8.7% in the control transfection group and 0.4% in the Slug
knockdown group (Fig. 3). No
significant difference in the percentage of CD133+ cells
was observed between the Snail overexpression, Snail knockdown and
control groups. These findings suggest that synergism between Slug
and CD133+ cells increases cell proliferation, migration
and invasion.
Coexpression of CD133 and Slug correlates
with tumor proliferation
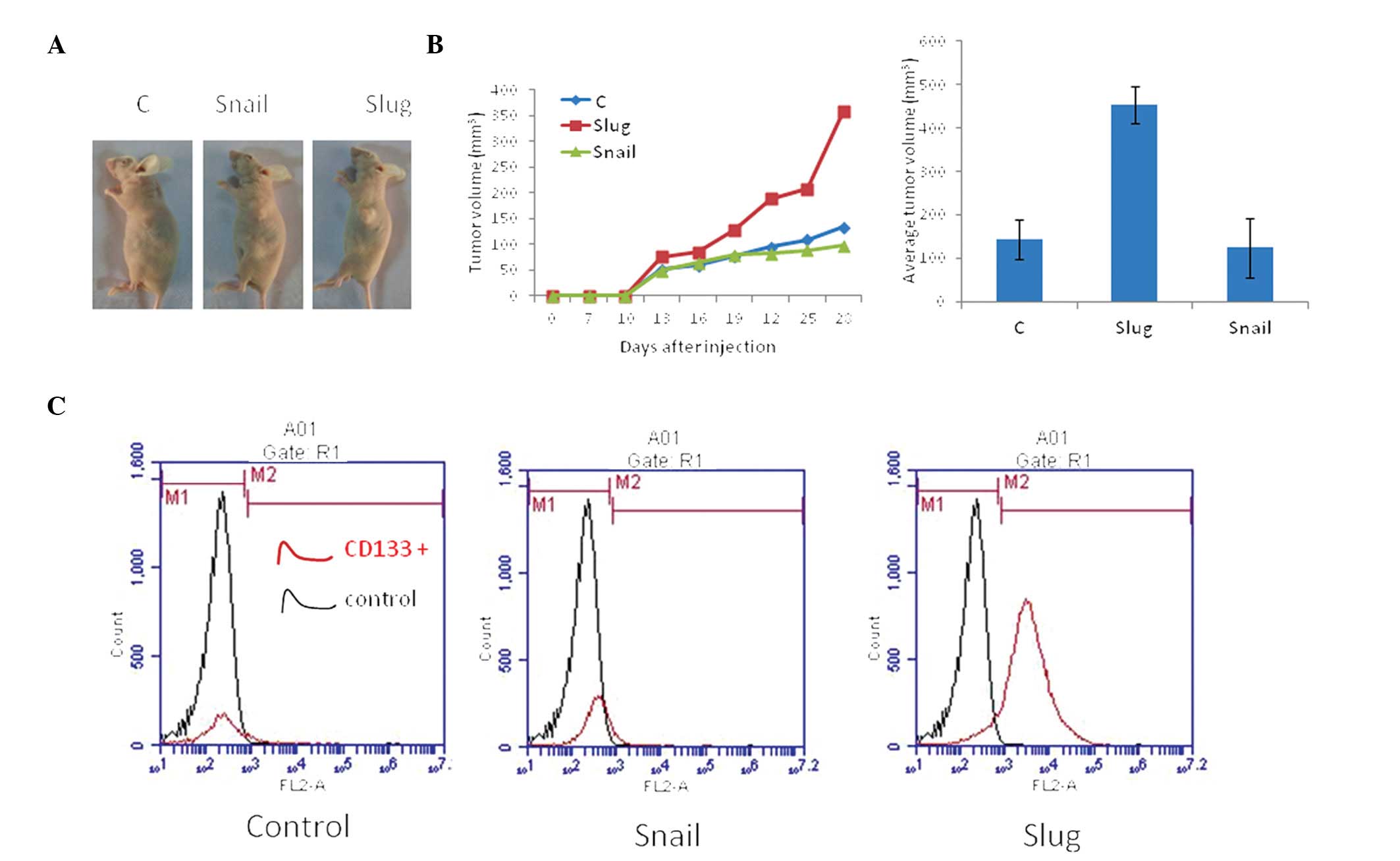
A murine xenograft model was used to examine the
in vivo effects of Slug and Snail expression on tumor
development. The HepG2 cells were utilized to establish xenografts
in nude mice. Nodule formation and growth (volume) were monitored
over 25 days. The Slug overexpression group displayed the highest
rate of tumor growth compared with the Snail overexpression (60%)
and control (50%) groups (Fig. 4).
The cells were collected from the tumor tissues and analyzed using
flow cytometry. The findings demonstrated a significant increase in
CD133+ cells in the Slug overexpression group compared
with the control group, but no significant difference was observed
between the Snail overexpression and the control groups (Fig. 4).
Discussion
HCC invasiveness is a key step that results in
metastasis and a poor prognosis (1,11);
therefore, the underlying molecular mechanisms are a focus of
investigation. EMT plays an important role in the development of
tissues during embryogenesis (13).
However, similar cell changes are recapitulated during pathological
processes, such as in cancer progression. Previous studies on EMT
have focused on cancer cell invasion and metastasis (6,14).
Several developmental genes that induce EMT act as E-cadherin
repressors. The first of these genes is the Snail family of
zinc-finger protein transcription factors, which is a DNA-binding
factor family that recognizes E-box motifs in target promoters,
such as E-cadherin. Slug is homologous to Snail and was the first
transcriptional repressor of E-cadherin to be described along with
other non-Snail transcriptional repressors of E-cadherin [such as,
E47, delta-crystallin/E2-box factor 1/zinc finger E-Box-binding
homeobox (Zeb) 1 and Smad-interacting protein 1/Zeb2] (15–18).
These repressors are tightly regulated at the transcriptional level
and/or by subcellular localization. Insights into the underlying
mechanisms of EMT regulation may provide novel chemotherapeutic and
antifibrotic therapies.
Tumor progression and invasion are complex
biological processes that involve the remodeling of stromal tissue
by invading cells (12,19–21).
Slug and Snail are members of an evolutionarily conserved family of
zinc-finger transcription factors. Slug and Snail are expressed in
the intermediate mesoderm and the metanephric mesenchyme during
renal development, and are downregulated prior to epithelial
differentiation (22,23). The kidneys developed normally in
mice with a loss-of-function mutation in Snail, which suggests the
functional redundancy of Snail and Slug (24,25).
However, the E-cadherin repressors, Snail, Slug and E47, produce
different genetic profiles when overexpressed in ovarian tumor
cells, suggesting differential regulation of these transcription
factors (8,26–28).
In the present study, the constitutive expression of Slug increased
invasion by inducing EMT and the results obtained following gene
knockdown were consistent with those of a previous study (21). Our findings suggest that Slug
induces EMT, increases the percentage of CD133+ cells,
promotes the clonigenicity of HCC cells and induces a stronger
stemness. These changes activate dormant developmental pathways in
invading tumor cells; therefore, suppressing invasion-related
molecules, such as Slug, may present an important mechanism to
suppress metastasis. Furthermore, increased Snail levels did not
significantly affect HCC cell proliferation, migration, invasion,
clonigenicity or the number of CD133+ cells. Thus, Snail
proteins may have a polarizing effect on HCC tissue growth.
In conclusion, our findings demonstrated that Slug
upregulation increased the number of CD133+ cells, which
is important for EMT and proliferation of ovarian cancer cells.
Therefore, Slug may be a potential new target for preventing tumor
invasion and metastasis.
References
|
1
|
Yang MH, Chen CL, Chau GY, et al:
Comprehensive analysis of the independent effect of twist and snail
in promoting metastasis of hepatocellular carcinoma. Hepatology.
50:1464–1474. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Hanazaki K, Matsushita A, Nakagawa K,
Misawa R and Amano J: Risk factors of long-term survival and
recurrence after curative resection of hepatocellular carcinoma.
Hepatogastroenterology. 52:552–557. 2005.PubMed/NCBI
|
|
3
|
Yeoman AD, Al-Chalabi T, Karani JB, et al:
Evaluation of risk factors in the development of hepatocellular
carcinoma in autoimmune hepatitis: Implications for follow-up and
screening. Hepatology. 48:863–870. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Hanazaki K, Matsushita A, Nakagawa K,
Misawa R and Amano J: Risk factors of intrahepatic recurrence after
curative resection of hepatocellular carcinoma.
Hepatogastroenterology. 52:580–586. 2005.PubMed/NCBI
|
|
5
|
Savagner P: The epithelial-mesenchymal
transition (EMT) phenomenon. Ann Oncol. 21(Suppl 7): vii89–vii92.
2010. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Bacac M and Stamenkovic I: Metastatic
cancer cell. Annu Rev Pathol. 3:221–247. 2008. View Article : Google Scholar
|
|
7
|
Kang Y and Massagué J:
Epithelial-mesenchymal transitions: twist in development and
metastasis. Cell. 118:277–279. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Smit MA, Geiger TR, Song JY, Gitelman I
and Peeper DS: A Twist-Snail axis critical for TrkB-induced
epithelial-mesenchymal transition-like transformation, anoikis
resistance, and metastasis. Mol Cell Biol. 29:3722–3737. 2009.
View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Hotz B, Arndt M, Dullat S, Bhargava S,
Buhr HJ and Hotz HG: Epithelial to mesenchymal transition:
expression of the regulators snail, slug, and twist in pancreatic
cancer. Clin Cancer Res. 13:4769–4776. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Šošić D, Richardson JA, Yu K, Ornitz DM
and Olson EN: Twist regulates cytokine gene expression through a
negative feedback loop that represses NF-kappaB activity. Cell.
112:169–180. 2003.PubMed/NCBI
|
|
11
|
McCawley LJ and Matrisian LM: Tumor
progression: defining the soil round the tumor seed. Curr Biol.
11:R25–R27. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Sun T, Zhao N, Zhao XL, et al: Expression
and functional significance of Twist1 in hepatocellular carcinoma:
its role in vasculogenic mimicry. Hepatology. 51:545–556. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Zhou C, Liu J, Tang Y and Liang X:
Inflammation linking EMT and cancer stem cells. Oral Oncol.
48:1068–1075. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Childs G and Segall JE: Twists and turns
of invasion. Nat Cell Biol. 14:337–339. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Firulli AB and Conway SJ:
Phosphoregulation of Twist1 provides a mechanism of cell fate
control. Curr Med Chem. 15:2641–2647. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Yang MH and Wu KJ: TWIST activation by
hypoxia inducible factor-1 (HIF-1): implications in metastasis and
development. Cell Cycle. 7:2090–2096. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Martin A and Cano A: Tumorigenesis: Twist1
links EMT to self-renewal. Nat Cell Biol. 12:924–925. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Pietras K and Ostman A: Hallmarks of
cancer: interactions with the tumor stroma. Exp Cell Res.
316:1324–1331. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Peng JY and Wang Y: Tumor stroma: A
determinant role in local recurrence of rectal cancer patients
receiving total mesorectal excision? Med Hypotheses. 75:442–444.
2010. View Article : Google Scholar
|
|
20
|
Erenpreisa J and Cragg MS: Cancer: a
matter of life cycle? Cell Biol Int. 31:1507–1510. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Sun T, Sun BC, Zhao XL, et al: Promotion
of tumor cell metastasis and vasculogenic mimicry by way of
transcription coactivation by Bcl-2 and Twist1: a study of
hepatocellular carcinoma. Hepatology. 54:1690–1706. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Timmerman LA, Grego-Bessa J, Raya A, et
al: Notch promotes epithelial-mesenchymal transition during cardiac
development and oncogenic transformation. Genes Dev. 18:99–115.
2004. View Article : Google Scholar
|
|
23
|
Alonso-Magdalena P, Brössner C, Reiner A,
et al: A role for epithelial-mesenchymal transition in the etiology
of benign prostatic hyperplasia. Proc Natl Acad Sci USA.
106:2859–2863. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Ikenouchi J, Matsuda M, Furuse M and
Tsukita S: Regulation of tight junctions during the
epithelium-mesenchyme transition: direct repression of the gene
expression of claudins/occludin by Snail. J Cell Sci.
116:1959–1967. 2003. View Article : Google Scholar
|
|
25
|
Carver EA, Jiang R, Lan Y, Oram KF and
Gridley T: The mouse snail gene encodes a key regulator of the
epithelial-mesenchymal transition. Mol Cell Biol. 21:8184–8188.
2001. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Smith JP, Pozzi A, Dhawan P, Singh AB and
Harris RC: Soluble HB-EGF induces epithelial-to-mesenchymal
transition in inner medullary collecting duct cells by upregulating
Snail-2. Am J Physiol Renal Physiol. 296:F957–F965. 2009.
View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Usami Y, Satake S, Nakayama F, et al:
Snail-associated epithelial-mesenchymal transition promotes
oesophageal squamous cell carcinoma motility and progression. J
Pathol. 215:330–339. 2008. View Article : Google Scholar
|
|
28
|
Medici D, Hay ED and Olsen BR: Snail and
Slug promote epithelial-mesenchymal transition through
beta-catenin-T-cell factor-4-dependent expression of transforming
growth factor-beta3. Mol Biol Cell. 19:4875–4887. 2008. View Article : Google Scholar : PubMed/NCBI
|