Introduction
Brain tumors account for 90% of all central nervous
system cancers in adults (1,2), and gliomas are the most common type of
malignant primary brain tumor. Glioma is a diffuse and
highly-invasive tumor, resulting in a dismal prognosis, and
long-term survivors are rare (3).
Gliomas are also characterized by high migratory and proliferative
abilities (4). However, the molecular
mechanism that triggers the development of gliomas is elusive and
requires additional investigation. Therefore, the treatment
strategies for glioma therapy and drugs for the treatment of the
tumors are also limited (5,6). Exploration of the specific mechanism of
the invasion and inhibition of glioma is required.
Previous studies have concluded that deregulation of
developmentally important genes contributes to malignant disease in
adults (7). For example, the
canonical Wnt/β-catenin pathway is required for normal embryonic
development, as well as controlled proliferation of adult stem
cells (8,9). However, mutations in the components of
the Wnt/β-catenin pathway have been associated with a number of
human cancers derived from breast, colorectal, ovarian and
neuroectodermal tissues (10,11), which is an important method for the
identification of novel therapeutic targets.
Sareddy et al (12) reported that the Wnt/β-catenin pathway
plays a major role in tumorigenesis and tumor progression in
N-ethyl-N-nitrosourea-induced rat gliomas. Pu et al
(13) demonstrated that the
downregulated expression of Wnt2 and β-catenin by small interfering
RNA (siRNA) suppressed malignant glioma cell growth. According to
the two aforementioned studies, it was hypothesized that the
Wnt/β-catenin pathway may participate in glioma progression and
development.
The Pygopus genes were initially identified as a
mechanism that controls Wnt/β-catenin signaling in Drosophila
melanogaster. There are two Pygopus genes that exist in
mammals, which code for the Pygopus 1 (Pygo1) and Pygopus 2 (Pygo2)
proteins (14,15). Previous studies have reported that
Pygo2 is essential for the growth of epithelial ovarian (16) and breast cancers (17). Therefore, the present study
investigated the role of Pygopus in glioma U251 cells, and may aid
in the identification of anti-glioma drugs for glioma therapy.
Materials and methods
Tissue samples
In the present study, 80 recently resected glioma
samples were obtained from patients that underwent primary surgery
at the Department of Neurosurgery, Pearl River Hospital, Nanfang
Medical University (Guangzhou, China). Additionally, five normal
brain tissue specimens, which were obtained from internal
decompression performed on patients with cerebral injury, or from
temporal lobe resection for the treatment of epilepsy, were also
included. All glioma samples were diagnosed by two independent
neuropathologists, and were paraffin-embedded and archived. The
patient protocol in the present study has been approved, and all
patients provided informed consent. The present study was approved
by the ethics committee of Jinan University (Jinan, China). All
specimens were collected according to approved protocols at Jilin
No. 1 Hospital. The paraffin-embedded tissue samples were cut into
4-µm thick sections for analysis.
Cell culture and transfection with
short hairpin (sh)RNA for Pygo2
Human glioblastoma U251 cells were purchased from
the Cell Center of the Institute of Biochemistry and Cell Biology,
Chinese Academy of Sciences (Shanghai, China). The U251 cells were
cultured in Dulbecco's modified Eagle's medium (DMEM; Gibco Life
Technologies, Carlsbad, CA, USA) supplemented with 10% fetal bovine
serum (Gibco Life Technologies) and 100 µg/ml streptomycin. The
cells were maintained at 37°C in a humidified atmosphere of 5%
CO2. The cells of the experimental group were treated
with Pygo2 shRNA (shRNA-Pygo2), the negative control group was
treated with scramble control shRNA (shRNA-Scr), and the control
group was treated with Lipofectamine alone (Con). All transfections
were performed using Lipofectamine 2000 according to the
manufacturer's instructions.
Immunohistochemical (IHC)
analysis
IHC analysis was performed according to the
manufacturer's instructions for streptavidin-peroxidase (SP;
Beijing Zhongshan Golden Bridge Biotechnology Co., Ltd., Beijing,
China), and immunochemistry was performed using a rabbit anti-human
Pygo2 polyclonal antibody (sc-74878; Santa Cruz Biotechnology Co.,
Ltd., Dallas, TX, USA) at a dilution of 1:75. Pygo2-positive
tissues were defined by the presence of intracellular staining.
Pygo2-negative tissues were defined by the absence of specific
intracellular staining, as observed in the negative control. The
expression levels of Pygo2 and cyclin D1 in each specimen were
scored between 0 and 4. The absence of staining or presence of weak
staining was scored as 0 and 1, respectively. Intensive staining of
25% of the tumor cells or moderately intensive staining of <80%
of the cells was scored as 2. Intensive staining of 25–50% of cells
or moderately intensive staining of >80% of the cells was scored
as 3, and intensive staining of >50% of the tumor cells was
scored as 4. The samples were independently evaluated under light
microscopy (CX31 microscope; Olympus Corporation, Tokyo, Japan) by
two pathologists without prior knowledge of the clinical data of
the patients.
Synthesis of shRNA
Two shRNA sequences were designed in the present
study. One shRNA sequence was designed using a specific pair of
oligonucleotides to yield shRNA-Pygo2, the Pygo2-targeting shRNA,
and the shRNA-Scr sequence was designed using the scramble control
sequences. shRNA-Pygo2 was designed and synthesized based on the
human Pygo2 cDNA sequences (GenBank accession no., NW_925683),
followed by insertion into the pSUPER vector (Oligoengine, Seattle,
WA, USA). The primers used to design the oligonucleotides are
listed in Table I. The sequence was
specific and no sequences were homologous with any relevant human
genes.
 | Table I.Primers for designing the
oligonucleotides. |
Table I.
Primers for designing the
oligonucleotides.
| Primers | Sequence |
|---|
| shRNA-Pygo2 |
|
|
Forward |
5′-GATCCCCTGTGAGGCCTCTTGTCAGAAATTCAAGAGATTTCTGACAAGAGGCCTCACATTTTTA-3′ |
|
Reverse |
5′-AGCTTAAAAATGTGAGGCCTCTTGTCAGAAATCTCTTGAATTTCTGACAAGAGGCCTCACAGGG-3′ |
| shRNA-Scr |
|
|
Forward |
5′-GATCCCCGTGGTTTCATCGCATCTGCTTCAAGAGAGCAGATGCGATGAAACCACTTTTTA-3′ |
|
Reverse |
5′-AGCTTAAAAAGTGGTTTCATCGCATCTGCTCTCTTGAAGCAGATGCGATGAAACCACGGG-3′ |
Reverse transcription-polymerase chain
reaction (RT-PCR)
The total RNA of the cultured U251 glioma cells was
isolated using the TRIzol reagent (Invitrogen, Groningen,
Netherlands). RT-PCR was performed according to the manufacturer's
instructions, and as previously described by Xu et al
(18). The primers for Pygo2 were as
follows: Forward, 5′-CCATTCTGTGTGAGGCCTCCT-3′; and reverse,
5′-CCATGCCCTCACGGATGT-3′. The primers produced a 165-bp fragment.
The primers for the internal control GAPDH were as follows:
Forward, 5′-ACCACAGTCCATGCCATCA-3′; and reverse,
5′-TCCACCACCCTGTTGCTGTA-3′. The GAPDH primers produced a 450-bp
fragment. The PCR reaction procedure was performed as previously
described by Chen et al (19).
Western blot analysis
The U251 cells were harvested at 48 h
post-transfection, and the total proteins were extracted with cell
lysis buffer containing protease inhibitor. The cell lysis buffer
was fixed as previously described by Xu et al (18). The western blot gels were then
incubated overnight at 4°C with mouse anti-human Pygo2 monoclonal
antibody (sc-98744; dilution, 1:1,000) or mouse anti-human cyclin
D1 (A12; dilution, 1:1,000) antibodies (Santa Cruz Biotechnology
Co., Ltd.). The specific western blot assay method was performed as
described by Wang et al (20).
β-actin (dilution, 1:2,000; Santa Cruz Biotechnology Co., Ltd.) was
employed as the internal control to quantify the protein expression
of the aforementioned proteins. MTT assays were performed by adding
MTT (Sigma-Aldrich, St. Louis, MO, USA) to the culture medium at a
final concentration of 5 mg/ml at 37°C. The reaction was terminated
by removal of the supernatant and addition of 200 µl DMSO to
dissolve the formazan product. The plates were read at 540 nm on a
micro-ELISA plate reader (Thermo MK3; Thermo Labsystems, Santa
Rosa, CA, USA). Each assay was performed in at least four wells.
The procedure used in this assay was performed as previously
described by Wang et al (21).
Luciferase reporter assay
The U251 cells were transiently transfected using
the basic nucleofector kit (Amaxa Biosystems GmbH, Cologne,
Germany), according to the manufacturer's instructions. Briefly,
the cells were trypsinized, counted using a hemocytometer
(15170–172, VWR International LLC, Radnor, USA) and centrifuged at
4,00 × g. Subsequently, ~5×106 cells were
resuspended in 100 µl of Basic Nucleofector Solution (Amaxa
Biosystems GmbH). The cells were transfected with 1 µg of reporter
plasmid, for both the transcription factor reporter plasmid
(pTOPflash) and the mutant control plasmid (pFOPflash), and 1 µg of
the internal control pRL-TK, with or without 4 µg of dominant
negative (DN)-TCF plasmids (program A33 onthe Nucleofector).
Subsequent to transfection, ~1×104 cells were
transferred to each well of a 24-well plate containing fresh,
pre-warmed DMEM and maintained for 48 h at 37°C in a 5%
CO2 atmosphere. The cells were then lysed in lysis
buffer, and 20 µl of each lysate was monitored for luciferase
activity using the Dual-Luciferase Reporter Assay System (Promega,
Madison, WI, USA). Light units were measured using Lmax II 384
(Molecular Devices LLC, Sunnyvale, CA, USA). The control reporter
pRL-TK contains a herpes simplex virus thymidine kinase promoter
driving a Renilla luciferase gene, and Renilla
luciferase activity was used to normalize the results for
transfection efficiency. The reporter activities were expressed as
the ratio of the luciferase activity of TOPflash to the luciferase
activity of FOPflash from experiments performed in triplicate.
Vascular mimicry assay
The formation of vascular mimicry in U251 cells was
tested using a reduced growth factor Matrigel matrix (BD
Biosciences, Franklin Lakes, NJ, USA). Eight-chamber slides were
pre-coated with 0.1 ml Matrigel per well and were incubated at 37°C
for 30 min. Floating spheres of glioma stem-like cells (GSLCs) were
resuspended in the endothelial cell basal medium-2 (Lonza, Basal,
Switzerland) containing 2% FCS, with or without 10 ng/ml VEGF
(PeproTech, Rocky Hill, NJ, USA). After 4 days, the tubules formed
by GSLCs were counted using light microscopy. The formation of
vascular mimicry was analyzed using Image Image Lab software,
version 4.1 (Bio-Rad, CA, USA).
The specific definition for vascular mimicry was as
follows: One closed and network-like lumen under the microscope
represented one typical vascular mimicry; one opened ring structure
represented an atypical vascular mimicry; and an unordered
structure demonstrated no vascular mimicry. The number of vascular
mimicries on 100 random fields was calculated.
Analysis of bromodeoxyuridine (BrdU)
incorporation
BrdU was added to the cells 48 h post-transfection
for 30 min, and the U251 cells were stained for BrdU using a BrdU
flow kit (BD Biosciences), according to the manufacturer's
instructions. The detailed processes were performed as described by
Wang et al (3).
Statistical analysis
Statistical analyses were performed using SPSS
software, version 19.0 (IBM, Armonk, NY, USA). The association
between the expression of the Pygo2 protein and the
clinicopathological characteristics of patients with glioma was
analyzed using Fisher's exact test. All data were expressed as the
mean ± standard error. Differences between groups were analyzed for
statistical significance using one-way analysis of variance.
P<0.05 was considered to indicate a statistically significant
difference.
Results
Pygo2 protein expression was enhanced
in human primary glioma samples
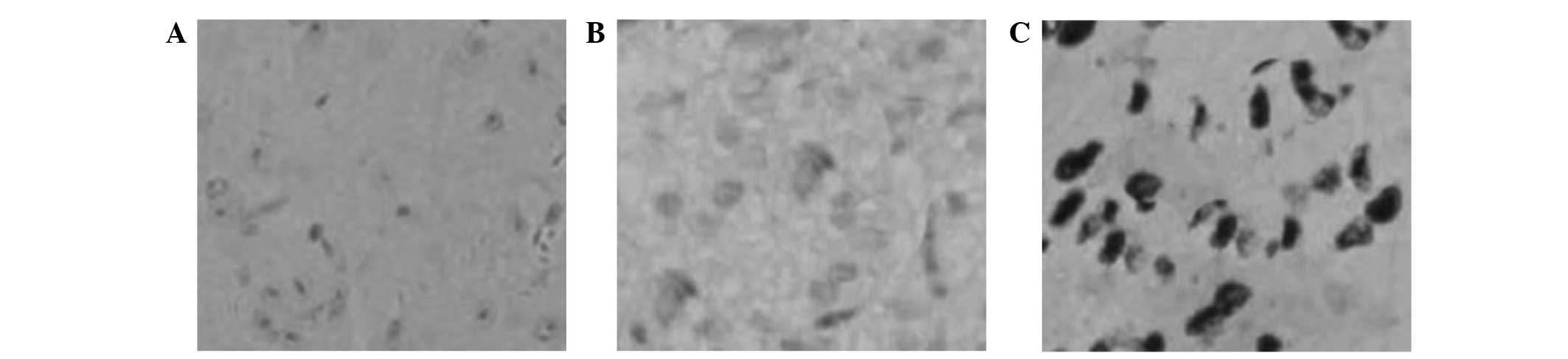
Immunochemistry was utilized to examine the Pygo2
protein in human brain glioma tissue. The results indicated that
the Pygo2 protein was not expressed in the normal brain tissue
samples (Fig. 1A). However, low
expression of the Pygo2 protein was observed in the grade II glioma
sections (Fig. 1B), and high
expression of the Pygo2 protein was observed in the grade IV glioma
tissues (Fig. 1C).
Expression of the Pygo2 protein was detected in 52
out of 80 glioma samples, and no Pygo2 expression was detected in
the normal brain tissues (Table II).
There were no significant differences between the incidence of
Pygo2 protein expression in males and females, and no significant
age association was observed (Table
II).
 | Table II.Association between the expression of
Pygo2 and the clinicopathological characteristics of brain glioma
patients. |
Table II.
Association between the expression of
Pygo2 and the clinicopathological characteristics of brain glioma
patients.
| Clinicopathological
characteristic | Total, n | Pygo2-positive, n
(%) | P-value |
|---|
| Gender |
|
|
|
| Male | 52 | 67.3 |
0.626 |
|
Female | 28 | 60.7 |
|
| Age, years |
|
|
0.121 |
|
<44 | 37 | 19 (51.4) |
|
| ≥44 | 43 | 33 (76.7) |
|
| Brain tissue
type |
|
|
|
|
Glioma | 80 | 52 (65.0) | <0.001 |
|
Normal | 5 | 0 (0.0) |
|
Specific shRNA inhibits Pygo2 protein
expression
In the present study, shRNA was revealed to target
the Pygo2 gene and downregulate the expression of the Pygo2
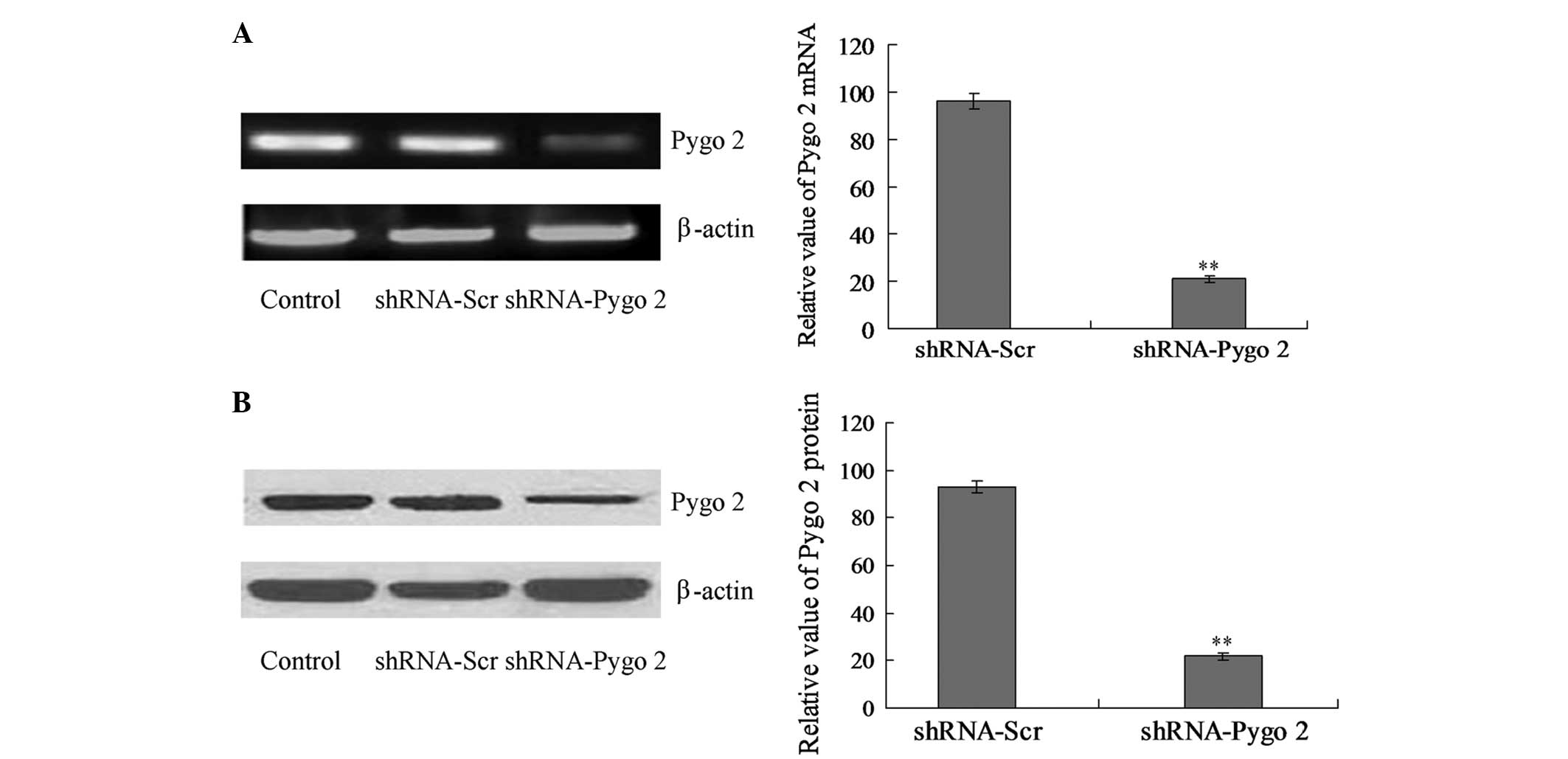
protein. The RT-PCR results revealed that shRNA-Scr did not affect
the transcription of Pygo2 mRNA (Fig.
2A). However, shRNA-Pygo2 significantly downregulated Pygo2
mRNA expression compared with the shRNA-Scr group (Fig. 2A; P<0.01). In addition, the
expression of the Pygo2 protein was also detected in the present
study. According to the results of the western blot analysis, the
Pygo2 protein levels in the cells treated with shRNA-Pygo2 were
significantly reduced compared with the shRNA-Scr treated cells
(Fig. 2B; P<0.01). The RT-PCR and
western blot analysis each illustrated that shRNA-Pygo2 was able to
efficiently and specifically inhibit the expression of Pygo2 in
U251 cells (Fig. 3B).
Transfection with shRNA-Pygo2 affects
the proliferation of U251 cells
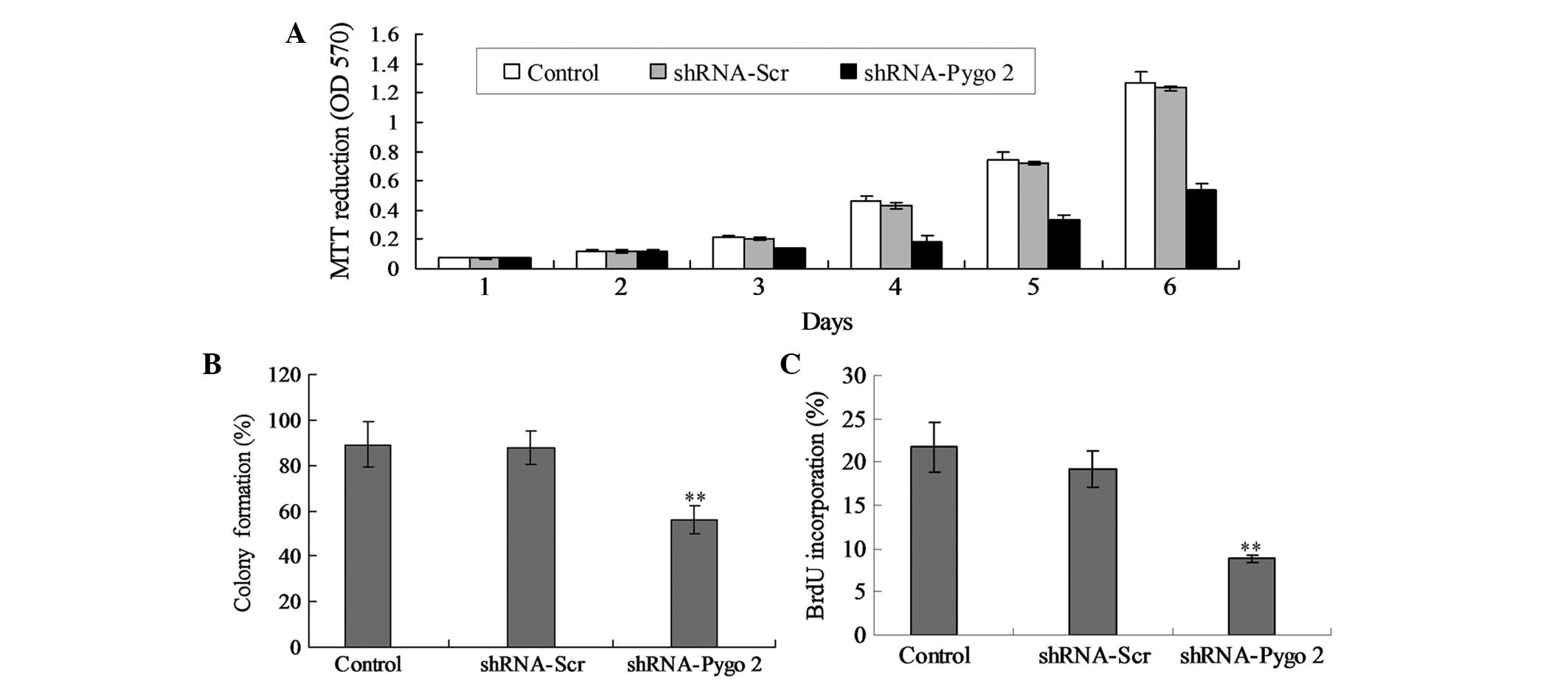
In order to identify whether shRNA-Pygo2 affected
the growth of glioma cells, MTT and plate colony formation assays
were employed to detect the proliferation and activity of the U251
cells. Transfection of cells with shRNA-Pygo2 significantly reduced
the cell proliferation compared with the shRNA-Scr treated cells
(Fig. 3A; P<0.05). In addition,
there were no significant differences between the control cells and
the cells treated with shRNA-Scr.
To determine the anti-proliferative effects of
shRNA-Pygo2, the ability of shRNA-Pygo2 to suppress colony
formation was examined. The results indicated that shRNA-Pygo2
induced significant inhibition of colony formation in U251
cells.
In addition, the BrdU method was also used to
confirm the effects of shRNA-Pygo2 on the U251 cell proliferation.
As indicated in Fig. 3C, treatment of
shRNA-Pygo2 may significantly decrease levels of BrdU
incorporation, which indicated that shRNA-Pygo2 may inhibit DNA
synthesis of the tumor cells.
shRNA-Pygo2 transfection inhibits
vascular mimicry
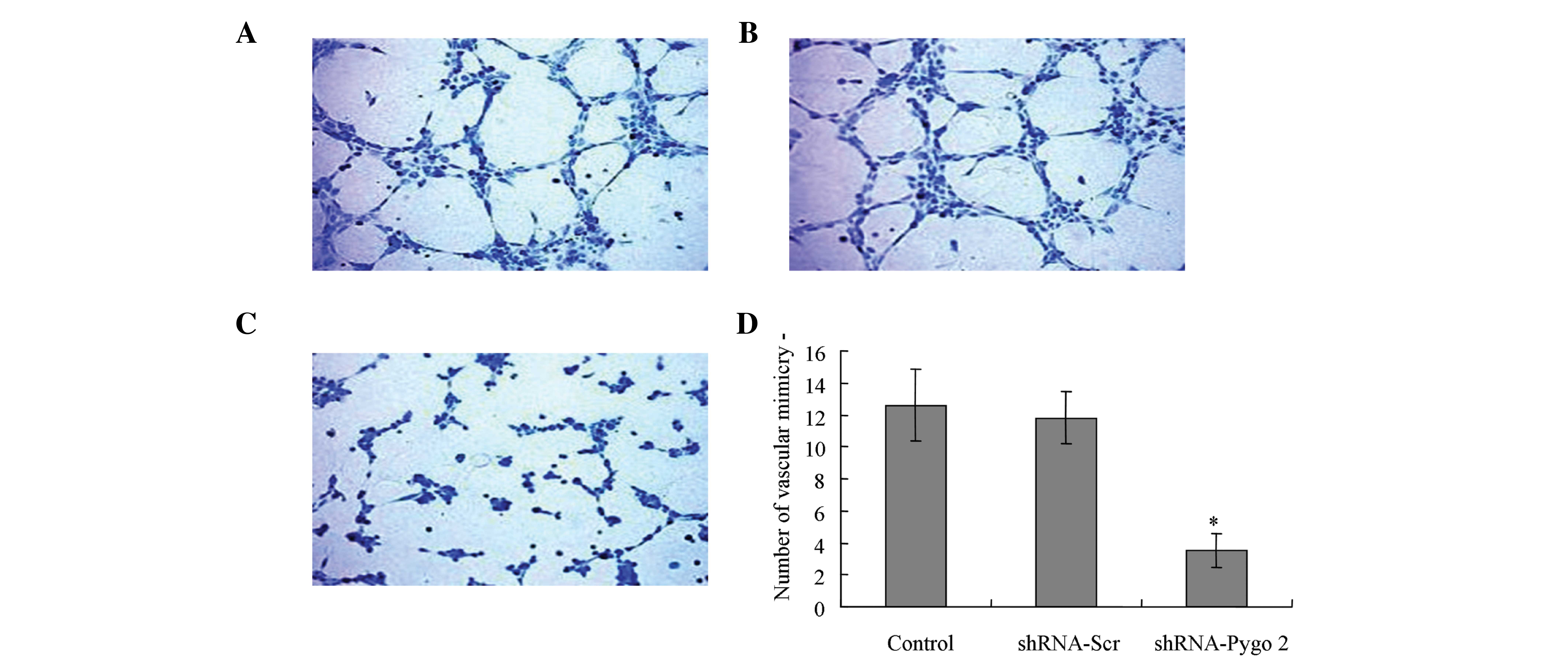
The findings of the vascular mimicry assay indicated
that there was no significant difference in vascular mimicry
between the control group and group transfected with shRNA-Scr
(Fig. 4A and B). However, shRNA-Pygo2
successfully disturbed the expression of Pygo2 and significantly
decreased the vascular-mimicry compared with the U251 cells
transfected with shRNA-Scr (P<0.01; Fig. 4C).
shRNA-Pygo2 downregulates the Wnt
signaling pathway
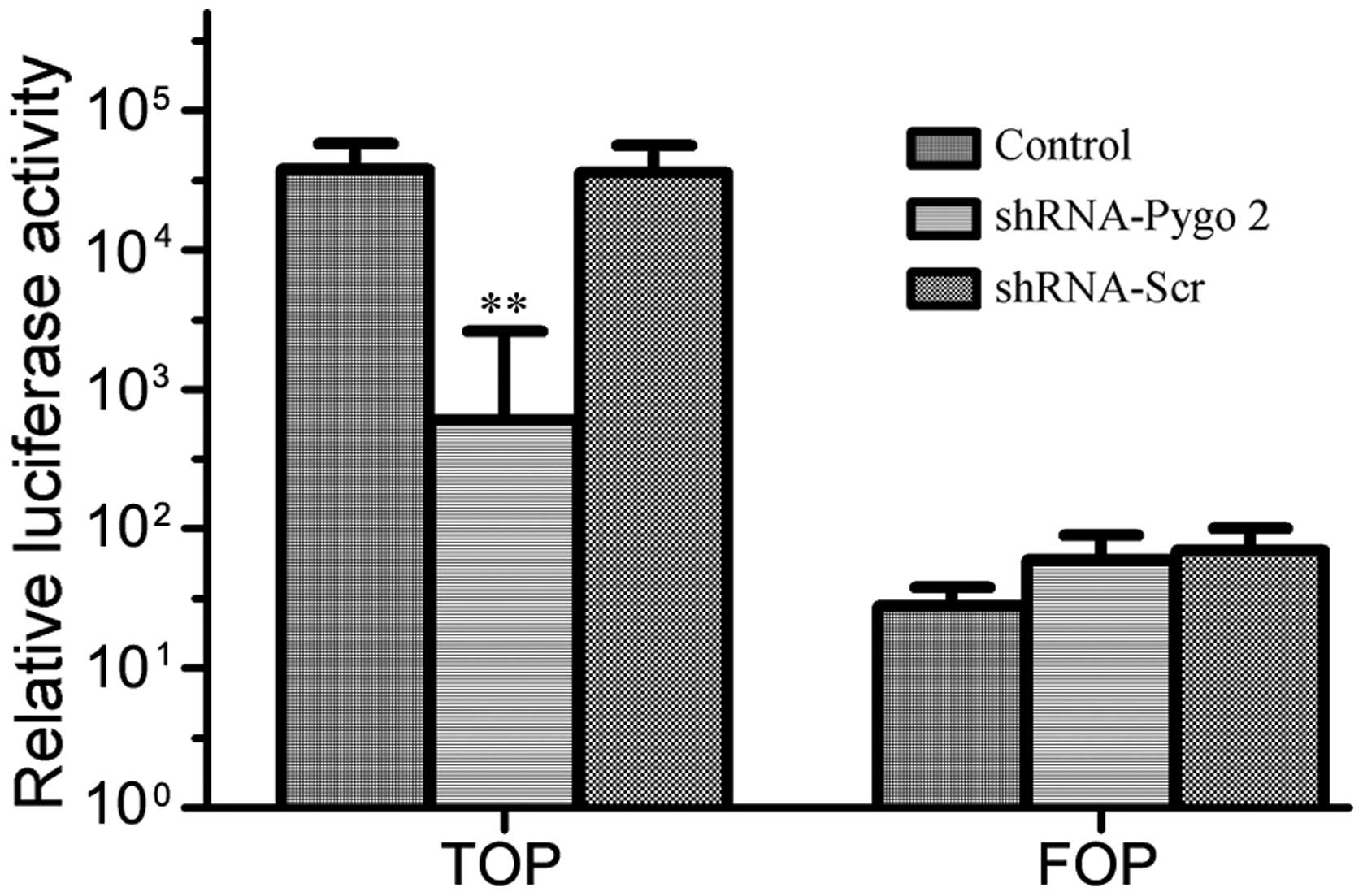
To obtain further evidence that the Wnt pathway is
involved in the vascular mimicry of U251 cells, a transcription
factor (TCF) reporter gene containing five optimal TCF-binding
sites, TOPflash, or the mutant control plasmid FOPflash was
transfected into the U251 cells. The results indicated that the
reporter activity of the Wnt pathway in the group treated with
shRNA-Pygo2 was significantly reduced compared with the group
treated with shRNA-Scr and the control group (P<0.05; Fig. 5).
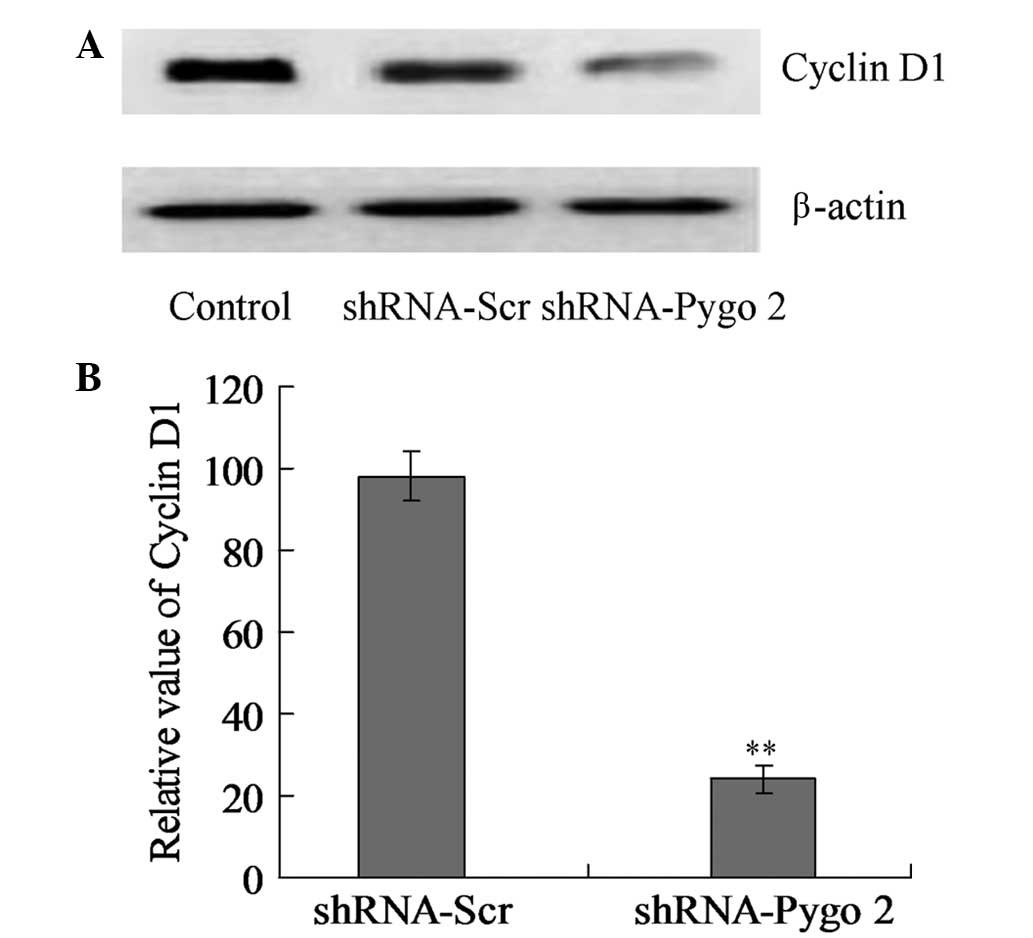
shRNA-Pygo2 triggers the
downregulation of cyclin D1 expression
To identify the specific pathway through which the
shRNA-Pygo2 inhibited cell proliferation, the Wnt/β-catenin pathway
component cyclin D1 was detected by western blot analysis. The
cyclin D1 protein levels were significantly decreased in the cells
treated with shRNA-Pygo2 compared with the cells treated with
shRNA-Scr and the control group cells (Fig. 6; P<0.01).
Discussion
Malignant glioma is known to be a highly invasive
tumor of the central nervous system that is challenging to treat.
Although therapeutic methods have been investigated for numerous
years (22–24), only a small amount of progress has
been achieved in patients with malignant gliomas. Therefore, the
identification of novel therapeutic targets or methods of treatment
is an extremely urgent and promising undertaking for the clinical
treatment of gliomas.
Numerous studies have reported that the Pygopus
protein plays an important role in developmental brain patterning
(25,26). In the present study, the association
between the expression of the Pygo2 protein and the clinical grade
of brain glioma was investigated. The present results indicated
that the expression level of Pygo2 increased with the clinical
grade. Therefore, it was confirmed that the tumor grade exhibited a
positive association with Pygo2 expression in brain glioma. The
present results were consistent with previous studies that
indicated that Pygo2 protein expression was enhanced in breast
(17) and epithelial ovarian cancers
(16). However, the two
aforementioned studies indicated that Pygo2 is necessary for the
proliferation of breast and epithelial ovarian cancer cells, and
the specific function of Pygo2 in the progress of gliomas also
remains elusive. Prior to the present study, it was hypothesized
that the role of Pygo2 in the development of gliomas may be
associated with certain signaling pathways.
Tumors may result from the interruption of the
developmental signaling pathways that regulate embryonic
development in form and structure (27). The Wnt/β-catenin signaling pathway is
a major developmental pathway that regulates central nervous system
development during embryogenesis, as well as during adulthood
(28). The Wnt/β-catenin signaling
pathway also plays an important role in carcinogenesis (29). However, the Wnt/β-catenin signaling
pathway-associated cascade genes in gliomas has not been
investigated in previous studies. Furthermore, the association
between the downstream genes in the Wnt/β-catenin signaling pathway
and the malignant progression of glioma is also poorly studied.
Thus, in the present study, specific shRNA against Pygo2 mRNA was
established and termed shRNA-Pygo2. The shRNA-Pygo2 was then
transfected into the U251 cells. The results indicated that Pygo2
knockdown inhibited cell proliferation, colony formation and BrdU
incorporation.
In the present study, the vascular mimicry assay
revealed that shRNA-Pygo2 significantly decreased the vascular
mimicry in U251 cells compared to treatment with shRNA-Scr. The
formation of vascular mimicry may be due to the blocking expression
of Pygo2. The decreased vascular mimicry also triggers the
inhibition of the differentiation and proliferation of U251 cells
in a later step.
The present study hypothesized that the vascular
mimicry induced by shRNA-Pygo2 may involve the Wnt signaling
pathway, and therefore the vascular mimicry assay was performed.
The results indicated that the Wnt pathway participated in the
development of vascular mimicry and cell proliferation induced by
shRNA-Pygo2.
Previous studies have also transfected the siRNA
sequences specific to Pygo protein mRNA, and inhibited the
TCF/LEF-mediated transcriptional activation of reporter genes
(30,31), which suggested that Pygo2 members may
be involved in TCF/β-catenin-driven transcription. In the present
study, the expression of cyclin D1 protein in the U251 cells
treated with shRNA-Pygo2 was investigated. The results revealed
that the expression of cyclin D1 was significantly reduced by Pygo2
shRNA, which was consistent with the decreased expression of cyclin
D1 in breast cancer cells following Pygo2 antisense RNA treatment
(17).
In conclusion, Pygo2 expression is associated with
the clinical grade of glioma malignancy. shRNA-Pygo2 effectively
suppresses the proliferation of glioma cells by inhibiting the
cyclin D1 expression of the canonical Wnt/β-catenin pathway for
brain glioma.
Acknowledgements
The present study was supported by the Medical
Scientific Research Projects of Guangdong Provincial Health Bureau
(grant no., A2012330) and the Scientific Cultivation and Innovation
Fund of Jinan University.
References
|
1
|
Zhang X, Zhang W, Cao WD, Cheng G and
Zhang YQ: Glioblastoma multiforme: Molecular characterization and
current treatment strategy (Review). Exp Ther Med. 3:9–14.
2012.PubMed/NCBI
|
|
2
|
An L, Liu Y, Wu A and Guan Y: microRNA-124
inhibits migration and invasion by down-regulating ROCK1 in glioma.
PLoS One. 8:e694782013. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Wang ZX, Chen YY, Li BA, Tan GW, Liu XY,
Shen SH, Zhu HW and Wang HD: Decreased pygopus 2 expression
suppresses glioblastoma U251 cell growth. J Neurooncol. 100:31–41.
2010. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Tu Y, Gao X, Li G, Fu H, Cui D, Liu H, Jin
W and Zhang Y: MicroRNA-218 inhibits glioma invasion, migration,
proliferation, and cancer stem-like cell self-renewal by targeting
the polycomb group gene Bmi1. Cancer Res. 73:6046–6055. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Ohgaki H and Kleihues P: Population-based
studies on incidence, survival rates, and genetic alterations in
astrocytic and oligodendroglial gliomas. J Neuropathol Exp Neurol.
64:479–489. 2005.PubMed/NCBI
|
|
6
|
Chen Z, Li J, Song X, Wang Z and Yue W:
Use of novel sonosensitizer in sonodynamic therapy of U251 glioma
cells in vitro. Exp Ther Med. 3:273–278. 2012.PubMed/NCBI
|
|
7
|
Reya T and Clevers H: Wnt signaling in
stem cells and cancer. Nature. 434:843–850. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Bienz M: Beta-catenin: A pivot between
cell adhersion and Wnt signaling. Curr Biol. 15:R64–R67. 2005.
View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Pinto D and Clevers H: Wnt, stem cells and
cancer in the intestine. Biol Cell. 97:185–196. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Lustig B and Behrens J: The Wnt signaling
pathway and its role in tumor development. J Cancer Res Clin Oncol.
129:199–221. 2003.PubMed/NCBI
|
|
11
|
Brown AM: Wnt signaling in breast cancer:
have we come full circle? Breast Cancer Res. 3:351–355. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Sareddy GR, Challa S, Panigrahi M and Babu
PP: Wnt/beta-catenin/Tcf signaling pathway activation in malignant
progression of rat gliomas induced by transplacental
N-ethyl-N-nitrosourea exposure. Neurochem Res. 34:1278–1288. 2009.
View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Pu P, Zhang Z, Kang C, Jiang R, Jia Z,
Wang G and Jiang H: Downregulation of Wnt 2 and beta-catenin by
siRNA suppresses malignant glioma cell growth. Cancer Gene Ther.
16:351–361. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Belenkaya TY, Han C, Standley HJ and Lin
X, Houston DW, Heasman J and Lin X: Pygopus endodes a nuclear
protein essential for wingless/Wnt signaling. Development.
129:4089–4101. 2002.PubMed/NCBI
|
|
15
|
Kramps T, Peter O, Brunner E, Nellen D,
Froesch B, Chatterjee S, Murone M, Züllig S and Basler K:
Wnt/wingless signaling requires BCL9/legless-mediated recruitment
of pygopus to the nuclear beta-catenin-TCF complex. Cell.
109:47–60. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Popadiuk CM, Xiong J, Wells MG, Andrews
PG, Dankwa K, Hirasawa K, Lake BB and Kao KR: Antisense suppression
of pygopus 2 results in growth arrest of epithelial ovarian cancer.
Clin Cancer Res. 12:2216–2223. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Andrews PG, Lake BB, Popadiuk C and Kao
KR: Requirement of Pygopus 2 in breast cancer. Int J Oncol.
30:357–363. 2007.PubMed/NCBI
|
|
18
|
Xu K, Liu XN, Zhang HB, An N, Wang Y,
Zhang ZC and Wang YN: Replication-defective HSV-1 effectively
targets trigeminal ganglion and inhibits viral pathopoiesis by
mediating interferon gamma expression in SH-SY5Y cells. J Mol
Neurosci. 53:78–86. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Chen YY, Li BA, Wang HD, Liu XY, Tan GW,
Ma YH, Shen SH, Zhu HW and Wang ZX: The role of Pygopus 2 in rat
glioma cell growth. Med Oncol. 28:631–640. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Wang X, Shi Q, Xu K, Gao C, Chen C, Li XL,
Wang GR, Tian C, Han J and Dong XP: Familial CJD associated PrP
mutants within transmembrane region induced Ctm-PrP retention in ER
and triggered apoptosis by ER stress in SH-SY5Y cells. PLoS One.
6:e146022011. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Wang X, Dong CF, Shi Q, Shi S, Wang GR,
Lei YJ, Xu K, An R, Chen JM, Jiang HY, Tian C, Gao C, et al:
Cytosolic prion protein induces apoptosis in human neuronal cell
SH-SY5Y via mitochondrial disruption pathway. BMB Rep. 42:444–449.
2009. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Peng B, Li D, Qin M, Luo D, Zhang X, Zhao
H and Hu S: MicroRNA218 inhibits glioma migration and invasion via
inhibiting glioma-associated oncogene homolog 1 expression at N
terminus. Tumor Biol. 35:3831–3837. 2014. View Article : Google Scholar
|
|
23
|
Jha MK and Suk K: Pyruvate dehydrogenase
kinase as a potential therapeutic target for malignant gliomas.
Brain Tumor Res Treat. 1:57–63. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Young JS, Kim JW, Ahmed AU and Lesniak MS:
Therapeutic cell carriers: a potential road to cure glioma. Expert
Rev Neurotehr. 14:651–660. 2014. View Article : Google Scholar
|
|
25
|
Lake BB and Kao KR: Pygopus is required
for embryonic brain patterning in Xenopus. Dev Biol. 261:132–148.
2003. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Li B, Rhéaume C, Teng A, Bilanchone V,
Munguia JE, Hu M, Jessen S, Piccolo S, Waterman ML and Dai X:
Developmental phenotypes and reduced Wnt signaling in mice
deficient for pygopus 2. Genesis. 45:318–325. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Morin PJ: Beta-catenin signaling and
cancer. Bioessays. 21:1021–1030. 1999. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Fogarty MP, Kessler JD and Wechsler-Reya
RJ: Morphing into cancer: The role of developmental signaling
pathways in brain tumor formation. J Neurobiol. 64:458–475. 2005.
View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Polakis P: Wnt signaling and cancer. Genes
Dev. 14:1837–1851. 2000.PubMed/NCBI
|
|
30
|
Thompson B, Townsley F, Rosin-Arbesfeld R,
Musisi H and Bienz M: A new nuclear component of the Wnt signaling
pathway. Nat Cell Biol. 4:367–373. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Shao H, Ma J, Guo T and Hu R: Triptolide
induces apoptosis of breast cancer cells via a mechanism associated
with the Wnt/β-catenin signaling pathway. Exp Ther Med. 8:505–508.
2014.PubMed/NCBI
|