Introduction
Hepatoblastoma (HB) is the most common type of
malignant liver tumor in infants and children. Although it accounts
for just 0.5–1.5 cases per million children per year, the mortality
rate is 35–50% in high-risk patients (1). Previous studies have suggested an
association with familial adenomatous polyposis (FAP) (2), both low and high birth weights (3), and constitutional trisomy 18 (4); however its etiology remains unknown.
Currently, alphafetoprotein levels, histological
analysis, tumor resectability and tumor metastasis are the only
prognosis factors for HB. Therefore, novel targets for early
detection and improved therapies and prognosis are required.
Epigenetic reprogramming has a crucial role in
tumorigenesis; one of the key mechanisms is DNA methylation. DNA
methylation, maintained by DNA methyltransferases, refers to the
addition of a methyl group to the carbon 5 of the cytosine ring in
CpG dinucleotides of gene promoter regions (5). DNA methylation is the only genetically
programmed DNA modification process in mammals that is involved in
the regulation of various biological processes, including gene
transcription, X-chromosome inactivation, genomic imprinting and
chromatin modification (6–8). It is generally recognized that DNA
methylation is a form of gene closure; the higher the level of
methylation, the more silent a gene is. Therefore, the
hypermethylation of suppressor genes or hypomethylation of
oncogenes may contribute to tumorigenesis. This hypothesis has been
investigated and supported in various studies on tumors (9), including HB (10).
Kallikrein-related peptidase 4 (KLK4), which is a
member of the KLK family, is considered to be an oncogene
associated with various types of cancer, including colorectal
adenocarcinoma and prostate cancer (11,12). In a
pilot study, we detected aberrant expression of KLK4 in HB and
normal tissues, and theorized if alterations in methylation status
affect its expression. Therefore, the present study aimed to
investigate KLK4 methylation in 10 HB samples in order to profile
genes that are differentially methylated in this disease.
Materials and methods
Patients and sample collection
The present study utilized 10 HB and adjacent
non-tumor tissues from patients who underwent partial hepatectomy
at the Children's Hospital of Fudan University (Shanghai, China).
The patients ranged in age from 7 to 30 months. Informed consent
was obtained retrospectively from clinical files and the diagnoses
were confirmed by the Department of Pathology following the
presence of >80% viable tumor cells (13). Clinical and pathological data are
listed in Table I. The Ethics
Committee of the Children's Hospital of Fudan University approved
the use of these human samples. Genomic DNA was extracted according
to standard procedures from 10 matched HB tumors and non-tumor
tissue sections that were stored at −80°C immediately after
surgery.
 | Table I.Clinical and pathological
characteristics of the study subjects. |
Table I.
Clinical and pathological
characteristics of the study subjects.
| Subject | Age
(months)a | Gender | Diagnosis type | Alpha-fetoprotein
(ng/ml) |
|---|
| 1 | 7 | Male | Mixed embryonal/fetal
subtype | 68490 |
| 2 | 23 | Male | Mixed embryonal/fetal
subtype | >121,000 |
| 3 | 11 | Female | Mixed embryonal/fetal
subtype | >121,000 |
| 4 | 10 | Female | Mixed embryonal/fetal
subtype | >121,000 |
| 5 | 20 | Female | Mixed embryonal/fetal
subtype | >121,000 |
| 6 | 7 | Male | Mixed embryonal/fetal
subtype | >121,000 |
| 7 | 30 | Male | Epithelial type | >121,000 |
| 8 | 14 | Male | Epithelial type | >121,000 |
| 9 | 19 | Male | Epithelial type | >121,000 |
| 10 | 7 | Female | Epithelial type | >121,000 |
DNA/RNA extraction and reverse
transcription quantitative-polymerase chain reaction (RT-qPCR)
DNA was extracted from 10 HB primary tumors and
adjacent non-tumor tissues. DNA samples were stored at −80°C until
subsequent use for mass spectrometry analysis. Total RNA was
extracted from 10 HB tumor and non-tumor pairs using TRIzol reagent
(Invitrogen; Thermo Fisher Scientific, Inc., Waltham, MA, USA),
and, reverse-transcribed into cDNA using a PrimeScript RT reagent
kit (Perfect Real Time) with gDNA Eraser (Takara Biotechnology Co.,
Ltd., Dalian, China) following the manufacturer's protocol. This
was then subjected to qPCR analysis to assess the levels of KLK4
mRNA expression using a SYBRGreen PCR kit (Takara Biotechnology
Co., Ltd.). The KLK4 primers used for qPCR were: Forward,
5′-GGACTCCTGCAACGGTGACTCT-3′ and reverse,
5′-TAGACACCTGGCACGCCAACTT-3′. The PCR cycling conditions were as
follows: 95°C for 30 sec, followed by 95°C for 5 sec and 60°C for
34 sec, for 45 cycles. For quantitative results, KLK4 expression
was represented as fold-change by the 2−ΔΔCq method and
statistically analyzed (14).
Primer design and PCR tagging for
EpiTYPER assay
Primers for the KLK4 gene were designed to cover the
regions with the most CpG sites (Fig.
1). Selected amplicons were 500 bp upstream of the promoter of
the KLK4 gene. Genomic DNA was bisulfite-treated, and its reverse
primer was supplemented with a T7-promoter tag for subsequent PCR
amplification.
Mass spectrometry
Following treatment with shrimp alkaline phosphatase
(SAP) in vitro transcription and uracil-specific cleavage,
the DNA samples were robotically dispensed onto silicon matrix
preloaded chips (SpectroCHIP; Sequenom, San Diego, CA, USA). Mass
spectra were collected using a MassARRAY Compact matrix-assisted
laser desorption/ionization time-of-flight system (Sequenom), and
the methylation ratios of the spectra were generated using EpiTYPER
software v1.0 (Sequenom).
Statistical methods
Statistical analyses and graphical depiction of data
were generated using GraphPad Prism 5.0 (GraphPad Software, Inc.,
La Jolla, CA, USA). Results are presented as the mean ± standard
error of the mean and were evaluated using a two-tailed Student's
t-test unless otherwise specified, in which instance unpaired
t-test or Pearson's correlation analysis were used. Statistical
analyses were also performed using SPSS 19.0 software (IBM SPSS,
Armonk, NY, USA) for Windows. P<0.05 was considered
statistically significant to indicate a statistically significant
difference.
Results
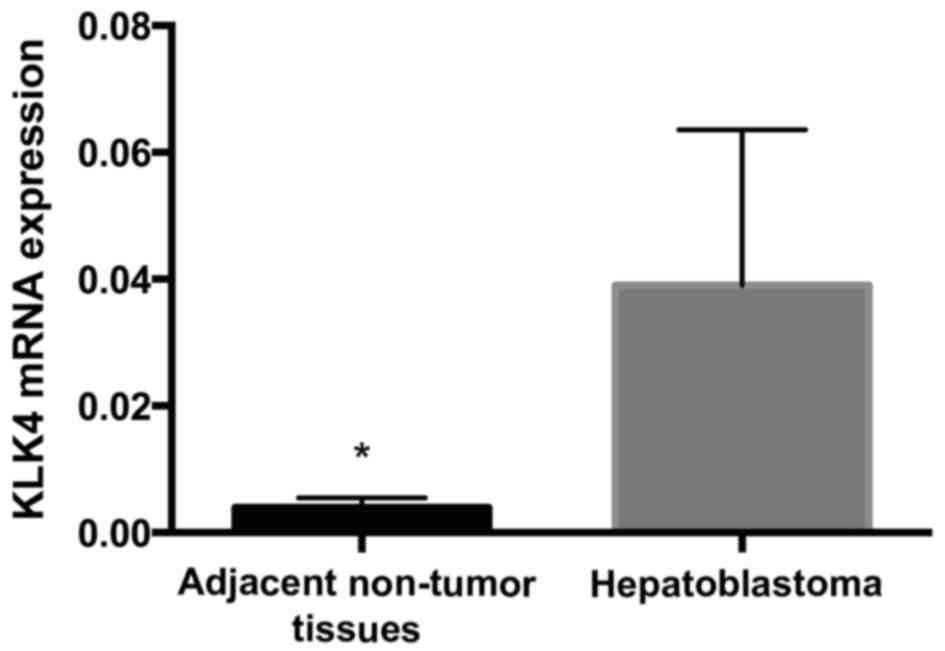
KLK4 mRNA expression levels are
elevated in HB
Expression levels of KLK4 mRNA were analyzed in the
10 pairs of tumor (n=10) and adjacent normal liver (n=10) tissues
using RT-qPCR. Expression levels of KLK4 in HB tissues were
significantly increased, as compared with matched non-tumor liver
tissues (0.039±0.0077 vs. 0.004±0.0005, respectively; P<0.01;
Fig. 2).
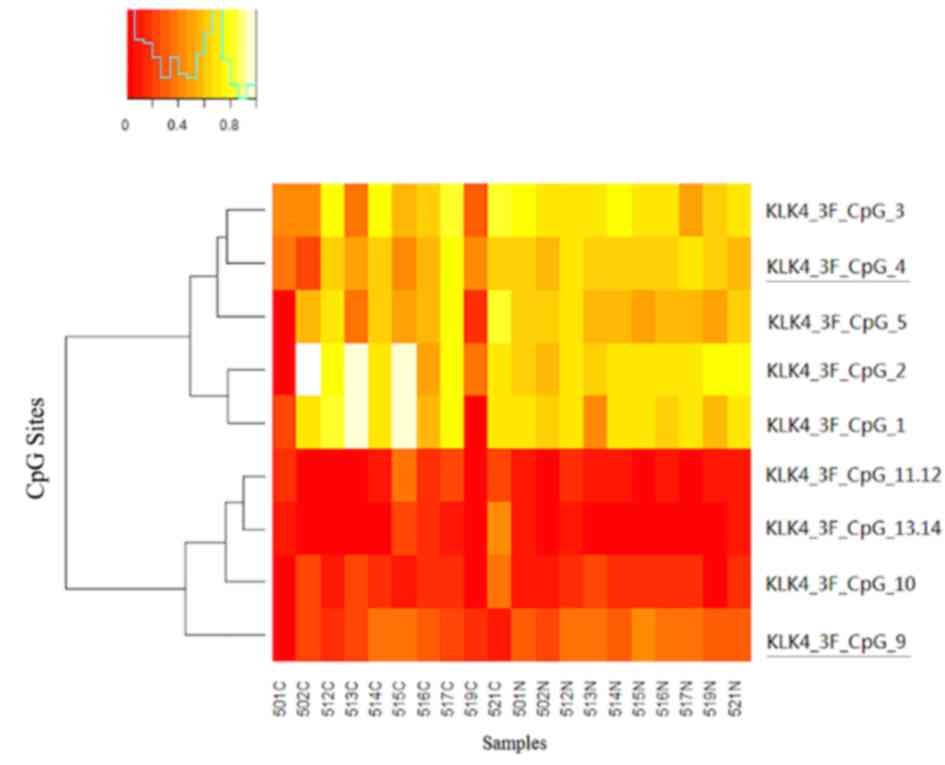
DNA methylation of KLK4 is reduced in
HB
Methylation patterns of KLK4 were verified in the 10
HB samples and match non-tumor controls. Enriched methylation
positions located at 500 bp upstream of the transcription
initiation site were analyzed; 11/22 sites in the amplicons were
amenable to analysis. Using two-way hierarchical cluster analysis,
two sites were found to be at a significantly lower degree of
methylation in the HB tissues, as compared with their non-tumor
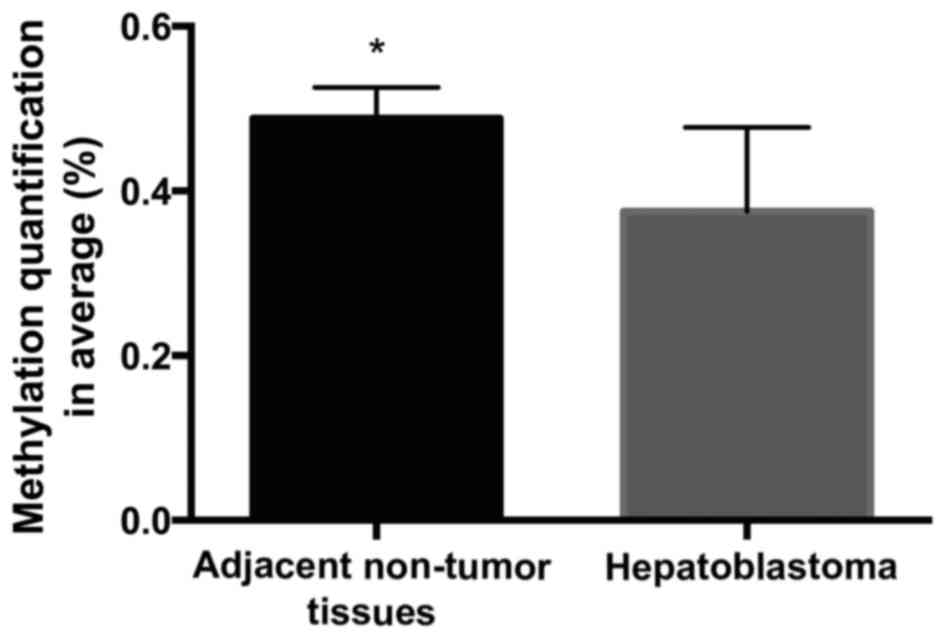
counterparts (Fig. 3). Furthermore,
these sites exhibited a significant difference in the mean
methylation levels of the KLK4 gene between HB and adjacent
non-tumor tissues (0.3755±0.03218 vs. 0.4885±0.01176, respectively;
P<0.01; Fig. 4).
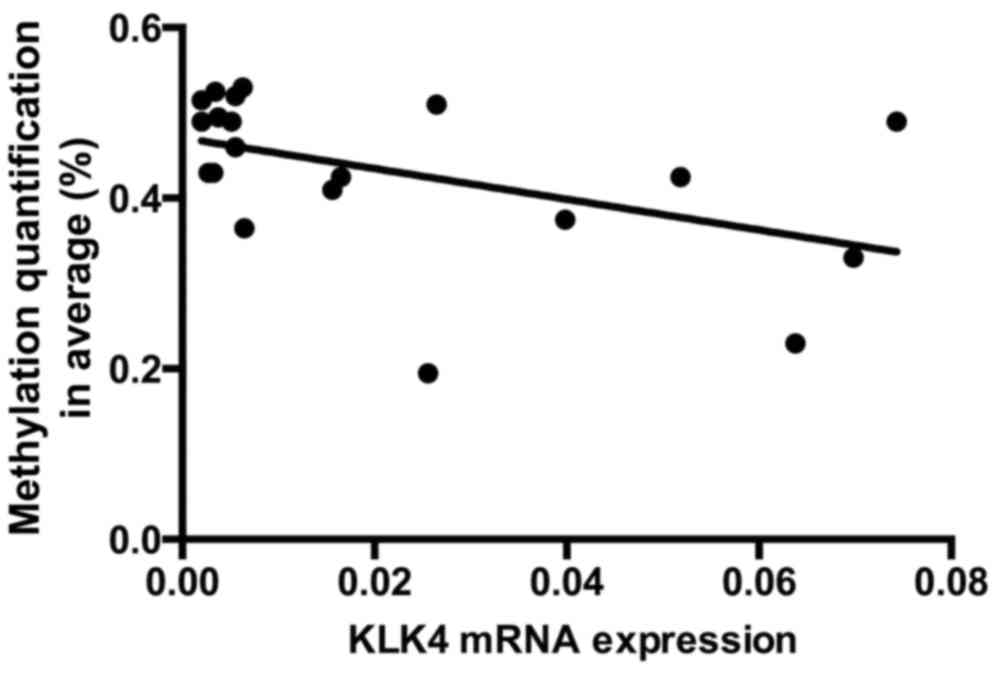
Correlation between hypomethylation
with KLK4 and expression of mRNA
Using linear Pearson's R correlation, the
correlations between KLK4 mRNA expression and DNA methylation
status of CpG4 and CpG9 sites harvested from the 10 samples were
analyzed. This analysis demonstrated that the expression of KLK4 is
negatively correlated with its methylation status (r=−0.47;
P=0.037; Fig. 5).
Discussion
Pediatric solid tumors develop after relatively
short latency periods, and thus are becoming one of the most common
reasons for child mortality (15,16). HB is
the most common liver tumor found in children, and its incidence is
increasing in North America and Europe (17). Although its pathogenesis and
progression have been extensively studied for the last two decades,
its etiology remains to be fully elucidated. Through a previous
study that involved sequencing of the HB exome, we recently
detected a novel oncogene (caprin family member 2) and three tumor
suppressors (speckle-type POZ protein, olfactory receptor family 5
subfamily I member 1 and cell division cycle 20B) that influence HB
cell growth (18). The present group
has also detected an association between long non-coding RNA
(lncRNA) and HB in a previous study (19).
Hypermethylation or hypomethylation of gene promoter
regions is recognized as one of the mechanisms that can silence or
activate the oncogenes (20). It has
recently been indicated that aberrant promoter methylation of the
RAS association domain family protein 1 may contribute to the
pathogenesis of HB and is considered to be a significant prognostic
indicator in HB (1,21).
KLK4 is a member of the KLK family. It has
previously been reported that KLK4 is overexpressed in endometrial
carcinoma (22), colorectal
adenocarcinoma, and prostate cancer; however, the mechanism remains
unclear. Detection of KLK4 mRNA expression levels in HB and normal
liver tissues by qPCR in the present study demonstrated a
significant difference between tissue types. KLK4 mRNA expression
levels in HB tissues were significantly higher than in normal liver
tissues. In an independent cohort of 10 adjacent HB-non-tumor
tissues pairs, we theorized whether there was a correlation between
KLK4 mRNA expression and DNA methylation. To further understand the
association between HB and KLK4 methylation, the methylation status
of KLK4 genes were analyzed in the 10 paired samples. Following
methylation quantification of the KLK4 gene using MALDI-TOF MS,
three CpG sites were detected per sample. Hypermethylation degrees
of two CpG sites in the promoter regions of the KLK4 gene for
normal liver tissues were significantly higher than those of HB
samples. Furthermore, correlation analysis indicated that
expression of KLK4 mRNA is significantly negatively correlated with
its methylation status. This finding supports previously published
data, which suggested that KLK4 may be an oncogene (23–26).
To the best of our knowledge, the present study is
the first to examine the methylation of KLK4 genes in HB, and an
association between the level of KLK4 mRNA and methylation was
successfully detected. This finding is consistent with the results
of previous studies, demonstrating that hypomethylation of promoter
regions of crucial genes is able to activate relevant gene
expression and may contribute to tumorigenesis (27–29). A
limitation of the present study is that only 10 pairs of patient
samples were used. Additional samples are required for
investigation in order to more accurately represent the
population.
In conclusion, the present findings indicate that
aberrant promoter methylation of KLK4 may contribute to the
tumorigenesis of HB, and that KLK4 may be a potential biomarker for
HB. However, since HB is an uncommon disease, its etiology,
pathophysiology, and molecular mechanisms remain unknown. Further
studies are required in order to fully understand and treat this
disease.
Acknowledgements
This study would like to thank the following for
providing financial support: The National Key Clinical Specialty
Construction Programs of China (2014–2016), the Shanghai ‘Non
key-in-key discipline’ Clinical medical centers (2014–2016), the
Shanghai Rising-Star Program (A type) (grant no. 15QA1400800), the
National Natural Science Foundation of China (grant no. 81572324)
and the Science Foundation of Shanghai (grant no. 15ZR1404200).
References
|
1
|
Honda S, Miyagi H, Suzuki H, Minato M,
Haruta M, Kaneko Y, Hatanaka KC, Hiyama E, Kamijo T, Okada T and
Taketomi A: RASSF1A methylation indicates a poor prognosis in
hepatoblastoma patients. Pediatr Surg Int. 29:1147–1152. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Kingston JE, Herbert A, Draper GJ and Mann
JR: Association between hepatoblastoma and polyposis coli. Arch Dis
Child. 58:959–962. 1983. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Ross JA: Hepatoblastoma and birth weight:
Too little, too big, or just right? J Pediatr. 130:516–517.
1997.PubMed/NCBI
|
|
4
|
Mamlok V, Nichols M, Lockhart L and Mamlok
R: Trisomy 18 and hepatoblastoma. Am J Med Genet. 33:125–126. 1989.
View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Herman JG and Baylin SB: Gene silencing in
cancer in association with promoter hypermethylation. N Engl J Med.
349:2042–2054. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Geiman TM and Robertson KD: Chromatin
remodeling, histone modifications, and DNA methylation-how does it
all fit together? J Cell Biochem. 87:117–125. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Reik W and Walter J: Genomic imprinting:
Parental influence on the genome. Nat Rev Genet. 2:21–32. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Chow J and Heard E: X inactivation and the
complexities of silencing a sex chromosome. Curr Opin Cell Biol.
21:359–366. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
O'Sullivan E and Goggins M: DNA
methylation analysis in human cancer. Methods Mol Biol.
980:131–156. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Esteller M: Epigenetics in cancer. N Engl
J Med. 358:1148–1159. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Kontos CK, Chantzis D, Papadopoulos IN and
Scorilas A: Kallikrein-related peptidase 4 (KLK4) mRNA predicts
short-term relapse in colorectal adenocarcinoma patients. Cancer
Lett. 330:106–112. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Lose F, Srinivasan S, O'Mara T, Marquart
L, Chambers S, Gardiner RA, Aitken JF; Australian Prostate Cancer
BioResource; Spurdle AB, Batra J and Clements JA: Genetic
association of the KLK4 locus with risk of prostate cancer. PloS
One. 7:e445202012. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Haas JE, Feusner JH and Finegold MJ: Small
cell undifferentiated histology in hepatoblastoma may be
unfavorable. Cancer. 92:3130–3134. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Johnson MR, Wang KS, Smith JB, Heslin MJ
and Diasio RB: Quantitation of dihydropyrimidine dehydrogenase
expression by real-time reverse transcription polymerase chain
reaction. Anal Biochem. 278:175–184. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Otte JB: Progress in the surgical
treatment of malignant liver tumors in children. Cancer Treat Rev.
36:360–371. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Tiao GM, Bobey N, Allen S, Nieves N,
Alonso M, Bucuvalas J, Wells R and Ryckman F: The current
management of hepatoblastoma: A combination of chemotherapy,
conventional resection, and liver transplantation. J Pediatr.
146:204–211. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
von Schweinitz D: Hepatoblastoma: Recent
developments in research and treatment. Semin Pediatr Surg.
21:21–30. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Jia D, Dong R, Jing Y, Xu D, Wang Q, Chen
L, Li Q, Huang Y, Zhang Y, Zhang Z, et al: Exome sequencing of
hepatoblastoma reveals novel mutations and cancer genes in the Wnt
pathway and ubiquitin ligase complex. Hepatology. 60:1686–1696.
2014. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Dong R, Jia D, Xue P, Cui X, Li K, Zheng
S, He X and Dong K: Genome-wide analysis of long noncoding RNA
(lncRNA) expression in hepatoblastoma tissues. PloS One.
9:e855992014. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Baylin SB and Ohm JE: Epigenetic gene
silencing in cancer-A mechanism for early oncogenic pathway
addiction? Nat Rev Cancer. 6:107–116. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Harada K, Toyooka S, Maitra A, Maruyama R,
Toyooka KO, Timmons CF, Tomlinson GE, Mastrangelo D, Hay RJ, Minna
JD and Gazdar AF: Aberrant promoter methylation and silencing of
the RASSF1A gene in pediatric tumors and cell lines. Oncogene.
21:4345–4349. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Zhang SQ, Cai B, Liu L, He YY, Yang YX and
Wan XP: Kallikrein 4 overexpression in endometrial carcinoma and
upregulation by estrogen via mitogen-activated protein kinase
signal pathway. Int J Gynecol Cancer. 19:1377–1383. 2009.
View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Klokk TI, Kilander A, Xi ZJ, Waehre H,
Risberg B, Danielsen HE and Saatcioglu F: Kallikrein 4 is a
proliferative factor that is overexpressed in prostate cancer.
Cancer Res. 67:5221–5230. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Xi ZJ, Kaern J, Davidson B, Klokk TI,
Risberg B, Tropé C and Saatcioglu F: Kallikrein 4 is associated
with paclitaxel resistance in ovarian cancer. Gynecol Oncol.
94:80–85. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Papachristopoulou G, Avgeris M and
Scorilas A: Expression analysis and study of KLK4 in benign and
malignant breast tumours. Thromb Haemost. 101:381–387.
2009.PubMed/NCBI
|
|
26
|
Mangé A, Desmetz C, Berthes ML, Maudelonde
T and Solassol J: Specific increase of human kallikrein 4 mRNA and
protein levels in breast cancer stromal cells. Biochem Biophys Res
Commun. 375:107–112. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Doi A, Park IH, Wen B, Murakami P, Aryee
MJ, Irizarry R, Herb B, Ladd-Acosta C, Rho J, Loewer S, et al:
Differential methylation of tissue-and cancer-specific CpG island
shores distinguishes human induced pluripotent stem cells,
embryonic stem cells and fibroblasts. Nat Genet. 41:1350–1353.
2009. View
Article : Google Scholar : PubMed/NCBI
|
|
28
|
Irizarry RA, Ladd-Acosta C, Wen B, Wu Z,
Montano C, Onyango P, Cui H, Gabo K, Rongione M, Webster M, et al:
The human colon cancer methylome shows similar hypo- and
hypermethylation at conserved tissue-specific CpG island shores.
Nat Genet. 41:178–186. 2009. View
Article : Google Scholar : PubMed/NCBI
|
|
29
|
Ogoshi K, Hashimoto SI, Nakatani Y, Qu W,
Oshima K, Tokunaga K, Sugano S, Hattori M, Morishita S and
Matsushima K: Genome-wide profiling of DNA methylation in human
cancer cells. Genomics. 98:280–287. 2011. View Article : Google Scholar : PubMed/NCBI
|