Introduction
Colon cancer is a digestive tract cancer that ranks
third among all cancers in incidence and mortality. Rapidly
changing life styles along with diet structure are prime reasons
for the rapid increase of this pathological state (1). The incidence and development of tumor
includes abnormal changes in multiple cancer and tumor suppressor
genes. The methylation of CpG island in the area of the tumor
suppressor gene promoter is one of the most important factors
responsible for cancer (2). APC
(adenomatous polyposis coli) is a tumor suppressor gene of the
length of 8535 bp and is located on the 5q21 chromosome. It plays
an important role in Wnt signal transduction pathway (3). APC gene encodes APC protein, which has
the ability to regulate cell differentiation, proliferation and
migration (4). Research has found
that the methylation of CpG island in the area of APC gene promoter
could lead to deletion of APC gene. This in turn, disturbs the
conduction of Wnt signal pathway leading to the incidence and
development of tumors (5). Research
has confirmed the hyper-methylation of APC gene and the phenomenon
of deletion of gene transcription during multiple cancers including
breast, gastric, esophagus, pancreatic and lung cancer (6). In addition, methylation of APC gene is
closely correlated with the incidence, development, invasion and
migration of malignant tumors (7,8).
The present study adopted the method of methylation
specific PCR (MSP) to examine the state of methylation of APC gene
in the tissues of colon cancer patients and SW1116 cells. 5-aza-dC
exerted demethylation by inhibiting methylase (9). Thus, in the present study we inhibited
the methylation of APC gene and then studied the effect of APC
methylation on the proliferation and invasion of SW1116 cells.
Clinico-pathological data of patients were statistically
summarized. The correlation between the methylation of APC gene and
clinicopathological features was analyzed. Finally, the correlation
between the methylation of APC gene and colon cancer is
discussed.
Materials and methods
Materials
Human colon cancer cells SW1116 were procured from
the Chinese Academy of Sciences Shanghai Cell Bank (Shanghai,
China); 5-aza-dC, MTT was obtained from Sigma (St. Louis, MO, USA).
RPMI-1640 culture medium and fetal bovine serum (FBS) were procured
from HyClone Laboratories, Inc. (Logan, UT, USA); Epi-Tect
Bisulfite kits agent was provided by Qiagen (Nordrhein-Westfalen,
Germany); DNA extraction kits were from Tiangen Biotech (Beijing)
Co., Ltd., (Beijing, China); Taq DNA polymerase, dNTP mixture, DNA
marker and primer synthesis were obtained from Takara Bio (Dalian)
Co., Ltd. (Dalian, China) and Transwell cabinet was obtained from
Corning, Inc. (New York, NY, USA).
Colon tissue samples of 60 colon cancer patients,
who were given surgical treatment in our hospital from January 2012
to December 2014 were collected. Colon cancer tissues were taken
from resected tumor samples. Normal tumor-adjacent tissues were
also excised from the part 10 cm away from tumor margin (no
infiltration of cancer cells were found under microscope). Tissue
samples were immediately preserved in liquid nitrogen after
collection. There were 32 male cases, 28 female cases and the
average patient age was 54.7±14.3 years. All patients were
confirmed with colon cancer by clinical and pathological diagnosis.
The patients were operated for the first time and had no history of
radiotherapy or chemotherapy. All participants or their families
signed a written informed consent form. Furthernore, the Ethics
Committee of Xuzhou Cental Hospital approved the present study.
Detection of the methylation of APC
gene in patient tissue samples by MSP
DNA extraction kits were used to extract the total
DNA in tissue samples. The ultraviolet spectrophotometer (U-3010;
Hitachi, Ltd., Tokyo, Japan) was used to detect the DNA content and
purity (A260/A280 >1.8 was considered qualified). The total DNA
extracted was modified by bisulfite according to the instructions
on the Epi-Tect Bisulfite kits. The modified DNA was respectively
given MSP and non-MSP (Table I). The
2% agarose gel was used for electrophoresis and ethidium bromide
was used for staining. Gel imaging system (UVP, LLC, Upland, CA,
USA) was utilized for R result analyses.
 | Table I.Primer sequences of APC gene
methylation and non-methylation. |
Table I.
Primer sequences of APC gene
methylation and non-methylation.
| Primer | Sequence | Product size
(bp) |
|---|
| APC-MF |
5′-TATTGCGGAGTGCGGGTC-3′ | 98 |
| APC-MR |
5′-TCGACGAACTCCCGACGA-3′ |
|
| APC-UF |
5′-GTGTTTTATTGTGGAGTGTGGGTT-3′ | 108 |
| APC-UR |
5′-CCAATCAACAAACTCCCAACAA-3′ |
|
Cell culture and group processing
SW1116 cells were cultured in RPMI-1640 culture
medium (including 10% FBS) in the incubator at 37°C with 5%
CO2. The medium was changed every 24 h. Digestion
passage was conducted after cell fusion. The experiment was divided
into the normal control group and the 5-aza-dC group. The control
group was cultured in a standard manner. The 5-aza-dC group was
given 5-aza-dC of 5 µmol/l terminal concentration to process SW1116
cells.
Detection of the methylation of APC
gene in cells
SW1116 cells at the logarithmic growth phase were
collected. Each well was added with 2 ml (2×105
cells/ml) SW1116 cell suspension. Cells were cultured in 6 well
plates for 24 h, then processed as described above. Cells were
collected after digestion by pancreatin for 24 h. DNA extraction
kits were applied to extract total DNA. The state of methylation of
APC gene in cells of each group was determined by MSP.
Detection of proliferation ability of
SW1116 cells by MTT
SW1116 cells at the logarithmic growth phase were
collected. The wells were added with 200 ml (1×104
cells/ml) SW1116 cell suspension. Cells were cultured in 96-well
plates for 24 h, and processed as described above. Each group was
set with 6 parallel wells and culture was continued for 24 h then,
the culture medium was discarded. The plates were then washed 3
times, followed by addition of 100 µl MTT (5 mg/ml) in each well
and the culture was continued for 4 h. The wells were then added
with 100 µl DMSO and the plate shaken for 10 min in the dark. The
absorbance value (OD value) was recorded at 570 nm. The cell
proliferation rate was calculated according to the following
formula: proliferation rate = OD value of the experiment group/OD
value of the normal group × 100%.
Detection of invasion ability of
SW1116 cells by Transwell
The upper chamber of Transwell was evenly added with
100 µl single cell suspension at 4×105/ml and 100 µl
culture medium. The lower chamber was added with 500 µl culture
medium with 30% FBS, 24 h later, 4% paraformaldehyde was added for
fixation and crystal violet for staining. After 15 min, images were
taken for analysis.
The correlation between the
methylation of APC gene and the pathological parameters of colon
cancer patients
Patients were divided into groups according to sex,
age, tumor size, differentiation degree, lymph node metastasis and
Dukes staging index. The correlation between parameters and the
methylation of APC gene was analyzed.
Statistical analysis
The data were processed by SPSS17.0 (IBM Corp.,
Armonk, NY, USA). Measurement data was expressed by mean ± standard
deviation and analyzed by single factor ANOVA. Comparison of
numeration data among different groups was by χ2
analysis. P≤0.05 values confirmed statistical significance.
Results
APC gene methylation in the tissues of
colon cancer patients
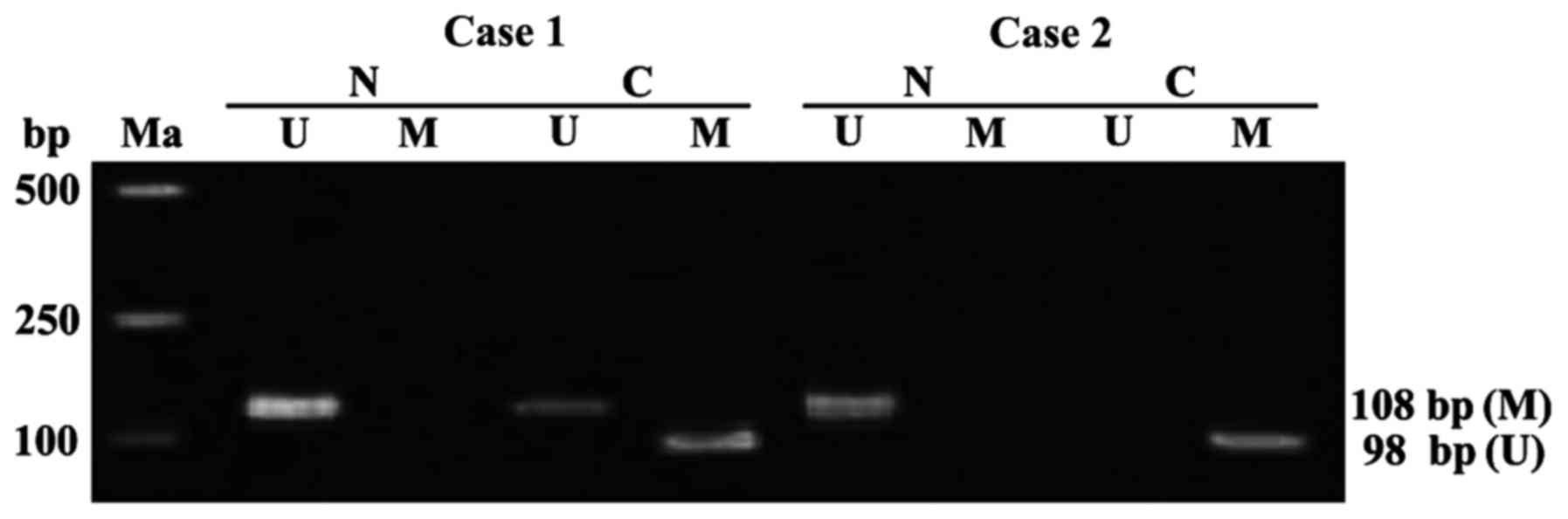
APC genes in the tumor-adjacent normal tissues of 60
colon cancer patients were free from methylation. The detection
rate of APC gene methylation in colon cancer tissues was 68.33%
(41/60), including 36 cases of complete methylation, 5 cases of
partial methylation (Fig. 1). The
state of APC gene methylation in the colon cancer was significantly
different from that in the tumor-adjacent normal tissues
(P<0.01).
Effect of 5-aza-dC on APC gene
methylation in SW1116 cells
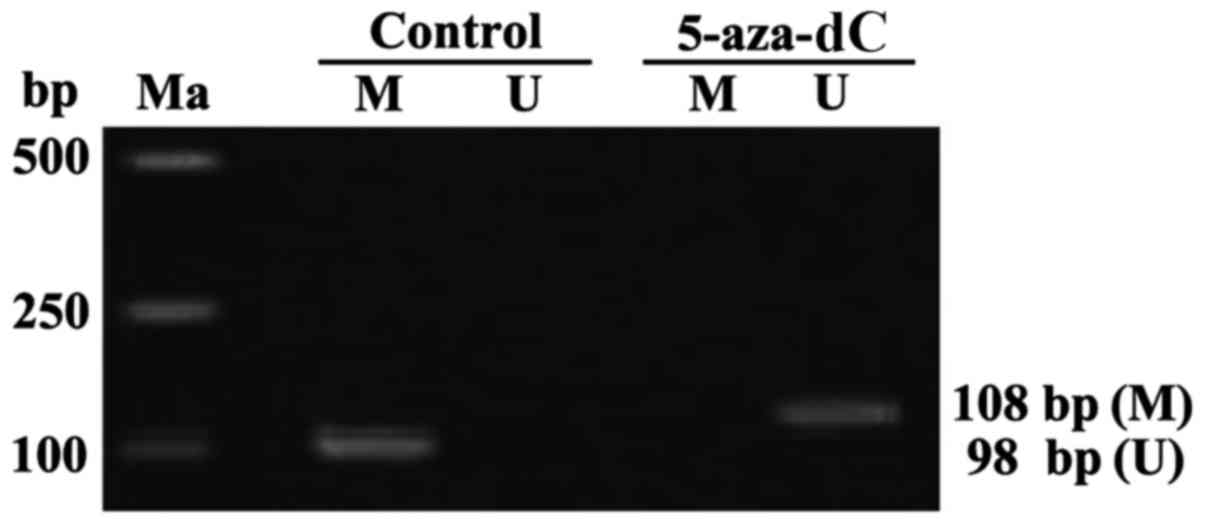
APC genes in the cells of the normal group showed
complete methylation. Whereas, the APC genes in the cells of the
5-aza-dC group revealed no methylation. thus, intervention with
5-aza-dC for 24 h, inhibited the APC gene methylation in SW1116
cells (Fig. 2).
Effect of 5-aza-dC induces inhibition
of APC gene methylation on the cell proliferation in SW1116
cells
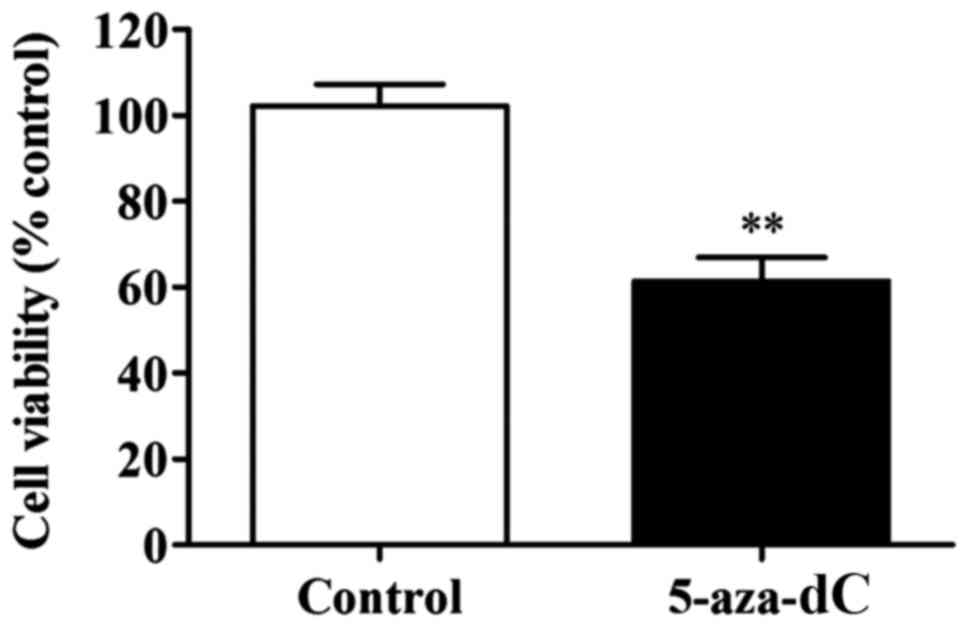
The results clearly showed that the 5-aza-dC-induced
inhibition of APC gene methylation significantly lowered the
proliferation rate in SW1116 cells in comparison to the cells of
normal group (P<0.01) (Fig. 3).
This observation confirmed that the inhibition of APC gene
methylation in the SW1116 cells reduced the proliferation of SW1116
cells.
Effect of 5-aza-dC inhibition of APC
gene methylation on cell invasion in the SW1116 cells
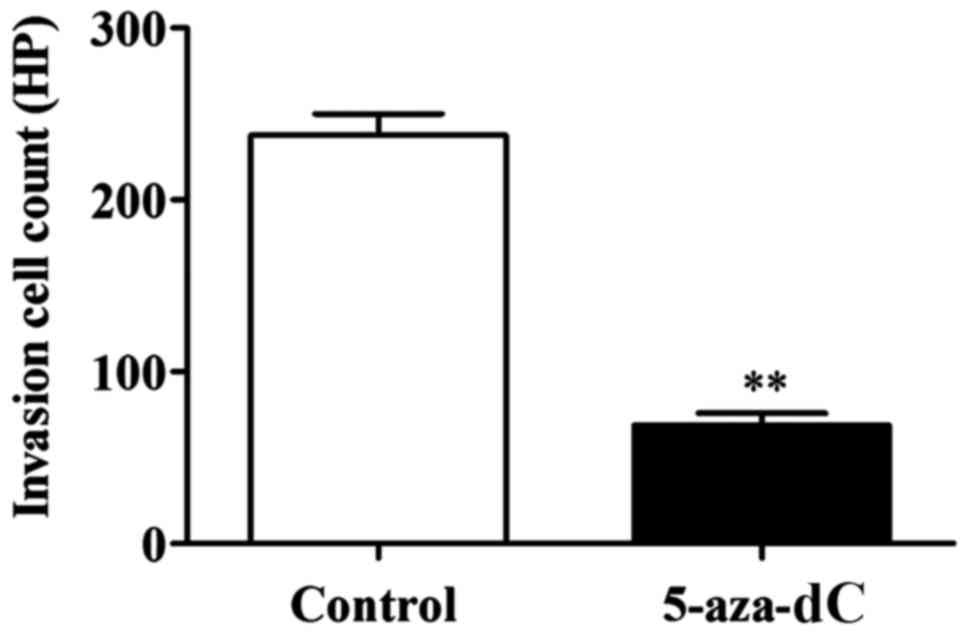
The number of invaded cells in the 5-aza-dC group
was significantly lower than that in the normal group (P<0.01).
This confirmed that the inhibition of APC gene methylation in the
SW1116 cells has the ability to inhibit the invasion ability of
SW1116 cells (Fig. 4).
The correlation between APC gene
methylation and clinicopathological parameters
Results of the correlation between
clinicopathological parameters of colon cancer and APC gene
methylation are shown in Table II.
The correlation test indicated that APC gene methylation was
correlated with tumor size, differentiation degree, lymph node
metastasis and Dukes staging (P<0.05, P<0.01, P<0.05 and
P<0.05), respectively. However, no correlation was observed with
age or sex (P>0.05).
 | Table II.The correlation between APC gene
methylation and clinicopathological parameters. |
Table II.
The correlation between APC gene
methylation and clinicopathological parameters.
|
|
| APC gene
methylation |
|
|---|
|
|
|
|
|
|---|
| Clinicopathological
parameters | Case no. | Methylation (%) | Non-methylation
(%) | P-value |
|---|
| Sex |
|
|
| >0.05 |
| Male | 32 | 21 (65.63) | 11 (34.37) |
|
|
Female | 28 | 20 (71.43) | 8
(28.57) |
|
| Age |
|
|
| >0.05 |
| ≥54
years | 36 | 28 (77.78) | 8
(22.22) |
|
| <54
years | 24 | 13 (54.17) | 11 (45.83) |
|
| Tumor size |
|
|
| <0.05 |
| ≥5
cm | 33 | 27 (81.82) | 6
(18.18) |
|
| <5
cm | 27 | 14 (51.85) | 13 (48.15) |
|
| Differentiation
degree |
|
|
| <0.01 |
| High | 37 | 30 (81.08) | 7
(18.92) |
|
| Low | 23 | 11 (47.83) | 12 (52.17) |
|
| Lymph node
metastasis |
|
|
| <0.05 |
| Yes | 31 | 25 (80.65) | 6
(19.35) |
|
| No | 29 | 16 (55.17) | 13 (44.83) |
|
| Dukes staging |
|
|
| <0.01 |
| A+B | 27 | 13 (48.15) | 14 (51.85) |
|
| C+D | 33 | 28 (84.85) | 5
(15.15) |
|
Discussion
The incidence of colorectal cancer is a complex
process that involves multiple factors. As the early clinical
symptoms of colon cancer patients are not obvious, most patients
are diagnosed at the advanced stage of colon cancer (10). The epigenetic changes also play a very
important role in the incidence and development of colon cancer
(11). DNA methylation is a common
epigenetic modification that could cause transcriptional silencing
of tumor suppressor genes (12).
Research has found that the methylation of tumor suppressor genes
is closely correlated with the differentiation and proliferation of
tumor cells (13,14). DNA methylation is an important event
occurring at the early stage of tumors. Therefore, detection of the
methylation of tumor-related genes could provide reference for the
diagnosis, therapy and prognosis of colorectal cancer (15). APC is a tumor suppressor gene which is
important in the Wnt signal pathway. This protein plays an
important role in cell proliferation, apoptosis and invasion
(16). Hyper-methylation of APC gene
has been found in malignant tumors such as gastric, liver and
pancreatic cancer (17,18).
In the present study, the methylation rate of APC
gene in colon cancer tissues was 68.33%. No methylation of APC gene
was observed in tumor-adjacent tissues. To confirm the effect of
APC gene methylation on colon cancer cells, it was perceived that
the inhibition of APC gene methylation resulted in the reduction of
proliferation and invasion of SW1116 cells. To further confirm the
clinical significance of APC gene methylation, the present study
analyzed the correlation between APC gene methylation and
clinicopathological parameters. Results confirmed that APC gene
methylation was correlated with tumor size, differentiation degree,
lymph node metastasis and Dukes staging.
Arnold et al (19) found that APC gene showed high
methylation rate in the tumor tissues of colon cancer patients.
Deng et al (20) revealed that
the abnormal APC gene methylation was correlated well with the
deactivation of APC protein functions. In the present study, APC
gene was found with high methylation in the tumor tissues of colon
cancer patients. The APC gene methylation is not only correlated
with the proliferation and invasion of colon cancer cells, but also
with tumor size, differentiation degree, lymph node metastasis and
Dukes staging of patients with colon cancer.
The present study concludes that the methylation of
APC gene is closely correlated with colon cancer, especially with
tumor size, differentiation degree, lymph node metastasis, and
Dukes staging of patients. Therefore, the application of
demethylation drugs to inhibit the methylation of APC gene is
likely to be useful in treatment of colon cancer.
References
|
1
|
Siegel R, Naishadham D and Jemal A: Cancer
statistics, 2013. CA Cancer J Clin. 63:11–30. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Grady WM, Willis J, Guilford PJ, Dunbier
AK, Toro TT, Lynch H, Wiesner G, Ferguson K, Eng C, Park JG, et al:
Methylation of the CDH1 promoter as the second genetic hit in
hereditary diffuse gastric cancer. Nat Genet. 26:16–17. 2000.
View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Jarrett CR, Blancato J, Cao T, Bressette
DS, Cepeda M, Young PE, King CR and Byers SW: Human APC2
localization and allelic imbalance. Cancer Res. 61:7978–7984.
2001.PubMed/NCBI
|
|
4
|
van Noort M, Meeldijk J, van der Zee R,
Destree O and Clevers H: Wnt signaling controls the phosphorylation
status of β-catenin. J Biol Chem. 277:17901–17905. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Zysman M, Saka A, Millar A, Knight J,
Chapman W and Bapat B: Methylation of Αdenomatous polyposis coli in
endometrial cancer occurs more frequently in tumors with
microsatellite instability phenotype. Cancer Res. 62:3663–3666.
2002.PubMed/NCBI
|
|
6
|
Dong SM, Kim HS, Rha SH and Sidransky D:
Promoter hypermethylation of multiple genes in carcinoma of the
uterine cervix. Clin Cancer Res. 7:1982–1986. 2001.PubMed/NCBI
|
|
7
|
Tamura G, Maesawa C, Suzuki Y, Ogasawara
S, Terashima M, Saito K and Satodate R: Primary gastric carcinoma
cells frequently lose heterozygosity at the APC and MCC genetic
loci. Jpn J Cancer Res. 84:1015–1018. 1993. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Chen J, Röcken C, Lofton-Day C, Schulz HU,
Müller O, Kutzner N, Malfertheiner P and Ebert MP: Molecular
analysis of APC promoter methylation and protein expression in
colorectal cancer metastasis. Carcinogenesis. 26:37–43. 2005.
View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Bender CM, Pao MM and Jones PA: Inhibition
of DNA methylation by 5-aza-2-deoxycytidine suppresses the growth
of human tumor cell lines. Cancer Res. 58:95–101. 1998.PubMed/NCBI
|
|
10
|
Gordon MB, Nakhle S and Ludlam WH:
Patients with acromegaly presenting with colon cancer: A case
series. Case Rep Endocrinol. 2016:1–4. 2016. View Article : Google Scholar
|
|
11
|
Lim S, Metzger E, Schüle R, Kirfel J and
Buettner R: Epigenetic regulation of cancer growth by histone
demethylases. Int J Cancer. 127:1991–1998. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Jubb AM, Bell SM and Quirke P: Methylation
and colorectal cancer. J Pathol. 195:111–134. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Guo Y, Guo W, Chen Z, Kuang G, Yang Z and
Dong Z: Hypermethylation and aberrant expression of Wnt-antagonist
family genes in gastric cardia adenocarcinoma. Neoplasma.
58:110–117. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Wu D, Wong P, Li W, Vogel CF and Matsumura
F: Suppression of WIF-1 through promoter hypermethylation causes
accelerated proliferation of the aryl hydrocarbon receptor (AHR)
overexpressing MCF10AT1 breast cancer cells. Toxicology.
285:97–103. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Prabhu JS, Korlimarla A, Banerjee A, Wani
SKP and Sahoo R: Gene-specific methylation: Potential markers for
colorectal cancer. Int J Biol Markers. 24:57–62. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Fearnhead NS, Britton MP and Bodmer WF:
The ABC of APC. Hum Mol Genet. 10:721–733. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Esteller M, Sparks A, Toyota M,
Sanchez-Cespedes M, Capella G, Peinado MA, Gonzalez S, Tarafa G,
Sidransky D, Meltzer SJ, et al: Analysis of Adenomatous polyposis
coli promoter hypermethylation in human cancer. Cancer Res.
60:4366–4371. 2000.PubMed/NCBI
|
|
18
|
Tsuchiya T, Tamura G, Sato K, Endoh Y,
Sakata K, Jin Z, Motoyama T, Usuba O, Kimura W, Nishizuka S, et al:
Distinct methylation patterns of two APC gene promoters in normal
and cancerous gastric epithelia. Oncogene. 19:3642–3646. 2000.
View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Arnold CN, Goel A, Niedzwiecki D, Dowell
JM, Wasserman L, Compton C, Mayer RJ, Bertagnolli MM and Boland CR:
APC promoter hypermethylation contributes to the loss of APC
expression in colorectal cancers with allelic loss on 5q. Cancer
Biol Ther. 3:960–964. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Deng G, Song GA, Pong E, Sleisenger M and
Kim YS: Promoter methylation inhibits APC gene expression by
causing changes in chromatin conformation and interfering with the
binding of transcription factor CCAAT-binding factor. Cancer Res.
64:2692–2698. 2004. View Article : Google Scholar : PubMed/NCBI
|