Introduction
Hox antisense intergenic RNA (HOTAIR), a lncRNA that
acts as an oncogenic molecule in various types of cancer, is
localized to the HOXC gene cluster. HOTAIR interacts with PRC2
(polycomb repressive complex 2) to enhance H3K27 trimethylation,
and thereby decreases the expression of a large number of genes.
Several groups, including our laboratory, have reported that high
HOTAIR expression is correlated with a poor prognosis in several
types of cancer, including breast (1), colorectal (2), non-small lung cell (3), and gastric cancer (4). Interestingly, recent report suggested
that effects of HOTAIR are strongly tissue-dependent and can even
differ within the same type of cancer (5). Thus, the underlying mechanism by which
HOTAIR is involved in malignant progression remains uncertain.
Hepatocellular carcinoma (HCC) is the fifth leading
cause of cancer and second leading cause of cancer-related
mortality worldwide. More than 600,000 deaths are associated with
HCC every year worldwide (6).
Previous report suggested that the expression of HOTAIR is
associated with tumor recurrence and poor prognosis in
hepatocellular carcinoma (7,8). Several in vitro analyses were
reported using HCC cell line, HepG2; HOTAIR is a FOXC1-activated
driver of malignancy, which acts in part through the repression of
miR-1 in HepG2 cells (9). HOTAIR
silence activates P16Ink4a and P14ARF
signaling by enhancing miR-218 expression and suppressing Bmi-1
expression, resulting in the suppression of tumorigenesis in HepG2
cells (10). Introduction of human
HOTAIR into HepG2 cells revealed that HOTAIR promoted more rapid
proliferation (7). Although different
intracellular signaling are expected among multiple HCC cell lines,
the research of HOTAIR using HCC cell lines except for HepG2 is not
fully performed.
The tumor microenvironment is known to play
important roles in cancer development and behavior (11). Macrophages are a component of the
microenvironment in tumors, which called tumor-associated
macrophages (TAMs). Four decades ago, TAMs from malignant
metastatic cancers were reported to promote tumor growth and
metastasis (12). Recently, TAMs can
also promote initiation of tumor cells, inhibit antitumor immune
responses mediated by T cells, and stimulate tumor angiogenesis and
subsequently tumor progression (13).
Myeloid-derived suppressor cells (MDSCs) are another
critical component in the microenvironment (14). MDSCs are a heterogeneous group of
immature myeloid cells and expanded in response to a variety of
tumor factors. An increased presence of MDSCs is associated with
tumor progression and poorer outcomes. In HCC patients, MDSCs
defined as CD14+HLA-DR−/low cells exert their
immunosuppressive function through the induction of
CD4+CD25+Foxp3+ regulatory T cells
(15).
Chemoattractants including chemokines such as CCL2
and CCL5, and cytokines (for example, CSF-1 and members of the VEGF
family) are important mediators of the recruitment and functional
polarization of TAMs. CCL2 is highly upregulated in HCC patients
(16), and inhibition of CCL2 could
be an effective therapeutic approach against hepatocellular
carcinoma (17). The CCL2 is required
for recruitment of monocytes/macrophages and is implicated in
various aspects of liver pathology, including HCC (16). Furthermore, recent study suggested
that microenvironment-derived CCL2 results in the accumulation of
MDSCs in glioma (18). Thus, CCL2 is
critical for immunosuppression to promote cancer metastasis. It is
well-known that tumor cells as well as stromal cells were thought
to be the source of CCL2 in established tumors (19). However, it is not still elucidated how
HCC cells regulate CCL2 production.
In this study, we examined whether HOTAIR-expressing
cancer cells exert the malignant phenotypes. We investigate the
effect of HOTAIR against both cancer cell itself and peripheral
blood monocyte cells (PBMCs) as immune cells in tumor
microenvironment.
Materials and methods
Ethics statement
The present study was conducted according to the
principles expressed in the Declaration of Helsinki and was
approved (MCC-AE-2016-7) by the Ethics Committees at the Miyagi
Cancer Center Research Institute (Natori, Japan). Experimental
protocols involving animals were approved by the Miyagi Cancer
Center Animal Care and Use Committee.
Cell lines and cell culture
Hepatoma cell line Li-7 and Hep3B was obtained from
RIKEN BioResource Center (Tsukuba, Japan) and Cell Resource Center
for Biomedical Research Cell Bank, Institute of Development, Aging,
and Cancer, Tohoku University (Sendai, Japan), respectively. These
cells were maintained in Dulbeccos modified Eagles medium (DMEM;
Gibco Life Technologies, Carlsbad, CA, USA) supplemented with 10%
fetal bovine serum (Euro-Clone, Milano, Italy) and 1%
penicillin-streptomycin (Gibco Life Technologies). Cells were
cultured in a humidified 5% CO2 incubator at 37°C.
Retroviral transfection
Human HOTAIR cDNA (obtained from Addgene, Cambridge,
MA, USA) was amplified by PCR and inserted into the pBabe-hygro
vector (pBabe-HOTAIR). The recombinant retrovirus was produced with
the Platinum-A packaging-cell line (Plat-A, kindly provided by
Prof. Kitamura, Tokyo University) as described previously (3). Briefly, Plat-A cells were transfected
with pBabe-HOTAIR or a pBabe-hygro empty vector (Empty). FugeneHD
(Roche Applied Science, Mannheim, Germany) and Opti-MEM I (Gibco
Life Technologies) were added following the manufacturers protocol.
The retrovirus-containing supernatant was collected 48 h after
transfection and passed through a 0.45-µm filter. Li7 and Hep3B
cells were infected with the recombinant retroviruses and then
selected with hygromycin.
Isolation of peripheral blood
mononuclear cells (PBMCs)
The heparinized blood was collected from healthy
donors, and isolated by density gradient centrifugation using
Lymphoprep (Axis-Shield PoC AS, Oslo, Norway) following the
manufacturers protocol.
Wound-healing assay
Cells were seeded in 24-well plates in normal
cell-growth medium, incubated until confluent, and a yellow pipette
tip was used to make a straight scratch, simulating a wound. After
a 24 h incubation, the area occupied by cells that had migrated
into the scratch area was measured using NIS-Elements software
(Nikon, Tokyo, Japan).
Co-incubation assay
A trans-membrane of 0.4 µm pore size (Corning Life
Science, Pittsburgh, PA, USA) was used for the analysis of soluble
factors secreted by HOTAIR overexpressing hepatoma cell lines or
HOTAIR-overexpressing hepatoma cell lines. The upper chamber
included PBMCs (1×106 cells/500 µl) from healthy donors,
and the lower chamber included HOTAIR-overexpressing hepatoma cell
lines (1.0×105 cells/500 µl). After 72 h co-incubation
at 37°C under an atmosphere of 5% CO2, multicolor FACS
analysis was performed.
Flow cytometry analysis
To determine the frequency of proportions of
macrophages and MDSCs, the cells were stained with specific
antibodies for 30 min at 4°C, washed twice, and subsequently
analyzed using a FACSCanto II (Becton Dickinson, CA). The
antibodies used: anti-human APC-CD14 (clone: HCD14), FITC-CD68
(clone: Y1/82A), Pacific Blue-HLA-DR (clone: L243), PE-CD33 (clone:
P67.6), APC/Cy7-CD11b (clone: ICRF44), and 7-AAD for dead cell
removal. All monoclonal antibodies used in this study were
purchased from BioLegend, Inc., (San Diego, CA, USA).
ELISA assay
Enzyme-linked immunosorbent assay (ELISA) kits for
CCL2 were purchased (R&D Systems, Inc., Minneapolis, MN, USA),
and ELISA was performed according to manufacturers instruction.
Microarray analysis
A microarray analysis was performed to search for
genes regulated by HOTAIR. We used the SurePrint G3 Human GE 8×60K
Microarray (Agilent Technologies, Santa Clara, CA, USA) to profile
the gene expression in control and HOTAIR-overexpressing Li-7
cells. The obtained gene expression data were expressed in
logarithmic scale, and heat maps were generated using the weighted
average difference (WAD) algorithm (20). The analyses were performed using R
software.
RNA preparation, cDNA synthesis, and
quantitative real-time RT-PCR
Total RNA was extracted from HCC cell lines by
RNeasy mini kit (Qiagen Inc., Valencia, CA, USA) according to the
manufacturers protocol. cDNAs were synthesized from 1.0 µg of total
RNA with the PrimeScript 1st Strand cDNA Synthesis kit (TaKaRa Bio,
Shiga, Japan) following the manufacturers protocol.
HOTAIR expression was quantified as previously
described (3). The primer sequences
used: for HOTAIR, F 5′-ggt aga aaa agc aac cac gaa gc-3′ and R
5′-aca taa acc tct gtc tgt gag tgc c-3′. HOTAIR expression was
normalized to RNA polymerase II (RPII) expression in each sample,
and calculated by ddCt methods (21)
(LighCycler480; Roche, Basel, Switzerland). All statistics were
performed at the ∆Ct stage in order to exclude potential bias due
to averaging of data transformed through the equation
2−∆∆Ct. Statistical analysis was undertaken using
Student t-test.
Statistical analysis
All statistical analyses were performed using JMP 11
(SAS Institute, Cary, NC, USA). P<0.05 indicated that the
difference was statistically significant using Students t-test.
Graphs show average and standard deviation unless otherwise
specified.
Results
Overexpression of HOTAIR in Li-7 cells
did not alter cell proliferation and migration
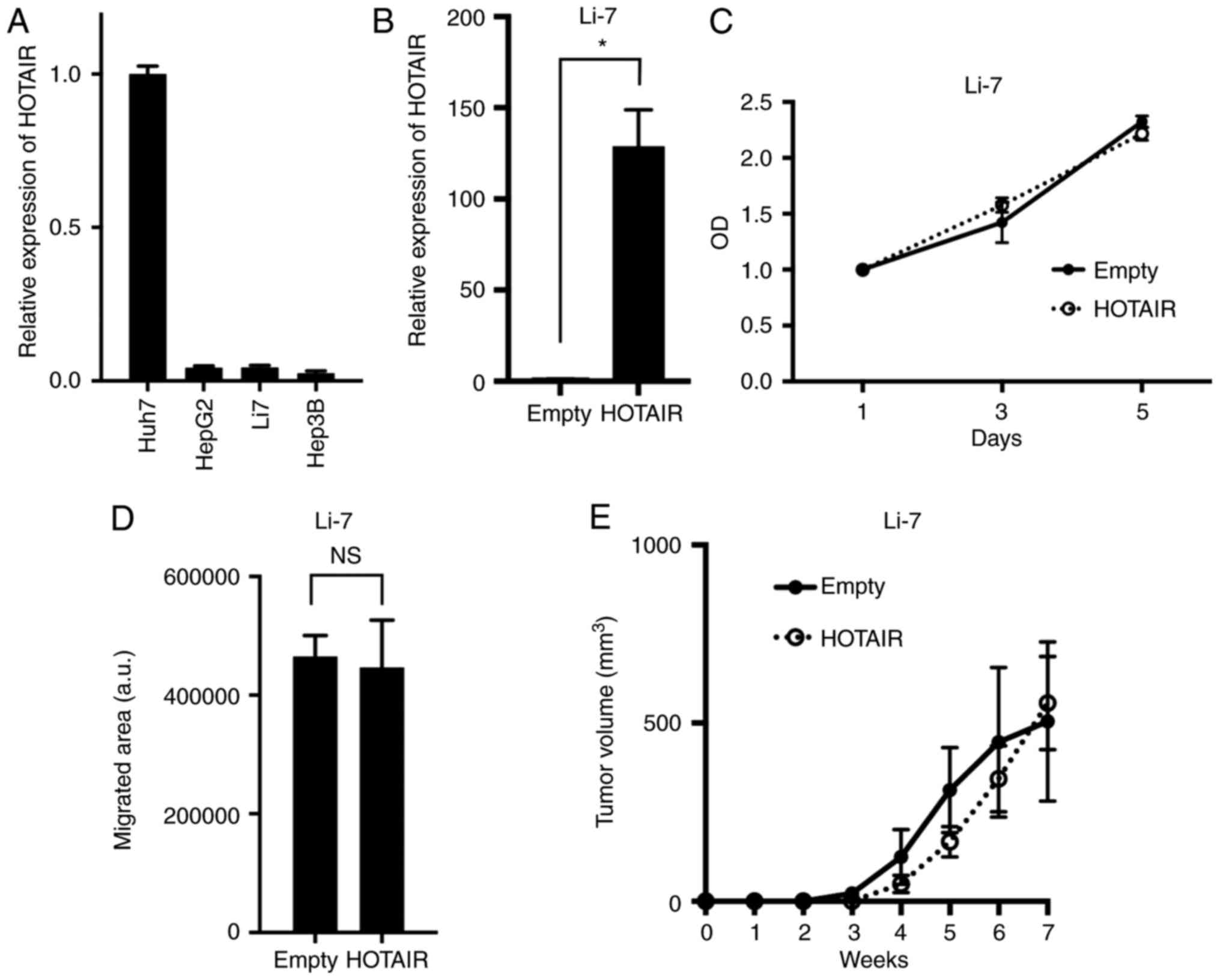
To evaluate the role of HOTAIR in HCC, we used human
HCC cell line Li-7 which expressed relatively lower HOTAIR
(Fig. 1A) to generate cells that
stably overexpressed HOTAIR (Fig.
1B). These HOTAIR-transduced cells overexpressed HOTAIR at
levels 128-fold higher (Li7-HOTAIR) than did empty
vector-transduced cells. We assayed the effect of HOTAIR on cell
proliferation using a MTT and found that HOTAIR overexpression in
Li-7 cells did not alter their proliferation compared to control
cells (Fig. 1C). We evaluated HOTAIRs
role in migratory capability with a wound-healing assay, and found
that the HOTAIR overexpression did not alter their migration
compared to control cells (Fig. 1D).
We further tested tumorigenicity of HOTAIR-overexpressing cells
using NOD/Shi-scid-IL2Rγnull (NOG) mice. Because NOG mice are
severely immunodificient, tumorigenicity assay using NOG mice can
reveal the proliferation and colonization capability of cancer
cell, and not evaluate the tumor immunity. No significant
difference was found between HOTAIR-overexpression and control
cells (Fig. 1E). These data indicated
that HOTAIR did not alter malignant phenotypes in Li-7 cells.
CCL2 secretion was induced by
HOTAIR
Since our results showed that HOTAIR did not
regulate cancer phenotypes, which is different from previous
studies using HepG2 cells, we screen the effect of HOTAIR in HCC
cell lines to identify the different role of HOTAIR in HCC cells.
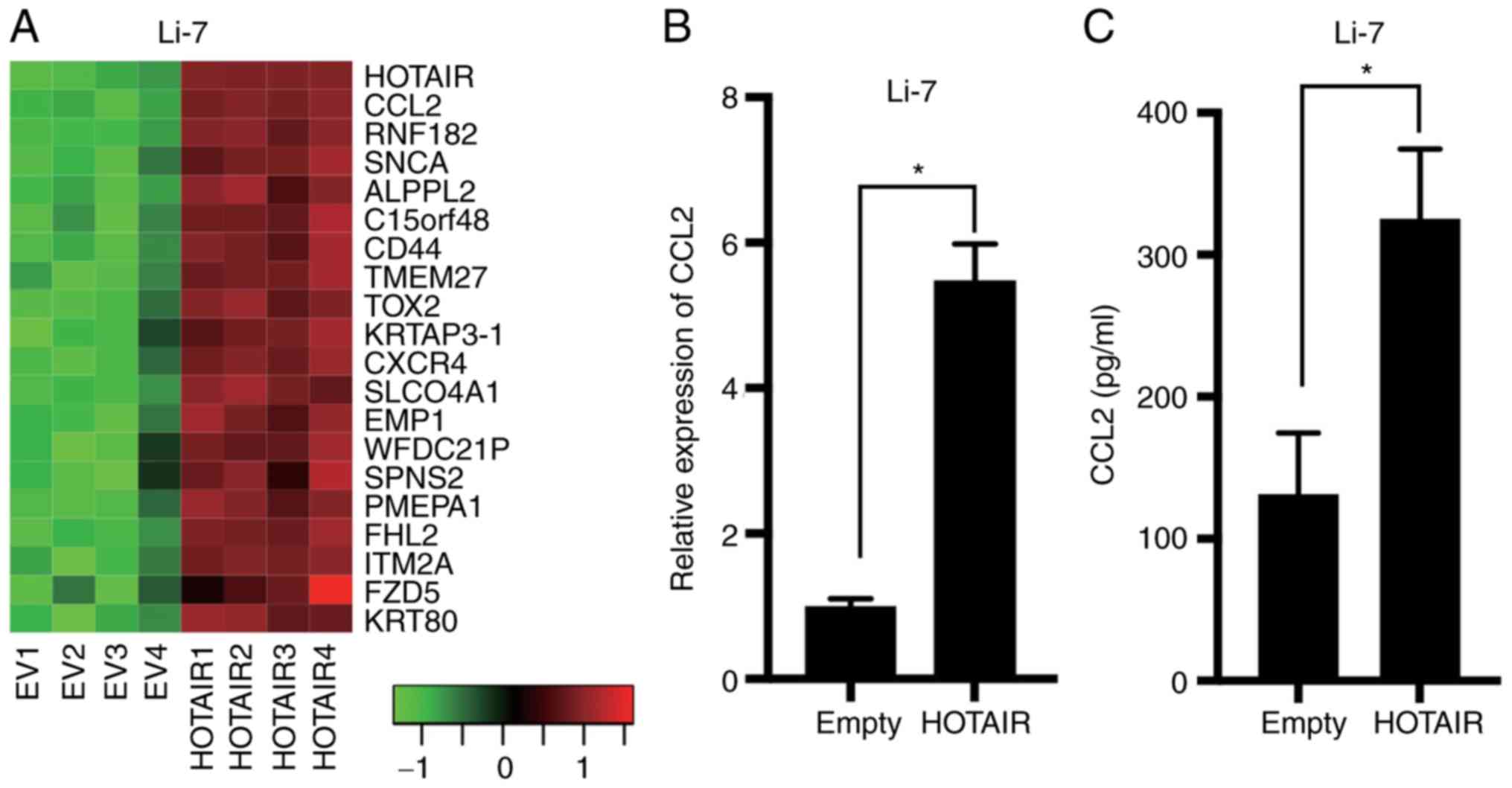
We compared gene expression profiles between HOTAIR-overexpression
and control Li-7 cells by microarray analysis using a weighted
average difference (WAD) algorithm (Fig.
2A). Analysis of the microarray data showed that CCL2, also
referred as monocyte chemoattractant protein 1, was differentially
expressed in the overexpressing and control Li-7 cells. Real-time
PCR and ELISA results confirmed that CCL2 was upregulated in the
Li-7-HOTAIR cells (Fig. 2B and C).
These results suggested that CCL2 acts downstream of HOTAIR in
HCC.
HOTAIR-overexpression induced
macrophage and MDSC proliferation in Li-7 and Hep3B cells
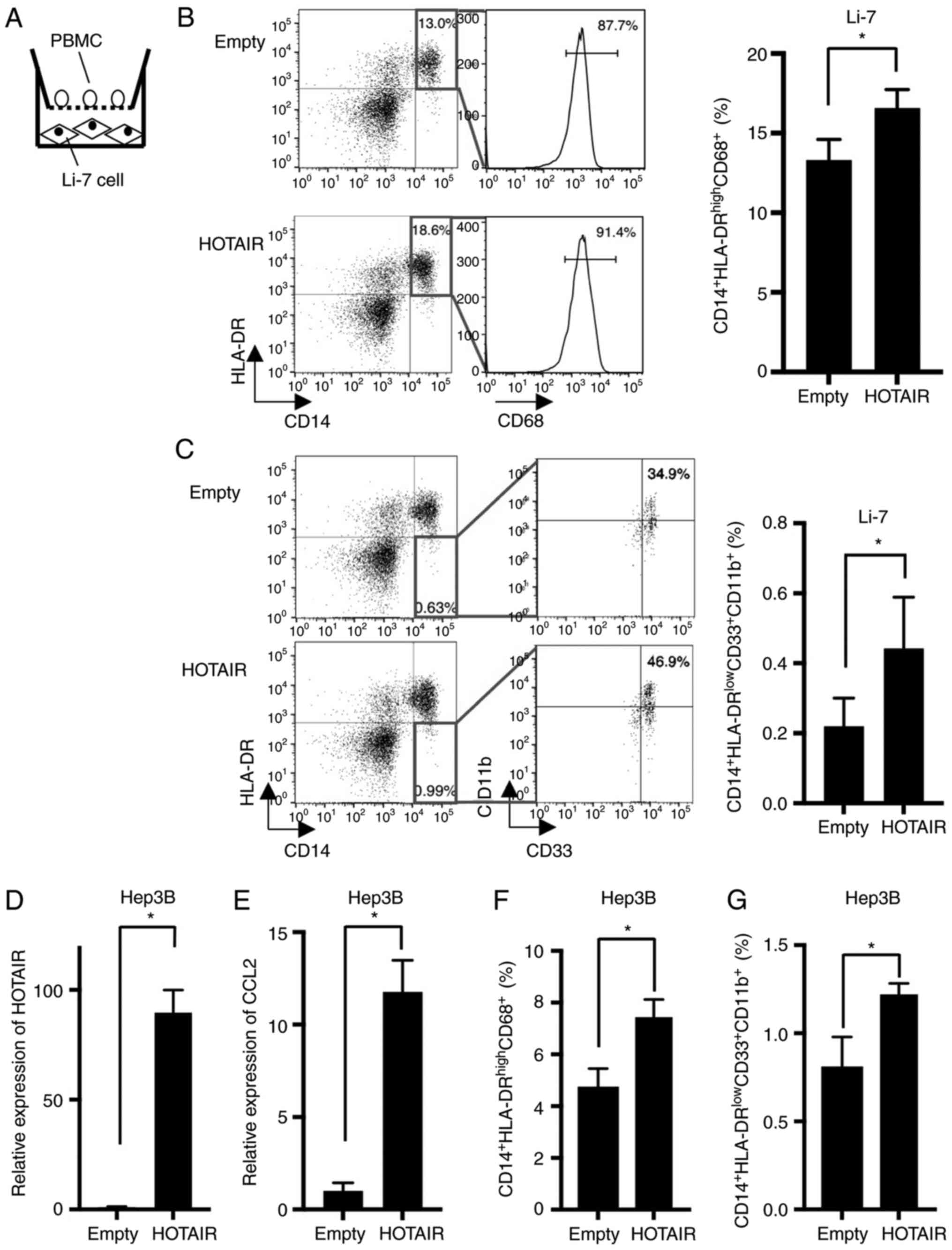
Because CCL2 is known to recruit TAMs and MDSCs, we
examined the effect of HOTAIR-overexpressing Li-7 cells against
PBMCs (Fig. 3A). The proportion of
CD14+HLA-DR+CD68+ macrophages were
increased in PBMCs co-cultured with HOTAIR-overexpressing cells
(Fig. 3B). Furthermore,
CD14+HLA-DR−CD33+CD11b+
MDSCs were also increased in co-cultured PBMCs (Fig. 3C).
To confirm the effect of HOTAIR against macrophages
and MDSCs, we established HOTAIR-overexpressing cells using another
HCC cell line, Hep3B. HOTAIR was expressed 89-fold higher in
HOTAIR-overexpressing Hep3B cells than in control cells (Fig. 3D). We measured the expression of CCL2,
and found that CCL2 was upregulated in the Hep3B-HOTAIR cells
(Fig. 3E). Then we investigated the
effect of HOTAIR-overexpressing Hep3B cells against PBMCs using
co-culture method, and found that the proportion of macrophages
(Fig. 3F) and MDSCs (Fig. 3G) were increased by co-cultured
HOTAIR-overexpressing Hep3B cells, which is compatible with the
data using Li-7 cells.
Collectively, these data suggested that HOTAIR in
HCC cell lines plays critical roles in the promotion of macrophage
and MDSC, through the secretion of cytokines and/or chemokines by
HCC cells.
Discussion
In this study, we demonstrated that
HOTAIR-overexpressing cells secreted higher CCL2 and promotes
macrophage/MDSC proliferation. Previous report indicated that CCL2
is required for recruitment of macrophage and accumulation of MDSCs
in the tumor microenvironment (16,18),
suggesting that high expression of HOTAIR in cancer cells recruit
TAMs/MDSCs through the secretion of CCL2, resulting in promoting
tumor growth and metastasis. To the best of our knowledge, this is
the first report that HOTAIR can affect the surrounding immune
cells through humoral factor such as CCL2. CCL2 is known to be
highly expressed in HCC, and believed to be a good target for
immunotherapy (16). To test whether
HOTAIR could be a specific target for immunotherapy against CCL2,
we analyzed the correlation between CCL2 and HOTAIR expression in
Gene Expression Omnibus (GEO) data set of HCC (GSE2109, n=40). We
found that the group consist of patients with high expression level
of both CCL2 and HOTAIR tended to associated with advanced clinical
stages (P=0.078, data not shown). This data could be compatible
with our findings in this study, although statistical significant
was not reached due to the small scale of cases. Further study will
be required to elucidate the role of HOTAIR in human tissue.
Several reports indicated the regulatory mechanisms
of CCL2 production: the transcription of the CCL2 gene is shown to
increase in response to stimuli, such as TNF or LPS, by the binding
of NF-kB dimers to two distal NF-kB binding sites in murine
fibroblasts (22). SP1 also regulates
the basal level transcription of the CCL2 gene by binding to a
GC-box located in the proximal region of the 5′-UTR (23). In this study, we demonstrated that
HOTAIR can regulate CCL2 production in a HCC cell line. Recent
study suggested that SP1 is a target of miR-326, and HOTAIR can
modulate the pathway of miR-326/SP1 pathway (24). Thus, we speculate that HOTAIR regulate
CCL2 through the transcriptional factors such as SP1.
Previous study suggested that introduction of human
HOTAIR into liver cancer cells revealed that HOTAIR promoted more
rapid proliferation compared to control HepG2 cells (7). Another report demonstrated that HOTAIR
knockdown dramatically inhibited cell viability and induced
G1-phase arrest in vitro and suppressed
tumorigenicity in vivo by promoting miR-218 expression in
HepG2 and Bel7404 cell lines. In this study, using Li-7 cell line,
no significant difference was found between HOTAIR-overexpressing
and control cells in cell proliferation, migration, and in
vivo tumorigenicity assay. Because the function of HOTAIR are
strongly cell type-dependent (5), we
speculate that the context of intracellular signaling is different
between HepG2 and other HCC cell lines. Further study will be
required to elucidate the cell type-dependent molecular function of
HOTAIR in types of HCC cell lines, and identify the critical
molecule(s) which cooperate with HOTAIR.
In conclusion, we newly identified CCL2 as a
downstream molecule of HOTAIR, which may be involved in the
recruitment of macrophages/MDSC in tumor microenvironment.
Acknowledgements
This work was supported by JSPS KAKENHI (grant nos.
15K09055 and 16K07132), Japan Agency for Medical Research and
Development, and Novartis Pharma Research Grants. This work was
partly supported by the Biomedical Research Core of Tohoku
University School of Medicine.
Glossary
Abbreviations
Abbreviations:
|
CCL2
|
chemokine (C-C motif) ligand 2
|
|
cDNA
|
complementary DNA
|
|
DMEM
|
Dulbeccos modified Eagles medium
|
|
ELISA
|
enzyme linked immunosorbent assay
|
|
FBS
|
fetal bovine serum
|
|
HOTAIR
|
HOX antisense intergenic RNA
|
|
HCC
|
hepatocellular carcinoma
|
|
lincRNA
|
large intervening non-coding RNA
|
|
MDSC
|
myeloid-derived suppressor cell
|
|
PBMC
|
peripheral blood mononuclear cell
|
|
PBS
|
phosphate buffered saline
|
|
PRC2
|
polycomb repressive complex 2
|
|
RPII
|
RNA polymerase II
|
|
TAM
|
tumor associated macrophage
|
References
|
1
|
Gupta RA, Shah N, Wang KC, Kim J, Horlings
HM, Wong DJ, Tsai MC, Hung T, Argani P, Rinn JL, et al: Long
non-coding RNA HOTAIR reprograms chromatin state to promote cancer
metastasis. Nature. 464:1071–1076. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Kogo R, Shimamura T, Mimori K, Kawahara K,
Imoto S, Sudo T, Tanaka F, Shibata K, Suzuki A, Komune S, et al:
Long noncoding RNA HOTAIR regulates polycomb-dependent chromatin
modification and is associated with poor prognosis in colorectal
cancers. Cancer Res. 71:6320–6326. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Nakagawa T, Endo H, Yokoyama M, Abe J,
Tamai K, Tanaka N, Sato I, Takahashi S, Kondo T and Satoh K: Large
noncoding RNA HOTAIR enhances aggressive biological behavior and is
associated with short disease-free survival in human non-small cell
lung cancer. Biochem Biophys Res Commun. 436:319–324. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Endo H, Shiroki T, Nakagawa T, Yokoyama M,
Tamai K, Yamanami H, Fujiya T, Sato I, Yamaguchi K, Tanaka N, et
al: Enhanced expression of long non-coding RNA HOTAIR is associated
with the development of gastric cancer. PLoS One. 8:e770702013.
View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Heubach J, Monsior J, Deenen R, Niegisch
G, Szarvas T, Niedworok C, Schulz WA and Hoffmann MJ: The long
noncoding RNA HOTAIR has tissue and cell type-dependent effects on
HOX gene expression and phenotype of urothelial cancer cells. Mol
Cancer. 14:1082015. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Chatterjee R and Mitra A: An overview of
effective therapies and recent advances in biomarkers for chronic
liver diseases and associated liver cancer. Int Immunopharmacol.
24:335–345. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Ishibashi M, Kogo R, Shibata K, Sawada G,
Takahashi Y, Kurashige J, Akiyoshi S, Sasaki S, Iwaya T, Sudo T, et
al: Clinical significance of the expression of long non-coding RNA
HOTAIR in primary hepatocellular carcinoma. Oncol Rep. 29:946–950.
2013. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Gao JZ, Li J, Du JL and Li XL: Long
non-coding RNA HOTAIR is a marker for hepatocellular carcinoma
progression and tumor recurrence. Oncol Lett. 11:1791–1798.
2016.PubMed/NCBI
|
|
9
|
Su DN, Wu SP, Chen HT and He JH: HOTAIR, a
long non-coding RNA driver of malignancy whose expression is
activated by FOXC1, negatively regulates miRNA-1 in hepatocellular
carcinoma. Oncol Lett. 12:4061–4067. 2016.PubMed/NCBI
|
|
10
|
Fu WM, Zhu X, Wang WM, Lu YF, Hu BG, Wang
H, Liang WC, Wang SS, Ko CH, Waye MM, et al: Hotair mediates
hepatocarcinogenesis through suppressing miRNA-218 expression and
activating P14 and P16 signaling. J Hepatol. 63:886–895. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Hanahan D and Weinberg RA: Hallmarks of
cancer: The next generation. Cell. 144:646–674. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Mantovani A: Effects on in vitro tumor
growth of murine macrophages isolated from sarcoma lines differing
in immunogenicity and metastasizing capacity. Int J Cancer.
22:741–746. 1978. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Noy R and Pollard JW: Tumor-associated
macrophages: From mechanisms to therapy. Immunity. 41:49–61. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Pyzer AR, Cole L, Rosenblatt J and Avigan
DE: Myeloid-derived suppressor cells as effectors of immune
suppression in cancer. Int J Cancer. 139:1915–1926. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Hoechst B, Ormandy LA, Ballmaier M, Lehner
F, Krüger C, Manns MP, Greten TF and Korangy F: A new population of
myeloid-derived suppressor cells in hepatocellular carcinoma
patients induces CD4(+)CD25(+)Foxp3(+) T cells. Gastroenterology.
135:234–243. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Li X, Yao W, Yuan Y, Chen P, Li B, Li J,
Chu R, Song H, Xie D, Jiang X and Wang H: Targeting of
tumour-infiltrating macrophages via CCL2/CCR2 signalling as a
therapeutic strategy against hepatocellular carcinoma. Gut.
66:157–167. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Teng KY, Han J, Zhang X, Hsu SH, He S,
Wani NA, Barajas JM, Snyder LA, Frankel WL, Caligiuri MA, et al:
Blocking the CCL2-CCR2 Axis Using CCL2-Neutralizing Antibody Is an
Effective Therapy for Hepatocellular Cancer in a Mouse Model. Mol
Cancer Ther. 16:312–322. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Chang AL, Miska J, Wainwright DA, Dey M,
Rivetta CV, Yu D, Kanojia D, Pituch KC, Qiao J, Pytel P, et al:
CCL2 produced by the glioma microenvironment is essential for the
recruitment of regulatory T cells and myeloid-derived suppressor
cells. Cancer Res. 76:5671–5682. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Yoshimura T: The production of monocyte
chemoattractant protein-1 (MCP-1)/CCL2 in tumor microenvironments.
Cytokine. pii:S1043–4666. 2017.
|
|
20
|
Kadota K, Nakai Y and Shimizu K: A
weighted average difference method for detecting differentially
expressed genes from microarray data. Algorithms Mol Biol. 3:82008.
View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Ping D, Boekhoudt GH, Rogers EM and Boss
JM: Nuclear factor-kappa B p65 mediates the assembly and activation
of the TNF-responsive element of the murine monocyte
chemoattractant-1 gene. J Immunol. 162:727–734. 1999.PubMed/NCBI
|
|
23
|
Ueda A, Okuda K, Ohno S, Shirai A,
Igarashi T, Matsunaga K, Fukushima J, Kawamoto S, Ishigatsubo Y and
Okubo T: NF-kappa B and Sp1 regulate transcription of the human
monocyte chemoattractant protein-1 gene. J Immunol. 153:2052–2063.
1994.PubMed/NCBI
|
|
24
|
Li J, Li S, Chen Z, Wang J, Chen Y, Xu Z,
Jin M and Yu W: miR-326 reverses chemoresistance in human lung
adenocarcinoma cells by targeting specificity protein 1. Tumour
Biol. 37:13287–13294. 2016. View Article : Google Scholar : PubMed/NCBI
|