Introduction
Lung cancer is one of the most common types of
cancer and the leading cause of cancer-associated mortality
worldwide (1,2). Non-small cell lung cancer (NSCLC) is the
main type of lung cancer and accounts for nearly 84% of cases.
Despite decades of intensive efforts in the cancer research field,
the prognosis of patients with NSCLC remains poor, with a 5-year
overall survival rate of ~10% (3).
Thus, further investigation of novel biomarkers and therapeutic
targets for NSCLC is urgently required (4).
The family of forkhead box (FOX) transcription
factors, which includes the FOXP subfamily, serves important roles
in oncogenesis. Notably, downregulation of FOXP4 markedly reduced
the NSCLC cell growth and invasion (5), while FOXP2 worked as a potential novel
tumor metastasis suppressor in breast cancer (6). In NSCLC, higher expression of FOXN1 has
been associated with improved survival times in patients (7). Using tissue arrays and
immunohistochemical analysis, FOXP2 has been observed to function
as a novel candidate tumor suppressor gene in different cancer
types. However, to our knowledge, the function of FOXN1 in cancer
remains unknown. In the present study, the aim was to evaluate the
role of FOXN1 in NSCLC.
EZH2 is overexpressed and associated with worse
overall survival rate in NSCLC patients (8). Nuclear β-catenin could serves as an
important prognosticator for patients with NSCLC (9).
Materials and methods
Cell lines and reagents
Normal human bronchial epithelial (HBE) cells, A549,
H1299, SPC-A1, H460, H520 and SK-MES-1 lung cancer cell lines were
purchased from the American Type Culture Collection (Manassas, VA,
USA). The cells were cultured at 37°C in a humidified atmosphere
with 5% CO2 in RPMI-1640 medium (Invitrogen; Thermo
Fisher Scientific, Inc., Waltham, MA, USA) supplemented with 10%
fetal bovine serum (FBS; Hyclone; Thermo Fisher Scientific, Inc.)
and 1% penicillin/streptomycin (Invitrogen; Thermo Fisher
Scientific, Inc.). Mouse anti-human FOXN1 polyclonal antibody (cat
no. sc-271256; 1:500), rabbit anti-human enhancer of zeste homolog
2 (EZH2) polyclonal antibody (cat no. sc-25383; 1:1,000), mouse
anti-human β-actin monoclonal antibody (cat. no. SC47778; 1:2,000)
horseradish peroxidase-labeled secondary antibodies including goat
anti-mouse IgG-horseradish peroxidase (cat no. sc-2005; 1:3,000)
and anti-rabbit IgG-horseradish peroxidase (cat no. sc-2004;
1:3,000) were purchased from Santa Cruz Biotechnology, Inc.
(Dallas, TX, USA).
Transfection for FOXN1
downregulation
Two different FOXN1-short hairpin RNA (shRNA)
ShFOXN1#1 TRCN0000013197
(CCGGCCTCCTGCTATGGGCAGACATCTCGAGATGTCTGCCCATAGCAGGAGGTTTTT)
ShFOXN1#2 TRCN0000013193
(CCGGGCTCACACTCATCCACACTTACTCGAGTAAGTGTGGATGAGTGTGAGCTTTTT) ShEZH2
TRCN0000040074
(CCGGGCTAGGTTAATTGGGACCAAACTCGAGTTTGGTCCCAATTAACCTAGCTTTTTG)
Shβ-catenin TRCN0000314991
(CCGGTTGTTATCAGAGGACTAAATACTCGAGTATTTAGTCCTCTGATAACAATTTTTG) or
non-targeting negative control shRNA was designed and synthesized
by Sigma-Aldrich (Merck KGaA; Darmstadt, Germany).
Transfection for FOXN1
overexpression
pcDNA3.1-FOXN1 was cloned by GenePharm Co., Ltd.
(Shanghai, China). pcDNA3.1 was used the control. Cultured cells
were subjected to transfection using Lipofectamine® 3000
(Invitrogen; Thermo Fisher Scientific, Inc.), according to the
manufacturer's protocol. A total of 20 µM relative shRNA or 5 µg
relative plasmid were used for transfection. Following incubation
at 37°C for 48–72 h, cells were collected and lysed to verify the
expression of target genes or proteins by reverse
transcription-quantitative polymerase chain reaction (RT-qPCR) and
western blot analysis, respectively.
Patients
A total of 60 NSCLC patients diagnosed and treated
at the People's Hospital of Guizhou Province (Guiyang, China) were
enrolled into the present study. Written informed consent was
obtained from each patient, and the current study was approved by
the Ethics Committee of the People's Hospital of Guizhou Province.
Tumor tissues and the adjacent normal tissues were attained during
surgery between February 2010 and December 2014. Tissue samples
were frozen and stored at −80°C for RT-PCR. Histological and
pathological diagnoses were performed by the pathology department.
The tissues were embedded in paraffin and cut into 2 µm sections.
Slides were incubated in three washes of xylene for 5 min each,
followed by two washes of 100% ethanol for 10 min, 95% ethanol for
10 min and ddH2O for 5 min. Then the samples were
blocked in 10% normal goat serum in PBS at room temperature for 30
min and incubated at 4°C overnight in primary antibody solution of
anti-FOXN1 (1:100) The EnVision Detection System kit (Dako; Agilent
Technologies, Inc., Santa Clara, CA, USA) was used to visualize the
3,3′-diaminobenzidine chromogen (room temperature for 20 min).
Samples were counterstained with hematoxylin (0.2%) at room
temperature for 5 min (Zhongshan Golden Bridge Biotechnology
Company) according to the Staging Lung Cancer's Revised
International System (10). Neutral
gum was used to cover the slides and they were dried at room
temperature. Staining was visualized under an Olympus optical
microscope (Olympus Corporation, Tokyo, Japan) at ×20 magnification
for analysis.
Western blot analysis
Lung cancer cells were harvested and lysed using
0.5% NP-40 cell lysis buffer (Sigma-Aldrich; Merck KGaA) for 20 min
on ice, and transparent protein lysates were obtained following
centrifugation at 12,000 × g for 10 min at 4°C. The Bradford assay
reagent (Thermo Fisher Scientific, Inc.) was then used to determine
the protein concentration in the lysates. Next, 30 µg protein from
each group was separated by 10% SDS-PAGE and then transferred onto
polyvinylidene difluoride membranes. Subsequently, 5% skim milk was
used to block nonspecific binding in the samples at room
temperature for 1 h, followed by addition of the primary antibodies
against FOXN1, EZH2 and β-actin for overnight incubation at 4°C.
β-actin served as the internal control. The membranes were then
washed with Tris-buffered saline/Tween 20 for 5 min and five times,
incubated with respective secondary antibodies goat anti-mouse
IgG-horseradish peroxidase (1:3,000) and anti-rabbit
IgG-horseradish peroxidase (1:3,000) at room temperature for 1 h
and visualized using an enhanced chemiluminescence-based methods
(western blotting detection system; Thermo Fisher Scientific,
Inc.).
RNA extraction and RT-qPCR
Total RNA was extracted from the tissues or cells
using TRIzol reagent (Invitrogen; Thermo Fisher Scientific, Inc.),
according to the manufacturer's protocol. The RNA concentration was
assessed using a nanophotometer UV/VIS spectrophotometer (Implen
GmbH, München, Germany). A total of 2 µg RNA was converted into
cDNA using Revert Aid™ First Strand cDNA Synthesis kit
(Fermentas; Thermo Fisher Scientific, Inc.). The qPCR reaction was
subsequently performed on a 7500 Real-Time Detection System
(Applied Biosystems; Thermo Fisher Scientific, Inc.) according to
the following conditions: Initial step, 95°C for 3 min; second
step, 95°C for 10 sec, 60°C for 30 sec and 72°C for 10 sec for a
total of 38 cycles. The obtained data were analyzed using the
2−ΔΔCq method, (11) with
GAPDH used as an internal control. The primers used were as
follows: GAPDH, forward 5′-GAGAAGTATGACAACAGCCTC-3′ and reverse
5′-ATGGACTGTGGTCATGAGTC-3′; FOXN1, forward
5′-GCTCCTCACACTATCAGTACC-3′ and reverse
5′-AAGATGAGGATGCTGTAGGA-3′.
Invasion assay
Transwell migration chambers (BD Biosciences,
Franklin Lakes, NJ, USA) were used to measure the invasive capacity
of cells. Briefly, 100 µl Matrigel (BD Biosciences) was coated onto
8-µm filter chambers and plated on 24-well plates. A total of
2×104 cells were resuspended using 100 µl Dulbecco's
modified Eagle's medium (DMEM; Invitrogen; Thermo Fisher
Scientific, Inc.) basal medium without FBS and seeded into the
upper chamber, while 500 µl DMEM with 10% FBS was added into the
lower chamber. Following incubation at 37°C in a 5% CO2
atmosphere for 12 h, the medium was removed and the cells were
washed twice with cool PBS to remove cells from the upper chamber.
Subsequent to fixing with 4% paraformaldehyde for 15 min, the
invading cells in the lower chamber were stained with 1% crystal
violet for 15 min. The number of cells was counted under an upright
light microscope in 10 different fields of view (Leica DM4B; Leica
Microsystems, Shanghai, China). At least three independent
experiments were performed.
MTT assay
Cell proliferation was determined using an MTT
proliferation kit (Beijing Solarbio Science & Technology Co.,
Ltd., Beijing, China). The transfected cells were subjected to
centrifugation at 800 × g for 5 min at 4°C, followed by
resuspension with complete medium. Next, the cells were seeded into
a 96-well plate at a density of 2×104 cells per well.
Following incubation for 24, 48 and 72 h, MTT reagent was added and
an MTT assay was performed following the manufacturer's protocol.
Each group was examined in three separate wells.
Quantitative chromatin
immunoprecipitation (qChIP) analysis
The qChIP assay was performed using the Chip-IT
Express kit (Active Motif, Carlsbad, CA, USA), according to the
manufacturer's protocol. qPCR was performed with 5 µl of the
immunoprecipitated target DNA, 1 µl primers and 9 µl mixture (1 µl
enzyme, 2 µl dNTP and 6 µl SYBR green solution buffer; all included
in the ChIP-IT kit). The primers used were as follows: EZH2,
forward, 5′-GAGGCATGAGAATCGCTTGA-3′, and reverse,
5′-GCCGGACCCGTTACTACTTT-3′; β-catenin, forward,
5′-CAGTTGGCATTACCACTTAT-3′, and reverse,
5′-TTCACTCATGGAGGTAGGAT-3′. The PCR amplification was performed at
95°C for 5 min, followed by 35 cycles of 95°C for 20 sec, 55°C for
20 sec and 72°C for 30 sec. The 2−ΔΔCq method of
quantification was used, with GAPDH used as an internal control, as
per the RT-qPCR section.
Dual-luciferase reporter assay
A dual-luciferase reporter gene system (Promega
Corporation, Madison, WI, USA) was used to verify the direct target
genes for FOXN1, according to the manufacturer's protocol. Briefly,
the full length of EZH2 promoter was synthesized by clonal
expansion and cloned into the pmirGLO luciferase gene vector
(Promega Corporation). FOXN1 and control plasmids were
co-transfected into the cells along with the luciferase reporter
vectors using Lipofectamine 2000 (Invitrogen; Thermo Fisher
Scientific, Inc.). After 36 h at 37°C, the expression of
Renilla (Takara Biotechnology Co., Ltd., Dalian, China) was
used as an internal reference to adjust the number of cells and the
transfection efficiency differences.
Statistical analysis
All experiments were performed at least three times.
The data were analyzed using SPSS version 17.0 (SPSS, Inc.,
Chicago, IL, USA). Data was expressed as the mean ± standard
deviation of three independent experiments. Differences between the
experimental and control groups were compared using Student's
t-test to compare differences between two groups. A two-way
analysis of variance was performed in cases where more than two
groups were compared. The post hoc test employed was Tukey's range
test. P<0.05 was considered to indicate a statistically
significant difference.
Results
FOXN1 is downregulated in NSCLC
tissues and cells
In order to understand the role of FOXN1 in NSCLC,
60 pairs of tissue samples were collected from NSCLC patients, and
the mRNA expression of FOXN1 was determined by RT-qPCR. As
illustrated in Fig. 1A, the
expression of FOXN1 was significantly downregulated in tumor
tissues when compared with the paired noncancerous tissues
(P<0.05). Furthermore, FOXN1 mRNA expression was detected in six
NSCLC cell lines, namely A549, H1299, SPC-A1, H460, H520 and
SK-MES-1, and normal human bronchial epithelial (HBE) cells. It was
observed that the mRNA level of FOXN1 was lower in the NSCLC cell
lines, with a significant effect observed for A549, H1299 and H520
cells, as compared with the HBE cells (P<0.05; Fig. 1B). A549 and H1299 cells were used for
subsequent experiments.
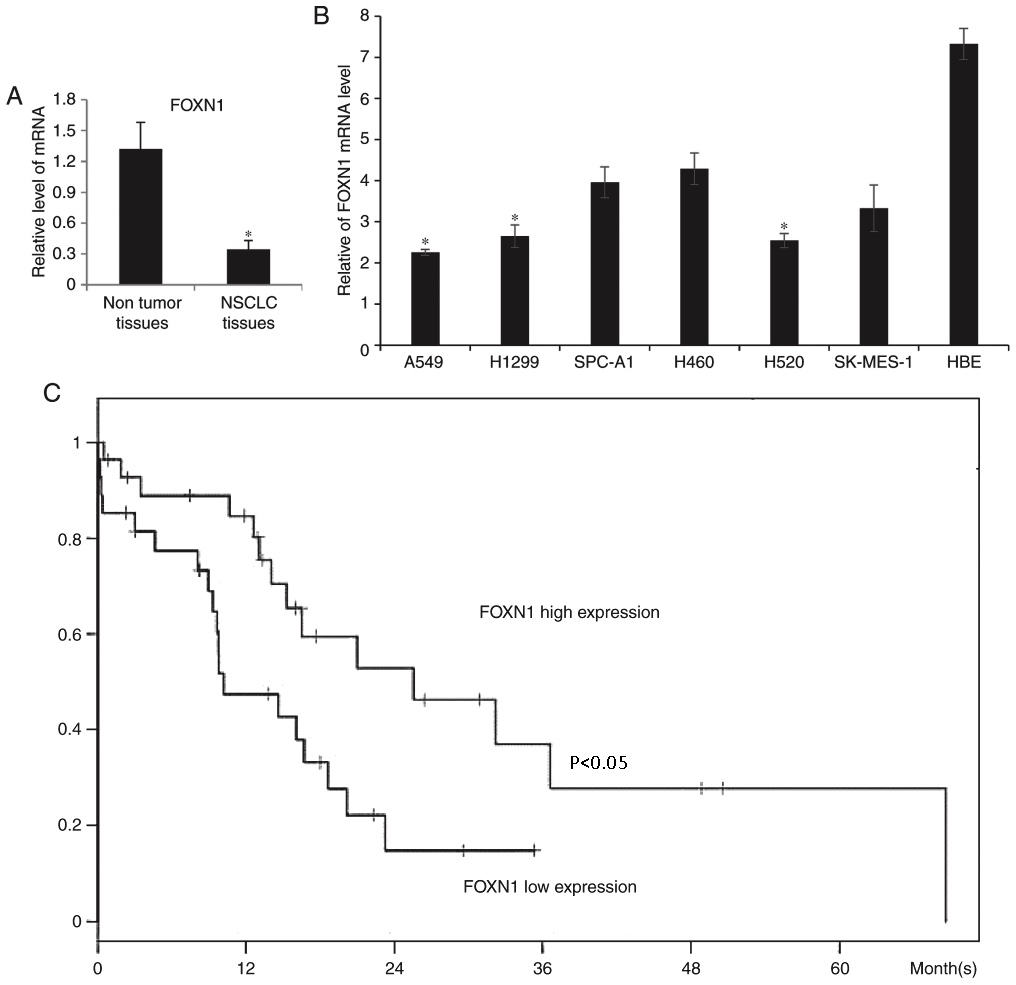 | Figure 1.Increased expression of FOXN1 is
associated with improved prognosis of NSCLC patients. (A) NSCLC
tissues and paired noncancerous lung tissues (n=60), as well as (B)
NSCLC cell lines (A549, H1299, SPC-A1, H460, H520 and SK-MES-1) and
normal HBE cells, were examined by reverse
transcription-quantitative polymerase chain reaction to measure the
FOXN1 mRNA levels. *P<0.05 vs. HBE cells. (C) Kaplan-Meier
curves were constructed to measure the patient survival rate, and
Cox log-rank test was used to measure the prognostic significance.
The x-axis represents the survival months, while the y-axis
represents the survival probability. (P<0.05). FOXN1, forkhead
box N1; NSCLC, non-small cell lung cancer; HBE, human bronchial
epithelial cells. |
Higher expression of FOXN1 is
associated with better prognosis in NSCLC patients
To further determine the function of FOXN1 during
NSCLC development and progression, the association of FOXN1
expression with the NSCLC patient clinicopathological
characteristics, including the gender, age, tumor size,
differentiation, metastasis and TNM staging, was analyzed. As shown
in Table I, a higher expression of
FOXN1 was associated with smaller tumor size, absence of metastasis
and lower TNM stage (P<0.05), which indicated an improved
prognosis. Furthermore, Kaplan-Meier curves were constructed to
analyze the NSCLC patient survival rate. As shown in Fig. 1C, a higher FOXN1 expression was
correlated with a better overall survival rate (P<0.05).
 | Table I.Clinicopathological variables in 60
non-small cell lung cancer patients. |
Table I.
Clinicopathological variables in 60
non-small cell lung cancer patients.
|
|
| FOXN1 protein
expression |
|
|---|
|
|
|
|
|
|---|
| Variables | Patient no.
(n=60) | Low (n=37) | High (n=23) | P-value |
|---|
| Age (years) |
|
|
| 0.887 |
|
<50 | 32 | 20 | 12 |
|
| ≥50 | 28 | 17 | 11 |
|
| Gender |
|
|
| 0.210 |
| Male | 33 | 18 | 15 |
|
|
Female | 27 | 19 | 8 |
|
| Tumor size |
|
|
| 0.017 |
| Small (≤3
cm) | 30 | 14 | 16 |
|
| Large (≥3
cm) | 30 | 23 | 7 |
|
| Metastasis |
|
|
| 0.002 |
|
Present | 36 | 28 | 8 |
|
|
Absent | 24 | 9 | 15 |
|
| TNM stage |
|
|
| <0.001 |
| I +
II | 28 | 10 | 18 |
|
| III +
IV | 32 | 27 | 5 |
|
| Differentiation |
|
|
| 0.580 |
|
Well/moderate | 34 | 22 | 12 |
|
|
Poor | 26 | 15 | 11 |
|
FOXN1 suppresses NSCLC cell
proliferation
In order to determine the role of FOXN1 in NSCLC
cells, the effect of FOXN1 overexpression on A549 and H1299 cell
proliferation was investigated. Initially, two specific shRNAs
targeted against FOXN1 (namely shFOXN1-1 and shFOXN1-2) were used
to reduce the expression of FOXN1 in NSCLC cells. In addition,
stable A549 or H1299 cell lines with overexpression of FOXN1 were
established by plasmid transfection. The cells transfected with
pcDNA3.1 plasmid was named as vector group, while the cells
transfected with FOXN1 plasmid was named as FOXN1. The successful
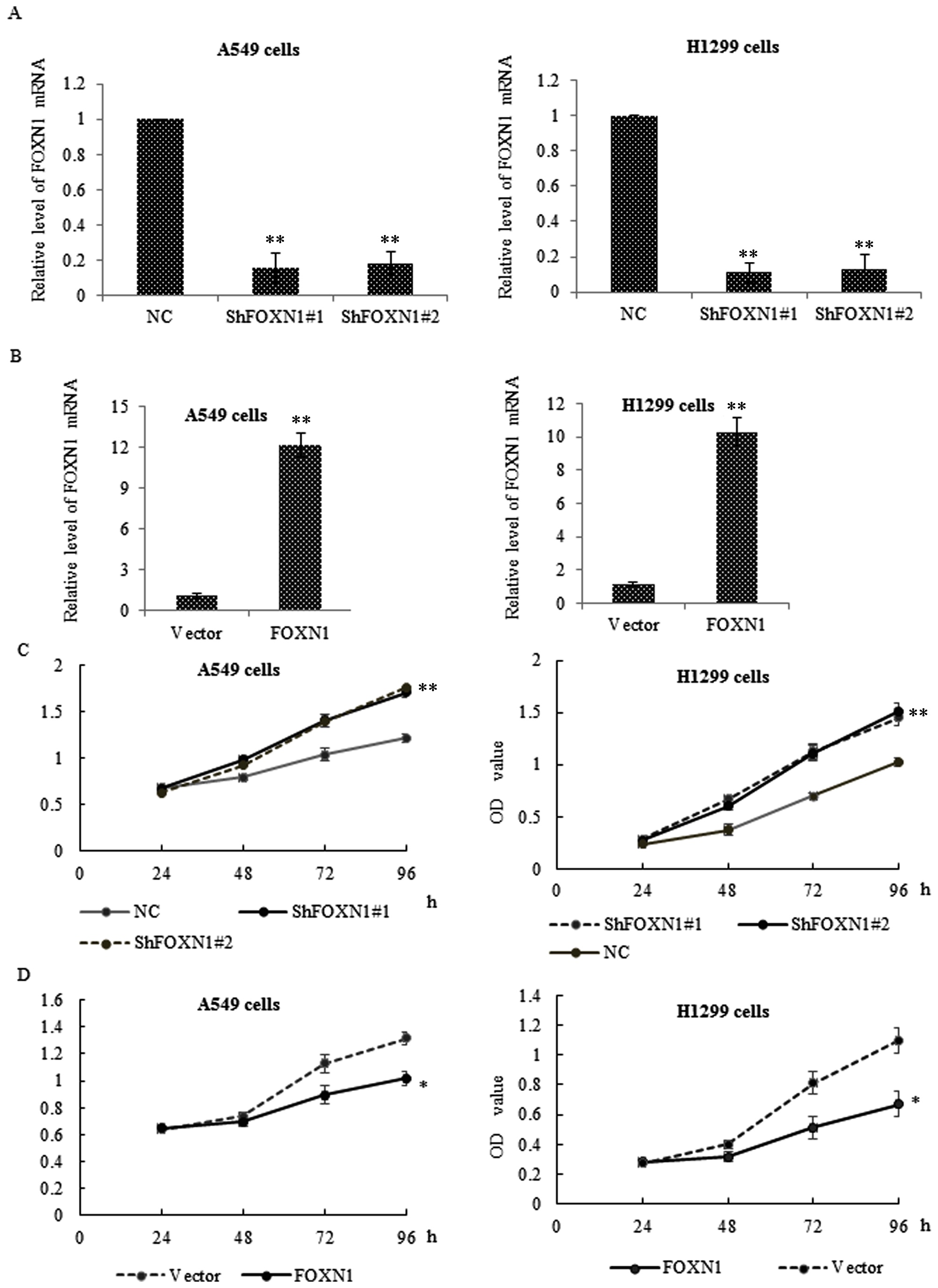
knockdown (Fig. 2A) and
overexpression (Fig. 2B) of FOXN1 in
the cells was confirmed by RT-qPCR. Subsequently, an MTT assay was
performed, which revealed that the cell growth was significantly
promoted in cells transfected with shFOXN1 for expression knockdown
as compared with the control cells (Fig.
2C). By contrast, ectopic overexpression of FOXN1 in the A549
and H1299 cells markedly suppressed the cell growth (Fig. 2D).
FOXN1 inhibits NSCLC cell
invasion
To examine whether FOXN1 exerts an effect on the
A549 and H1299 cell invasive ability, a Transwell assay was
performed. In this assay, the number of cells that migrated from
the upper to the lower side of the membrane in the chambers was
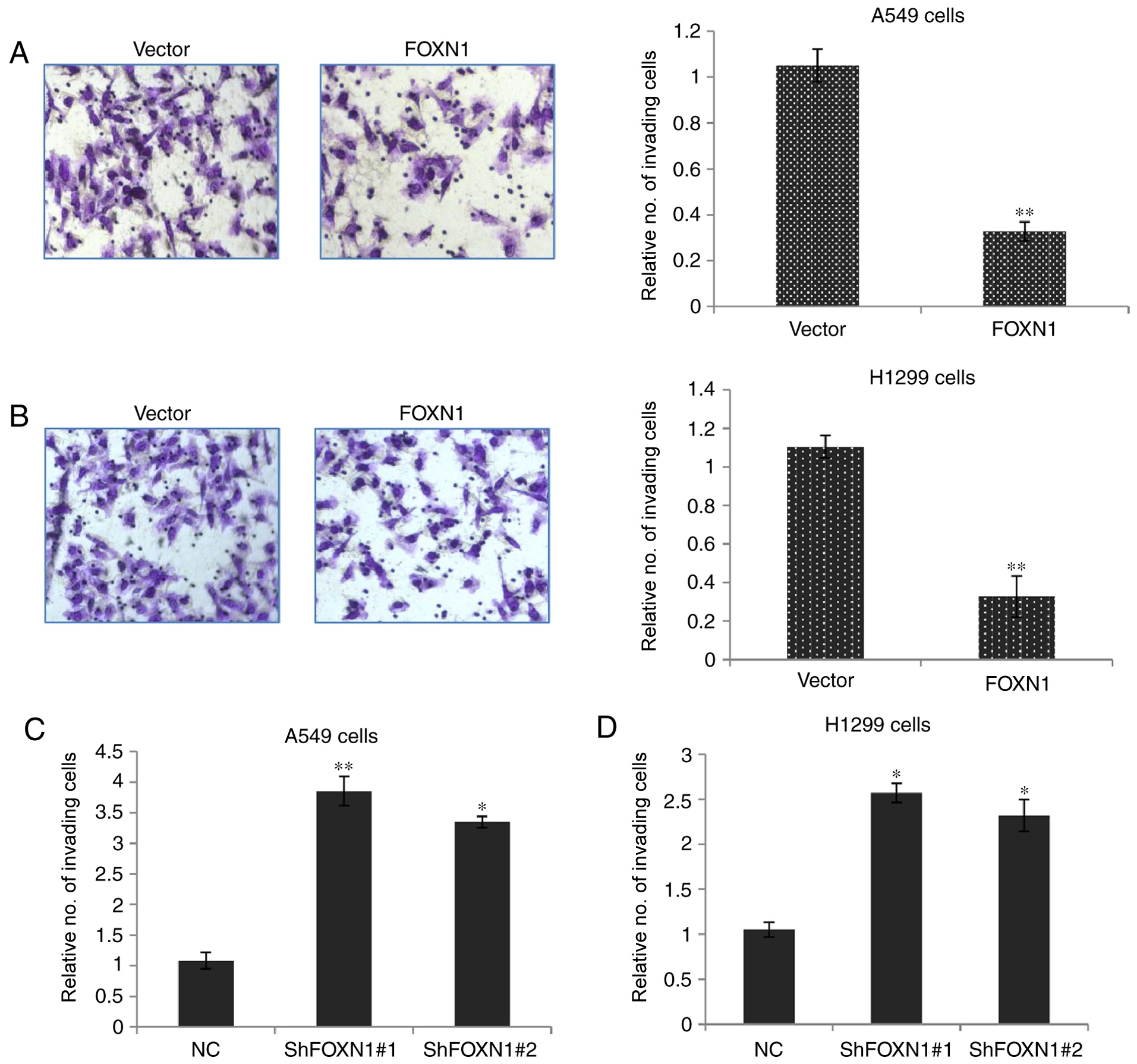
calculated. As shown in Fig. 3A and
B, the number of invading cells was markedly decreased in the
FOXN1 overexpression plasmid-transfected cells as compared with the
control group. By contrast, a reduced expression of FOXN1
demonstrated the potential to promote cell invasion in A549
(Fig. 3C) and H1299 cells (Fig. 3D), respectively. These results
suggested that the overexpression of FOXN1 clearly suppressed the
invasion ability of A549 and H1299 cells.
EZH2 and β-catenin are novel target
genes of FOXN1
In order to understand the underlying mechanisms of
FOXN1 in suppressing NSCLC, a quantitative chromatin
immunoprecipitation (ChIP) assay was performed in A549 and H1299
cells, with normal IgG used as a negative control. Different genes
involved in proliferation and invasion were selected for
investigation, including E-cadherin, N-cadherin, r-catenin,
β-catenin, caspase-3, caspase-7, hypoxia-inducible factor 1α
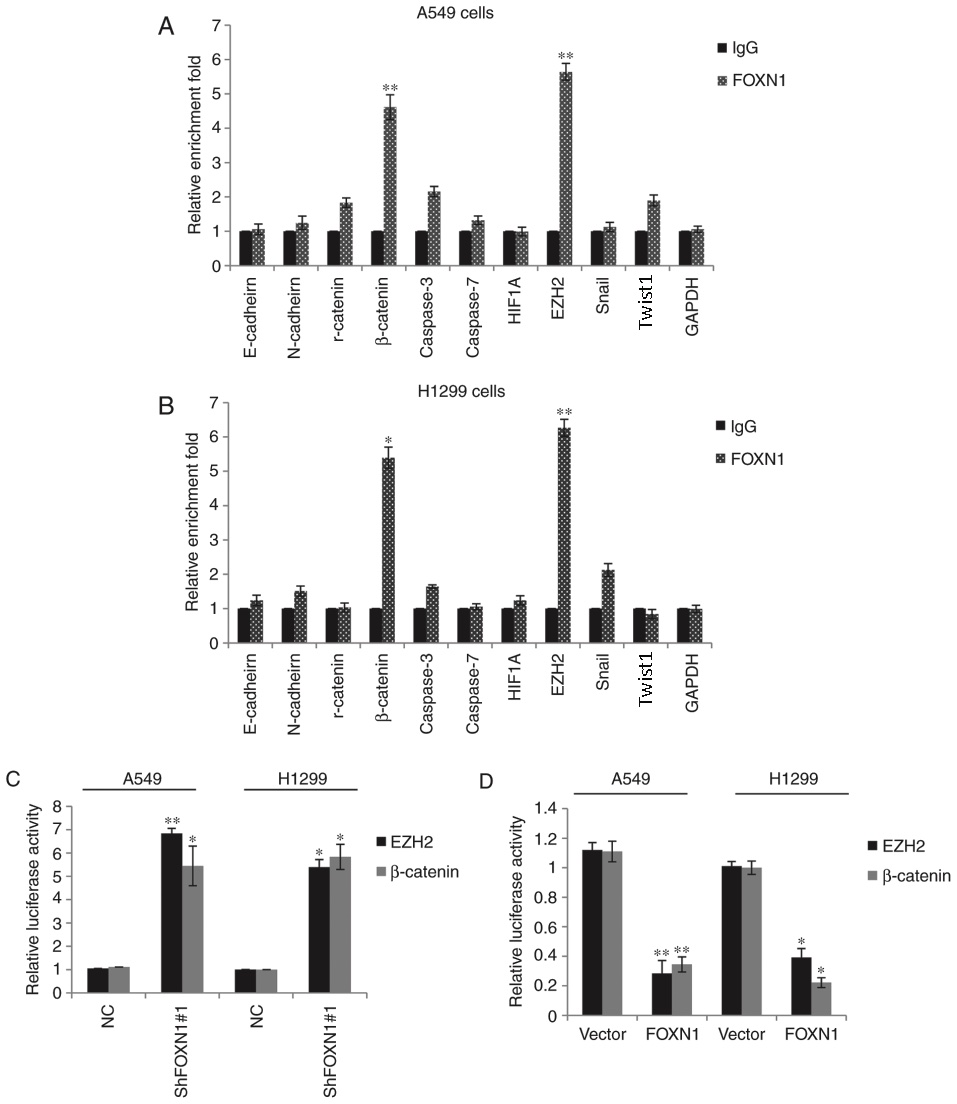
(HIF1A), EZH2, Snail and Twist1. As shown in Fig. 4A, in A549 cells, the bindings of FOXN1
to E-cadherin and β-catenin promoters was were significantly higher
when compared with that of the normal IgG. Similar binding
enrichment was also detected in H1299 cells (Fig. 4B). FOXN1 is known to typically
function as a transcription repressor factor (12); thus, to further support this
observation, a dual-luciferase reporter assay was conducted in A549
and H1299 cells. Cells co-transfected with shFOXN1 along with EZH2
or β-catenin promoter-driven luciferase reporters exhibited a
moderately enhanced luciferase activity, as compared with cells
transfected with Renilla and pcDNA3.1 plasmids (Fig. 4C). By contrast, in the FOXN1
overexpression group, reduced luciferase activity was detected
(Fig. 4D).
Inhibition of the EZH2/β-catenin
cascade is involved in the suppressive effect of FOXN1 on
NSCLC
Consistent with the results of the dual-luciferase
reporter assay, in A459 and H1299 cells subjected to FOXN1
knockdown, significantly increased mRNA levels of EZH2 and
β-catenin were confirmed by RT-qPCR assay. By contrast, ectopic
overexpression of FOXN1 in A549 and H1299 cells markedly suppressed
the EZH2 and β-catenin mRNA levels (Fig.
5A). Furthermore, as shown in Fig.
5B, shFOXN1 transfection increased the protein levels of EZH2
and β-catenin, while overexpression of FOXN1 decreased the protein
levels of EZH2 and β-catenin.
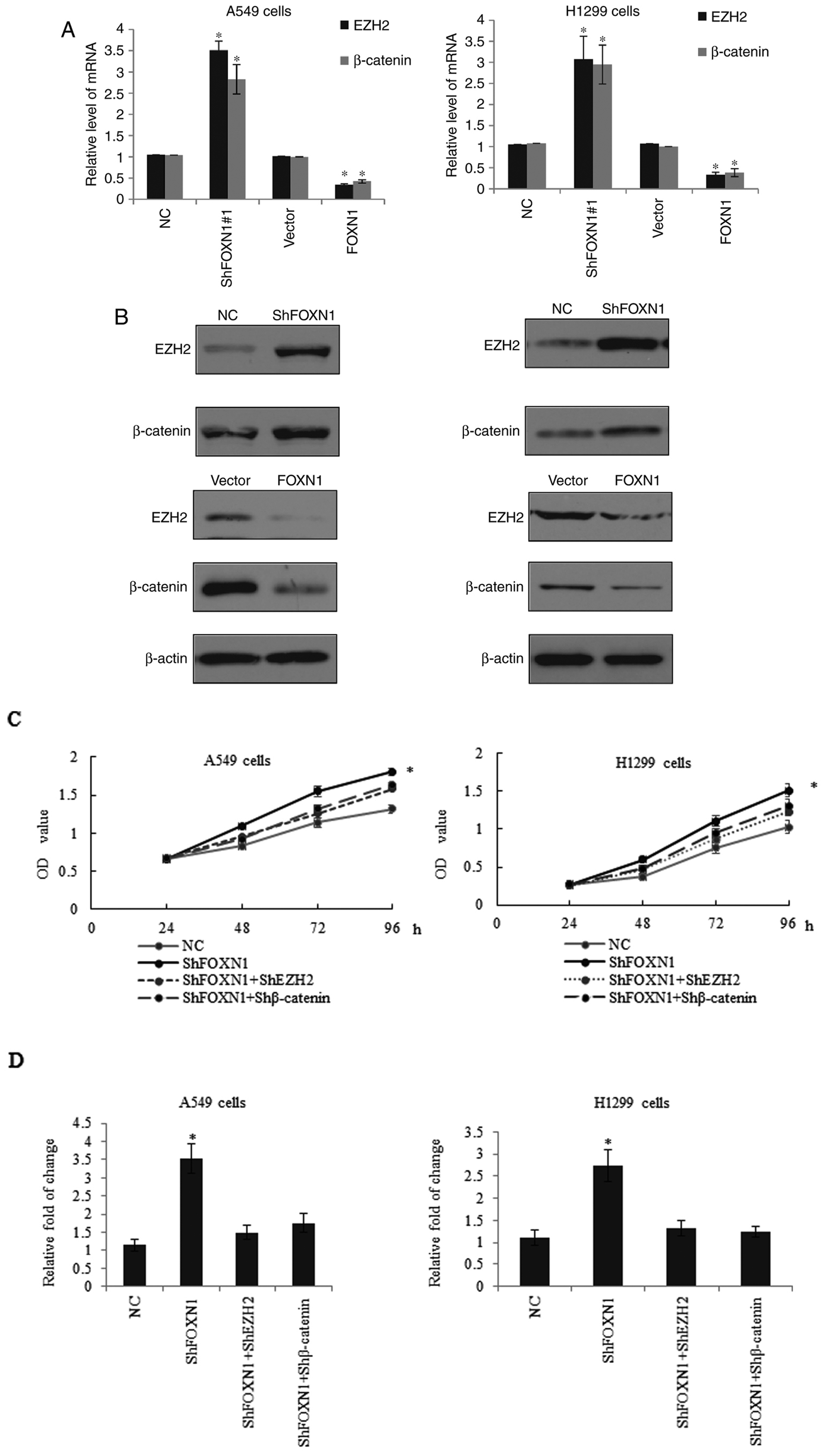 | Figure 5.Inhibition of the EZH2/β-catenin
cascade is involved in the suppressive effect of FOXN1 on NSCLC.
A549 or H1299 cells were transfected with NC shRNA, shRNA targeting
FOXN1, FOXN1 overexpression construct or a control vector. (A) EZH2
and β-catenin mRNA levels were measured by reverse
transcription-quantitative polymerase chain reaction, while (B) the
relative protein levels were detected by western blot analysis. (C)
MTT and (D) Transwell assays were performed in A549 cells or H1299
cells transfected with NC, shFOXN1, shFOXN1 + shEZH2, or shFOXN1 +
shβ-catenin. Data are represented as the mean ± standard deviation
of three independent experiments. *P<0.05 vs. corresponding
control group. FOXN1, forkhead box N1; shRNA, short hairpin RNA;
NC, negative control shRNA; EZH2, enhancer of zeste homolog 2; OD,
optical density. |
To establish further insight into the mechanism, the
study subsequently examined whether EZH2 and β-catenin were
involved in the suppressive effect of FOXN1 on NSCLC cell
proliferation and invasion. An MTT assay was performed, and
revealed that the additional knockdown of EZH2 or β-catenin by
shRNA transfection was observed to partially rescue the effect of
FOXN1 knockdown on the proliferation potential of A549 and H1299
cells, as compared with the cells with FOXN1 knockdown alone
(Fig. 5C). In the Transwell assay,
A549 or H1299 cells co-transfected with shFOXN1 + shEZH2 or shFOXN1
+ shβ-catenin demonstrated partial alleviation of the effect of
FOXN1 knockdown on the NSCLC cell invasive potential (Fig. 5D).
Discussion
Cell proliferation and metastasis are two important
malignant characteristics in NSCLC, which are accompanied by the
dynamic changes in the gene expression (13). The results of the present study
demonstrated that FOXN1 expression regulated the NSCLC cell growth
and invasion. Overexpression of FOXN1 could inhibit the NSCLC
progression. In the NSCLC patients, higher expression of FOXN1 is
associated with better prognosis. To our knowledge, this is the
first report about the function of FOXN1 in NSCLC.
The current study further revealed that FOXN1 served
major roles in NSCLC invasion by directly targeting EZH2 and
β-catenin therefore suppressing EZH2 and β-catenin expression. By
epigenetically silencing tumor suppressor genes, EZH2 has been
reported to promote carcinogenesis, and overexpression of EZH2 was
correlated with NSCLC cell migration (14). Furthermore, increased expression of
EZH2 has been correlated with lymph node metastasis, tumor size and
TNM stage in NSCLC (15). It has also
been reported that the expression of EZH2 was significantly higher
in NSCLC brain metastases (16). A
previous study reported that inhibition of EZH2 sensitized BRG1 and
epidermal growth factor receptor mutant lung tumors to
topoisomerase II inhibitors, while knockdown of EZH2 reversed the
cisplatin-resistance of NSCLC and gastric cancer cells (17). The regulation of EZH2 has been
revealed by numerous studies in recent years; for instance,
microRNA-138 inhibited tumor growth through repression of EZH2 in
NSCLC and osteosarcoma cells (18,19).
Furthermore, the long non-coding RNA MALAT1 enhanced the oncogenic
activities of EZH2 in castration-resistant prostate cancer
(20). These aforementioned findings
regarding EZH2 were consistent with the observations of the current
study. Furthermore, the present results reported that FOXN1 may be
a novel regulator of EZH2 in NSCLC. The inhibition of EZH2 by FOXN1
is of importance in the NSCLC suppression.
It has previously been reported that overexpression
of β-catenin was associated with poor overall survival rates in
patients with Stage IA-IIA squamous cell lung cancer irrespective
of adjuvant chemotherapy (21). By
activating the β-catenin signaling pathway, DEAD-box helicase 5
promoted the proliferation and tumorigenesis of NSCLC (22). In addition, through promoting
β-catenin degradation, Shisa3 was associated with prolonged
survival times in lung cancer (23).
The majority of these studies focused on the negative regulation of
β-catenin; however, transcriptional repression may also be another
important mechanism for β-catenin inhibition (24,25). The
present study reported that FOXN1 was a negative regulator of
β-catenin, thus contributing toward NSCLC repression. Notably, it
was observed that FOXN1 served major roles in NSCLC proliferation
and invasion by directly targeting EZH2 and β-catenin, which
suggested that FOXN1 may be a multifunctional regulator in
NSCLC.
In conclusion, the present study revealed that FOXN1
inhibited the NSCLC progression and served as a tumor suppressor.
EZH2 and β-catenin may be potential targets for the development of
anti-proliferation and anti-invasion in the treatment of NSCLC.
However, further investigation is required in order to determine
whether the current results may be translated into the development
of valuable NSCLC prognosis biomarkers.
Acknowledgements
Not applicable.
Funding
No funding was received.
Availability of data and materials
The datasets used and/or analyzed during the current
study are available from the corresponding author on reasonable
request.
Authors' contributions
XJ conceived and designed the work. WW analyzed the
data provided technical assistance for the transwell assay. XX and
YJ analyzed the data. XJ wrote the paper. All authors read and
approved the final manuscript.
Ethics approval and consent to
participate
All human tissues are collected under Institutional
Review Committee (IRB) and Health Insurance Portability and
Accountability Act (HIPAA) approved protocols.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Jemal A, Siegel R, Ward E, Hao Y, Xu J,
Murray T and Thun MJ: Cancer statistics, 2008. CA Cancer J Clin.
58:71–96. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Van Z, Wijk N and Fong KM: Update in lung
cancer: Prologue to a modern review series. Respirology.
20:183–184. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
O'Dowd EL and Baldwin DR: Early diagnosis
pivotal to survival in lung cancer. Practitioner. 258(21–24): 2–3.
2014.
|
|
4
|
Sharma SP: New therapeutic target for
non-small-cell lung cancer. Lancet Oncol. 15:e5332014. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Yang T, Li H, Thakur A, Chen T, Xue J, Li
D and Chen M: FOXP4 modulates tumor growth and independently
associates with miR-138 in non-small cell lung cancer cells. Tumour
Biol. 36:8185–8191. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Cuiffo BG and Karnoub AE: Silencing FOXP2
in breast cancer cells promotes cancer stem cell traits and
metastasis. Mol Cell Oncol. 3:e10190222015. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Feng J, Zhang X, Zhu H, Wang X, Ni S and
Huang J: High expression of FoxP1 is associated with improved
survival in patients with non-small cell lung cancer. Am J Clin
Pathol. 138:230–235. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Liu H, Li W, Yu X, Gao F, Duan Z, Ma X,
Tan S, Yuan Y, Liu L, Wang J, et al: EZH2-mediated Puma gene
repression regulates non-small cell lung cancer cell proliferation
and cisplatin-induced apoptosis. Oncotarget. 7:56338–56354.
2016.PubMed/NCBI
|
|
9
|
Jin J, Zhan P, Katoh M, Kobayashi SS, Phan
K, Qian H, Li H, Wang X, Wang X and Song Y: written on behalf of
the AME Lung Cancer Collaborative Group: Prognostic significance of
β-catenin expression in patients with non-small cell lung cancer: A
meta-analysis. Transl Lung Cancer Res. 6:97–108. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Zell JA, Ou Ignatius SH, Ziogas A and
Anton-Culver H: Validation of the proposed International
Association for the Study of Lung Cancer non-small cell lung cancer
staging system revisions for advanced bronchioloalveolar carcinoma
using data from the California Cancer Registry. J Thorac Oncol.
2:1078–1085. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Palamaro L, Romano R, Fusco A, Giardino G,
Gallo V and Pignata C: FOXN1 in organ development and human
diseases. Int Rev Immunol. 33:83–93. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Mittal V: Epithelial mesenchymal
transition in aggressive lung cancers. Adv Exp Med Biol. 890:37–56.
2016. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Xu C, Hou Z, Zhan P, Zhao W, Chang C, Zou
J, Hu H, Zhang Y, Yao X, Yu L and Yan J: EZH2 regulates cancer cell
migration through repressing TIMP-3 in non-small cell lung cancer.
Med Oncol. 30:7132013. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Geng J, Li X, Zhou Z, Wu CL, Dai M and Bai
X: EZH2 promotes tumor progression via regulating VEGF-A/AKT
signaling in non-small cell lung cancer. Cancer Lett. 359:275–287.
2015. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Behrens C, Solis LM, Lin H, Yuan P, Tang
X, Kadara H, Riquelme E, Galindo H, Moran CA, Kalhor N, et al: EZH2
protein expression associates with the early pathogenesis, tumor
progression, and prognosis of non-small cell lung carcinoma. Clin
Cancer Res. 19:6556–6565. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Zhou W, Wang J, Man WY, Zhang QW and Xu
WG: siRNA silencing EZH2 reverses cisplatin-resistance of human
non-small cell lung and gastric cancer cells. Asian Pac J Cancer
Prev. 16:2425–2430. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Zhu Z, Tang J, Wang J, Duan G, Zhou L and
Zhou X: MiR-138 acts as a tumor suppressor by targeting EZH2 and
enhances cisplatin-induced apoptosis in osteosarcoma cells. PLoS
One. 11:e01500262016. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Zhang HJ, Zhang H, Zhao M, Lv Z, Zhang X,
Qin X, Wang H, Wang S, Su J, Lv X, et al: MiR-138 inhibits tumor
growth through repression of EZH2 in non-small cell lung cancer.
Cell Physiol Biochem. 31:56–65. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Wang D, Ding L, Wang L, Zhao Y, Sun Z,
Karnes RJ, Zhang J and Huang H: LncRNA MALAT1 enhances oncogenic
activities of EZH2 in castration-resistant prostate cancer.
Oncotarget. 6:41045–41055. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Kim Y, Jin D, Lee BB, Cho EY, Han J, Shim
YM, Kim HK and Kim DH: Overexpression of β-catenin and cyclin D1 is
associated with poor overall survival in patients with stage IA-IIA
squamous cell lung cancer irrespective of adjuvant chemotherapy. J
Thorac Oncol. 11:2193–2201. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Wang Z, Luo Z, Zhou L, Li X, Jiang T and
Fu E: DDX5 promotes proliferation and tumorigenesis of
non-small-cell lung cancer cells by activating β-catenin signaling
pathway. Cancer Sci. 106:1303–1312. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Chen CC, Chen HY, Su KY, Hong QS, Yan BS,
Chen CH, Pan SH, Chang YL, Wang CJ, Hung PF, et al: Shisa3 is
associated with prolonged survival through promoting β-catenin
degradation in lung cancer. Am J Respir Crit Care Med. 190:433–444.
2014. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Moon RT, Kohn AD, De Ferrari GV and Kaykas
A: WNT and beta-catenin signalling: Diseases and therapies. Nat Rev
Genet. 5:691–701. 2004. View
Article : Google Scholar : PubMed/NCBI
|
|
25
|
Tseng RC, Lin RK, Wen CK, Tseng C, Hsu HS,
Hsu WH and Wang YC: Epigenetic silencing of AXIN2/betaTrCP and
deregulation of p53-mediated control lead to wild-type beta-catenin
nuclear accumulation in lung tumorigenesis. Oncogene. 27:4488–4496.
2008. View Article : Google Scholar : PubMed/NCBI
|