Introduction
Brain injury, such as stroke and traumatic brain
injury (TBI), is one of the main causes of mortality worldwide. It
is a substantial cause of mortality and disability among young and
middle-aged individuals worldwide (1,2).
Brain injury includes molecular cascades that disseminate to
neighboring tissues, a condition known as secondary brain injury.
Primary injury in the brain is irreversible; however, secondary
damage that subsequently develops is responsive to therapeutic
interventions (3,4).
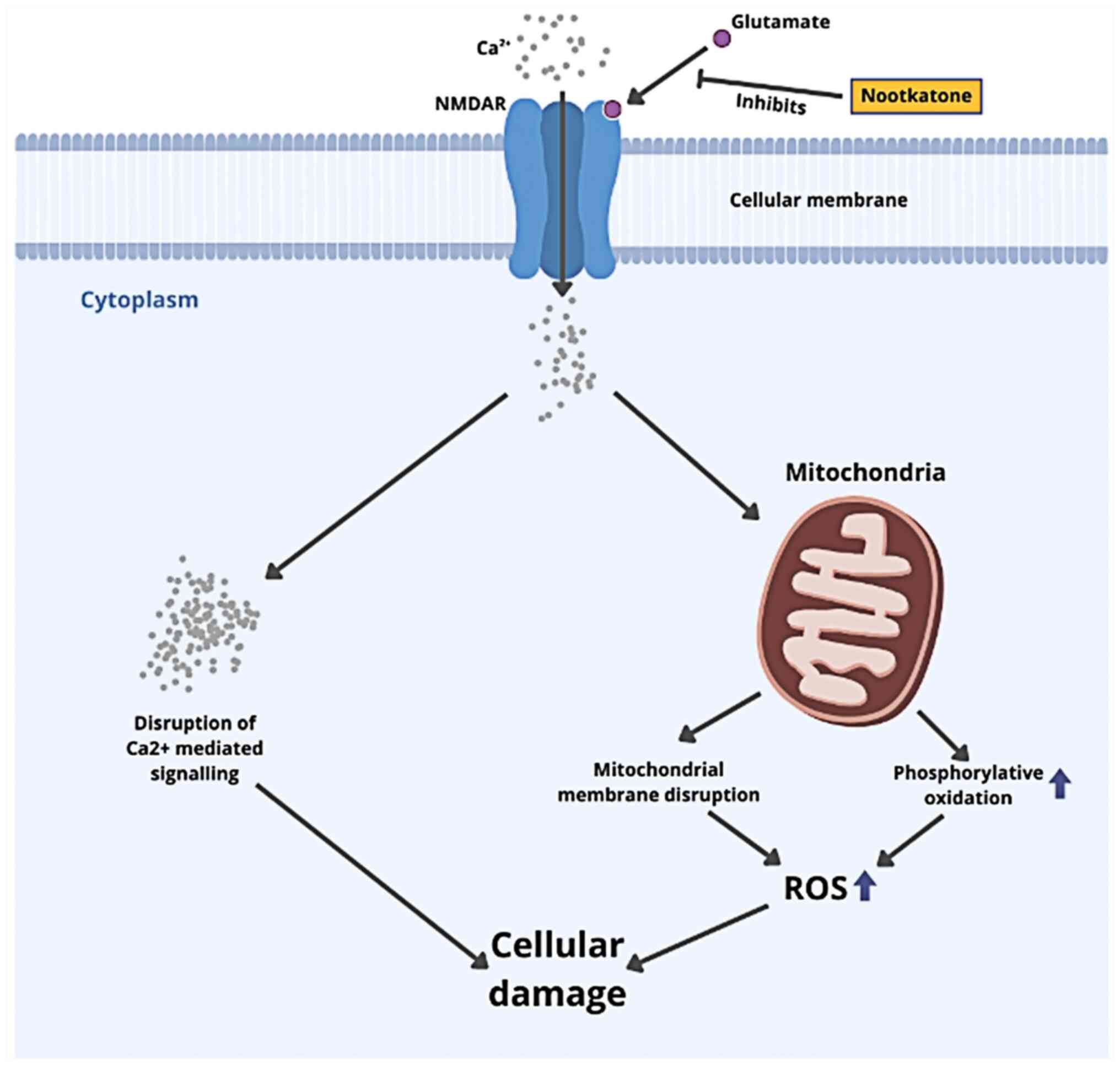
Brain injury is a molecular pathophysiological
sequence linked to excitotoxicity, cell death and oxidative stress.
Brain injury destroys the blood-brain barrier (BBB), causing
excessive glutamate release, which activates N-methyl-D-aspartate
(NMDA) receptors and induces neuronal depolarization. However,
shear and strain forces from a head injury may activate NMDA
receptors. Excessive NMDA receptor activation causes excessive
Ca2+ and Na+ entry into the cell. Excessive
Ca2+ influx in the cytosol leads to mitochondrial
dysfunction and reactive oxygen species (ROS) generation.
Activating mitochondrial protein apoptosis-inducing factor (AIF)
and cytochrome c leads to apoptotic cell death (4-6).
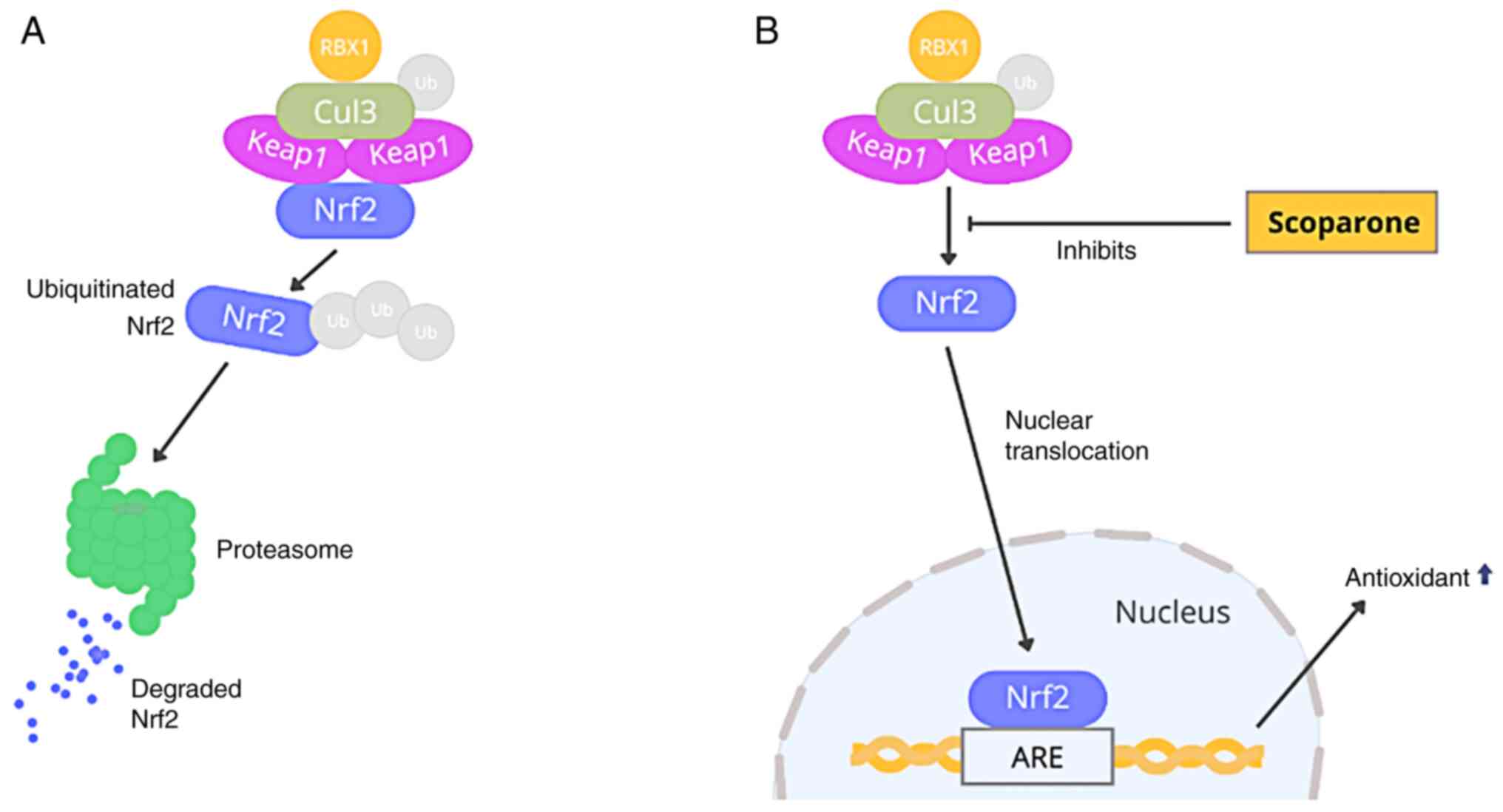
Brain injury-induced oxidative radicals reduce
nuclear factor erythroid 2-related factor 2 (Nrf2) signaling,
interfering with Kelch-like ECH-associated protein-1 (Keap1)-Nrf2
connections. Keap1 is a Cullin3 (Cul3)-based ubiquitin E3 ligase
substrate adaptor component. Nrf2 is a transcription factor that
regulates antioxidant genes and protects against oxidative damage.
Keap1 interacts with Nrf2 in the cytoplasm. The Cul3-Rbk1 complex
ubiquitinates Nrf2, inducing proteasome degradation. Consequently,
oxidative stress alters the structure of Keap1, affecting
Keap1-Nrf2 interactions and limiting Keap1-mediated ubiquitination
(7,8). To activate downstream signaling
pathways encoding detoxification enzymes and antioxidant proteins,
such as nicotinamide adenine dinucleotide phosphate (NADPH), NADPH
quinone dehydrogenase 1 (NQO1), heme oxygenase 1 (HO-1), superoxide
dismutase (SOD) and glutathione peroxidase, newly released Nrf2
from the Keap1-Nrf2 complex translocates from the cytoplasm into
the nucleus where it sequentially binds to the antioxidant response
element (9,10). These findings suggest that the
interaction of the Keap1-Nrf2 complex may be a downstream target
for therapeutic agents in their role as protective factors against
oxidative stress.
Orange (Citrus sinensis) is a major fruit
commodity which is widely consumed, although the peel is generally
regarded as waste. Previous studies have demonstrated that the
antioxidant content of orange peel is higher than that of the juice
bag (11,12). The antioxidant function of orange
peel is mainly due to its high content of flavonoids, polyphenols
and other bioactive components (13-16).
Previous studies on the antioxidant effects of
orange peel have been carried out. Orange peel extract has been
reported to exert antioxidant effects against chemotherapy-induced
toxicity, promote the upregulation of the Nrf2 signaling pathway in
arthritis models, and attenuate seizure-induced oxidative stress
(12-14).
However, the identified specific compounds that play a critical
role in antioxidant signaling are still lacking, and the process of
developing clinical compounds takes longer. In this case, screening
potential compounds as drug candidates through molecular docking is
an option for predicting the interaction mechanisms between ligands
and target proteins (17). The
present study aimed to predict the interaction of potential
compounds in tangerine peel (Citrus sinensis) with the
target protein, Keap1, in the upregulation of the Nrf2 signaling
pathway and NMDA receptor against brain injury-induced oxidative
stress and neuronal apoptosis.
Materials and methods
Citrus sinensis extraction
The samples used were Citrus sinensis peels
obtained from fruit suppliers in Malang, East Java, Indonesia. The
peels were dried in an oven at 60˚C for 24 h and then ground into a
fine powder using a mill. The resulting powder was then diluted
with 96% ethanol in a 12.5:100 (sample:solvent) ratio with a total
volume of 1,300 µl. The samples were then vortexed at 2,000 rpm for
2 min. The supernatants were taken, filtered using a syringe filter
of 0.22 µm, and injected into the liquid chromatography/high
resolution mass spectrometry (LC/HRMS) instrument (Thermo Fisher
Scientific, Inc.).
Analysis of active compounds of citrus
sinensis peels
Compounds inside ethanol extract of Citrus
sinensis peel were analyzed with high-performance liquid
chromatography (HPLC) using the Dionex Ultimate 3000 system (Thermo
Fisher Scientific, Inc.) without any additional structural analysis
or protein profiling analysis. The solvents used were 0.1% formic
acid in water and 0.1% formic acid in acetonitrile. The Hypersil
GOLD column (Thermo Fisher Scientific, Inc.) with a flow speed of
40 µl/min was used for the analysis. HRMS was performed using Q
Exactive Orbitrap (Thermo Fisher Scientific, Inc.). Data were
processed using Compound Discoverer software with mzCloud MS/MS
Library (Thermo Fisher Scientific, Inc.). The results of the active
compounds obtained have been uploaded to the Metabolights database
(Study Number: MTBLS5785).
Active compounds and ADME of Citrus
sinensis peels
Compounds were selected by filtering Lipinski's
criteria and analyzing the BBB permeability of compounds found by
LC-HRMS analysis. A total of 16 compounds were found to have good
BBB permeability, as shown in Table
SI. Three-dimensional structures were obtained from PubChem
(pubchem.ncbi.nlm.nih.gov). Additionally,
the pharmacokinetics of these drugs were evaluated for compliance
with Lipinski's rule of five and ADME using the SwissADME website
(swissadme.ch). The toxicity of each compound was
also evaluated each using the OSIRIS website (18,19).
Retrieval of protein samples
The target proteins in the present study were NMDA,
a glutamate receptor, and Keap1. Keap1 is a transcription factor
involved in synthesizing antioxidants, such as SOD1, SOD2 and
glutathione, as well as neurotrophins, such as brain-derived
neurotrophic factor (20). The PDB
protein code for the NMDA protein used was 4KFQ, whereas that of
Keap1 protein was 5WIY from the Protein Data Bank (PDB) (rcsb.org). The three-dimensional structure of this
protein is available for download in.pdb format. The residual water
molecules and existing control ligands were subsequently separated
from the proteins using PyMOL 2.0 (https://pymol.org/2/). Molecular docking was performed
on a specific site, and grid box dimensions were established by
fixing x, y, and z, as shown in Table
I.
 | Table IGrid box docking dimension of the
study. |
Table I
Grid box docking dimension of the
study.
| Protein target | Center_X | Center_Y | Center_Z | Size_X (Ao) | Size_Y (Ao) | Size_Z (Ao) |
|---|
| Keap1 | 432.216 | 74.894 | 112.592 | 10 | 10 | 10 |
| NMDA | 26.917 | 35.357 | 46.858 | 10 | 10 | 10 |
Molecular docking simulation and
visualization
The molecular docking procedure was performed using
PyRx 0.9 software (https://pyrx.sourceforge.io/) on a personal computer
with the Windows 10 operating system, 8GB RAM, 500GB NVME SSD and
an AMD Athlon 3150U CPU. Molecular docking revealed the site of
interaction and binding affinity interaction between the ligands
and the target proteins. The interaction of the amino acid residues
of the target protein with the ligands was visualized using Pymol
and Discovery Studio 2021 software (https://discover.3ds.com/discovery-studio-visualizer-download)
to discover the binding location and the type of molecular
interactions that occur.
Molecular dynamics simulation
Molecular dynamics simulation was performed using
WebGro webserver (https://simlab.uams.edu/ProteinWithLigand/index.html).
Before the simulation process, ligand preparation was performed
using the PRODRG2 webserver (http://davapc1.bioch.dundee.ac.uk/cgi-bin/prodrg/submit.html).
Subsequently, the prepared proteins and ligands were uploaded to
Webgro with GROMOS96 43a1 as force field parameters, 0.15M NaCl
solvent, 5,000 steps energy minimization, 310 K temperature with 1
bar pressure, MD integrator leapfrog, 100 nsec simulation time, and
limitations frames per MD simulation fixed at 1,000.
Results
Compound analysis of Citrus sinensis
peel extract
Several compounds were found in the LC-HRMS
analysis. Compound intensities were ranked based on peak area. Data
for retention time, compound formula (Table SI), 2D structure, and 2D chemical
formulas were also obtained (Fig.
S1). Based on LC-HRMS findings, the pharmacokinetic profiles of
each compound were further analyzed. Several compounds with good
pharmacokinetic profiles based on Lipinski's rule of five and BBB
permeability were identified, as shown in Table II.
 | Table IIPharmacokinetic profile of chosen
compounds in Citrus sinensis peels extract. |
Table II
Pharmacokinetic profile of chosen
compounds in Citrus sinensis peels extract.
| Compound Name | MW (Da) | Hydrogen donor | Hydrogen
acceptor | LogP | MR | BBB
permeability |
|---|
| Stachydrine | 143.18 | 0 | 2 | -1.10 | 41.35 | No |
| Choline | 104.17 | 1 | 1 | -1.38 | 29.69 | No |
| Nobiletin | 402.39 | 0 | 8 | 3.02 | 106.87 | No |
| Tangeretin | 372.37 | 0 | 7 | 3.02 | Yes | Yes |
| Proline | 115.13 | 2 | 3 | -0.92 | 32.52 | No |
| Pipecolic acid | 129.16 | 2 | 3 | -0.61 | 37.33 | No |
|
Triethanolamine | 149.19 | 3 | 4 | -0.66 | 37.34 | No |
| Cetrimonium | 284.54 | 0 | 0 | 3.51 | 95.82 | No |
| Hesperidin | 610.56 | 8 | 15 | -0.72 | 141.41 | No |
| Scoparone | 206.19 | 0 | 4 | 1.84 | Yes | Yes |
| Alminoprofen | 219.28 | 2 | 2 | 2.57 | Yes | Yes |
| Limonin | 470.51 | 0 | 8 | 2.55 | 11.17 | No |
| Naringin | 580.53 | 8 | 14 | -0.79 | 134.91 | No |
| Linoleic acid | 280.45 | 1 | 2 | 5.45 | Yes | Yes |
| Nootkatone | 218.33 | 0 | 1 | 3.58 | Yes | Yes |
| Chanoclavine | 256.34 | 3 | 2 | 2.39 | Yes | Yes |
Pharmacokinetic profile of selected
compounds
According to Lipinski's rule of five, substances
with good pharmacokinetics, particularly oral administration, have
a molecular weight of <500 Daltons, a hydrogen acceptor value of
<10, a hydrogen donor value of <5, a water partition
coefficient (logP) <5, and a molar refractivity of
40-130(21). Compounds that meet
Lipinski's rule of five have good pharmacokinetic and
bioavailability profiles as they are similar to drugs. In the
present study, the pharmacokinetic investigation of selected
compounds found in Citrus sinensis peel extract exhibited
good results. Nootkatone, alminoprofen, linoleic acid,
chanoclavine, scoparone and tangeretin all fulfill Lipinski's rule
of five criteria and exhibit adequate BBB permeability, allowing
them to cross the BBB, as shown in Table II. Nootkatone (CID: 1268142),
alminoprofen (CID: 2097), linoleic acid (CID: 5280450),
chanoclavine (CID: 5281381), tangeretin (CID: 68077) and scoparone
(CID: 8417). The three-dimensional structures of these chemicals
were retrieved in sdf format. It was also found that the selected
compounds had good pharmacokinetics and bioavailability when
administered orally. The parameter of physicochemistry
lipophilicity must be evaluated, while designing new
pharmaceuticals, since it has been found to significantly influence
a number of pharmacokinetic features such as absorption,
distribution, permeability and drug clearance processes (22).
Toxicity analysis
Toxicity identification is an essential parameter in
determining compound safety for human use. The present study used
the OSIRIS website to predict toxicity in some parameters. There
were a mutagenic, tumorigenic, irritant and reproductive effects.
Mutagenic and tumorigenic parameters predict the effect of a
compound to become mutagenic and induce tumor (18). Toxicity analysis revealed various
risk profiles for each compound that had been selected based on its
good pharmacokinetic profile. The only compound with a high
toxicity risk was tangeretin, with a predicted high risk in
mutagenic and tumorigenic properties. Scoparone, chanoclavine and
alminoprofen were predicted to have a medium toxicity risk in the
reproductive system. Nootkatone and linoleic acid were predicted to
have no toxicity risk in the reproductive system, mutagenic and
tumorigenic properties, or as an irritant (Table SII).
Molecular interaction
The present study analyzed nootkatone, alminoprofen,
linoleic acid, chanocalvine, scoparone and tangeretin, since they
have good BBB permeability. The molecular docking analysis
identified that scoparone bound to Keap1 with the highest affinity
compared to the control ligand and other ligands, whereas
nootkatone attached to the NMDA receptor with the highest affinity
compared to other ligands, but had a similar affinity to the
control ligand. Scoparone exhibited a binding affinity of -5.0
Kcal/mol for NMDA receptors, whereas nootkatone had a binding
affinity of -7.8 Kcal/mol (Table
III). The molecule with the lowest bond energy will have
constant temperature and pressure, and this is known as a stable
molecule (19,23,24).
The amino acid residues affect the binding domain of the target
protein, as well as the sort of chemical interplays in the binding
domain.
 | Table IIIBinding affinity of ligands and
protein receptor. |
Table III
Binding affinity of ligands and
protein receptor.
| | Binding affinity
(Kcal/mol) |
|---|
| Compounds | Keap1 | NMDA receptor |
|---|
| Nootkatone | 2.9 | -7.8a |
| Alminoprofen | -2.4 | -6.9 |
| Linoleic acid | -1.6 | -6,3 |
| Chanoclavine | 3.0 | -7.5 |
| Scoparone | -5.0a | -7.0 |
| Tangeretin | -2.1 | -0.4 |
|
4-Amino-1,7-dihydro-6H-pyrazolo[3,4-d]pyrimidine-6-thione
(control) | -4.2 | |
|
1-Sulfanyl[1,2,4]triazolo[4,3-a]quinoxalin-4(5H)-one
(control) | | -7.8 |
Molecules interact when a bond is formed between the
ligand and the protein. These bonds are located on certain amino
acids to affect a particular reaction or function (25,26).
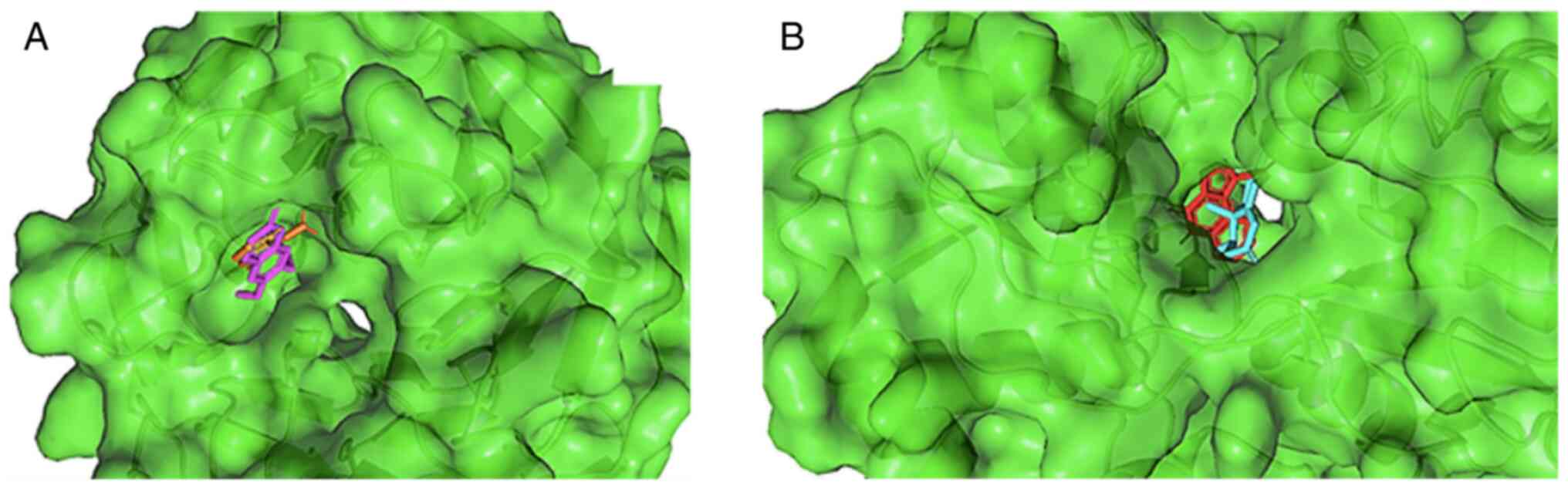
Scoparone and nootkatone attach to the same binding pocket as the
control ligand, as illustrated in Fig.
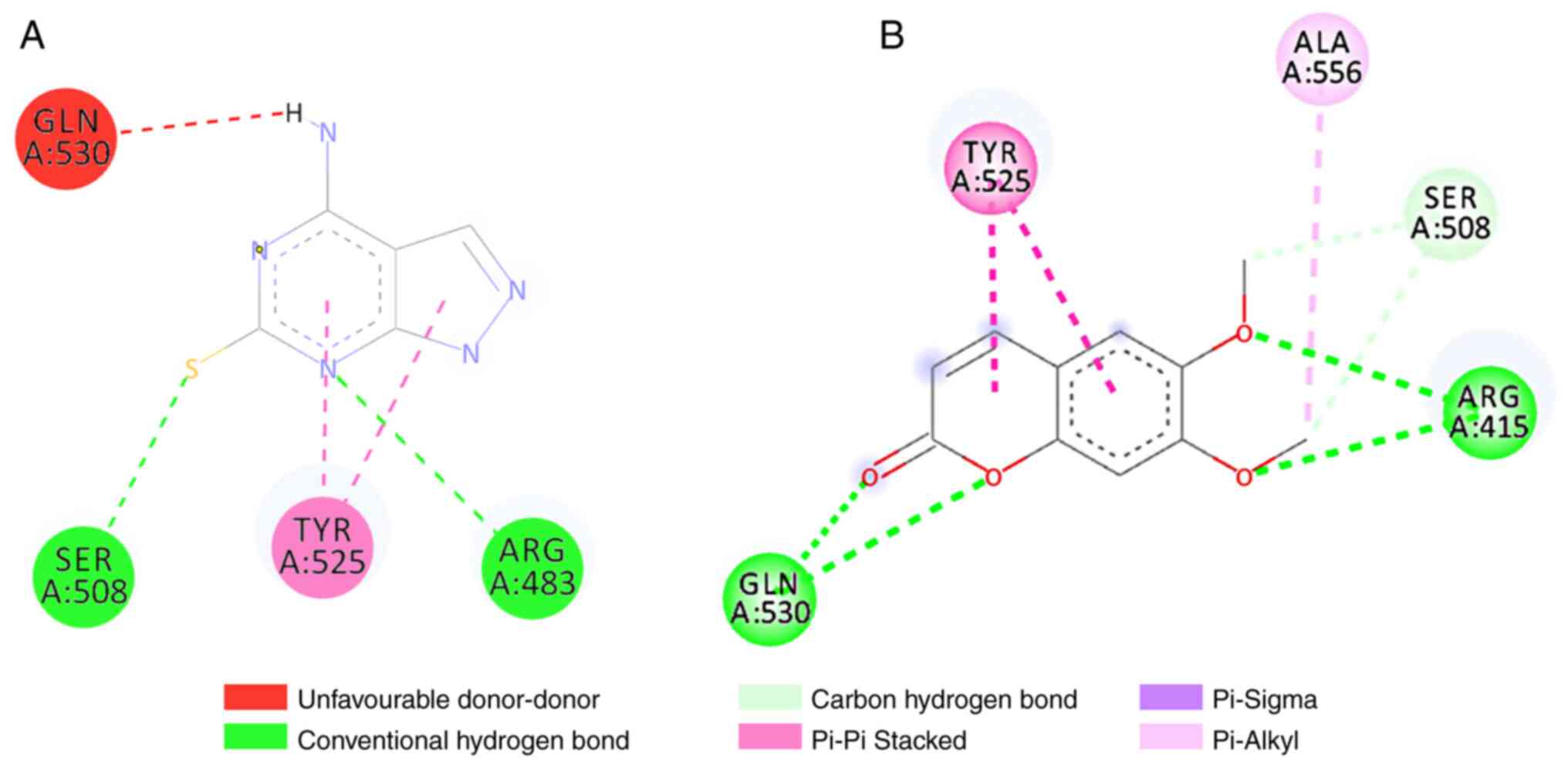
1. Scoparone binds to Keap1 by forming five bond interactions,
three hydrogen bonds and two hydrophobic bonds. These interactions
occur via hydrogen bonds in ARG415, SER508 and GLN530, whereas they
occur via hydrophobic bonds in TYR525 and ALA556 (Fig. 2). Moreover, nootkatone interacted
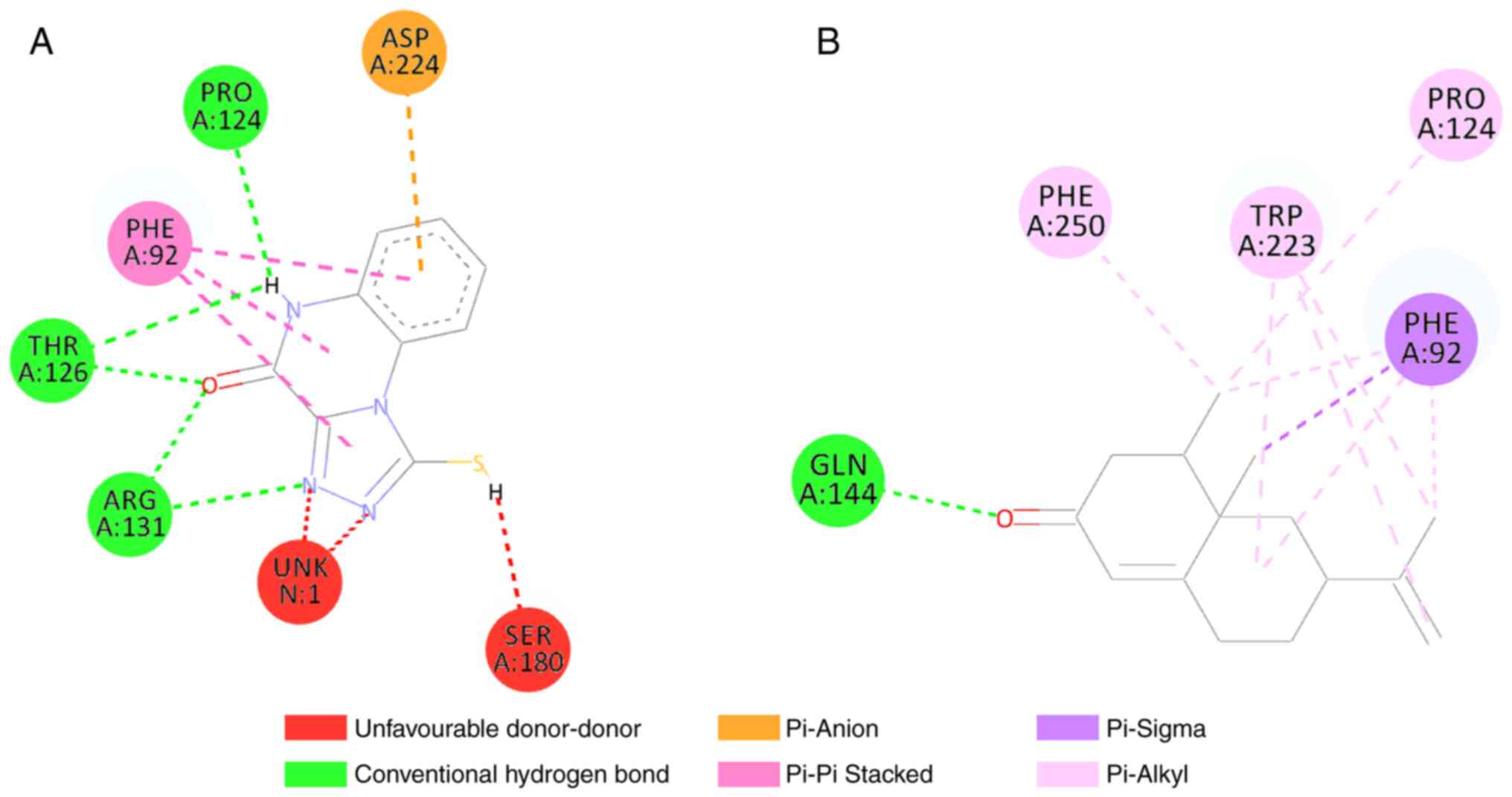
at five amino acid residues and formed two types of bonds.
Nootkatone formed hydrogen bonds with GLN144, and hydrophobic bonds
with the PRO124, PHE250, TRP223 and PHE92 amino acids (Fig. 3). Compared to the control, some
amino acids in Keap1 interacted with scoparone were similar
compared to control ligand (TYR525 and SER508). Interaction between
NMDA receptor and nootkatone also showed similar interaction of
amino acids compared to control ligand (PRO124 and PHE92). This
suggested that scoparone interaction with Keap1 and nootkatone
interaction with NMDA receptor might have similar inhibitory
results compared to control ligands.
Molecular dynamics simulation
Molecular dynamics simulation is a technique to
simulate the whole protein-ligand system due to the course of a
certain time to analyze the confirmation changes. These parameters
are the radius of gyration, root-mean-square deviation (RMSD),
root-mean-square fluctuation (RMSF) and a number of hydrogen bonds
(27).
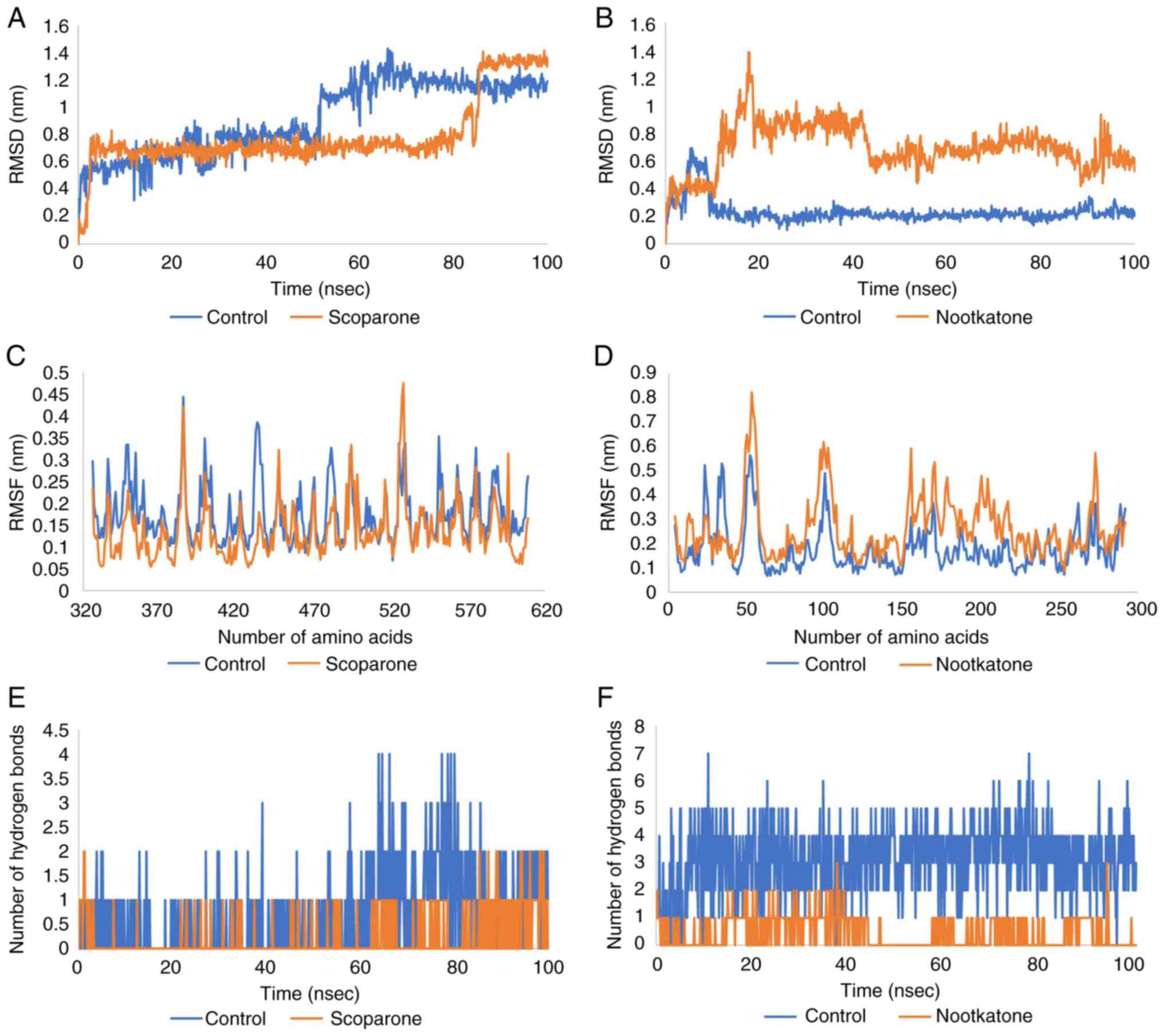
The average RMSD fluctuations for the protein and
ligand in the Keap1 complex were 0.7 nm, with an equilibrium since
4 nsec. The RMSD of the scoparone-Keap1 complex was found to be
stable, first from 4 to 85 nsec and later from 86 to 100 nsec.
Similarly, large deviations in RMSD were observed for the
control-Keap1 complex, indicating an unstable nature of the complex
thus formed. Furthermore, the RMSD value of the nootkatone-NMDA
complex was 0.6 with equilibrium since 22 nsec. The RMSD of
control-NMDA was more stable than that of nootkatone, as shown in
Fig. 4A and B.
RMSF was measured to further compute the residual
flexibility over a period of 100 nsec. The RMSF scoparone-Keap1
complex was <0.45 nm for each residue, the same as that of the
control. But the mean of RMSF Scoparone-Keap1 was more stable than
the control. Moreover, the RMSF of the nootkatone-NMDA complex was
<0.81 nm for each residue, although that of the control was more
stable, with a score of <0.6 nm for each residue (Fig. 4C and D).
Hydrogen bonds are regarded as a powerful
dipole-dipole interaction. There are two varieties: Conventional
and non-conventional. Hydrogen and elements other than N, O and F
may form conventional hydrogen bonds. For the protein-ligand
complex, hydrogen bonding and their number are more crucial. In
terms of the average number of hydrogen bonds, the stability of the
ligand in the active binding cavity of a protein is determined. The
shorter the duration of engagement, the more effective it will be.
The number of hydrogen bonds in the scoparone-Keap1 and
nootkatone-NMDA complex was less than that of the control, as shown
in Fig. 4E and F.
Discussion
The molecular docking method aimed to assign to the
antioxidant herbal compounds found in the peels of Citrus
sinensis. Through the use of various bioinformatics techniques,
anti-inflammatory drugs can be identified and modified, which may
lead to the discovery of novel aspects of anti-inflammatory
disorders that are less well-known in society, thereby revealing
human indifference to these issues. It is essential to be well
informed and to conduct in-depth research on the proteins involved
in anti-inflammation and the traditional herbal medications used to
treat this condition. It is necessary to compile and discuss the
vast amount of knowledge gained synergistically.
Keap1 protein and NMDA receptor play a crucial role
in the process of increased oxidative stress in brain injury
conditions. The presence of mechanical trauma to the brain causes
damage to the BBB, leading to an increase in glutamate release and
the induction of NMDA receptors (3,6,9).
This induction causes an increase in extracellular Ca2+
ions, thus increasing oxidative stress conditions in the cell and
disrupting the process of releasing the Keap1-Nrf2 protein,
consequently inactivating the Nrf2 protein (20,28).
In the present study, LC/HRMS revealed that the scoparone and
nootkatone compounds in orange peel extract (Citrus
sinensis) had good pharmacokinetic potential (the property of
penetrating the BBB and meeting Lipinski criteria), molecular
docking, and molecular dynamics.
The scoparone ligand binds to five amino acids in
Keap1 protein: Arginine415, glycine530, serine508, tyrosine525 and
alanine556, with a higher binding affinity value than the control
ligand. The bonds between Keap1 protein and scoparone ligands are
six hydrogen bonds and three hydrophobic bonds. The bond between
the scoparone ligands on the amino acids arginine415 and serine508
is a polar area (an electrostatic interaction occurs), and the
amino acid tyrosine525 is a non-polar area (a hydrophobic bond
occurs) (20). However, in the
control ligand there are only two hydrogen bonds in the amino acids
arginine483 and serine508 and two hydrophobic bonds in the amino
acid tyrosine525; thus, the scoparone ligand has a higher level of
stability and binding affinity compared to the control ligand
(20,27). The results of molecular dynamics
simulation revealed that the scoparone ligands had a better
conformational stability level (less fluctuation, RMSF parameters)
and a greater docking fit rate (average of scoparone, 0.7 nm; and
control average, 1 nm; RMSD parameters) than the control ligands
(29). These results indicate that
the scoparone ligand has a better and more stable inhibitory
potential against Keap1 protein than the control ligand. Therefore,
the in silico findings demonstrated that scoparone is a
potential inhibitor of Keap1, which may lead to the enhanced
release of Nrf2 and its subsequent activation of the antioxidative
enzyme pathway.
Previous studies have specifically demonstrated the
antioxidant effects of scoparone in various tissues, such as
neuronal and myocardial tissues. The study by Wu et al
(30) in 2019 demonstrated
decreased levels of ROS and malondialdehyde (MDA), and increased
levels of glutathione peroxidase and SOD in an in vitro
model of oxygen glucose deprivation/reoxygenation injury using
hippocampal neurons treated with 25, 50 and 100 µM scoparone. The
study by Lyu et al (31) in
2021 demonstrated that mice with myocardial hypertrophy treated
with scoparone at 60 mg/kg body weight daily via oral gavage
exhibited an alleviation of cardiac hypertrophy and fibrosis, and
ROS production via the inhibition of ras-related C3 botulinum toxin
substrate 1 that stimulates oxidative stress (31).
It has been shown that the Nrf2 pathway is a
critical pathway in the body's response against oxidative stress.
Keap1 is known as a Nrf2 regulator by targeting Nrf2 in the
ubiquitination process. Keap1 is also known to have a
stress-related receptor. When activated under stress conditions
such as oxidative stress, Nrf2 is released from the ubiquitination
process, translocates to the cell nucleus, and stimulates
transcription of various antioxidants and neurotrophins in the
brain, as illustrated in Fig. 5
(20,28).
The increased extracellular glutamate and aspartate
concentrations in the rat hippocampus following transient cerebral
ischemia (stroke) have been the focus of intense investigation for
over two decades. Considering this hypothesis, investigations using
different animal models have found that NMDA receptor antagonists
may improve outcomes after TBI and stroke, paving the way for
large-scale placebo-controlled human trials in TBI and stroke
(32).
The results of molecular docking of the nootkatone
compound to the NMDA receptor protein revealed the presence of
bonds in five amino acids, namely phenylalanin92, proline124,
glysine144, tryptopan223 and phenylalanin250, with one hydrogen
bond and nine hydrophobic bonds, as shown in Fig. 3. Binding to the amino acids
phenylalanine92, proline124, glysine144 and phenylalanine250 are
receptor residues of the NMDA protein on control ligands and have
been shown to act as antagonists in vitro studies (33). These results indicate that
nootkatone ligands have good potential as NMDA receptor inhibitors.
The binding affinity results showed that the nootkatone and control
ligands had the same value, namely -7.8 Kcal/mol. The results of
molecular dynamics show that the control ligand has a better
conformational stability level (less fluctuation, RMSF parameters)
and has a greater docking fit (control ligand 0.2 nm average and
nootkatone ligand 0.6 nm, RMSD parameter) than the nootkatone
ligand. However, the RMSD results of the nootkatone ligand were
still acceptable because the value was <3 nm (34). Control ligands have better
potential to act as NMDA receptor inhibitors than nootkatone
ligands.
NMDA receptor is required for neurodevelopment,
central nervous system tissue remodeling and plasticity, and
dendritic spine formation (memory formation). The primary mechanism
underlying subsequent brain damage following TBI is the increased
NMDA receptor activation. The ion channel activity of the NMDA
receptor at the postsynaptic membrane regulates Ca2+
redistribution. The NMDA receptor-induced Ca2+ excess
contributes to neuronal damage and synaptic plasticity disruption.
NMDA receptor activation also causes nitrative stress and
mitochondrial dysfunction following TBI. This is linked to NMDA
receptor activity modulating neuronal nitric oxides synthase
(35). This is in line with the
findings of the present study that NMDA has a strong affinity for
nootkatone, comparable to the control ligand. Citrus
sinensis peel may lower glutamate excitotoxicity, oxidative
stress and apoptosis, as shown in Fig.
6 (35,36).
Previous research has demonstrated the antioxidant
effects of nootkatone in neuronal and renal tissues. The study by
Yan et al (37) revealed a
significant decrease in MDA levels in the hippocampal neurons of
mice with D-galactosamine-induced liver injury treated with
nootkatone at 5 and 10 mg/kg body weight intragastrically. Park
et al (38) demonstrated a
significant decrease in ROS production and an increase in the
levels of antioxidant enzymes, such as HO-1 and NQO1 in mice
exposed to lipopolysaccharide (LPS) and in LPS-stimulated microglia
model treated with nootkatone. The study by Chen et al
(39) revealed a significant
increase in the levels of catalase and SOD in mice with unilateral
ureteral obstruction treated orally with nootkatone at 10 mg/kg
body weight.
The importance of Keap1 and Nrf2 in the
antioxidative process and their subsequent anti-inflammatory and
various beneficial effects can be applied in various pathological
conditions, such as TBI. The pathological process of secondary
brain injury caused by delayed neurochemical reaction following
initial mechanical trauma involves neuronal cell death (40). Various clinical and preclinical
studies have demonstrated that the type of neuronal cell death
involved in various parts of the brain, such as the hippocampus,
cortex and thalamus is mainly apoptotic (38-40).
The release of ROS, particularly mitochondrial ROS
in ischemia-related brain pathologies, such as stroke and TBI, is
one of the triggers for neuronal apoptosis (41-43).
Previous studies have indicated that increased ROS generation in
neuronal cytosol leads to the release of various pro-apoptotic
proteins, such as cytochrome c (44,45).
This increase in apoptosis suggests that oxidative stress plays a
key role in apoptosis. The molecular docking analysis performed
herein demonstrated that Citrus sinensis peel compounds with
a high binding affinity to Keap1, such as scoparone, had a higher
binding affinity than the control ligand. This finding suggests
Citrus sinensis peel has potential for use as an oxidative
stress inhibitor for the treatment of in brain injury via the
inhibition of Keap1 binding to Nrf2.
In conclusion, some compounds, notably scoparone and
nootkatone, in Citrus sinensis peel exhibit promising
oxidative stress therapeutic potential by exerting inhibitory
effects on the Keap1 and NMDA pathways in brain injury. However,
the present study utilized the LC/HRMS and in silico
computational methods, which were insufficient to analyze practical
efficacy and various factors in clinical conditions. Hence, further
in vitro and in vivo studies are warranted to
validate the therapeutic potential of ethanol extract of Citrus
sinensis peel in brain injury.
Supplementary Material
Results of active compound analysis
from Citrus sinensis peel.
LC-HRMS analysis of Citrus
sinensis peel extract.
Toxicity analysis of chosen compounds
in Citrus sinensis peels extract.
Acknowledgements
Not applicable.
Funding
Funding: No funding was received.
Availability of data and materials
The datasets generated and/or analyzed during the
current study are available in the Figshare repository (https://doi.org/10.6084/m9.figshare.20552073.v1) and
on the MetaboLights Compound Database (https://www.ebi.ac.uk/metabolights/MTBLS5785). Data
are available under the terms of the Creative Commons Attribution
4.0 International license (CC-BY 4.0).
Authors' contributions
MFRS and WMS were involved in the conception and
design of the study, in data collection and analysis, and in the
writing, revising and reviewing of the manuscript. MFRS, GFAP, RAV,
AAAM, WMS, LDF, FLS and MH were involved in the conception and
design of the study, and in the revising and reviewing of the
manuscript. MFRS, GFAP, AAAM, and RAV confirm the authenticity of
all the raw data. All authors have read and approved the final
manuscript.
Ethics approval and consent to
participate
Not applicable.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Maas AIR, Menon DK, Adelson PD, Andelic N,
Bell MJ, Belli A, Bragge P, Brazinova A, Büki A, Chesnut RM, et al:
Traumatic brain injury: Integrated approaches to improve
prevention, clinical care, and research. Lancet Neurol.
16:987–1048. 2017.PubMed/NCBI View Article : Google Scholar
|
|
2
|
Xiong C, Hanafy S, Chan V, Hu ZJ, Sutton
M, Escobar M, Colantonio A and Mollayeva T: Comorbidity in adults
with traumatic brain injury and all-cause mortality: A systematic
review. BMJ Open. 9(e029072)2019.PubMed/NCBI View Article : Google Scholar
|
|
3
|
Dorsett CR, McGuire JL, DePasquale EAK,
Gardner AE, Floyd CL and McCullumsmith RE: Glutamate
neurotransmission in rodent models of traumatic brain injury. J
Neurotrauma. 34:263–272. 2017.PubMed/NCBI View Article : Google Scholar
|
|
4
|
Thapa K, Khan H, Singh TG and Kaur A:
Traumatic brain injury: Mechanistic insight on pathophysiology and
potential therapeutic targets. J Mol Neurosci. 71:1725–1742.
2021.PubMed/NCBI View Article : Google Scholar
|
|
5
|
Khatri N, Thakur M, Pareek V, Kumar S,
Sharma S and Datusalia AK: Oxidative stress: Major threat in
traumatic brain injury. CNS Neurol Disord Drug Targets. 17:689–695.
2018.PubMed/NCBI View Article : Google Scholar
|
|
6
|
Kurniawan DB, Syaban MFR, Mufidah A,
Zulfikri MUR and Riawan W: Protective effect of Saccharomyces
cerevisiae in Rattus norvegicus Ischemic Stroke Model. Res J Pharm
Technol. 14:5785–5789. 2021.
|
|
7
|
Kansanen E, Kuosmanen SM, Leinonen H and
Levonen AL: The Keap1-Nrf2 pathway: Mechanisms of activation and
dysregulation in cancer. Redox Biol. 1:45–49. 2013.PubMed/NCBI View Article : Google Scholar
|
|
8
|
Baird L and Yamamoto M: The molecular
mechanisms regulating the KEAP1-NRF2 pathway. Mol Cell Biol.
40:e00099–20. 2020.PubMed/NCBI View Article : Google Scholar
|
|
9
|
Bhowmick S, D'Mello V, Caruso D and
Abdul-Muneer PM: Traumatic brain injury-induced downregulation of
Nrf2 activates inflammatory response and apoptotic cell death. J
Mol Med (Berl). 97:1627–1641. 2019.PubMed/NCBI View Article : Google Scholar
|
|
10
|
Liang J, Wu S, Xie W and He H: Ketamine
ameliorates oxidative stress-induced apoptosis in experimental
traumatic brain injury via the Nrf2 pathway. Drug Des Devel Ther.
12:845–853. 2018.PubMed/NCBI View Article : Google Scholar
|
|
11
|
Rafiq S, Kaul R, Sofi SA, Bashir N, Nazir
F and Ahmad Nayik G: Citrus peel as a source of functional
ingredient: A review. J Saudi Soc Agric Sci. 17:351–358. 2018.
|
|
12
|
Saini RK, Ranjit A, Sharma K, Prasad P,
Shang X, Gowda KGM and Keum YS: Bioactive compounds of citrus
fruits: A review of composition and health benefits of carotenoids,
flavonoids, limonoids, and terpenes. Antioxidants (Basel).
11(239)2022.PubMed/NCBI View Article : Google Scholar
|
|
13
|
Singh B, Singh JP, Kaur A and Singh N:
Phenolic composition, antioxidant potential and health benefits of
citrus peel. Food Res Int. 132(109114)2020.PubMed/NCBI View Article : Google Scholar
|
|
14
|
Abdelghffar EA, El-Nashar HAS,
AL-Mohammadi AGA and Eldahshan OA: Orange fruit (Citrus sinensis)
peel extract attenuates chemotherapy-induced toxicity in male rats.
Food Funct. 12:9443–9455. 2021.PubMed/NCBI View Article : Google Scholar
|
|
15
|
Li X, Xie P, Hou Y, Chen S, He P, Xiao Z,
Zhan J, Luo D, Gu M and Lin D: Tangeretin inhibits oxidative stress
and inflammation via upregulating Nrf-2 signaling pathway in
collagen-induced arthritic rats. Pharmacology. 104:187–195.
2019.PubMed/NCBI View Article : Google Scholar
|
|
16
|
Sahu DR, Chowdhury B and Sahoo BM:
Anti-convulsant action and attenuation of oxidative stress by
citrus limon peel extracts in PTZ and MES induced convulsion in
albino rats. Cent Nerv Syst Agents Med Chem. 20:177–185.
2020.PubMed/NCBI View Article : Google Scholar
|
|
17
|
Pinzi L and Rastelli G: Molecular Docking:
Shifting paradigms in drug discovery. Int J Mol Sci.
20(4331)2019.PubMed/NCBI View Article : Google Scholar
|
|
18
|
Sander T, Freyss J, von Korff M, Reich JR
and Rufener C: OSIRIS, an entirely in-house developed drug
discovery informatics system. J Chem Inf Model. 49:232–246.
2009.PubMed/NCBI View Article : Google Scholar
|
|
19
|
Rahman PA, Syaban MFR, Anoraga SG and
Sabila FL: Molecular docking analysis from Bryophyllum pinnatum
Compound as A COVID-19 cytokine storm therapy. Open Access Maced J
Med Sci. 10:779–784. 2022.
|
|
20
|
Zhong M, Lynch A, Muellers SN, Jehle S,
Luo L, Hall DR, Iwase R, Carolan JP, Egbert M, Wakefield A, et al:
Interaction energetics and druggability of the protein-protein
interaction between Kelch-like ECH-Associated Protein 1 (KEAP1) and
nuclear factor erythroid 2 like 2 (Nrf2). Biochemistry. 59:563–581.
2020.PubMed/NCBI View Article : Google Scholar
|
|
21
|
Lipinski CA: Lead- and drug-like
compounds: The rule-of-five revolution. Drug Discov Today Technol.
1:337–341. 2004.PubMed/NCBI View Article : Google Scholar
|
|
22
|
Daina A and Zoete V: A BOILED-Egg to
predict gastrointestinal absorption and brain penetration of small
molecules. ChemMedChem. 11:1117–1121. 2016.PubMed/NCBI View Article : Google Scholar
|
|
23
|
Kharisma VD, Widyananda MH, Ansori ANM,
Nege AS, Naw SW and Nugraha AP: Tea catechin as antiviral agent via
apoptosis agonist and triple inhibitor mechanism against HIV-1
infection: A bioinformatics approach. J Pharm Pharmacogn Res.
9:435–445. 2021.
|
|
24
|
Yueniwati Y, Rizki Syaban MF, Faratisha
IFD, Yunita KC, Kurniawan DB, Putra GFA and Erwan NE: Molecular
docking approach of natural compound from herbal medicine in java
against severe acute respiratory syndrome coronavirus-2 receptor.
Open Access Maced J Med Sci. 9:1181–1186. 2021.PubMed/NCBI View Article : Google Scholar
|
|
25
|
Nugraha RY, Faratisha IF, Mardhiyyah K,
Ariel DG, Putri FF, Nafisatuzzamrudah Winarsih S, Sardjono TW and
Fitri LE: Antimalarial properties of isoquinoline derivative from
streptomyces hygroscopicus subsp. Hygroscopicus: An in silico
approach. BioMed Res Int. 2020(6135696)2020.PubMed/NCBI View Article : Google Scholar
|
|
26
|
Yueniwati Y, Syaban MFR, Erwan NE, Putra
GFA and Krisnayana AD: Molecular docking analysis of ficus
religiosa active compound with anti-inflammatory activity by
targeting tumour necrosis factor alpha and vascular endothelial
growth factor receptor in diabetic wound healing. Open Access Maced
J Med Sci. 9:1031–1036. 2021.
|
|
27
|
Yuniwati Y, Syaban MFR, Anoraga SG and
Sabila FL: Molecular docking approach of bryophyllum pinnatum
compounds as atherosclerosis therapy by targeting adenosine
monophosphate-activated protein kinase and inducible nitric oxide
synthase. Acta Inform Med. 30:91–95. 2022.PubMed/NCBI View Article : Google Scholar
|
|
28
|
Yao W, Lin S, Su J, Cao Q, Chen Y, Chen J,
Zhang Z, Hashimoto K, Qi Q and Zhang JC: Activation of BDNF by
transcription factor Nrf2 contributes to antidepressant-like
actions in rodents. Transl Psychiatry. 11(140)2021.PubMed/NCBI View Article : Google Scholar
|
|
29
|
Vishvakarma VK, Singh MB, Jain P, Kumari K
and Singh P: Hunting the main protease of SARS-CoV-2 by
plitidepsin: Molecular docking and temperature-dependent molecular
dynamics simulations. Amino Acids. 54:205–213. 2022.PubMed/NCBI View Article : Google Scholar
|
|
30
|
Wu C, Li T, Zhu B, Zhu R, Zhang Y, Xing F
and Chen Y: Scoparone protects neuronal cells from oxygen glucose
deprivation/reoxygenation injury. RSC Adv. 9:2302–2308.
2019.PubMed/NCBI View Article : Google Scholar
|
|
31
|
Lyu L, Chen J, Wang W, Yan T, Lin J, Gao
H, Li H, Lv R, Xu F, Fang L and Chen Y: Scoparone alleviates Ang
II-induced pathological myocardial hypertrophy in mice by
inhibiting oxidative stress. J Cell Mol Med. 25:3136–3148.
2021.PubMed/NCBI View Article : Google Scholar
|
|
32
|
Shohami E and Biegon A: Novel approach to
the role of NMDA receptors in traumatic brain injury. CNS Neurol
Disord Drug Targets. 13:567–573. 2014.PubMed/NCBI View Article : Google Scholar
|
|
33
|
Ugale VG and Bari SB: Structural
exploration of quinazolin-4(3H)-ones as Anticonvulsants: Rational
design, synthesis, pharmacological evaluation, and molecular
docking studies. Arch Pharm (Weinheim). 349:864–880.
2016.PubMed/NCBI View Article : Google Scholar
|
|
34
|
Martínez L: Automatic identification of
mobile and rigid substructures in molecular dynamics simulations
and fractional structural fluctuation analysis. PLoS One.
10(e0119264)2015.PubMed/NCBI View Article : Google Scholar
|
|
35
|
Luo P, Li X, Wu X, Dai S, Yang Y, Xu H,
Jing D, Rao W, Xu H, Gao X, et al: Preso regulates NMDA
receptor-mediated excitotoxicity via modulating nitric oxide and
calcium responses after traumatic brain injury. Cell Death Dis.
10(496)2019.PubMed/NCBI View Article : Google Scholar
|
|
36
|
Carvajal FJ, Mattison HA and Cerpa W: Role
of NMDA receptor-mediated glutamatergic signaling in chronic and
acute neuropathologies. Neural Plast. 2016(2701526)2016.PubMed/NCBI View Article : Google Scholar
|
|
37
|
Yan T, Li F, Xiong W, Wu B, Xiao F, He B
and Jia Y: Nootkatone improves anxiety- and depression-like
behavior by targeting hyperammonemia-induced oxidative stress in
D-galactosamine model of liver injury. Environ Toxicol. 36:694–706.
2021.PubMed/NCBI View Article : Google Scholar
|
|
38
|
Park JE, Park JS, Leem YH, Kim DY and Kim
HS: NQO1 mediates the anti-inflammatory effects of nootkatone in
lipopolysaccharide-induced neuroinflammation by modulating the AMPK
signaling pathway. Free Radic Biol Med. 164:354–368.
2021.PubMed/NCBI View Article : Google Scholar
|
|
39
|
Chen CM, Lin CY, Chung YP, Liu CH, Huang
KT, Guan SS, Wu CT and Liu SH: Protective effects of nootkatone on
renal inflammation, apoptosis, and fibrosis in a unilateral
ureteral obstructive mouse model. Nutrients.
13(3921)2021.PubMed/NCBI View Article : Google Scholar
|
|
40
|
Akamatsu Y and Hanafy KA: Cell death and
recovery in traumatic brain injury. Neurotherapeutics. 17:446–456.
2020.PubMed/NCBI View Article : Google Scholar
|
|
41
|
Kaya SS, Mahmood A, Li Y, Yavuz E, Göksel
M and Chopp M: Apoptosis and expression of p53 response proteins
and cyclin D1 after cortical impact in rat brain. Brain Res.
818:23–33. 1999.PubMed/NCBI View Article : Google Scholar
|
|
42
|
Clark RS, Chen J, Watkins SC, Kochanek PM,
Chen M, Stetler RA, Loeffert JE and Graham SH: Apoptosis-suppressor
gene bcl-2 expression after traumatic brain injury in rats. J
Neurosci. 17:9172–9182. 1997.PubMed/NCBI View Article : Google Scholar
|
|
43
|
Conti AC, Raghupathi R, Trojanowski JQ and
McIntosh TK: Experimental brain injury induces regionally distinct
apoptosis during the acute and delayed post-traumatic period. J
Neurosci. 18:5663–5672. 1998.PubMed/NCBI View Article : Google Scholar
|
|
44
|
Sugawara T, Lewén A, Gasche Y, Yu F and
Chan PH: Overexpression of SOD1 protects vulnerable motor neurons
after spinal cord injury by attenuating mitochondrial cytochrome c
release. FASEB J. 16:1997–1999. 2002.PubMed/NCBI View Article : Google Scholar
|
|
45
|
Kirkland RA, Windelborn JA, Kasprzak JM
and Franklin JL: A Bax-induced pro-oxidant state is critical for
cytochrome c release during programmed neuronal death. J Neurosci.
22:6480–6490. 2002.PubMed/NCBI View Article : Google Scholar
|