Introduction
Esophageal cancer is the eighth most common type of
cancer and the sixth leading cause of cancer-associated mortality
worldwide (1–3). Esophageal cancer incidence in men is
higher compared with that in women, and its occurrence increases
with age, and mortality rate is ~90% for all cases (1,2,4). Esophageal squamous cell carcinoma
(ESCC) is the predominant histological type amongst Chinese
populations, resulting in 150,000 cases of mortality annually
(5). Despite advances in medical and
surgical techniques, the prognosis for ESCC remains poor, and
long-term survival is in the range of 26.2–49.4% due to local or
distant recurrences (6). Therefore,
it is critical to identify novel molecular mechanisms to elucidate
oncogenesis and metastasis in ESCC.
MicroRNAs (miRNAs/miRs) are small (18–22 nucleotide)
endogenous non-coding RNAs that serve crucial roles in various
biological processes (7,8). Mature miRNAs usually bind to the
3′-untranslated regions of target genes to downregulate the
expression of target genes at post-transcriptional levels through
promotion of mRNA degradation or repression of the translation of
target genes (7,9). Since miRNA-lin-4 was initially
identified in 1993, numerous miRNAs have been ascertained to be
involved in various physiological and pathological processes,
including carcinogenesis (10).
Several miRNAs such as miR-21, miR-34a and miR-155 have been found
to be associated with carcinogenesis by targeting oncogenes or
anti-oncogenes (11–13).
Recently, miR-486-5p has been reported to function
as a tumor suppressor in non-small-cell lung cancer (NSCLC)
(14), breast cancer (15) and hepatocellular carcinoma (16). However, the function and clinical
significance of miR-486-5p in ESCC has yet to be elucidated
(15). In the present study, the
expression levels of miR-486-5p in ESCC tissues were determined,
and the function of miR-486-5p in ESCC cells were investigated by
cell migration, proliferation and apoptosis assays. The results
indicated that miR-486-5p was downregulated in ESCC tissues and
functioned as an anti-oncogene in ESCC by affecting cellular
migration, proliferation and apoptosis.
Materials and methods
ESCC tissue sample collection
All ESCC and adjacent normal tissues used in the
present study were collected in Dongnan Affiliated Hospital of
Xiamen University (Zhangzhou, China). Written informed consent was
obtained from all patients. Ethical approval for the collection and
use of all samples was approved by the Ethics Committee of Dongnan
Affiliated Hospital of Xiamen University. Fresh tissues were
immersed in RNAlater (Qiagen GmbH, Hilden, Germany) in 30 min after
resection and subsequently stored at −80°C for future use.
Cell culture and transfection
Human ESCC cell lines Eca109 and TE-1 were purchased
from the Shanghai Institute of Biochemistry and Cell Biology
(Shanghai, China). Eca109 and TE-1 were cultured in RPMI 1640
(Invitrogen) supplemented with 10% fetal bovine serum (Invitrogen),
100 U/ml penicillin and 100 g/ml streptomycin (Gibco; Thermo Fisher
Scientific, Inc.), at 37°C for 24 h in a humidified incubator
containing 5% CO2. For the restoration of miR-486-5p in
ESCC tissues with endogenously downregulated miR-486-5p,
synthesized miR-486-5p mimics (GenePharma Co., Ltd., Shanghai,
China) was transfected into cells using Lipofectamine 2000
(Invitrogen) according to the manufacturer's protocol. The cells
were trypsinized, and total RNA was extracted using TRIzol reagent
(Invitrogen; Thermo Fisher Scientific, Inc., Waltham, MA, USA) 24 h
after transfection.
RNA isolation and reverse
transcription-quantitative polymerase chain reaction (RT-qPCR)
assays
Total RNA was extracted from 36 ESCC tissue samples
and adjacent normal esophageal tissues, or from the trypsinized
ESCC cell lines Eca109 and TE-1, using TRIzol reagent (Invitrogen),
and purified using an RNeasy Maxi Kit (Qiagen GmbH) according to
the manufacturer's protocol. To obtain the cDNA templates, 1 µg
total RNA of each sample was used for reverse transcription using
an miScript Reverse Transcription kit (Qiagen GmbH). This reaction
was performed at 37°C for 60 min, then 95°C for 5 min. The qPCR
reaction of miR-486-5p was performed on an ABI PRISM 7000
Fluorescent Quantitative PCR System (Applied Biosystems; Thermo
Fisher Scientific, Inc., Waltham, MA, USA) using miScript SYBR
Green PCR Kit (Qiagen GmbH). The 20 µl reaction mixture contained 1
µl cDNA template, 2 µl 10X miScript Universal Primer, 0.4 µl of
each of the specific miRNA primers, 10 µl 2X QuantiTect SYBR Green
PCR Master mix and 6.6 µl RNase-free water. Primer sequences were
as follows: Forward, 5′-TCCTGTACTGAGCTGCCCCGAG-3′ [the reverse
primer was provided by the miScript SYBR Green PCR Kit (Qiagen
GmbH)] for miR-486-5p; and forward, 5′-CTCGCTTCGGCAGCACA-3′ and
reverse, 5′-ACGCTTCACGAATTTGCGT-3′, for U6 (U6 was used as an
endogenous control in the present study). Amplification conditions
were set as follows: 95°C for 2 min, followed by 95°C for 15 sec,
58°C 30 sec and 72°C for 30 sec, for 40 cycles. This experiment was
repeated 3 times, with accompanying no cDNA and no reverse
transcriptase controls. The expression of miR-486-5p was analyzed
using the ΔΔCq method (17),
normalizing to U6 expression.
Migration assay
A wound scratch assay was used to assess the
migratory ability of Eca109 and TE-1 cells in vitro. Cells
(~150,000) were seeded into a 12-well dish and transfected with
miR-486-5p mimics (60 pmol) or the negative, scrambled control (60
pmol) (GenePharma Co., Ltd.) 24 h later. A sterile 200 µl pipette
tip was used to scrape a clear line through the cell monolayer 5 h
post-transfection. The cells were then rinsed three times with
phosphate-buffered saline (PBS) and cultured in an incubator at
37°C. Images of the wound scratches were acquired with an inverted
light microscope (DM IRB; Leica Microsystems GmbH, Wetzlar,
Germany) at 0 and 24 h after the wounds were made. The migration
distance (µm) was measured with a standard caliper and the
experiments were performed in triplicate and analyzed by at least
two observers.
MTT assay
The cell proliferation of Eca109 and TE-1 cells was
determined with MTT assay kit (Sigma-Aldrich, St. Louis, MO, USA).
To determine cell growth, ~5,000 cells were seeded into the wells
of 96-well plates and transfected with miR-486-5p mimics (5 pmol)
or the negative, scrambled control (5 pmol). MTT (20 µl; 5 mg/ml;
Sigma-Aldrich) was added to each well at 0, 24, 48 and 72 h after
transfection. Subsequent to incubation for 4 h, the MTT medium was
removed and 150 µl dimethyl sulfoxide was added. After shaking for
15 min at room temperature, the optical density (OD) of each sample
was determined with an Enzyme Immunoassay Instrument (Model 680
microplate reader; Bio-Rad Laboratories, Inc., Hercules, CA, USA)
at a wavelength of 490/630 nm.
Flow cytometry
For apoptosis assays, Eca109 and TE-1 cells were
cultured in 6-well plates at 37°C to a confluence of ~65% and
transfected with miR-486-5p mimics or a negative control. After 48
h of transfection, Eca109 and TE-1 cells were harvested and washed
twice with cold PBS, then resuspended in 10 µl 1X binding buffer
(Invitrogen). Annexin V-FITC (5 µl; Invitrogen) and 10 µl propidium
iodide was added to each sample. According to the manufacturer's
protocol, the fluorescence of stained cells was then assessed by
flow cytometer (Beckman Coulter, Inc., Brea, CA, USA) using 488 nm
excitation within 30 min.
Statistical analysis
Statistical analysis was performed using the SPSS
statistical software package (version 17.0; SPSS, Inc., Chicago,
IL, USA). Statistical significance was determined by paired and
Student's t-tests. P<0.05 and P<0.01 were considered to
indicate a statistically significant difference.
Results
miR-486-5p expression levels were
downregulated in ESCC tissues
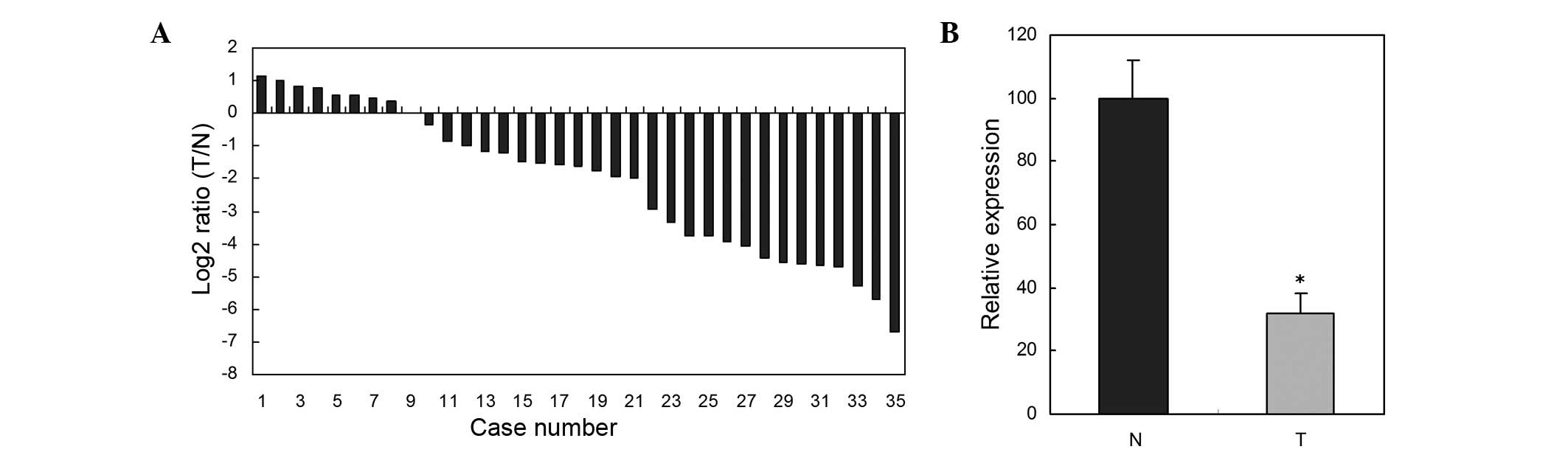
RT-qPCR was used to determine expression levels of
miR-486-5p in 35 paired ESCC tissues and adjacent normal tissues.
The relative expression of miR-486-5p in the 35 paired ESCC tissues
and adjacent normal tissues are shown in Fig. 1A. miR-486-5p expression in ESCC
tissues was significantly downregulated compared with those of the
paired adjacent normal tissues (P<0.05) (Fig. 1B).
Transfection efficiency
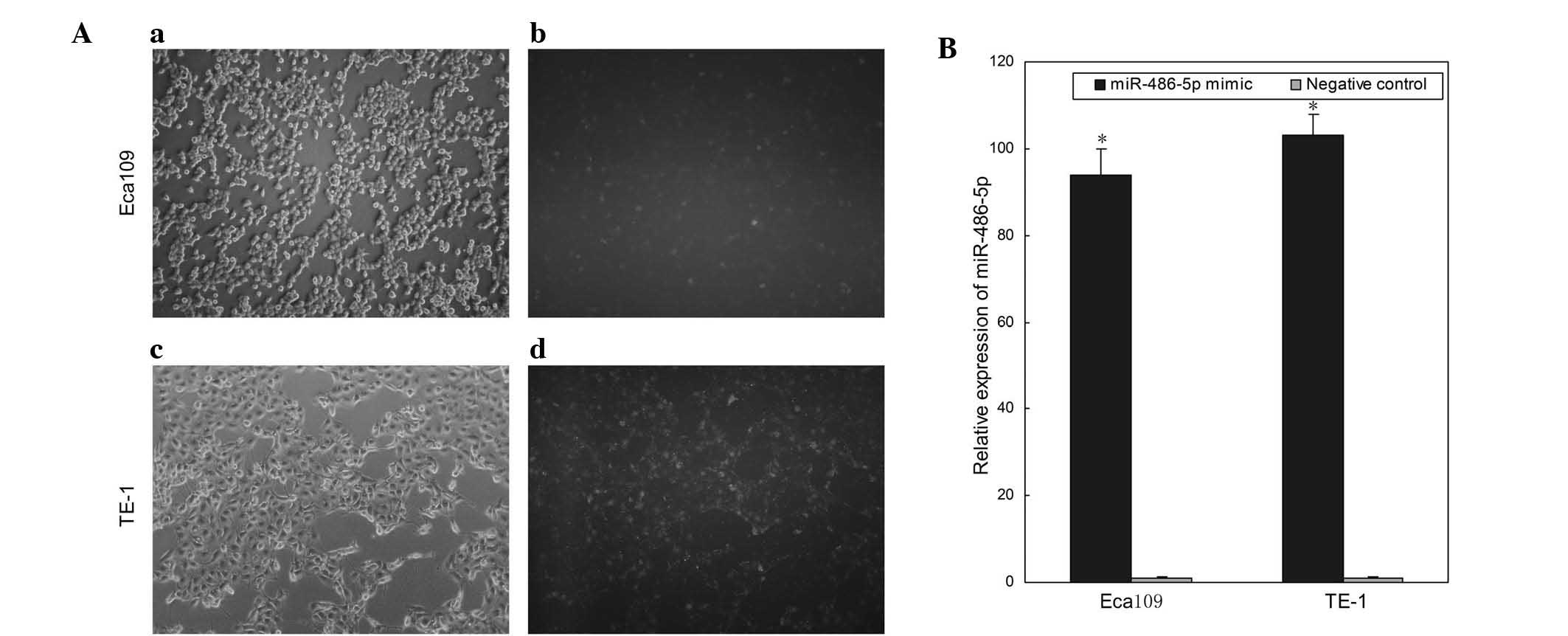
To analyze the function of miR-486-5p in ESCC,
miR-486-5p mimics and negative controls were transfected into the
ESCC cell lines, Eca109 and TE-1. Images of cells transfected with
Fam-labeled negative control were obtained 6 h after transfection.
As shown in Fig. 2A, the
transfection efficiency was ~80 and ~85% in Eca109 and TE-1 cells,
respectively. Compared with the negative control, the relative
expression levels of miR-486-5p in Eca109 and TE-1 cells
transfected with miR-486-5p mimics were 94- and 103-fold,
respectively (P<0.05; Fig. 2B).
The results demonstrated that miR-486-5p mimics were effective in
upregulating the expression of miR-486-5p.
Overexpression of miR-486-5p
suppressed ESCC cell migration in vitro
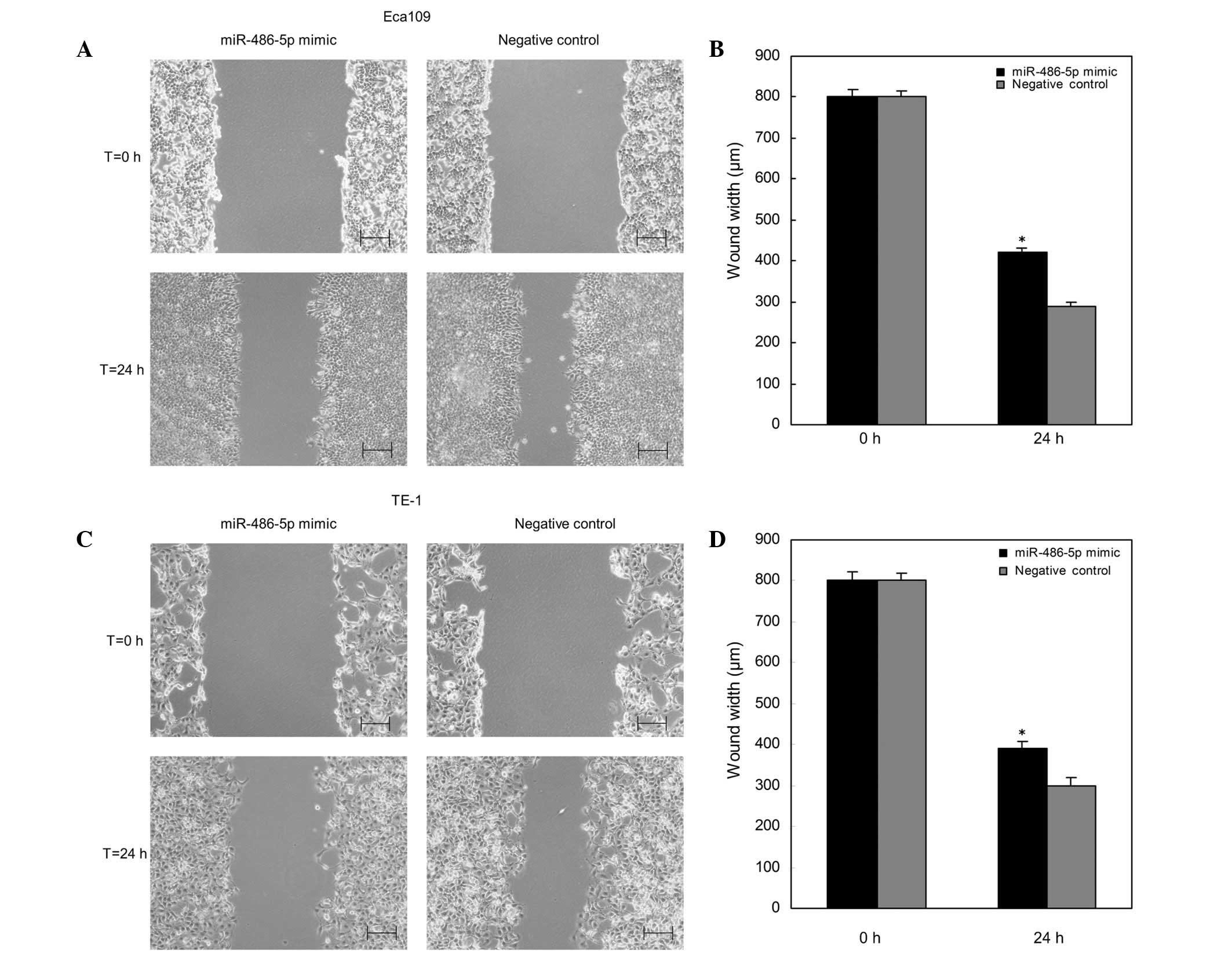
The effects of overexpression of miR-486-5p on cell
migration of ESCC cells in vitro was determined by the use
of a wound scratch assay. As shown in Fig. 3, the wound widths of Eca109 and TE-1
cells transfected with miR-486-5p mimics were wider (P<0.05)
compared with those of the negative control group at 24 h. Thus, it
was indicated that upregulation of miR-486-5p inhibited the cell
migration of ESCC cells.
miR-486-5p mimics inhibited cell
proliferation
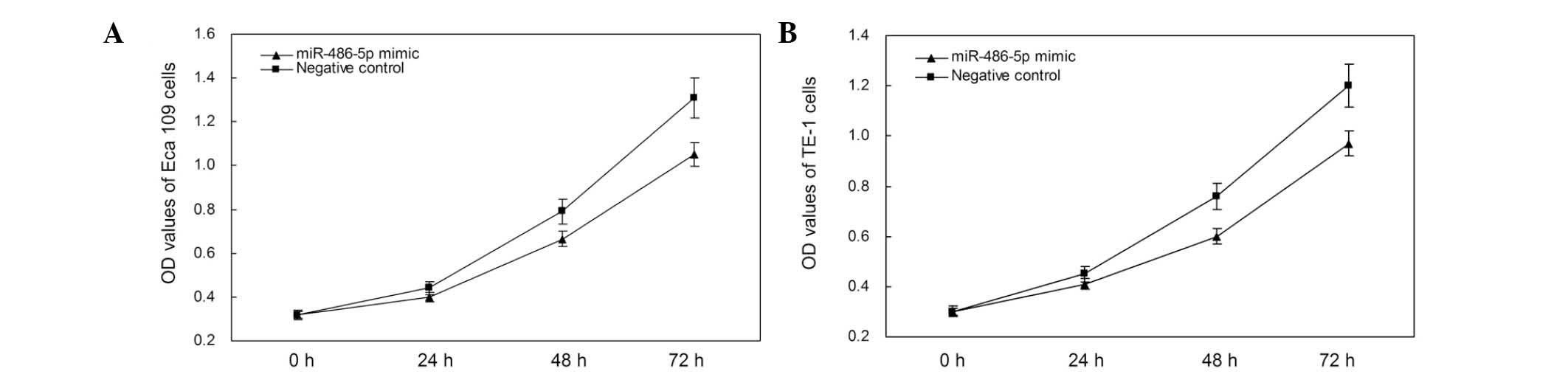
The impact of miR-486-5p on cell proliferation in
ESCC cells was analyzed using an MTT assay. The OD values of the
miR-486-5p mimic and negative control groups were measured at 0,
24, 48 and 72 h after transfection. The results showed that the
proliferation of Eca109 cells decreased by 9.09, 15.71 and 19.84%
(all P<0.05) at the respective aforementioned time-points, while
the proliferation of TE-1 cells decreased by 8.89, 14.47 and 19.17%
(all P<0.05; Fig. 4). These
results suggest that the upregulation of miR-486-5p by mimics
suppressed proliferation of ESCC cells in vitro.
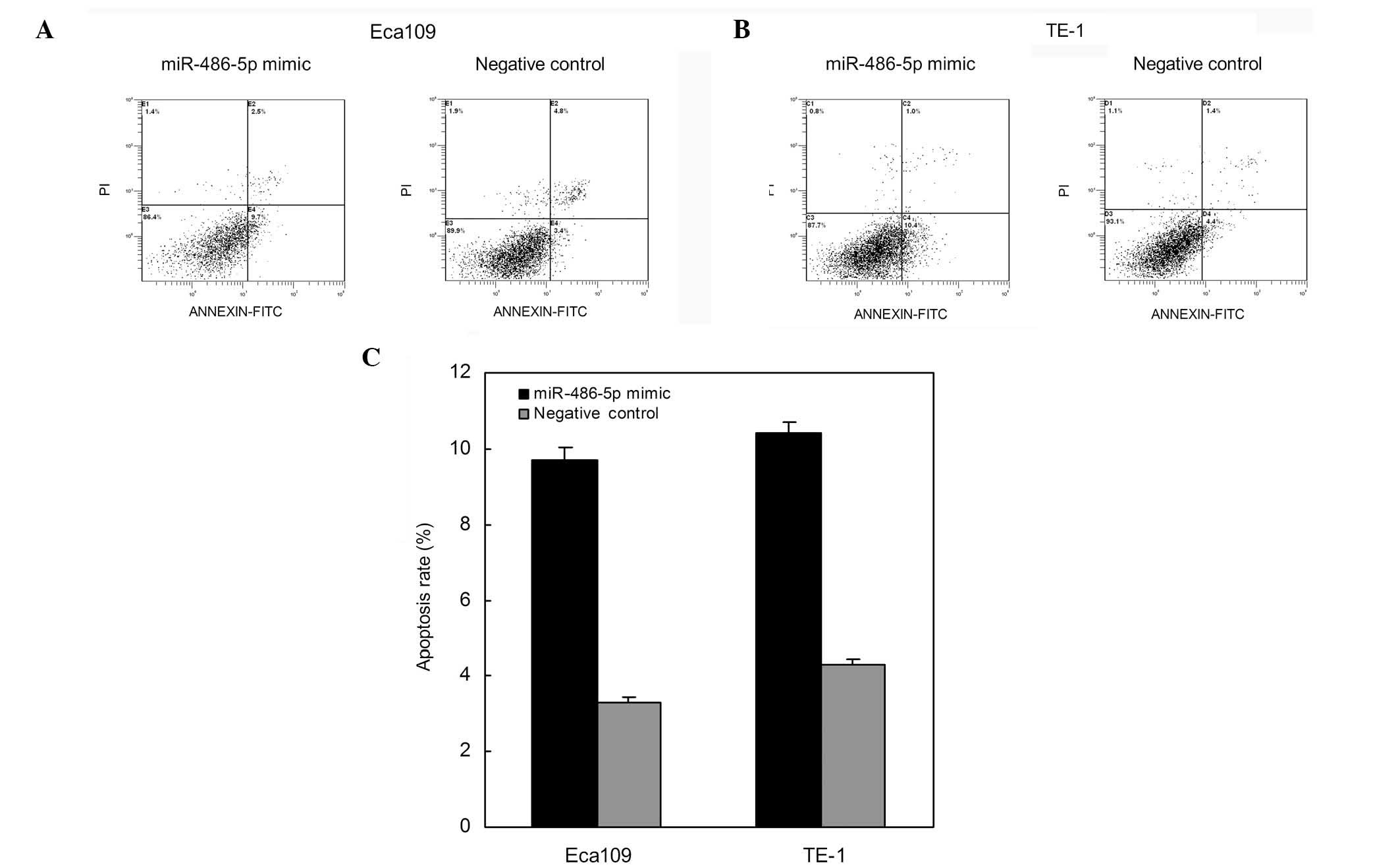
Restoration of miR-486-5p induced ESCC
cell apoptosis
To determine the effects of miR-486-5p on ESCC cell
apoptosis, flow cytometry was used to determine the apoptosis rates
after transfection. As shown in Fig.
5, apoptosis rates of Eca109 cells transfected with miR-486-5p
mimics and those of the negative control were 9.7 and 3.4%,
respectively (P<0.01) 48 h after transfection. The apoptosis
rates of TE-1 cells were 10.4 and 4.4%, respectively (P<0.01)
subsequent to transfection (Fig. 5).
Thus suggesting that the restoration of miR-486-5p expression
levels induced ESCC cell apoptosis.
Discussion
Carcinogenesis involves the activation of numerous
oncogenes and anti-oncogenes. In the complex network involving the
regulation of oncogenes and anti-oncogenes, miRNAs are associated
with gene regulation at the transcriptional and translational level
through base-pairing to complementary mRNA sequences in their
target genes (18,19). A miRNA may fulfil the role of an
anti-oncogene or an oncogene by regulating the levels of oncogenes
or anti-oncogenes (9). miRNAs serve
important roles in diverse cellular processes, including cell
proliferation, cellular differentiation, apoptosis, motility,
invasion and morphogenesis (7,20–25).
Numerous miRNAs have been found to be upregulated in ESCC,
including miR-10b (26), miR-21
(27–29), miR-192, miR-93 and miR-194 (29), miR-23a, miR-26a, miR-27b, miR-96,
miR-128b and miR-129 (30), and
miR-205 (31). By contrast,
downregulation of miR-375 (27),
miR-205, miR-203, miR-125b, miR-100 and miR-27b (29) have been detected in ESCC.
Furthermore, numerous miRNAs were found to serve oncogenic or
anti-oncogenic roles, including the facilitation of ESCC growth by
miR-21 through targeting PTEN and PDCD4 (28,32), in
addition to the ability of miR-145, miR-133a and miR-133b to
converge to target Fascin 1, reducing cell growth and invasion
(33). Furthermore, miR-210 targets
FGFRL1, exerting a negative effect on the cell cycle and
proliferation (34), while miR-296
contributes to ESCC growth by targeting cell CCND1 and p27
(35). Finally, miR-593 may
contribute to carcinogenesis through serine/threonine-protein
kinase (36).
The downregulation of miR-486-5p is a frequent
molecular event in certain human malignances (14–16,37–40).
Furthermore, miR-486-5p may function as tumor suppressor through
contributing to the progression and metastasis of NSCLC by
targeting ARHGAP5 (14), in addition
to the fact that miR-486-5p exerts its antiproliferative function
by directly downregulating PIM-1 expression in breast cancer cells
(15), miR-486-5p suppresses tumor
growth by targeting PIK3R1 in hepatocellular carcinoma (16). However, the expression and role of
miR-486-5p in ESCC has yet to be elucidated.
To determine the expression and role of miR-486-5p
in ESCC, RT-qPCR was used to quantify miR-486-5p expression levels
in 35 cases of ESCC tissues and paired normal tissues. The present
study showed that miR-486-5p expression levels were significantly
downregulated in ESCC tissues, compared with the expression levels
in paired normal esophageal tissues. The effects of miR-486-5p on
ESCC cell migration, proliferation and apoptosis were then analyzed
by transfection of ESCC cell lines with synthetic miR-486-5p
mimics. Transfection of miR-486-5p mimics into the ESCC cell lines
Eca109 and TE-1, inhibited cellular proliferation, migration and
induced apoptosis, compared with the negative control group. The
data indicates that miR-486-5p may be characterized as an
anti-oncogene in ESCC by inhibiting cellular proliferation and
migration, and promoting cellular apoptosis. Further identification
of miR-486-5p target genes is warranted to clarify the mechanism of
action of miR-486-5p in ESCC.
In conclusion, the present study revealed that
miR-486-5p was downregulated in ESCC and served a vital
anti-oncogenic role in ESCC by affecting cellular migration,
proliferation and apoptosis. Further studies are required to
determine the mechanism of action of miR-486-5p in ESCC.
References
|
1
|
Enzinger PC and Mayer RJ: Esophageal
cancer. N Engl J Med. 349:2241–2252. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Kollarova H, Machova L, Horakova D,
Janoutova G and Janout V: Epidemiology of esophageal cancer-an
overview article. Biomed Pap Med Fac Univ Palacky Olomouc Czech
Repub. 151:17–20. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Bohanes P, Yang D, Chhibar RS, Labonte MJ,
Winder T, Ning Y, Gerger A, Benhaim L, Paez D, Wakatsuki T, et al:
Influence of sex on the survival of patients with esophageal
cancer. J Clin Oncol. 30:2265–2272. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Steyerberg EW, Neville B, Weeks JC and
Earle CC: Referral patterns, treatment choices and outcomes in
locoregional esophageal cancer: A population-based analysis of
elderly patients. J Clin Oncol. 25:2389–2396. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Zhao P, Dai M, Chen W and Li N: Cancer
trends in China. Jpn J Clin Oncol. 40:281–285. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Liu J, Xie X, Zhou C, Peng S, Rao D and Fu
J: Which factors are associated with actual 5-year survival of
oesophageal squamous cell carcinoma? Eur J Cardiothorac Surg.
41:e7–e11. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Jiang L, Liu X, Chen Z, Jin Y, Heidbreder
CE, Kolokythas A, Wang A, Dai Y and Zhou X: MicroRNA-7 targets
IGF1R (insulin-like growth factor 1 receptor) in tongue squamous
cell carcinoma cells. Biochem J. 432:199–205. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Soeda S, Ohyashiki JH, Ohtsuki K, Umezu T,
Setoguchi Y and Ohyashiki K: Clinical relevance of plasma miR-106b
levels in patients with chronic obstructive pulmonary disease. Int
J Mol Med. 31:533–539. 2013.PubMed/NCBI
|
|
9
|
Xiong Y, Zhang L, Holloway AK, Wu X, Su L
and Kebebew E: MiR-886-3p regulates cell proliferation and
migration and is dysregulated in familial non-medullary thyroid
cancer. PLoS One. 6:e247172011. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Ha TY: MicroRNAs in human diseases: From
cancer to cardiovascular disease. Immune Netw. 11:135–154. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Shibuya H, Iinuma H, Shimada R, Horiuchi A
and Watanabe T: Clinicopathological and prognostic value of
microRNA-21 and microRNA-155 in colorectal cancer. Oncology.
79:313–320. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Akao Y, Noguchi S, Iio A, Kojima K, Takagi
T and Naoe T: Dysregulation of microRNA-34a expression causes
drug-resistance to 5-FU in human colon cancer DLD-1 cells. Cancer
Lett. 300:197–204. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Tili E, Michaille JJ, Wernicke D, Alder H,
Costinean S, Volinia S and Croce CM: Mutator activity induced by
microRNA-155 (miR-155) links inflammation and cancer. Proc Natl
Acad Sci USA. 108:4908–4913. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Wang J, Tian X, Han R, Zhang X, Wang X,
Shen H, Xue L, Liu Y, Yan X, Shen J, et al: Downregulation of
miR-486-5p contributes to tumor progression and metastasis by
targeting protumorigenic ARHGAP5 in lung cancer. Oncogene.
33:1181–1189. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Zhang G, Liu Z, Cui G, Wang X and Yang Z:
MicroRNA-486-5p targeting PIM-1 suppresses cell proliferation in
breast cancer cells. Tumour Biol. 35:11137–11145. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Huang XP, Hou J, Shen XY, Huang CY, Zhang
XH, Xie YA and Luo XL: MicroRNA-486-5p, which is downregulated in
hepatocellular carcinoma, suppresses tumor growth by targeting
PIK3R1. FEBS J. 282:579–594. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(−Delta Delta C(T)) Method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Jing Q, Huang S, Guth S, Zarubin T,
Motoyama A, Chen J, Di Padova F, Lin SC, Gram H and Han J:
Involvement of microRNA in AU-rich element-mediated mRNA
instability. Cell. 120:623–634. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Guo S, Bai H, Megyola CM, Halene S, Krause
DS, Scadden DT and Lu J: Complex oncogene dependence in
microRNA-125a-induced myeloproliferative neoplasms. Proc Natl Acad
Sci USA. 109:16636–16641. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Bhattacharyya S, Balakathiresan NS,
Dalgard C, Gutti U, Armistead D, Jozwik C, Srivastava M, Pollard HB
and Biswas R: Elevated miR-155 promotes inflammation in cystic
fibrosis by driving hyperexpression of interleukin-8. J Biol Chem.
286:11604–11615. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Lucotti S, Rainaldi G, Evangelista M and
Rizzo M: Fludarabine treatment favors the retention of miR-485-3p
by prostate cancer cells: Implications for survival. Mol Cancer.
12:522013. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Sarver AL, French AJ, Borralho PM,
Thayanithy V, Oberg AL, Silverstein KA, Morlan BW, Riska SM,
Boardman LA, Cunningham JM, et al: Human colon cancer profiles show
differential microRNA expression depending on mismatch repair
status and are characteristic of undifferentiated proliferative
states. BMC Cancer. 9:4012009. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Zhai Q, Zhou L, Zhao C, Wan J, Yu Z, Guo
X, Qin J, Chen J and Lu R: Identification of miR-508-3p and
miR-509-3p that are associated with cell invasion and migration and
involved in the apoptosis of renal cell carcinoma. Biochem Biophys
Res Commun. 419:621–626. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Bentwich I, Avniel A, Karov Y, Aharonov R,
Gilad S, Barad O, Barzilai A, Einat P, Einav U, Meiri E, et al:
Identification of hundreds of conserved and nonconserved human
microRNAs. Nat Genet. 37:766–770. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Liu X, Yu J, Jiang L, Wang A, Shi F, Ye H
and Zhou X: MicroRNA-222 regulates cell invasion by targeting
matrix metalloproteinase 1 (MMP1) and manganese superoxide
dismutase 2 (SOD2) in tongue squamous cell carcinoma cell lines.
Cancer Genomics Proteomics. 6:131–139. 2009.PubMed/NCBI
|
|
26
|
Tian Y, Luo A, Cai Y, Su Q, Ding F, Chen H
and Liu Z: MicroRNA-10b promotes migration and invasion through
KLF4 in human esophageal cancer cell lines. J Biol Chem.
285:7986–7994. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Mathé EA, Nguyen GH, Bowman ED, Zhao Y,
Budhu A, Schetter AJ, Braun R, Reimers M, Kumamoto K, Hughes D, et
al: MicroRNA expression in squamous cell carcinoma and
adenocarcinoma of the esophagus: Associations with survival. Clin
Cancer Res. 15:6192–6200. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Hiyoshi Y, Kamohara H, Karashima R, Sato
N, Imamura Y, Nagai Y, Yoshida N, Toyama E, Hayashi N, Watanabe M
and Baba H: MicroRNA-21 regulates the proliferation and invasion in
esophageal squamous cell carcinoma. Clin Cancer Res. 15:1915–1922.
2009. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Feber A, Xi L, Luketich JD, Pennathur A,
Landreneau RJ, Wu M, Swanson SJ, Godfrey TE and Litle VR: MicroRNA
expression profiles of esophageal cancer. J Thorac Cardiovasc Surg.
135:255–260. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Ogawa R, Ishiguro H, Kuwabara Y, Kimura M,
Mitsui A, Katada T, Harata K, Tanaka T and Fujii Y: Expression
profiling of micro-RNAs in human esophageal squamous cell carcinoma
using RT-PCR. Med Mol Morphol. 42:102–109. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Matsushima K, Isomoto H, Kohno S and Nakao
K: MicroRNAs and esophageal squamous cell carcinoma. Digestion.
82:138–144. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Ma WJ, Lv GD, Tuersun A, Liu Q, Liu H,
Zheng ST, Huang CG, Feng JG, Wang X, Lin RY, et al: Role of
microRNA-21 and effect on PTEN in Kazakh's esophageal squamous cell
carcinoma. Mol Biol Rep. 38:3253–3260. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Kano M, Seki N, Kikkawa N, Fujimura L,
Hoshino I, Akutsu Y, Chiyomaru T, Enokida H, Nakagawa M and
Matsubara H: miR-145, miR-133a and miR-133b: Tumor-suppressive
miRNAs target FSCN1 in esophageal squamous cell carcinoma. Int J
Cancer. 127:2804–2814. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Tsuchiya S, Fujiwara T, Sato F, Shimada Y,
Tanaka E, Sakai Y, Shimizu K and Tsujimoto G: MicroRNA-210
regulates cancer cell proliferation through targeting fibroblast
growth factor receptor-like 1 (FGFRL1). J Biol Chem. 286:420–428.
2011. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Hong L, Han Y, Zhang H, Li M, Gong T, Sun
L, Wu K, Zhao Q and Fan D: The prognostic and chemotherapeutic
value of miR-296 in esophageal squamous cell carcinoma. Ann Surg.
251:1056–1063. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Ito T, Sato F, Kan T, Cheng Y, David S,
Agarwal R, Paun BC, Jin Z, Olaru AV, Hamilton JP, et al: Polo-like
kinase 1 regulates cell proliferation and is targeted by miR-593*
in esophageal cancer. Int J Cancer. 129:2134–2146. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Shen J, Liu Z, Todd NW, Zhang H, Liao J,
Yu L, Guarnera MA, Li R, Cai L, Zhan M and Jiang F: Diagnosis of
lung cancer in individuals with solitary pulmonary nodules by
plasma microRNA biomarkers. BMC Cancer. 11:3742011. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Tan X, Qin W, Zhang L, Hang J, Li B, Zhang
C, Wan J, Zhou F, Shao K, Sun Y, et al: A 5-microRNA signature for
lung squamous cell carcinoma diagnosis and hsa-miR-31 for
prognosis. Clin Cancer Res. 17:6802–6811. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Ragusa M, Majorana A, Statello L, Maugeri
M, Salito L, Barbagallo D, Guglielmino MR, Duro LR, Angelica R,
Caltabiano R, et al: Specific alterations of microRNA transcriptome
and global network structure in colorectal carcinoma after
cetuximab treatment. Mol Cancer Ther. 9:3396–3409. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Bansal A, Lee IH, Hong X, Anand V, Mathur
SC, Gaddam S, Rastogi A, Wani SB, Gupta N, Visvanathan M, et al:
Feasibility of mcroRNAs as biomarkers for Barrett's Esophagus
progression: A pilot cross-sectional, phase 2 biomarker study. Am J
Gastroenterol. 106:1055–1063. 2011. View Article : Google Scholar : PubMed/NCBI
|