Introduction
Cutaneous squamous cell carcinoma (CSCC) is one of
the most common types of cancer affecting the skin, and is more
aggressive in comparison with other skin carcinomas (1,2). The
incidence of CSCC has been increasing in recent years, and the
prognosis of patients with advanced CSCC is unsatisfactory
(2,3). Therefore, the development of effective
treatment strategies for CSCC is urgently required. Investigation
on the molecular mechanism underlying CSCC growth and metastasis
may help to identify potential therapeutic candidates and targets
for CSCC.
MicroRNAs (miRs), a type of small noncoding RNAs
with a length of 18–25 nucleotides, have been demonstrated to
function as key regulators of gene expression via directly binding
to the seed sequence in the 3′-untranslated region (3′UTR) of their
target mRNAs, consequently causing translation repression or mRNA
degradation (4,5). Through regulating the protein
expression of their target genes, miRs participate in various
cellular biological processes, including cell proliferation,
apoptosis, migration, invasion and tumorigenesis (6–8). In
recent years, certain miRs have been implicated in the development
and malignant progression of CSCC (9,10). For
instance, miR-365 serves a promoting role in CSCC through targeting
nuclear factor I/B (9). In addition,
miR-125b directly targets matrix metallopeptidase 13 and inhibits
the CSCC cell proliferation, migration, and invasion (10). Among these miRs, miR-34a has been
reported to be significantly downregulated in the skin and oral SCC
tissues (11). However, the exact
role of miR-34a in the regulation of CSCC cells remains
unclear.
High-mobility group box 1 (HMGB1), a member of the
HMGB superfamily, is a non-histone, nuclear DNA-binding protein
that participates in the regulation of DNA organization and gene
transcription (12,13). It has been demonstrated that HMGB1
serves an oncogenic role in different types of human cancer,
including CSCC (14). Sun et
al (14) reported that HMGB1
promotes tumor metastasis in CSCC via the phosphoinositide
3-kinase/protein kinase B and mitogen-activated protein kinase
signaling pathways. However, the regulatory mechanism underlying
HMGB1 expression in CSCC remains unknown.
Therefore, the present study aimed to investigate
the regulatory mechanism of miR-34a underlying CSCC growth and
metastasis, as well as the involvement of HMGB1.
Materials and methods
Tissue collection
The present study was approved by the Ethics
Committee of Plastic Surgery Hospital, Chinese Academy of Medical
Sciences, Peking Union Medical College (Beijing, China). A total of
72 pairs of CSCC tissues and adjacent non-tumor tissues were
collected from patients, subsequent to obtaining informed consents.
The clinical information of the patients participating in the study
is summarized in Table I. Following
surgical resection, the tissues were immediately frozen in liquid
nitrogen and stored at −80°C until further use.
 | Table I.Association between miR-34a expression
and clinicopathological characteristics of cutaneous squamous cell
carcinoma patients. |
Table I.
Association between miR-34a expression
and clinicopathological characteristics of cutaneous squamous cell
carcinoma patients.
|
|
| miR-34a
expression |
|
|---|
|
|
|
|
|
|---|
| Variables | No. of cases
(n=72) | Low (n=39) | High (n=33) | P-value |
|---|
| Age (years) |
|
|
| 0.813 |
|
<60 | 33 | 17 | 16 |
|
| ≥60 | 39 | 22 | 17 |
|
| Sex |
|
|
| 0.154 |
| Male | 40 | 25 | 15 |
|
|
Female | 32 | 14 | 18 |
|
| Histopathological
grade |
|
|
| 0.016a |
| Well to
moderate | 43 | 18 | 25 |
|
| Poor | 29 | 21 | 8 |
|
| Lymph node
metastasis |
|
|
| 0.012a |
|
Present | 25 | 19 | 6 |
|
|
Absent | 47 | 20 | 27 |
|
Cell culture
The human normal skin cell line HaCaT and the CSCC
cell lines A431 and SCL-1 were purchased from the Cell Bank of Type
Culture Collection, Chinese Academy of Sciences (Shanghai, China).
Cells were cultured in Dulbecco's modified Eagle's medium (DMEM)
with 10% fetal bovine serum (FBS; both from Thermo Fisher
Scientific, Inc., Waltham, MA, USA) in a 37°C humidified atmosphere
with 5% CO2.
Cell transfection
Lipofectamine 2000 reagent (Thermo Fisher
Scientific, Inc.) was used to perform cell transfection, in
accordance with the manufacturer's protocol. Briefly, SCL-1 cells
were transfected with the negative control scrambled miRNA
(miR-NC), miR-34a mimic, miR-34a inhibitor, co-transfected with
miR-34a mimic and blank pcDNA3.1 vector (miR-34a + blank), or
co-transfected with miR-34a mimic and pcDNA3.1-HMGB1 open reading
frame (ORF) plasmid (miR-34a + HMGB1), respectively. Transfection
was for 48 h at 37°C.
Reverse transcription-quantitative
polymerase chain reaction (RT-qPCR) assay
Total RNA was extracted from the tissues and
transfected cells using TRIzol reagent (Thermo Fisher Scientific,
Inc.), according to the manufacturer's protocol. The RNA
concentration was measured using Qubit 3.0 (Thermo Fisher
Scientific, Inc.). Next, the total RNA was converted into cDNA
using a High Capacity cDNA Reverse Transcription kit (Thermo Fisher
Scientific, Inc.), according to the manufacturer's instructions.
The miR expression was then examined by qPCR using a
PrimeScript® miRNA RT-PCR kit (Takara Biotechnology Co.,
Ltd., Dalian, China) on an ABI 7500 thermocycler (Thermo Fisher
Scientific, Inc.), with U6 used as an internal reference. The mRNA
expression was determined using the standard SYBR-Green RT-PCR kit
(Takara Biotechnology Co., Ltd.) on an ABI 7500 thermocycler, with
GAPDH used as an internal reference. These procedures were
performed according to the manufacturer's protocols. The primer
sequences used in qPCR were as follows: HMGB1 forward,
5′-TATGGCAAAAGCGGACAAGG-3′ and reverse,
5′-CTTCGCAACATCACCAATGGA-3′; GAPDH forward,
5′-ACAACTTTGGTATCGTGGAAGG-3′ and reverse,
5′-GCCATCACGCCACAGTTTC-3′. The primers for miR expression detection
were purchased from Fulgene (Guangzhou, China). The reaction was
conducted at 95°C for 5 min, followed by 40 cycles of denaturation
at 95°C for 30 sec and an annealing/elongation step at 60°C for 30
sec. The relative expression was analyzed by the 2−ΔΔCq
method (15).
Western blot analysis
Cells were lysed with ice-cold
radioimmunoprecipitation assay buffer (Thermo Fisher Scientific,
Inc.). The concentration of protein was determined with a BCA
Protein Assay kit (Thermo Fisher Scientific, Inc.), according to
the manufacturer's protocol. Protein was separated by 10% SDS-PAGE
and then transferred onto a polyvinylidene difluoride (PVDF)
membrane (Thermo Fisher Scientific, Inc.). Next, the PVDF membrane
was incubated with phosphate-buffered saline (PBS) containing 5%
non-fat milk (Yili Industrial Group Co., Ltd., Beijing, China) for
3 h at room temperature. Following washing with PBS for three
times, the PVDF membrane was incubated with rabbit anti-human HMGB1
antibody (1:50; ab18256) or rabbit anti-human GAPDH antibody
(1:100; ab9485) (both from Abcam, Cambridge, MA, USA) at 37°C for 3
h. Subsequent to washing with PBS for three times, the PVDF
membrane was then incubated with goat anti-rabbit IgG secondary
antibody (1:5,000; ab7090; Abcam) at room temperature for 1 h. A
Chemiluminescent Substrate kit (Thermo Fisher Scientific, Inc.) was
then used to detect the signals, according to the manufacturer's
protocol. The relative protein expression was analyzed by Image-Pro
Plus software (version 6.0; Media Cybernetics, Inc., Rockville, MD,
USA). GAPDH was used as the internal reference.
Cell proliferation analysis
SCL-1 cells (2×104 cells/ml) were seeded
into 96-well plates (5,000 cells/well) and incubated at 37°C for
24, 48 and 72 h. At each time point, 0.5% MTT solution was added,
and the cells were then incubated at 37°C for 4 h. Subsequently,
cells were centrifuged at 1,500 × g for 3 min at room temperature,
the cell supernatant was discarded, and 150 µl dimethyl sulfoxide
was added to dissolve the formazan. The optical density was then
measured using a microplate reader (Bio-Rad Laboratories, Inc.,
Hercules, CA, USA) at a wavelength of 570 nm.
Cell migration analysis
SCL-1 cells were cultured to full confluence in
6-well plates (500,000 cells/well), and then a plastic scraper was
used to create wounds (~1 mm in width). Subsequently, the cells
were washed with PBS and then incubated in DMEM containing 10% FBS
at 37°C for 24 h. Images of the cells were captured under a light
microscope and the number of cells migrating into the scratched
area was recorded.
Cell invasion analysis
Cell invasion was determined using transwell
chambers pre-coated with Matrigel (BD Biosciences, Franklin Lakes,
NJ, USA), according to the manufacturer's protocol. Briefly, SCL-1
cell suspension (1×106 cells/ml) was prepared in DMEM.
Next, 300 µl of the SCL-1 cell suspension was added into the upper
chamber, while 300 µl DMEM with 10% FBS was added into the lower
chamber. Following incubation at 37°C for 24 h, cells that had not
migrated through the pores were wiped out using a cotton-tipped
swab. The filters were then fixed in 90% alcohol at room
temperature for 10 min, and stained by crystal violet
(Sigma-Aldrich; Merck, Darmstadt, Germany). Images of the invading
cells were obtained under an inverted microscope (Olympus Corp.,
Tokyo, Japan). The invading cells were counted.
Bioinformatics prediction
TargetScan version 7.1 online software (www.targetscan.org) was used to predict the potential
targets of miR-34a, according to the manufacturer's instructions.
Briefly, ‘human’ was selected as the target species, and ‘miR-34a’
was inserted as the investigated miR. MiR-34a was predicted to be
able to directly bind to the seeding sequences of the 3′-UTR of
HMGB1.
Luciferase reporter gene assay
The wild type (WT) and mutant type (MT) of HMGB1
3′UTR luciferase reporter gene plasmids were generated by Yearthbio
(Changsha, China). The SCL-1 cells were then co-transfected with
miR-34a mimic or miR-NC, and with the WT- or MT-HMGB1-3′UTR
reporter plasmid using Lipofectamine 2000 reagent. Subsequent to
incubation at 37°C for 48 h, the luciferase activity was examined
using a Dual-Luciferase Reporter Assay system (Promega Corp.,
Madison, WI, USA), according to the manufacturer's protocol.
Statistical analysis
Data are expressed as the mean value ± standard
deviation. SPSS version 19.0 software (IBM Corp., Armonk, NY, USA)
was used to perform statistical analysis. The contingency data was
analyzed by χ2 test. Data were also analyzed using
Student's t-test for two-group comparisons or one-way analysis of
variance for comparison of more than two groups. A P-value of
<0.05 was considered to indicate a difference that was
statistically significant.
Results
miR-34a is downregulated in CSCC
tissues and cell lines
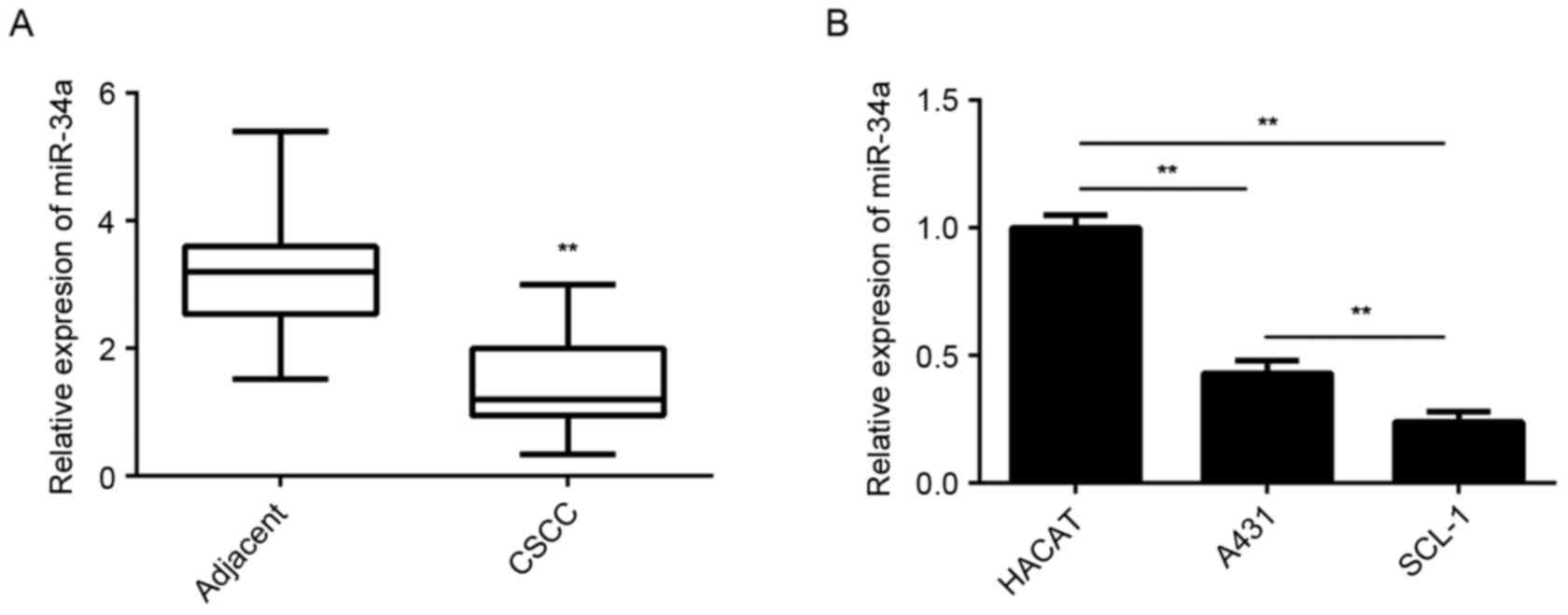
The miR-34a levels were examined using RT-qPCR in
CSCC and adjacent non-tumor tissues. The results demonstrated that
the expression levels of miR-34a were significantly lower in CSCC
tissues compared with the adjacent non-tumor tissues (Fig. 1A). In addition, it was observed that
miR-34a was significantly downregulated in CSCC cell lines,
including A431 and SCL-1, when compared with the expression in
normal skin HaCaT cells (Fig.
1B).
Furthermore, the clinical significance of miR-34a
expression in CSCC was examined based on the correlation of this
expression with the clinicopathological characteristics of
patients. These CSCC patients were divided into low miR-34a
expression group and high miR-34a expression group, based on the
mean value (1.64) of miR-34a expression levels as the cut-off
value. No significant association was detected between miR-34a
expression and the age or gender of patients (Table I). However, the low miR-34a
expression group had more CSCC patients with poor differentiation
degree and lymph node metastasis, when compared with the high
miR-34a expression group, indicating that low expression of miR-34a
was found to be significantly associated with poor differentiation
and presence of node metastasis (Table
I). Based on these findings, it is suggested that miR-34a
downregulation may contribute to the malignant progression of
CSCC.
Restoration of miR-34a expression
inhibits SCL-1 cell proliferation, migration and invasion
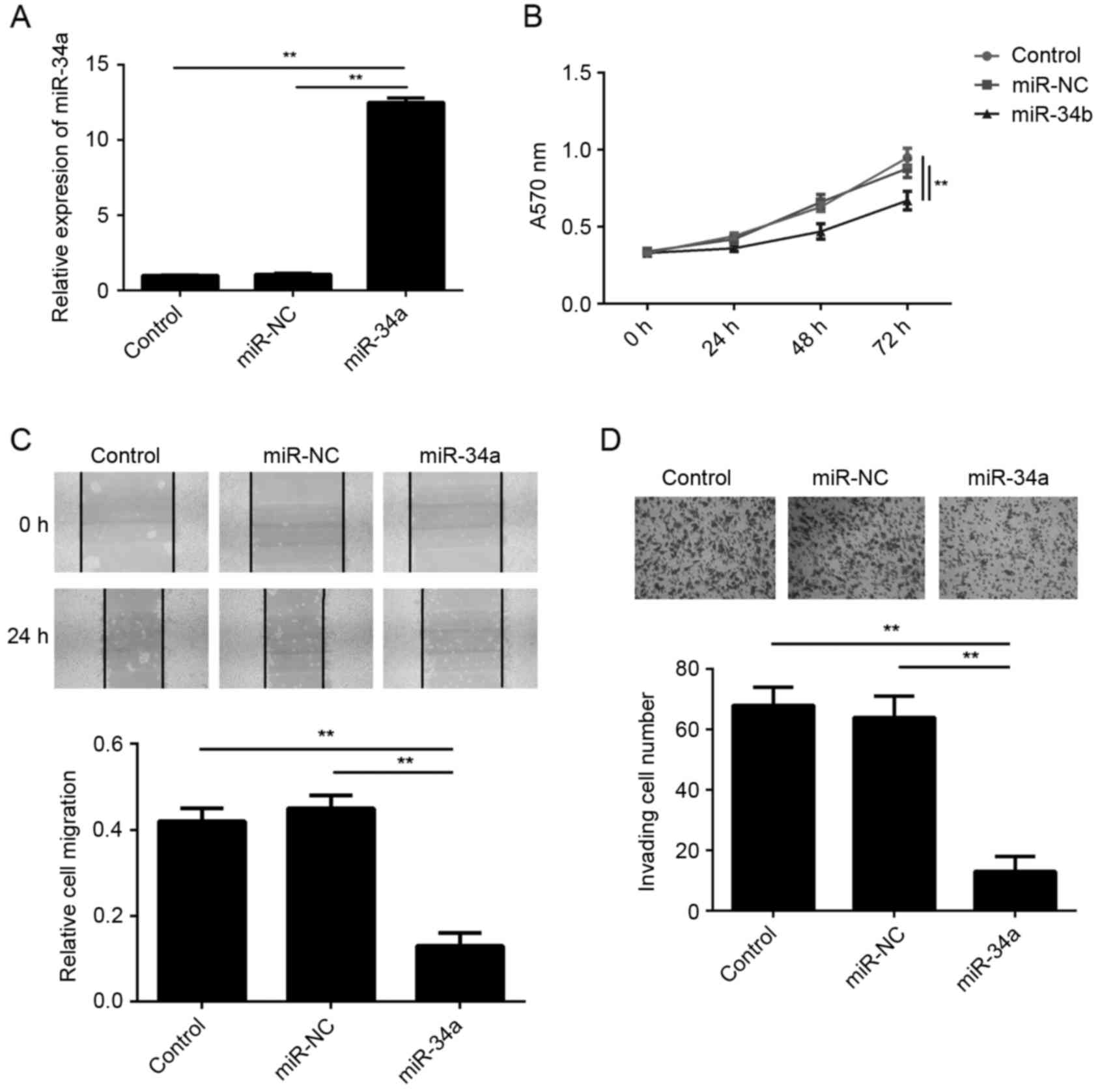
In order to restore the expression levels of miR-34a
in the CSCC cells, SCL-1 cells were transfected with an miR-34a
mimic. Following transfection, the miR-34a levels were
significantly increased in the miR-34a group compared with the
miR-NC group (Fig. 2A). In addition,
the results of MTT, wound healing and transwell assays further
demonstrated that the proliferation, migration and invasion of
SCL-1 cells, respectively, were significantly reduced in the
miR-34a group compared with the miR-NC group (Fig. 2B-D). These findings suggest that
miR-34a exerted suppressive effects on the malignant phenotypes of
SCL-1 cells.
HMGB1 is identified as a novel target
gene of miR-34a in SCL-1 cells
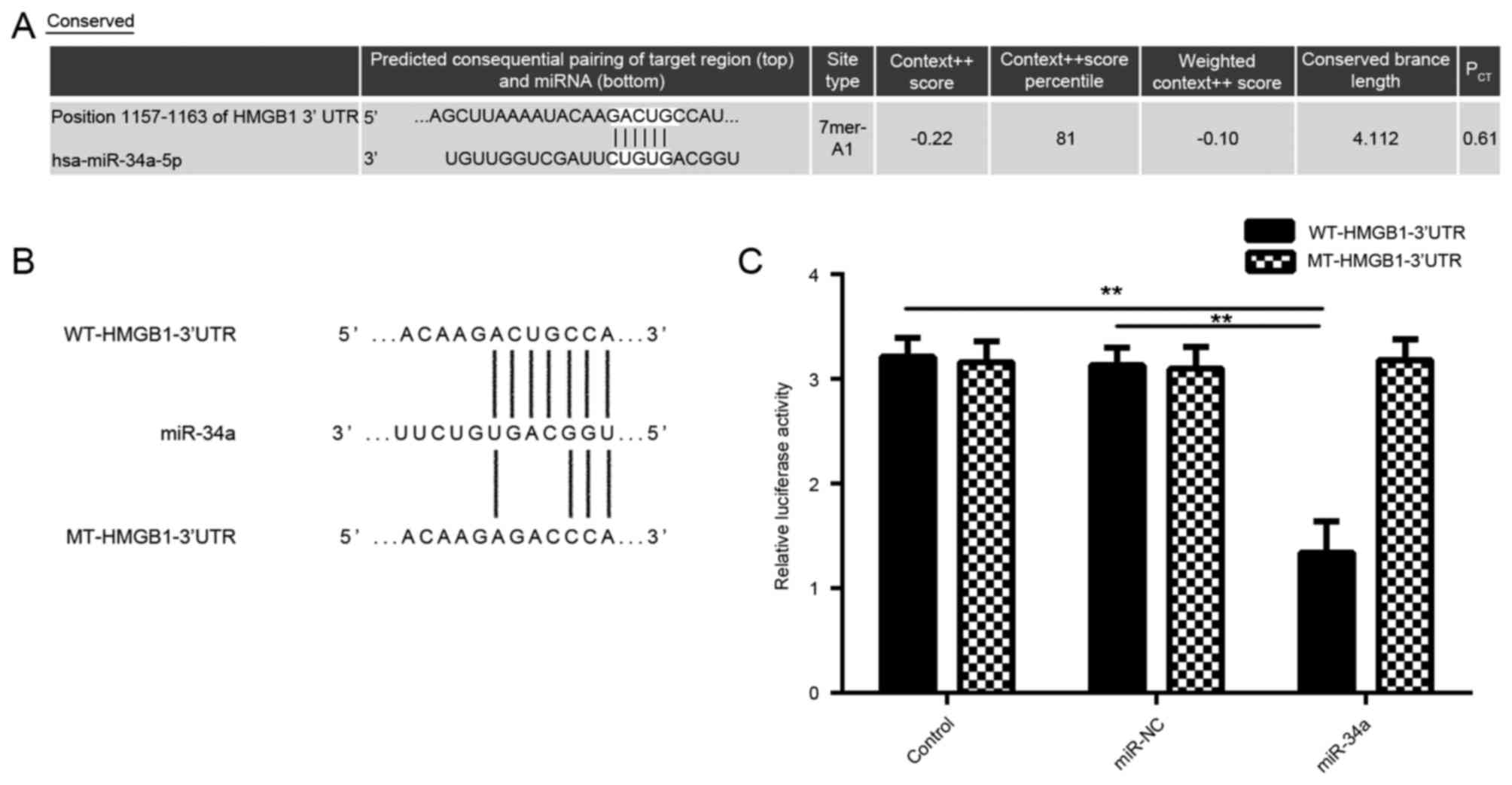
The study then examined the putative target of
miR-34a in CSCC cells using bioinformatics analysis. As shown in
Fig. 3A, bioinformatics prediction
results showed that miR-34a was able to directly bind to the
seeding sequences of the 3′UTR of HMGB1. Subsequently, the WT and
MT HMGB1 3′-UTR reporter gene plasmids were generated (Fig. 3B). A luciferase reporter gene assay
was then performed to confirm whether HMGB1 was a target gene of
miR-34a. As shown in Fig. 3C, the
luciferase activity was significantly decreased in SCL-1 cells
co-transfected with miR-34a mimic and WT-HMGB1-3′UTR reporter gene
plasmid, when compared with the control group; however, no
significant difference was observed upon co-transfection with
miR-34a mimic and the MT-HMGB1-3′UTR reporter gene plasmid. These
data indicate that HMGB1 is indeed a target gene of miR-34a.
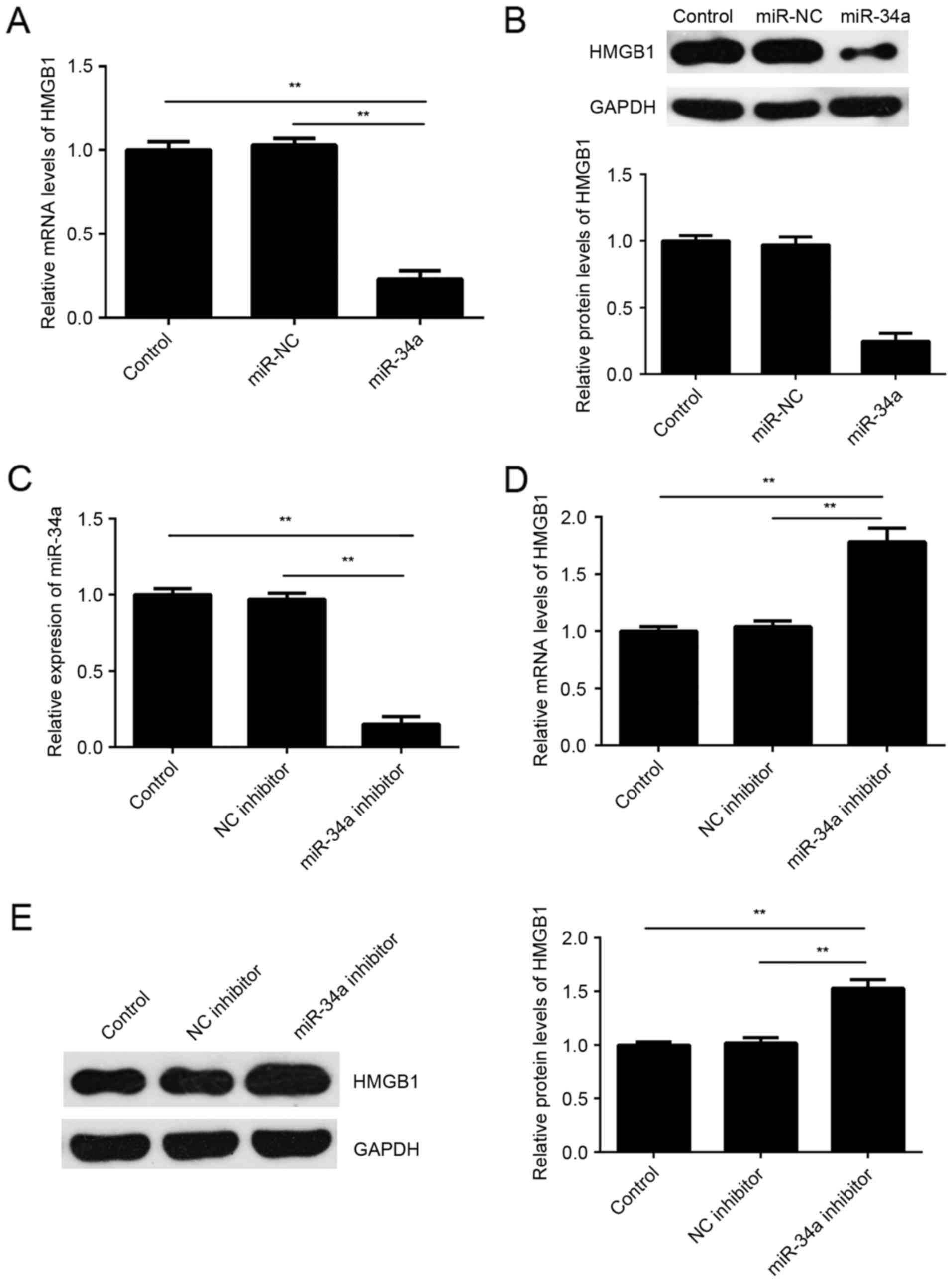
As miRs generally function as a negative regulator
of the expression of their target genes, the present study then
investigated the effects of miR-34a on the expression of HMGB1 in
SCL-1 cells. The results indicated that the mRNA and protein
expression levels of HMGB1 were significantly lower in the miR-34a
group compared with the miR-NC group (Fig. 4A and B). To further confirm these
findings, SCL-1 cells were transfected with miR-34a inhibitor or
with NC inhibitor, serving as the control group. Following
transfection, the miR-34a levels were significantly reduced
(Fig. 4C). Furthermore, it was
observed that the mRNA and protein expression levels of HMGB1 were
significant higher in the miR-34a inhibitor group compared with the
NC inhibitor group (Fig. 4D and E).
These aforementioned data demonstrated that miR-34a negatively
regulated the expression of HMGB1 by directly bind to its mRNA
3′UTR.
HMGB1 is upregulated in CSCC tissues
and cell lines
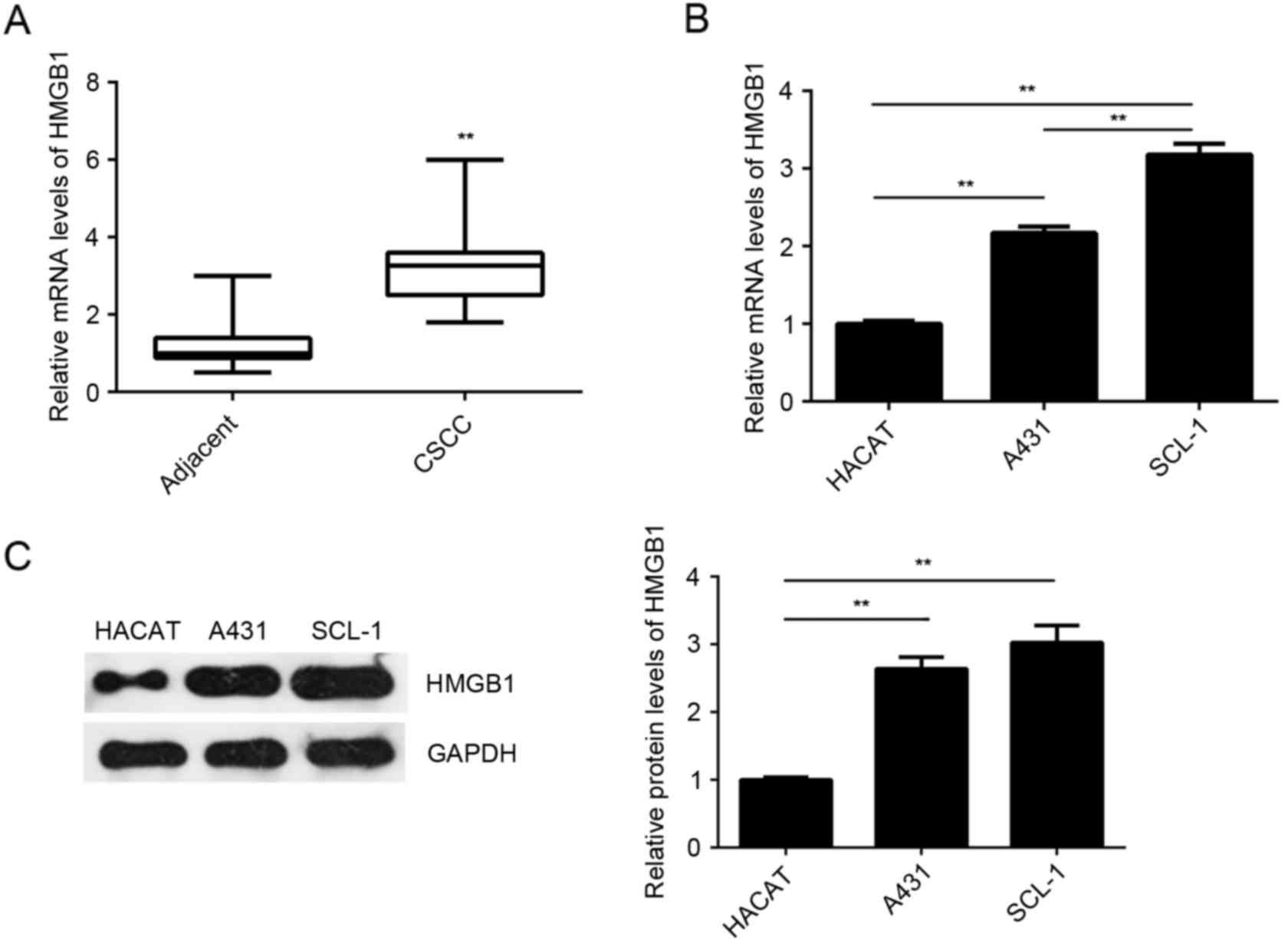
The expression of HMGB1 was further examined in the
CSCC tissues and cell lines. As indicated in Fig. 5A, the mRNA levels of HMGB1 were
significantly higher in the CSCC tissues compared with the adjacent
non-tumor tissues. The mRNA and protein levels of HMGB1 were also
increased in the examined CSCC cell lines (A431 and SCL-1) compared
with the normal HaCaT cells (Fig. 5B and
C). Based on these data, it is suggested that upregulation of
HMGB1 may be involved in the CSCC development and progression.
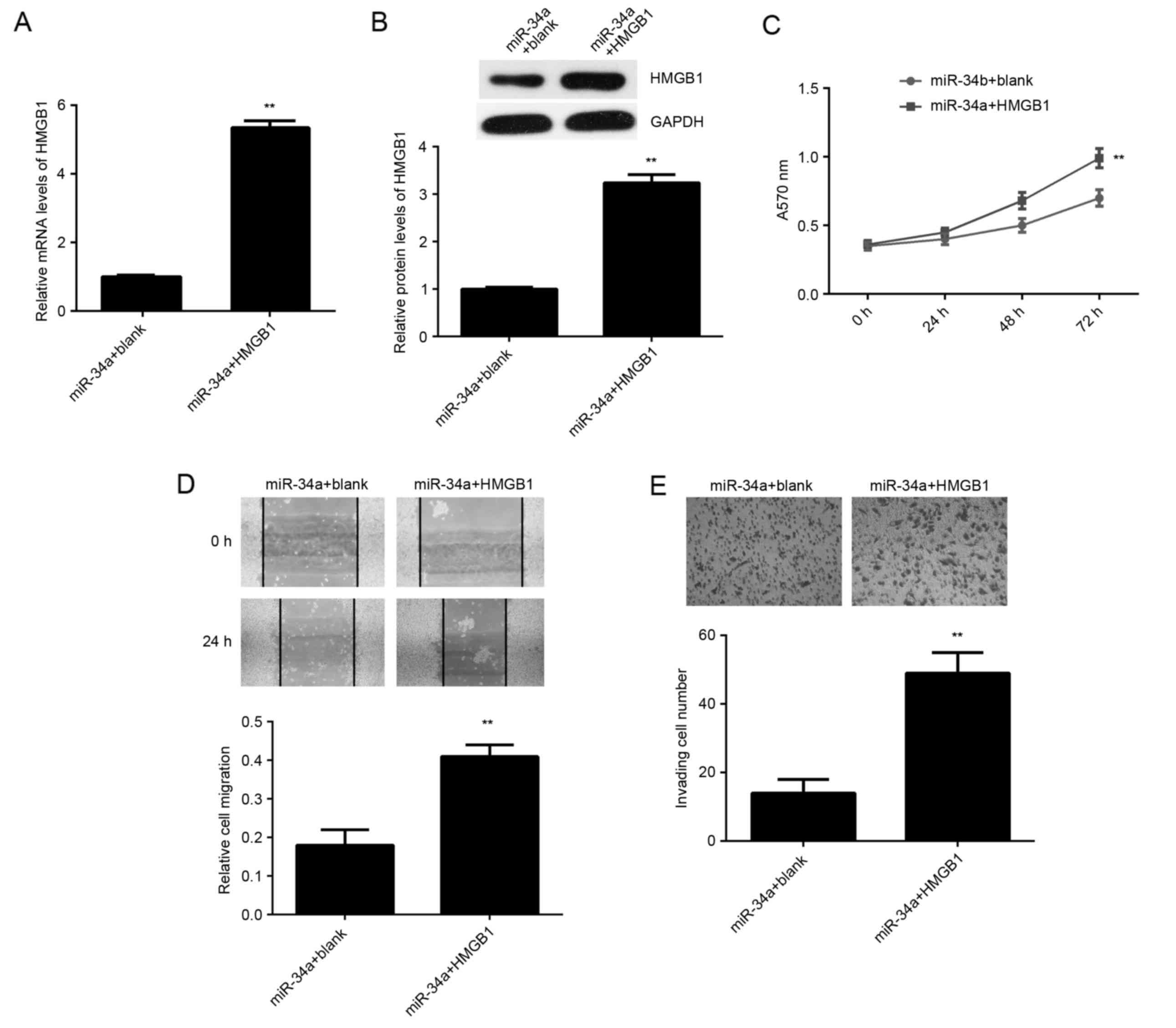
Overexpression of HMGB1 impairs the
suppressive effects of miR-34a on SCL-1 cells
Based on the aforementioned data, it was speculated
that HMGB1 may be involved in the miR-34a-mediated proliferation,
migration and invasion of SCL-1 cells. To clarify this speculation,
miR-34a-overexpressing SCL-1 cells were transfected with
pcDNA3.1-HMGB1 ORF plasmid or with blank pcDNA3.1 vector, serving
as the control. As shown in Fig. 6A and
B, the mRNA and protein levels of HMGB1 were significantly
increased in the miR-34a + HMGB1 group, when compared with the
miR-34a + blank group. The MTT, wound healing and transwell assays
further indicated that the proliferation, migration and invasion of
SCL-1 cells were increased in the miR-34a + HMGB1 group compared
with the miR-34a + blank group (Fig.
6C-E). These findings suggest that miR-34a serves a suppressive
role in the regulation of SCL-1 cell proliferation, migration and
invasion, at least partly, through targeting miR-34a.
Discussion
The regulatory mechanism of miR-34a in CSCC remains
largely unclear. In the present study, it was revealed that miR-34a
was significantly downregulated in CSCC tissues and cell lines,
while low miR-34a expression was associated with the aggressive
progression of CSCC. Restoration of the reduced miR-34a
significantly suppressed the proliferation, migration and invasion
of CSCC SCL-1 cells. HMGB1 was subsequently identified as a target
gene of miR-34a in SCL-1 cells, and its expression was
significantly upregulated in CSCC tissues and cell lines.
Furthermore, the protein expression of HMGB1 was negatively
regulated by miR-34a in SCL-1 cells, and overexpression of HMGB1
impaired the inhibitory effects of miR-34a on SCL-1 cells.
miR-34a has been demonstrated to serve tumor
promoting or suppressive roles in different types of human cancer
(16–18). For instance, Ma et al
(16) reported that miR-34a
inhibited the proliferation and promoted the apoptosis of non-small
cell lung cancer cells by targeting transforming growth factor β
receptor 2. Besides, miR-34a inhibited osteosarcoma cell
proliferation by reducing the expression of ether-à-go-go 1
(17). By contrast, upregulation of
miR-34a has been implicated in invasive cervical cancer (18). Recently, Dotto and Karine (11) reported that miR-34a was significantly
downregulated in skin and oral SCC tissues. In the present study,
it was also observed that miR-34a was significantly downregulated
in CSCC tissues and cell lines, when compared with the adjacent
non-tumor tissues and normal skin cells, respectively. The results
further demonstrated that the reduced expression of miR-34a was
significantly associated with advanced clinical stage and lymph
node metastasis. These findings suggest that downregulation of
miR-34a is implicated in CSCC progression. Further investigation
revealed that restoration of the miR-34a expression significantly
inhibited the proliferation, migration and invasion of CSCC cells,
suggesting that miR-34a may has suppressive effects on the CSCC
growth and metastasis.
As miRs generally function through the inhibition of
the expression of their targets (19), the potential target genes of miR-34a
were subsequently analyzed in the present study using
bioinformatics prediction, and HMGB1 was predicted to be a target
gene of miR-34a. Recently, miR-34a was found to suppress the
proliferation, migration and invasion of human cervical and
colorectal cancer cells via downregulation of HMGB1 (20). However, the association between
miR-34a and HMGB1 in other types of human cancer, including CSCC,
has not been previously reported, to the best of our knowledge.
HMGB1 is generally upregulated in human cancer and functions as an
oncogene (21,22). For instance, Pang et al
(21) reported that HMGB1 was
significantly upregulated in cervical carcinoma, and promoted cell
invasion and migration in vitro. Chen et al (22) revealed that HMGB1 promoted
hepatocellular carcinoma progression through miR-21-mediated matrix
metalloproteinase activity. To verify the association between
miR-34a and HMGB1 in CSCC, a luciferase reporter gene assay was
conducted in the current study, and the data confirmed that HMGB1
was indeed a target gene of miR-34a in CSCC cells. Recently, Sun
et al (14) reported that the
levels of HMGB1 were higher in the CSCC cell supernatant compared
with the human epidermoid carcinoma A431 cell supernatant. In the
present study, HMGB1 was observed to be upregulated in CSCC tissues
and cell lines, when compared with the adjacent non-tumor tissues
and normal skin cells, respectively. Furthermore, overexpression of
miR-34a caused a decrease in HMGB1 expression, while knockdown of
miR-34a increased the HMGB1 expression in CSCC cells. Therefore,
the upregulation of HMGB1 in CSCC may be due to the reduced
expression of miR-34a.
Based on the aforementioned data, it was speculated
that HMGB1 was involved in the suppressive effects of miR-34a on
CSCC cells. Further investigation revealed that overexpression of
HMGB1 impaired the suppressive effects of miR-34a on the CSCC cell
proliferation, migration and invasion, which confirms this
speculation. In addition, certain other miRs have also been
demonstrated to directly target HMGB1, including miR-142 (23), miR-126 (24), miR-181 (25), miR-218 (26), miR-129 (27), miR-24 (28) and miR-141 (29). Therefore, the findings of the present
study further expand the understanding of the regulatory mechanism
of miRs/HMGB1 signaling in human cancer.
In conclusion, the present study demonstrated for
the first time that miR-34a is downregulated in CSCC, and that it
exerts suppressive effects on the proliferation, migration and
invasion of CSCC cells, at least partly, via targeting HMGB1.
Therefore, the present study suggests that miR-34a may be a
potential therapeutic candidate for the treatment of CSCC.
References
|
1
|
Skulsky SL, O'Sullivan B, McArdle O,
Leader M, Roche M, Conlon PJ and O'Neill JP: Review of high-risk
features of cutaneous squamous cell carcinoma and discrepancies
between the American Joint Committee on Cancer and NCCN clinical
practice guidelines in oncology. Head Neck. 39:578–594. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Cheng J and Yan S: Prognostic variables in
high-risk cutaneous squamous cell carcinoma: A review. J Cutan
Pathol. 43:994–1004. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Sherwood V and Leigh IM: WNT signaling in
cutaneous squamous cell carcinoma: A future treatment strategy? J
Invest Dermatol. 136:1760–1767. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Zhu K, He Y, Xia C, Yan J, Hou J, Kong D,
Yang Y and Zheng G: MicroRNA-15a inhibits proliferation and induces
apoptosis in CNE1 nasopharyngeal carcinoma cells. Oncol Res.
24:145–151. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Wang G, Fu Y, Liu G, Ye Y and Zhang X:
miR-218 inhibits proliferation, migration, and EMT of gastric
cancer cells by targeting WASF3. Oncol Res. 25:355–364. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Lv H, Zhang Z, Wang Y, Li C, Gong W and
Wang X: MicroRNA-92a promotes colorectal cancer cell growth and
migration by inhibiting KLF4. Oncol Res. 23:283–290. 2016.
View Article : Google Scholar
|
|
7
|
Liu X, Li J, Yu Z, Sun R and Kan Q:
MiR-935 promotes liver cancer cell proliferation and migration by
targeting SOX7. Oncol Res. 25:427–435. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Ji S, Zhang B, Kong Y, Ma F and Hua Y:
MiR-326 inhibits gastric cancer cell growth through down regulating
NOB1. Oncol Res. 25:853–861. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Zhou M, Zhou L, Zheng L, Guo L, Wang Y,
Liu H, Ou C and Ding Z: miR-365 promotes cutaneous squamous cell
carcinoma (CSCC) through targeting nuclear factor I/B (NFIB). PLoS
One. 9:e1006202014. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Xu N, Zhang L, Meisgen F, Harada M,
Heilborn J, Homey B, Grandér D, Ståhle M, Sonkoly E and Pivarcsi A:
MicroRNA-125b down-regulates matrix metallopeptidase 13 and
inhibits cutaneous squamous cell carcinoma cell proliferation,
migration, and invasion. J Biol Chem. 287:29899–29908. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Dotto GP and Karine L: miR-34a/SIRT6 in
squamous differentiation and cancer. Cell Cycle. 13:1055–1056.
2014. View
Article : Google Scholar : PubMed/NCBI
|
|
12
|
Ding J, Cui X and Liu Q: Emerging role of
HMGB1 in lung diseases: Friend or foe. J Cell Mol Med.
21:1046–1057. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Anggayasti WL, Mancera RL, Bottomley S and
Helmerhorst E: The self-association of HMGB1 and its possible role
in the binding to DNA and cell-membrane receptors. FEBS Lett.
591:282–294. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Sun Y, Tu Y, He LI, Ji C and Cheng BO:
High mobility group box 1 regulates tumor metastasis in cutaneous
squamous cell carcinoma via the PI3K/AKT and MAPK signaling
pathways. Oncol Lett. 11:59–62. 2016.PubMed/NCBI
|
|
15
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Ma ZL, Hou PP, Li YL, Wang DT, Yuan TW,
Wei JL, Zhao BT, Lou JT, Zhao XT, Jin Y and Jin YX: MicroRNA-34a
inhibits the proliferation and promotes the apoptosis of non-small
cell lung cancer H1299 cell line by targeting TGFβR2. Tumour Biol.
36:2481–2490. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Wu X, Zhong D, Gao Q, Zhai W, Ding Z and
Wu J: MicroRNA-34a inhibits human osteosarcoma proliferation by
downregulating ether à go-go 1 expression. Int J Med Sci.
10:676–682. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Ribeiro J, Marinho-Dias J, Monteiro P,
Loureiro J, Baldaque I, Medeiros R and Sousa H: miR-34a and
miR-125b expression in HPV infection and cervical cancer
development. Biomed Res Int. 2015:3045842015. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Ambros V: The functions of animal
microRNAs. Nature. 431:350–355. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Chandrasekaran KS, Sathyanarayanan A and
Karunagaran D: Downregulation of HMGB1 by miR-34a is sufficient to
suppress proliferation, migration and invasion of human cervical
and colorectal cancer cells. Tumour Biol. 37:13155–13166. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Pang X, Zhang Y and Zhang S: High-mobility
group box 1 is overexpressed in cervical carcinoma and promotes
cell invasion and migration in vitro. Oncol Rep. 37:831–840. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Chen M, Liu Y, Varley P, Chang Y, He XX,
Huang H, Tang D, Lotze MT, Lin J and Tsung A: High-mobility group
box 1 promotes hepatocellular carcinoma progression through
miR-21-mediated matrix metalloproteinase activity. Cancer Res.
75:1645–1656. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Wang Y, Ouyang M, Wang Q and Jian Z:
MicroRNA-142-3p inhibits hypoxia/reoxygenation-induced apoptosis
and fibrosis of cardiomyocytes by targeting high mobility group box
1. Int J Mol Med. 38:1377–1386. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Tang ST, Wang F, Shao M, Wang Y and Zhu
HQ: MicroRNA-126 suppresses inflammation in endothelial cells under
hyperglycemic condition by targeting HMGB1. Vascul Pharmacol.
88:48–55. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Liu Y, Hu X, Xia D and Zhang S:
MicroRNA-181b is downregulated in non-small cell lung cancer and
inhibits cell motility by directly targeting HMGB1. Oncol Lett.
12:4181–4186. 2016.PubMed/NCBI
|
|
26
|
Gu J, Xu R, Li Y, Zhang J and Wang S:
MicroRNA-218 modulates activities of glioma cells by targeting
HMGB1. Am J Transl Res. 8:3780–3790. 2016.PubMed/NCBI
|
|
27
|
Liu Z, Dou C, Yao B, Xu M, Ding L, Wang Y,
Jia Y, Li Q, Zhang H, Tu K, et al: Methylation-mediated repression
of microRNA-129-2 suppresses cell aggressiveness by inhibiting high
mobility group box 1 in human hepatocellular carcinoma. Oncotarget.
7:36909–36923. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Yang J, Chen L, Ding J, Fan Z, Li S, Wu H,
Zhang J, Yang C, Wang H, Zeng P and Yang J: MicroRNA-24 inhibits
high glucose-induced vascular smooth muscle cell proliferation and
migration by targeting HMGB1. Gene. 586:268–273. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Zhu H, Huang L, Zhu S, Li X, Li Z, Yu C
and Yu X: Regulation of autophagy by systemic admission of
microRNA-141 to target HMGB1 in l-arginine-induced acute
pancreatitis in vivo. Pancreatology. 16:337–346. 2016. View Article : Google Scholar : PubMed/NCBI
|