Introduction
Enterobacteriaceae, such as Escherichia coli,
Klebsiella pneumoniae, Citrobacter freundii and
Enterobacter cloacae, are the most common pathogenic
bacteria for nosocomial infection (1). According to the results of national
bacterial resistance monitoring over the years, Enterobacteriaceae
accounted for 60–70% of all Gram-negative bacteria. Carbapenems,
such as imipenem, meropenem and ertapenem, are the most effective
antibiotics for the clinical treatment of infection caused by
Enterobacteriaceae, especially multi-drug-resistant strains
producing extended-spectrum β-lactamases (ESBLs) and AmpC enzyme
(2). However, carbapenem-resistant
Enterobacteriaceae (CRE) are emerging as carbapenems have been
widely used in clinical practice. For example, the resistance rates
of E. coli and K. pneumoniae have risen from 0 to 1%,
and those of some other members of the Enterobacteriaceae family
have already reached 7.0%. Some non-fermenters also have the
resistance rates of 20–30% (3).
Since most carbapenem-resistant bacteria also resist many commonly
used antibiotics, they have become pan-resistant strains, posing a
great threat to the life of patients and challenging the clinical
application (4). Therefore, it is
imperative to clarify the resistance mechanism of CRE strains and
to effectively control the resulting nosocomial infection.
Production of carbapenemases is one of the main
mechanisms for the resistance of CRE strains. Besides, these
strains can also produce ESBLs and/or AmpC enzyme as well as lose
outer membrane porin (OMP) proteins (5). Carbapenemases are β-lactamases using
carbapenems as hydrolysis substrates, including Ambler classes A, B
and C enzymes. For Enterobacteriaceae, KPC, as a class A
carbapenemase, has been highlighted. KPC enzymes can hydrolyze
almost all β-lactams including penicillins, cephalosporins and
carbapenems, but can be easily inhibited by clavulanic acid and
tazobactam. Meanwhile, they are resistant to other types of
antibiotics (6). From 2006 to 2016,
KPC-2-producing K. pneumoniae, C. freundii and
Serratia marcescens were isolated from the Zhejiang province
of China, indicating that KPC enzymes had been spread to bacteria
of Enterobacteriaceae other than K. pneumoniae (7).
The permeability of outer membrane is vital to drug
entrance into Gram-negative bacteria. OMP proteins can mainly be
classified into OmpC with high molecular weights and OmpF with low
molecular weights. Either deletion or decrease of OMP proteins
leads to bacterial drug resistance. As to Enterobacteriaceae, the
resistance to carbapenems may be ascribed to OMP protein deletion
or reduction in the presence/absence of production of β-lactamases
(e.g., AmpC enzymes and ESBLs). The OMP proteins of K.
pneumoniae are mainly OmpA and OmpK. Carbapenemase-producing
‘super bacteria’, especially NDM-1 and its variants, has become a
major public health concern worldwide. NDM-1 can hydrolyze a wide
range of β-lactam antibiotics including carbapenems, as the last
resort of antibiotics for treating infections induced by resistant
bacteria (8).
Thereby motivated, we aimed to study the incidence
of carbapenemase-producing strains in clinical isolates of
Enterobacteriaceae and the mechanism by which CRE strains resisted
carbapenems. Carbapenemases, ESBLs and AmpC enzyme genes in 78
clinically isolated strains were tested by β-lactamase detection
and DNA sequencing. The types and genotypes of these enzymes were
confirmed. OMP proteins in CRE strains were detected to clarify the
relationship between carbapenem resistance and deletion of these
proteins.
Materials and methods
Experimental and quality control
strains
A total of 6,584 Enterobacteriaceae strains
clinically isolated from December 2011 to December 2015 in our
hospital were selected, from which 78 strains that resisted any
carbapenem were screened. All strains were identified by API or ATB
(BioMérieux, Craponne, France). E. coli ATCC 25922 was used
as the quality control strain for the drug susceptibility test, and
clinically isolated C. freundii known to produce IMP-8
metalloenzyme was utilized as the positive quality control strain
in modified Hodge test.
E. coli producing TEM and SHV β-lactamases
and E. coli producing CTX-M and TOHO ESBLs were used as
positive control strains for ESBLs detection by polymerase chain
reaction (PCR). Clinically isolated strains were used as the
sensitive controls for SDS-PAGE of OMP proteins, and Lambda Marker
was employed as the standard for bacterial typing using
pulsed-field gel electrophoresis. The bacteria were all preserved
by our hospital.
Screening of CRE strains by the KB
test for drug susceptibility
Bacterial suspension at the concentration of
1.5×108/ml was uniformly coated onto the entire surface
of Mueller-Hinton agar culture medium and dried at room temperature
for several min. Then antibiotics-loaded filter paper discs were
uniformly stuck onto the agar surface by a sterile tweezer and
cultured in a 35°C incubator for 18–24 h. Afterwards, the
inhibitory circle diameter (mm) including the disc diameter was
measured by a vernier caliper. The experiment was performed and the
drug susceptibility was determined according to the standards (2010
edition) of the Clinical Laboratory Standards Institute (CLSI). The
susceptibilities to imipenem, meropenem and ertapenem were detected
by the disc diffusion method (Table
I).
 | Table I.Standards for determining drug
susceptibility using the Kirby-Bauer method (mm). |
Table I.
Standards for determining drug
susceptibility using the Kirby-Bauer method (mm).
| Drug | Drug content of disc
(µg) | Susceptible (S) | Intermediate (I) | Resistant (R) |
|---|
| Imipenem | 10 | ≥16 | 14–15 | ≤13 |
| Meropenem | 10 | ≥16 | 14–15 | ≤13 |
| Ertapenem | 10 | ≥19 | 16–18 | ≤15 |
Detection of carbapenemases by
modified Hodge test
The experiment was conducted according to the method
recommended by CLSI. E. coli ATCC 25,922 with a 0.5
McFarland turbidity standard was diluted and spread on an MH plate
with a paper strip of ertapenem (10 µg) stuck in the middle.
Bacteria were inoculated by an inoculating loop through streaking
from the outer edge of the paper strip to the edge of the plate,
during which particular attention should be paid not to scratch the
plate surface. The bacteria were then cultured overnight at 35°C.
On the next day, the bacteria exhibiting sagittal growth in the
inhibition zone of ertapenem were identified to produce
carbapenemases.
Detection of OMP proteins in 78 CRE
strains by SDS-PAGE
The strains stored at −80°C were thawed at room
temperature, inoculated on MH agar plates, and incubated at 37°C
for 18–20 h. A single colony was picked up, inoculated in LB
medium, cultured while shaking for 20 h, and centrifuged at 10,000
rpm and 4°C for 10 min to obtain the bacterial bodies that were
then washed twice with PBS, resuspended, ultrasonically fragmented,
and centrifuged at 20,000 rpm and 4°C for 10 min. After the
supernatant was discarded, the precipitate was suspended in PBS
containing 2% sodium dodecylamine and centrifuged at 16,000 rpm and
room temperature for 30 min, and OMP proteins were resuspended in
100 µl of PBS solution. The gel was fixed in an electrophoresis
device and added Tris-glycine electrophoresis buffer. The samples
were added in a predetermined order using a Hamilton microinjector,
15 µl for each sample. Attention should be paid to positive and
negative electrodes when the electrophoresis device was connected
with the power supply. Firstly, 50 V of voltage was applied, which
was then increased to 80 V after the front edge of dye entered the
separating gel. The electrophoresis was continued until the
indicating line of bromophenol blue reached the bottom of the
separating gel (about 2–3 h). The gel was thereafter removed,
soaked in a 5-fold volume of Coomassie brilliant blue staining
solution, and stained by gentle shaking on a horizontal shaker for
over 4 h at room temperature. Afterwards, the gel was taken out,
soaked in a destaining solution, and destained by gentle shaking
for 4–8 h, during which the destaining solution was refreshed 3–4
times. After destaining, the gel was photographed by Bio-Rad
GelDocXR gel imaging system (Bio-Rad Laboratories, Inc., Hercules,
CA, USA), and the relative expressions of OMP protein bands in
sensitive and resistant strains were measured by Quantity One image
analysis software (Bio-Rad Laboratories, Inc.).
Detection of carbapenemases, ESBLs and
plasmid AmpC enzyme genes in 78 CRE strains by PCR
Extraction of bacteria by boiling: A colony of
bacteria purely cultured overnight was suspended in 500 µl of TE
buffer, heated at 100°C for 13 min and centrifuged at 10,000 rpm
for 3 min. Then 2 µl of the supernatant was used as the template
for PCR reaction. The reaction system was 50 µl, including 5 µl of
10X buffer, 4 µl of dNTP, 1 µl of each primer, 0.25 µl of Tag
enzyme, 37 µl of ddH2O and 2 µl of template. Reaction
conditions: Pre-denaturation at 94°C for 5 min, 30 cycles of
reaction at 55°C for 45 sec and at 72°C for 1 min, and finally
extension at 72°C for 10 min. The primer sequences for PCR are
listed in Table II. The
electrophoresis of PCR products was conducted using 1.5% agarose
gel at 130 V for 35 min. Subsequently, the gel was stained with
ethidium bromide for 20 min and ultimately photographed by the gel
imaging system.
 | Table II.Primer sequences for PCR. |
Table II.
Primer sequences for PCR.
| Primer | Sequence (5′-3′) |
|---|
| KPC-R |
TCGCTAAACTCGAACAGG |
| KPC-F |
TTACTGCCCGTTGACGCCCAATCC |
| IMP-R |
CTACCGCAGCAGAGTCTTTG |
| IMP-F |
AACCAGTTTTGCCTTACCAT |
| IMI-R |
ATAGCCATCTTGTTTAGCTC |
| IMI-F |
TCTGCGATTACTTTATCCTC |
| NMC-R |
TGCAGCTTAATTATTTTCAGATTAG |
| NMC-F |
ATTTTTTTCATGATGAAGTTAAGCC |
| GIM-F |
AGAACCTTGACCGAACGCAG |
| GIM-R |
ACTCATGACTCCTCACGAGG |
| VIM-F |
TCTACATGACCGCGTCTGTC |
| VIM-R |
TGTGCTTTGACAACGTTCGC |
| SEM-IRS1 |
AACGGCTTCATTTTTGTTTAG |
| SEM-IRS2 |
GCTTCCGCAATAGTTTTATCA |
| SEM-IRS5 |
AGATAGTAAATTTTATAG |
| SEM-IRS6 | CTCTAACGCTAATAG |
| IND-R |
GCCCAGGTTAAAGATTTTGTAAT |
| IND-F |
CATGGCCACCGCCTTTCCATTC |
| IND-R1 |
GGTTTGCATATCTATCTGCC |
| IND-F1 |
ATCCAAAGAGAGGCTGGAGT |
| OXA-69-F |
CTAATAATTGATCTACTCAAG |
| OXA-69-R |
CCAGTGGATGGATGGATAGATTATC |
| OXA-55-F |
CATCTACCTTTAAAATTCCC |
| OXA-55-R |
AGCTGTTCCTGCTTGAGCAC |
| OXA-48-F |
TTGGTGGCATCGATTATCGG |
| OXA-48-R |
GAGCACTTCTTTTGTGATGGC |
| OXA-50-F |
AATCCGGCGCTCATCCATC |
| OXA-50-R |
GGTCGGCGACTGAGGGGG |
| OXA-60-F |
AAAGGAGTTGTCTCATGCTGTCTCG |
| OXA-60-R |
AACCTACAGGCGCGCGTCTCACGGTG |
Statistical analysis
All data were analyzed by GraphPad (GraphPad
Software, Inc., La Jolla, CA, USA). All experiments were performed
in triplicate, and the results were expressed as number and
percentage.
Results
Distribution of 78 CRE strains
The 78 CRE strains were isolated from urine,
respiratory tract, blood, cerebrospinal fluid and wound samples,
with the isolation rates of 55.1 (43/78), 41.0 (32/78), 1.3
(12/78), 1.3 (12/78) and 1.3% (12/78) respectively.
Besides, 71.8% (56/78) of the CRE strains were
isolated from wards in neurosurgery department, and the remaining
was from ICU [10.3% (8/78)], internal medicine department [6.4%
(5/78)], VIP wards [5.1% (4/78)], outpatient department [2.6%
(2/78)], foreigner department [2.6% (2/78)] and surgery department
[1.3% (1/78)].
CRE strains could be isolated from all patients
hospitalized in the same bed at different times from 2011 to 2015.
For example, three C. freundii strains and one K.
pneumoniae strain resisting carbapenems were isolated from four
patients hospitalized in January 2012, May and September 2013 and
February 2015. For the same patient, CRE strains with different
species could also be isolated. In the beginning, K.
pneumoniae strains were susceptible to carbapenems, but became
resistant after treatment for a period of time.
Susceptibility of CRE strains to
antibacterial agents
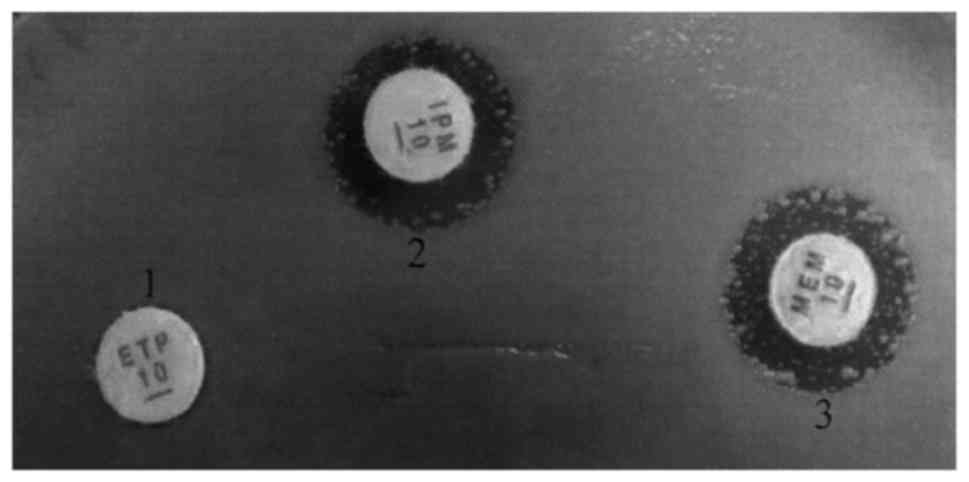
Drug susceptibility test (Fig. 1 and Table III) showed that 83.3% (63/78) of
the CRE strains were resistant to imipenem, meropenem and ertapenem
simultaneously.
 | Table III.Drug susceptibility test results
using the Kirby-Bauer method (n%). |
Table III.
Drug susceptibility test results
using the Kirby-Bauer method (n%).
| Drug | Number of
strains | % |
|---|
| Imipenem | 71 | 91.0 |
| Meropenem | 70 | 89.7 |
| Ertapenem | 72 | 92.3 |
| Imipenem +
meropenem | 66 | 84.6 |
| Imipenem +
ertapenem | 65 | 83.3 |
| Meropenem +
ertapenem | 67 | 85.9 |
| Imipenem +
meropenem + ertapenem | 63 | 83.0 |
Detection results of carbapenemase
genes
PCR showed that the 78 CRE strains had negative
detection results of VIM-2, SPM, NMC, IMI, IND, OXA-48, OXA-50,
OXA-55, OXA-60 and OXA-24 like carbapenemase genes. As shown in
Fig. 2A, 33.3% (26/78) of the
strains produce KPC-2 carbapenemase, of which 7.7 (2/26), 7.7
(2/26) and 3.8% (1/26) simultaneously produce VIM-1, IMP-2 and
IMP-1 carbapenemases respectively. In addition, 6.4 (5/78), 5.1
(4/78), 3.8 (3/78), 6.4 (5/78) and 12.8% (10/78) of the strains
produced VIM-1, IMP-1, IMP-2, GIM and OXA-69 carbapenemases
respectively (Fig. 2B). As to OXA
carbapenemases, 20.5% (16/78) of the strains produced OXA-23 like,
OXA-51 like or OXA-58 like carbapenemases.
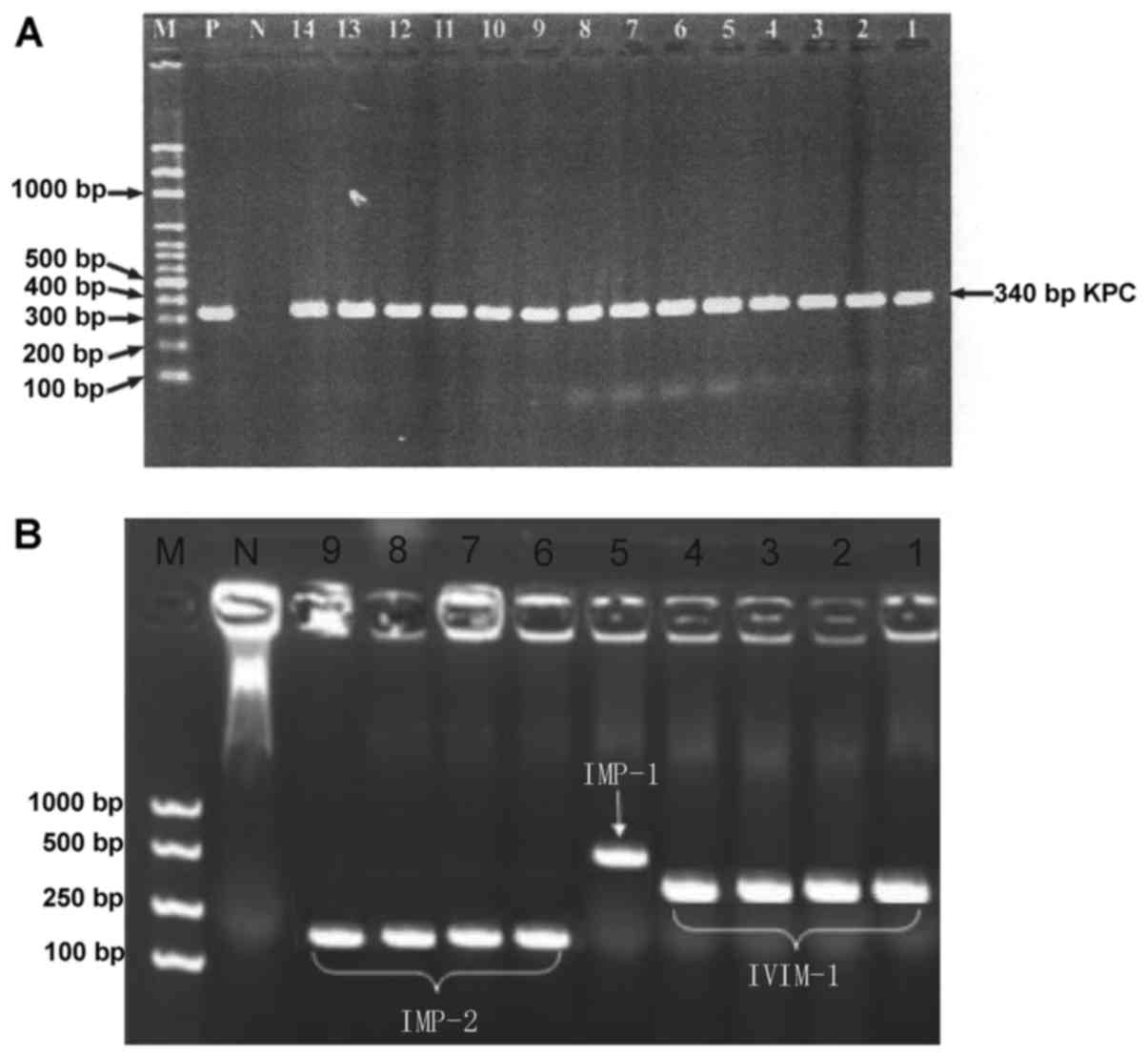 | Figure 2.(A) PCR results of CRE strains
producing KPC carbapenemases. 1–6, K. pneumoniae; 7–14,
C. freundii; N, negative control E. coli ATCC 25922;
P, positive control KPN2528; M, molecular weight marker. (B) PCR
results of CRE strains producing metallocarbapenemases. 1–4, K.
pneumoniae (VIM-1); 5, 1551K. pneumoniae (IMP-1); 6–9,
C. freundii (IMP-2); N, negative control E. coli ATCC
25922; M, molecular weight marker. PCR, polymerase chain reaction;
CRE, carbapenem-resistant Enterobacteriaceae; K. pneumoniae,
Klebsiella pneumoniae; C. freundii, Citrobacter
freundii; E. coli, Escherichia coli. |
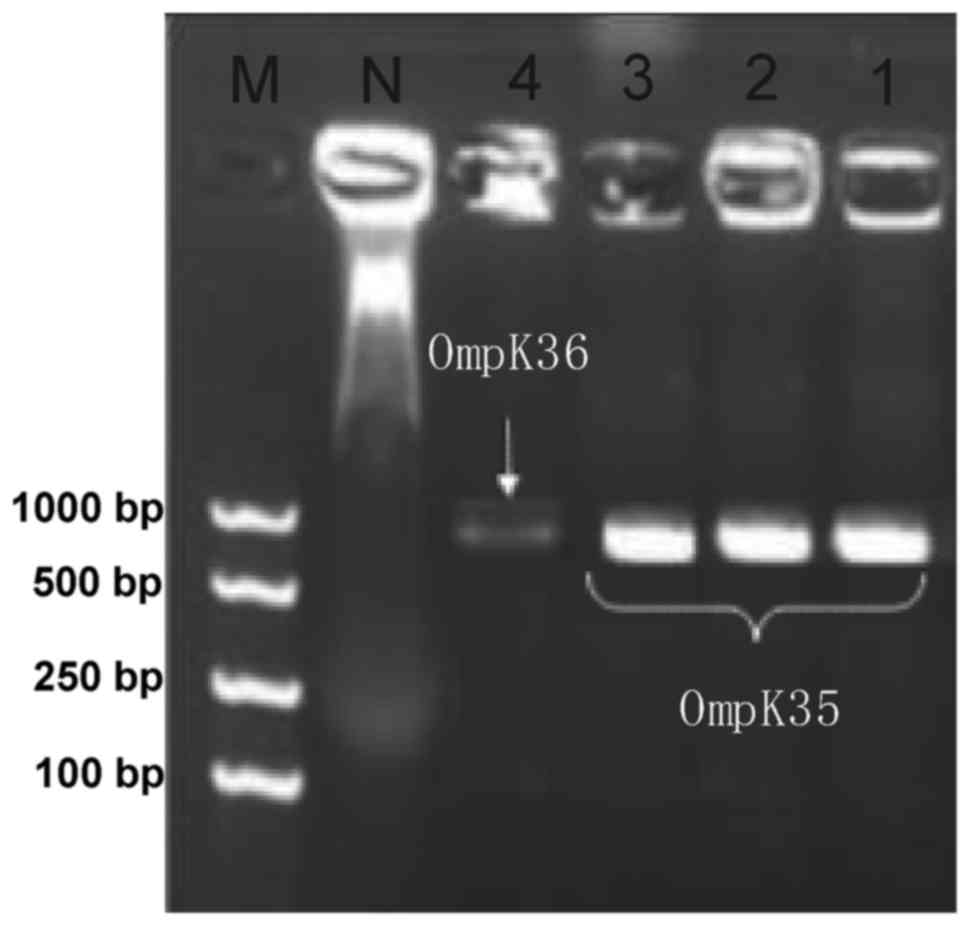
PCR results of OMP genes
PCR revealed that the detection rates of OmpK35,
OmpK36 and Ompk37 in the 78 strains were 37.2 (29/78), 6.4 (5/78)
and 3.8% (3/78) respectively, but 61.5% of the strains had negative
detection results for these three genes (Fig. 3 and Table
IV).
 | Table IV.Polymerase chain reaction results of
OMP genes. |
Table IV.
Polymerase chain reaction results of
OMP genes.
| Type | Number of
strains | % |
|---|
| All negative | 48 | 61.5 |
|
OmpK35-positive | 23 | 29.5 |
|
OmpK37-positive | 1 | 1.3 |
| OmpK35- and
OmpK36-positive | 4 | 5.1 |
| OmpK35- and
OmpK37-positive | 1 | 1.3 |
| OmpK35-, OmpK36-
and OmpK37-positive | 1 | 1.3 |
| Total | 78 | 100.0 |
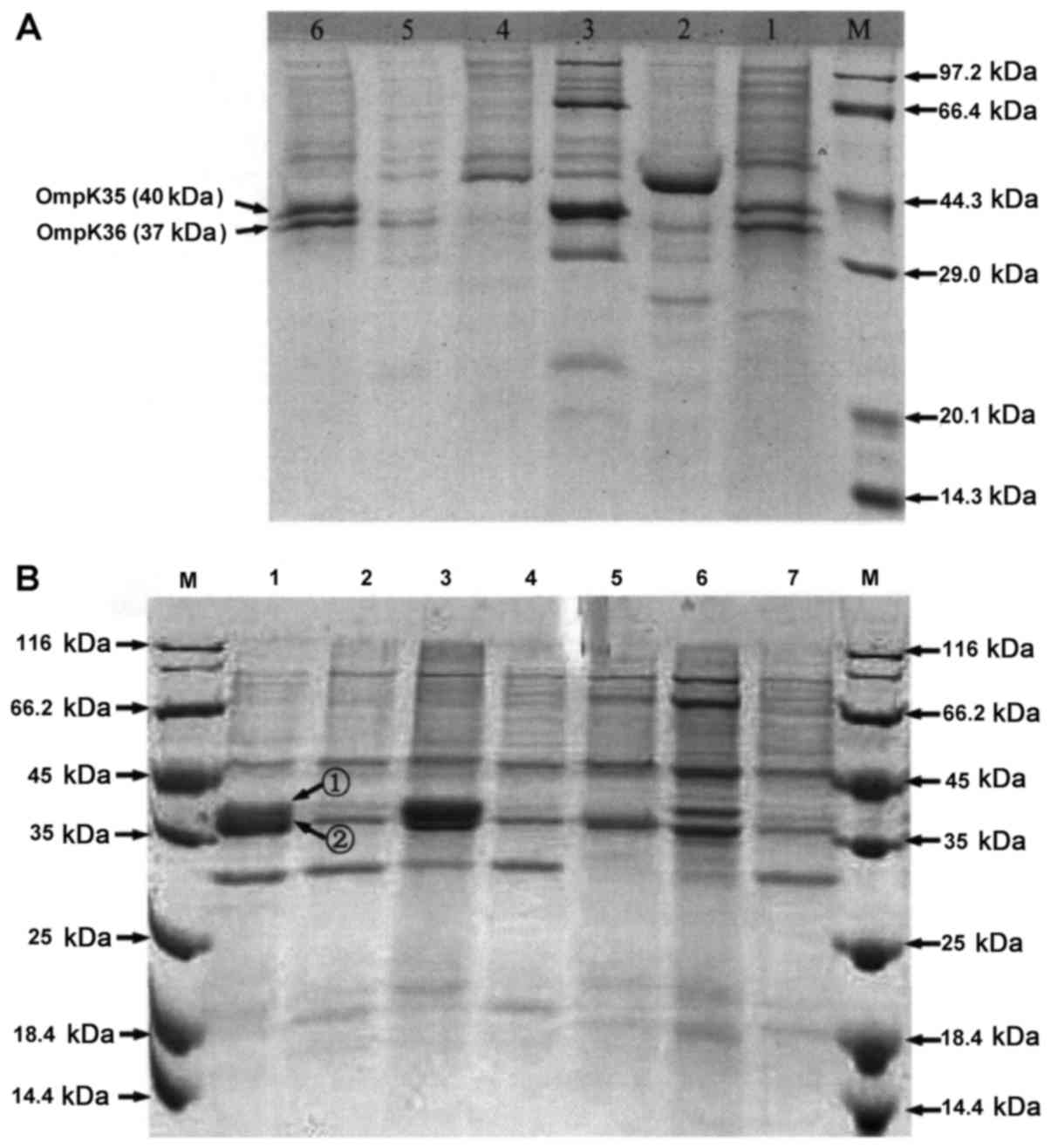
SDS-PAGE results of OMP proteins
SDS-PAGE showed that 40.9% of 44 K.
pneumoniae strains had normal OMP protein expressions, and the
remaining 59.1% had downregulated or deleted expression of OmpK35
or OmpK36 (Fig. 4 and Table V). Of 34 C. freundii strains,
only 2.9% had normal OMP protein expression, and the remaining
97.1% had downregulated or deleted expression of OmpC or OmpF. The
OMP proteins in most CRE strains were deleted or decreased compared
with those in the sensitive ones, indicating that they played
crucial roles in the resistant of CRE strains to carbapenems.
 | Table V.SDS-PAGE results of OMP proteins. |
Table V.
SDS-PAGE results of OMP proteins.
| A,
Klebsiella (44 strains) |
|---|
|
|---|
| Protein change | Number of
strains | % |
|---|
| Normal | 18 | 40.9 |
| Downregulated
OmpK35 + deleted OmpK36 | 5 | 11.4 |
| Downregulated
OmpK35 + OmpK36 | 3 | 6.8 |
| Normal OmpK35 +
downregulated OmpK36 | 3 | 6.8 |
| Deleted OmpK35 +
normal OmpK36 | 1 | 2.3 |
| Normal OmpK35 +
deleted OmpK36 | 4 | 9.1 |
| Deleted OmpK35 +
OmpK36 | 4 | 9.1 |
| Downregulated
OmpK35 + normal OmpK36 | 3 | 6.8 |
| Deleted OmpK35 +
downregulated OmpK36 | 3 | 6.8 |
|
| B, Other
Enterobacteriaceae (34 strains) |
|
| Protein
change | Number of
strains | % |
|
| Downregulated OmpC
+ OmpF | 21 | 61.8 |
| Normal OmpC +
OmpF | 1 | 2.9 |
| Deleted OmpC +
downregulated OmpF | 5 | 14.7 |
| Normal OmpC +
downregulated OmpF | 1 | 2.9 |
| Deleted OmpC +
OmpF | 3 | 8.8 |
| Downregulated OmpC
+ deleted OmpF | 3 | 8.8 |
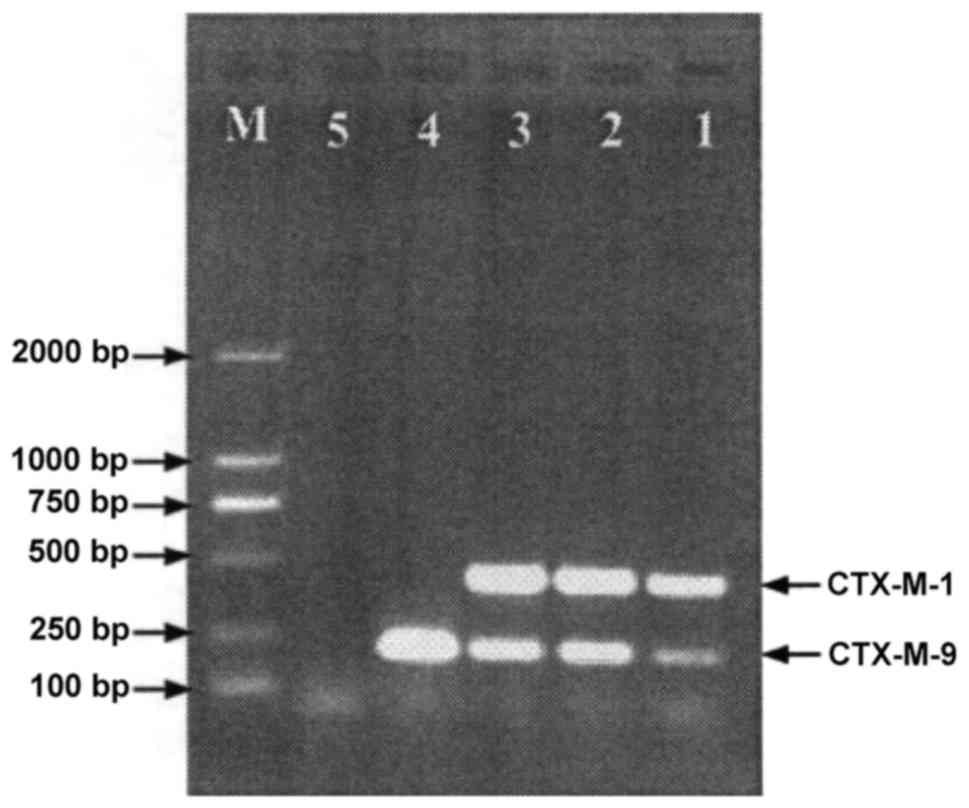
PCR results of ESBLs and plasmid AmpC
enzyme genes
PCR revealed that 83.3% (65/78) of the strains
produced CTX-M ESBLs (Fig. 5 and
Table VI), and 1.3 (1/78), 70.5
(55/78) and 25.6% (20/78) produced CTX-M-2, CTX-M-14 and CTX-M-15
ESBLs respectively. Moreover, 39.7% (31/78) of the strains produced
DHA-1 or CMY-2 plasmid AmpC enzyme (Fig.
6). Although carbapenemase genes were not detected in 39.7%
(31/78) of the strains, 87.9% (29/33) produced CTX-M ESBLs
(CTX-M-15, 48.3%; CTX-M-14, 72.4%; CTX-M-14L and CTX-M-15
simultaneously, 20.7%). Of all CRE strains, 35.9% (28/78) at least
lose one band of OMP protein.
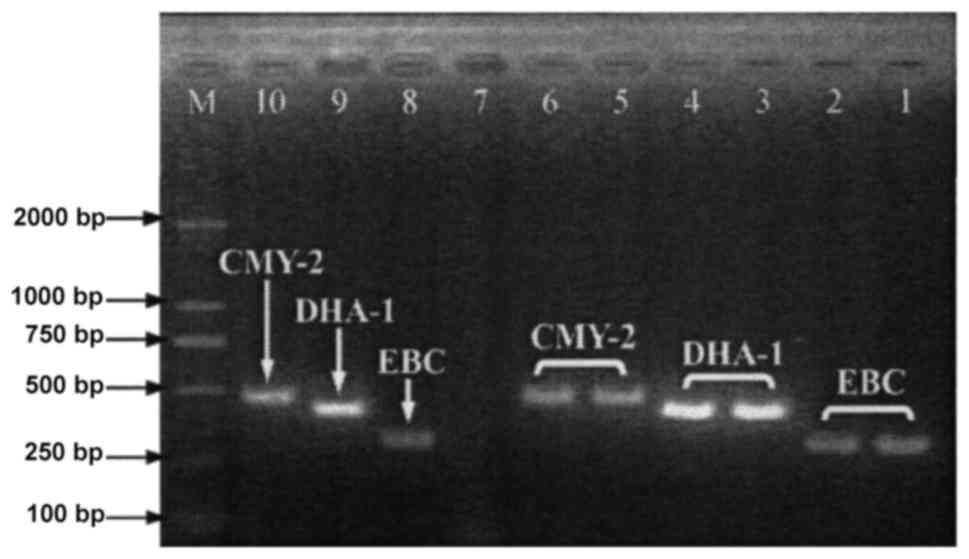 | Figure 6.PCR results of plasmid AmpC enzyme in
E. coli and K. pneumoniae. 1 and 2, EBC positive
control strains; 3 and 4, DHA-1 gene-positive; 5 and 6, CMY-2
gene-positive; 7, negative control; 8–10, EBC, DHA-1 gene and CMY-2
gene positive controls; M, molecular weight marker. PCR, polymerase
chain reaction; E. coli, Escherichia coli; K.
pneumoniae, Klebsiella pneumoniae. |
 | Table VI.ESBL genes in 78 CRE strains. |
Table VI.
ESBL genes in 78 CRE strains.
|
| KPN (43
strains) | CFR (25
strains) | Others (10
strains) | All strains |
|---|
| Resistant
genes | Number of
strains | % | Number of
strains | % | Number of
strains | % | Number of
strains | % |
|---|
| Negative | 0 |
0.0 | 0 |
0.0 | 3 |
30.0 | 3 |
3.8 |
| ESBLs | 16 |
37.2 | 2 |
8.0 | 2 |
20.0 | 20 |
25.6 |
| ESBLs + AmpC | 4 |
9.3 | 4 |
16.0 |
|
| 8 |
10.3 |
| Carbapenemase | 7 |
16.3 | 1 |
4.0 | 4 |
40.0 | 12 |
15.4 |
| Carbapenemase +
ESBLs | 10 |
23.3 | 3 |
12.0 |
|
| 13 |
16.7 |
| Carbapenemase +
ESBLs + AmpC | 6 |
14.0 | 15 |
60.0 | 1 |
10.0 | 22 |
28.2 |
| Total | 43 | 100.0 | 25 | 100.0 | 10 | 100.0 | 78 | 100.0 |
Based on the modified Hodge test, 93.6% (73/78) of
the strains produced carbapenemases, of which 35.6% (26/73) did not
contain carbapenemase genes.
Discussion
Enterobacteriaceae are Gram-negative, facultative
anaerobic bacilli or coccobacilli widely distributed in the
environment, mainly including Escherichia,
Klebsiella, Enterobacter, Citrobacter,
Serratia and Salmonella (9,10). As
important pathogenic bacteria for community-acquired infections and
nosocomial infections, Enterobacteriaceae can infect the
respiratory tract and urethra. Of all clinically isolated
Gram-negative bacilli, Enterobacteriaceae account for 80%, and they
also account for 50% of the bacteria detected in clinical
laboratories. Additionally, approximately 50% of septicemia and
over 70% of urinary infections are caused by Enterobacteriaceae
(11–13). Carbapenems (imipenem, meropenem and
ertapenem), which have broad antibacterial spectra as well as high
antibacterial activities and stability to β-lactamases, are potent
antibiotics for treating infections induced by Gram-negative
bacteria (14). However,
Enterobacteriaceae have become increasingly resistant to
carbapenems owing to extensive and irrational use in clinical
practice. As a result, the treatment outcomes are affected,
increasing the economic burden of patients. CRE directly endanger
human life and health by elevating the mortality rate. Recently,
humans have been threatened by KPC enzyme-producing Gram-negative
bacilli (4). Steinmann et al
reported that from July 2010 to January 2011, four patients died of
KPC-2-producing K. pneumoniae in a German hospital (15). Hu et al detected KPC
carbapenemases in 77 Enterobacteriaceae strains from Huashan
Hospital, Fudan University (16). In
March 2011, NDM-1-producing E. coli was isolated in Hong
Kong, China (17). Of 46
Enterobacteriaceae strains with decreased susceptibility to
carbapenems collected from 2004 to 2008 (18), 8 produced IMP-4 and 2 produced IMP-8.
Antibiotics enter bacterial cells via OMPs as porous channels. When
these proteins are deleted or reduced, the outer membrane
permeability is decreased, leading to drug resistance by hindering
their entrance. Enterobacteriaceae (e.g., E. cloacae and
Enterobacter aerogenes) not only have OmpK35 and OmpK36
analogous to OmpF and OmpC, but also express OmpD (19). One of the mechanisms for bacterial
drug resistance is that antibacterial agents fail to permeate into
cells. For Gram-negative bacteria, the permeability of OMPs is key
to the entrance of exit of antibiotics. OMPs related to the drug
resistance of Enterobacteriaceae mainly function through synthesis
reduction or deletion, mutation, blockage of channels and
replacement by other proteins. In this study, PCR and SDS-PAGE
showed that 60.3% (47/78) of the 78 strains produced
carbapenemases, and 33.3% (26/78) produced KPC-2 carbapenemase.
Accordingly, carbapenemases may be responsible for the resistance
of CRE strains to carbapenems. Compared with sensitive strains,
OMPs of most CRE strains were deleted or decreased. Of the 44
Klebsiella strains, 59.1% (26/44) did not express or
expressed less OmpK35 or OmpK36. Among the 34 strains of other
enterobacteria, 97.1% (33/34) did not express or expressed less
OmpC or OmpF. Of all CRE strains, 35.9% (28/78) lost at least one
OMP protein, suggesting that the strains resisted carbapenems also
by producing ESBLs and/or plasmid AmpC enzyme as well as by losing
OMP proteins.
The resistance of clinically isolated CRE strains
can primarily be ascribed to production of carbapenemases, also
involving deletion of OMP proteins or mutation of OMP genes. In
clinical practice, it is of great significance to monitor and to
control carbapenemase-producing CRE strains by PCR. In addition,
the emergence of resistant bacterial strains is directly associated
with irrational use of antibiotics, so it is crucial to timely find
them by predicting the probability of drug resistance and
performing drug susceptibility test when carbapenems are used. In
summary, severe infections can be prevented by rationally using
antibiotics and avoiding long-term contact between antibiotics and
bacteria.
References
|
1
|
Kumarasamy KK, Toleman MA, Walsh TR,
Bagaria J, Butt F, Balakrishnan R, Chaudhary U, Doumith M, Giske
CG, Irfan S, et al: Emergence of a new antibiotic resistance
mechanism in India, Pakistan, and the UK A molecular, biological,
and epidemiological study. Lancet Infect Dis. 10:597–602. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Barbier F, Pommier C, Garrouste-Orgeas M,
Schwebel C, Ruckly S, Dumenil AS, Lemiale V, Mourvillier B, Clec'h
C, Darmon M, et al: Extended-spectrum beta-lactamase-producing
Enterobacteriaceae in critically iii patients: Impact of carriage
and infection on carbapenem consumption, duration of Icu stay, and
mortality. Intensive Care Med Exp. 3 Suppl 1:A12015. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Mushtaq S, Woodford N, Hope R, Adkin R and
Livermore DM: Activity of BAL30072 alone or combined with
β-lactamase inhibitors or with meropenem against
carbapenem-resistant Enterobacteriaceae and non-fermenters. J
Antimicrob Chemother. 68:1601–1608. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Nordmann P, Dortet L and Poirel L:
Carbapenem resistance in Enterobacteriaceae: Here is the storm!
Trends Mol Med. 18:1–272. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Satlin MJ, Chen L, Patel G, Gomez-Simmonds
A, Weston G, Kim AC, Seo SK, Rosenthal ME, Sperber SJ, Jenkins SG,
et al: Multicenter clinical and molecular epidemiological analysis
of bacteremia due to carbapenem-resistant enterobacteriaceae (CRE)
in the CRE epicenter of the United States. Antimicrob Agents
Chemother. 61:pii: e02349–16. 2017. View Article : Google Scholar
|
|
6
|
Blair JM, Webber MA, Baylay AJ, Ogbolu DO
and Piddock LJ: Molecular mechanisms of antibiotic resistance. Nat
Rev Microbiol. 13:42–51. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Liu YY, Wang Y, Walsh TR, Yi LX, Zhang R,
Spencer J, Doi Y, Tian G, Dong B, Huang X, et al: Emergence of
plasmid-mediated colistin resistance mechanism MCR-1 in animals and
human beings in China: A microbiological and molecular biological
study. Lancet Infect Dis. 16:161–168. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Khan AU, Maryam L and Zarrilli R:
Structure, genetics and worldwide spread of New Delhi
metallo-β-lactamase (NDM): A threat to public health. BMC
Microbiol. 17:1012017. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Diene SM and Rolain JM: Carbapenemase
genes and genetic platforms in Gram-negative bacilli:
Enterobacteriaceae, Pseudomonas and Acinetobacter species. Clin
Microbiol Infect. 20:831–838. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Vasoo S, Barreto JN and Tosh PK: Emerging
issues in gram-negative bacterial resistance: An update for the
practicing clinician. Mayo Clin Proc. 90:pp. 395–403. 2015;
View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Brust K, Evans A and Plemmons R:
Favourable outcome in the treatment of carbapenem-resistant
Enterobacteriaceae urinary tract infection with high-dose
tigecycline. J Antimicrob Chemother. 69:2875–2876. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Gupta N, Limbago BM, Patel JB and Kallen
AJ: Carbapenem-resistant Enterobacteriaceae: Epidemiology and
prevention. Clin Infect Dis. 53:60–67. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Surgers L, Boyd A, Boelle PY, Lalande V,
Jolivot PA, Girard PM, Arlet G, Cambier C, Homor A, Decre D and
Meynard JL: Clinical and microbiological determinants of severe and
fatal outcomes in patients infected with Enterobacteriaceae
producing extended-spectrum β-lactamase. Eur J Clin Microbiol
Infect Dis. 36:1261–1268. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Paul M, Carmeli Y, Durante-Mangoni E,
Mouton JW, Tacconelli E, Theuretzbacher U, Mussini C and Leibovici
L: Combination therapy for carbapenem-resistant Gram-negative
bacteria. J Antimicrob Chemother. 69:2305–2309. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Steinmann J, Kaase M, Gatermann S, Popp W,
Steinmann E, Damman M, Paul A, Saner F, Buer J and Rath P: Outbreak
due to a Klebsiella pneumoniae strain harbouring KPC-2 and VIM-1 in
a German university hospital, July 2010 to January 2011. Euro
Surveill. 16:pii: 199442011.
|
|
16
|
Hu F, Chen S, Xu X, Guo Y, Liu Y, Zhu D
and Zhang Y: Emergence of carbapenem-resistant clinical
Enterobacteriaceae isolates from a teaching hospital in Shanghai,
China. J Med Microbiol. 61:132–136. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Ho PL, Lo WU, Yeung MK, Lin CH, Chow KH,
Ang I, Tong AH, Bao JY, Lok S and Lo JY: Complete sequencing of
pNDM-HK encoding NDM-1 carbapenemase from a multidrug-resistant
Escherichia coli strain isolated in Hong Kong. PLoS One.
6:e179892011. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Yang Q, Wang H, Sun H, Chen H, Xu Y and
Chen M: Phenotypic and genotypic characterization of
Enterobacteriaceae with decreased susceptibility to carbapenems:
Results from large hospital-based surveillance studies in China.
Antimicrob Agents Chemother. 54:573–577. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Lee JY, Hong YK, Lee H and Ko KS: High
prevalence of non-clonal imipenem-nonsusceptible Enterobacter spp.
isolates in Korea and their association with porin down-regulation.
Diagn Microbiol Infect Dis. 87:53–59. 2017. View Article : Google Scholar : PubMed/NCBI
|