Introduction
Asthma is a heterogeneous disease that is influenced
by a variety of factors, which may be hereditary or environmental.
Asthma is often accompanied by airway remodeling (AR), which may
cause difficulties in controlling the symptoms of asthma and
seriously damage human health. The epithelial to mesenchymal
transition (EMT) has been indicated to be associated with the
process of AR (1–3). Following damage by allergens and other
factors, cells may lose their epithelial characteristics, and may
undergo morphological changes from regular oval-shaped epithelial
cells to irregularly spindle-shaped stromal cells, lose epithelial
cell protein markers (such as E-cadherin) and obtain interstitial
cell protein markers, including vimentin and α-smooth muscle actin
(α-SMA) (4). Transformed epithelial
cells participate in the process of AR through cellular activities
characteristic of their nascent myofibroblast phenotype (5). In vitro cell experiments have
confirmed that EMT-associated changes in airway epithelial cells
are mainly regulated by transforming growth factor (TGF)-β1
(6); however, this regulatory
process is not only affected by the inflammatory response but also
by the expression of a variety of genes in vivo.
Disintegrin and metalloproteinase domain-containing
protein 33 (ADAM33), a susceptibility gene for asthma first
identified in 2002, is expressed in fibroblasts, smooth muscle
cells and other stromal cells in lung tissue, but is not expressed
in epithelial cells or T lymphocytes (7). It has been indicated that the
expression level of ADAM33 is inversely proportional to the percent
forced expiratory volume in 1 sec, a measure that is proportional
to the severity of asthma, suggesting that ADAM33 may be involved
in the AR process of asthma; however, the underlying mechanism has
remained elusive (8). Therefore, the
present study aimed to investigate the effects of TGF-β1 on ADAM33
expression in airway epithelial cells, to evaluate the association
between ADAM33 expression and TGF-β1-induced EMT, and to further
explore the mechanisms of ADAM33 in asthma-induced AR.
Materials and methods
Materials
The normal human bronchial epithelial cell line HBE
was purchased from the American Type Culture Collection (Manassas,
VA, USA). Dulbecco's modified Eagle's medium (DMEM), fetal calf
serum (FCS) and penicillin-streptomycin were from Gibco (Thermo
Fisher Scientific, Inc., Waltham, MA, USA). TGF-β1 was purchased
from Peprotech Co. (Rocky Hill, NJ, USA). The primary antibodies
against E-cadherin (cat. no. sc-71007), vimentin (cat. no.
sc-73258), ADAM33 (cat. no. sc-514055) and β-actin (cat. no.
sc-47778) and goat anti-mouse IgG-horseradish peroxidase (cat. no.
sc-2005), were purchased from Santa Cruz Biotechnology, Inc.
(Dallas, TX, USA). TRIzol reagent and the reverse-transcription
polymerase chain reaction (RT-PCR) detection kit (cat. no.
11746100) were from Invitrogen (Thermo Fisher Scientific,
Inc.).
Cell culture and treatment
HBE cells were cultured in DMEM supplemented with
10% FCS in a humidified atmosphere of 95% air and 5% CO2
in an incubator at 37°C. Cells were passaged when they reached 90%
confluence. When third-generation cells reached 80% confluence, the
cells were synchronized with high-glucose DMEM containing 1% FCS
for 24 h. Subsequently, the cells were treated with various
concentrations of TGF-β1 (0, 10, 20 or 30 ng/ml). After treatment
for 72 h, the cells were used in the subsequent experiments. Cells
were also transfected with small interfering (si)RNA targeting
ADAM33 (siADAM33) in the experimental group, or nonsense siRNA
(GenScript Biotech Corporation, Nanjing, China) in the negative
control (NC) group, using Lipofectamine™ 2000 (Thermo
Fisher Scientific, Inc.) following the manufacturer's protocol, and
treated with various concentrations of TGF-β1 (0, 10, 20 and 30
ng/ml) for use in the western blot analysis. The sequences of the
siRNAs used were as follows: siADAM33 forward,
5′-GGACUCUACCGUUCACCUATT-3′ and reverse,
5′-UAGGUGAACGGUAGAGUCCTT-3′; and nonsense siRNA forward,
5′-UUCUCCGAACGUGUCACGUTT-3′ and reverse,
5′-ACGUGACACGUUCGGAGAATT-3′.
Cell morphology observation
HBE cells were incubated with TGF-β1 for 72 h under
standard cell culture conditions. Subsequently, the morphology of
the cells was observed under a light microscope (magnification,
×200; Olympus CKX41; Olympus, Tokyo, Japan).
Immunohistochemistry
HBE cells on coverslips were fixed with 4%
paraformaldehyde for 15 min and permeabilized with 0.5% Triton
X-100 for 20 min, prior to incubation with 3%
H2O2 for 15 min and blocking in bovine serum
albumin (Biosharp, Hefei, China). The cells were then incubated
with the primary antibody against E-cadherin (sc-71007; 1:500
dilution; Santa Cruz Biotechnology, Inc.) at room temperature for 1
h, followed by detection with biotin-labeled secondary antibodies
(cat. no. sc-2040; 1:2,000; Santa Cruz Biotechnology, Inc.) at room
temperature for 30 min and a streptavidin biotin complex kit (cat.
no. GTX30965; GeneTex, Inc., Irvine, CA, USA) at 37°C for 20 min.
The coverslips were developed with diaminobenzidine for 15 min and
counterstained with 1% hematoxylin at room temperature for 1 min.
Finally, coverslips were mounted and observed under a
microscope.
Western blot analysis
After treatment for 72 h, total protein from the
cells was collected with radioimmunoprecipitation assay buffer. A
bicinchoninic acid assay was performed to detect the protein
concentration in the cell lysates. Protein samples (30 µg/lane)
were separated by 10% SDS-PAGE and then transferred onto a
polyvinylidene difluoride membrane (Merck KGaA, Darmstadt,
Germany), which was blocked with 5% bovine serum albumin for 2 h at
room temperature. Membranes were incubated with the primary
antibodies (1:1,000 dilution) overnight at 4°C, washed three times
with Tris-buffered saline containing Tween 20 and then incubated
with horseradish peroxidase-conjugated secondary antibodies
(1:2,000 dilution) for 2 h at room temperature. The membranes were
developed with an enhanced chemiluminescence kit (cat. no. P0018;
Beyotime Institute of Biotechnology, Haimen, China) according to
the manufacturer's protocol. The Tanon-5200 (Tanon Science and
Technology Co., Ltd., Shanghai, China) automatic chemiluminescence
image analysis system was used for gray scale scanning and
quantitative analysis. β-actin was used as the internal
control.
RT-qPCR analysis
Total RNA was isolated with TRIzol regent according
to the manufacturer's instructions. Complementary (c)DNA was
generated using the RT-PCR detection kit following the
manufacturer's protocol, and real-time PCR was then performed to
amplify the synthesized cDNA using the SYBR Green qPCR Mix
(Beyotime Institute of Biotechnology) following the manufacturer's
protocol. GAPDH was used as the internal control. The amplification
conditions were as follows: Initial activation at 95°C for 5 min,
followed by 40 amplification cycles of denaturation at 95°C for 30
sec, annealing at 56°C for 45 sec and extension at 72°C for 30 sec.
The primer sequences used in the real-time PCR analysis are listed
in Table I. The expression levels of
the target genes were analyzed using the 2−ΔΔCq method
(9).
 | Table I.Primer sequences for polymerase chain
reaction. |
Table I.
Primer sequences for polymerase chain
reaction.
| Gene/direction | Sequences
(5′-3′) |
|---|
| ADAM33 |
|
|
Forward |
TGCCTAGAGGCCGAGGAG |
|
Reverse |
CCAGAGCAGCAGCAGTAGTAG |
| E-cadherin |
|
|
Forward |
GCCCCGCCTTATGATTCTCTGC |
|
Reverse |
CTCGCCGCCTCCGTACATGTC |
| Vimentin |
|
|
Forward |
CTCTTCCAAACTTTTCCTCCC |
|
Reverse |
AGTTTCGTTGATAACCTGTCC |
| GAPDH |
|
|
Forward |
5′CCACATCGCTCAGACACCAT |
|
Reverse |
5′ACGGTGCCATGGAATTTGCC |
Wound healing assay
HBE cells were seeded in 6-well plates at a density
of 5×104/well and cultured for 24 h, after which a
straight scratch wound was made across the cell monolayer using a
200-µl pipette tip. Wells were washed with PBS and serum-free DMEM
was applied to the cells. Images were captured after 48 h.
Cell invasion assay
The invasion potential of HBE cells was assessed
using a Transwell assay. The Transwell inserts (Corning
Incorporated, Corning, NY, USA) were pre-coated with Matrigel.
Cells (5×104) in serum-free medium were seeded into the
upper chamber, while medium containing 10% FBS was added to the
lower chamber. After incubation for 48 h at 37°C, cells that had
invaded to the lower chamber were stained with crystal violet at
room temperature for 30 min and observed under a microscope.
Statistical analysis
SPSS 16.0 statistical software (SPSS, Inc., Chicago,
IL, USA) was used for all statistical analyses. Values are
expressed as the mean ± standard deviation and a Shapiro-Wilk
normal distribution test and a Levene variance homogeneity test
were performed. Analysis of variance was used for comparisons
between multiple groups. A Student Newman Keuls t-test was used to
compare the differences between two groups. Pearson's linear
correlation analysis was used to analyze different indexes.
P<0.05 was considered to indicate a statistically significant
difference.
Results
Effect of TGF-β1 on HBE cell
morphology
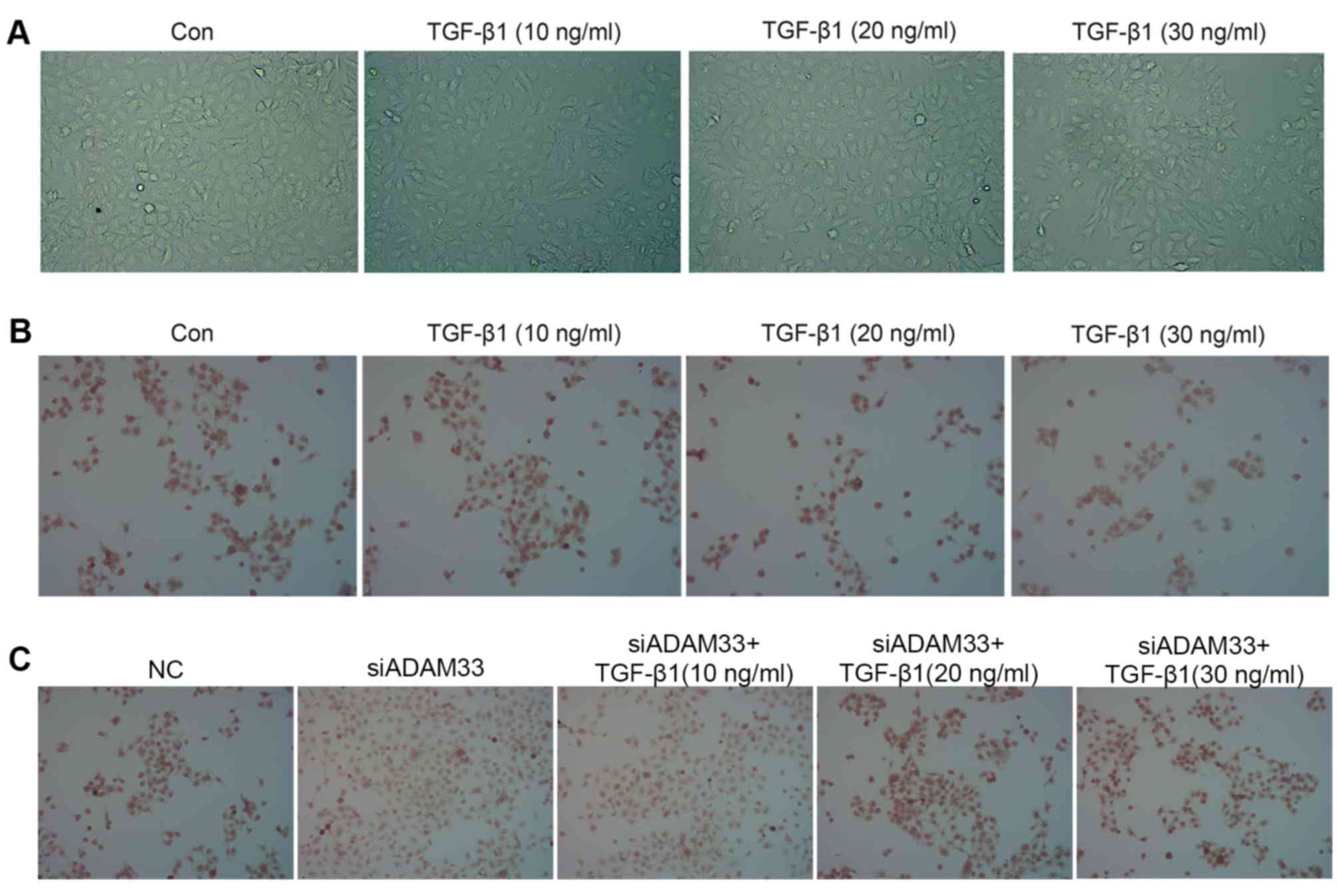
Following stimulation with different concentrations
of TGF-β1, HBE cell morphology exhibited changes characteristic for
EMT: Cell clusters were scattered with increased numbers of free
cells, and cells exhibited a spindle-shaped, fibroblast-like
morphology. With increasing TGF-β1 concentration, the cell
morphology changes became more obvious, such that the percentage of
cells exhibiting the fibroblast-like form was highest when the
TGF-β1 concentration was 30 ng/ml. By contrast, the cells in the
control group still displayed an oval-shaped, epithelial morphology
(Fig. 1A).
Effect of TGF-β1 on the protein
expression of ADAM33, E-cadherin and vimentin in HBE cells
Immunohistochemical analysis indicated that TGF-β1
inhibited E-cadherin protein expression in a dose-dependent manner
(Fig. 1B) and siADAM33 relieved this
inhibition (Fig. 1C). Following
treatment with various concentrations of TGF-β1 (10, 20 or 30
ng/ml), the protein expression levels of ADAM33 were significantly
increased in the HBE cells, which suggested that TGF-β1 promoted
the expression of ADAM33 in HBE cells (Fig. 2A and B). In addition, TGF-β1
inhibited E-cadherin protein expression in a dose-dependent manner
(Fig. 2A and C), whereas the protein
expression of vimentin was significantly increased in HBE cells
after stimulation with various concentrations of TGF-β1 (Fig. 2A and D).
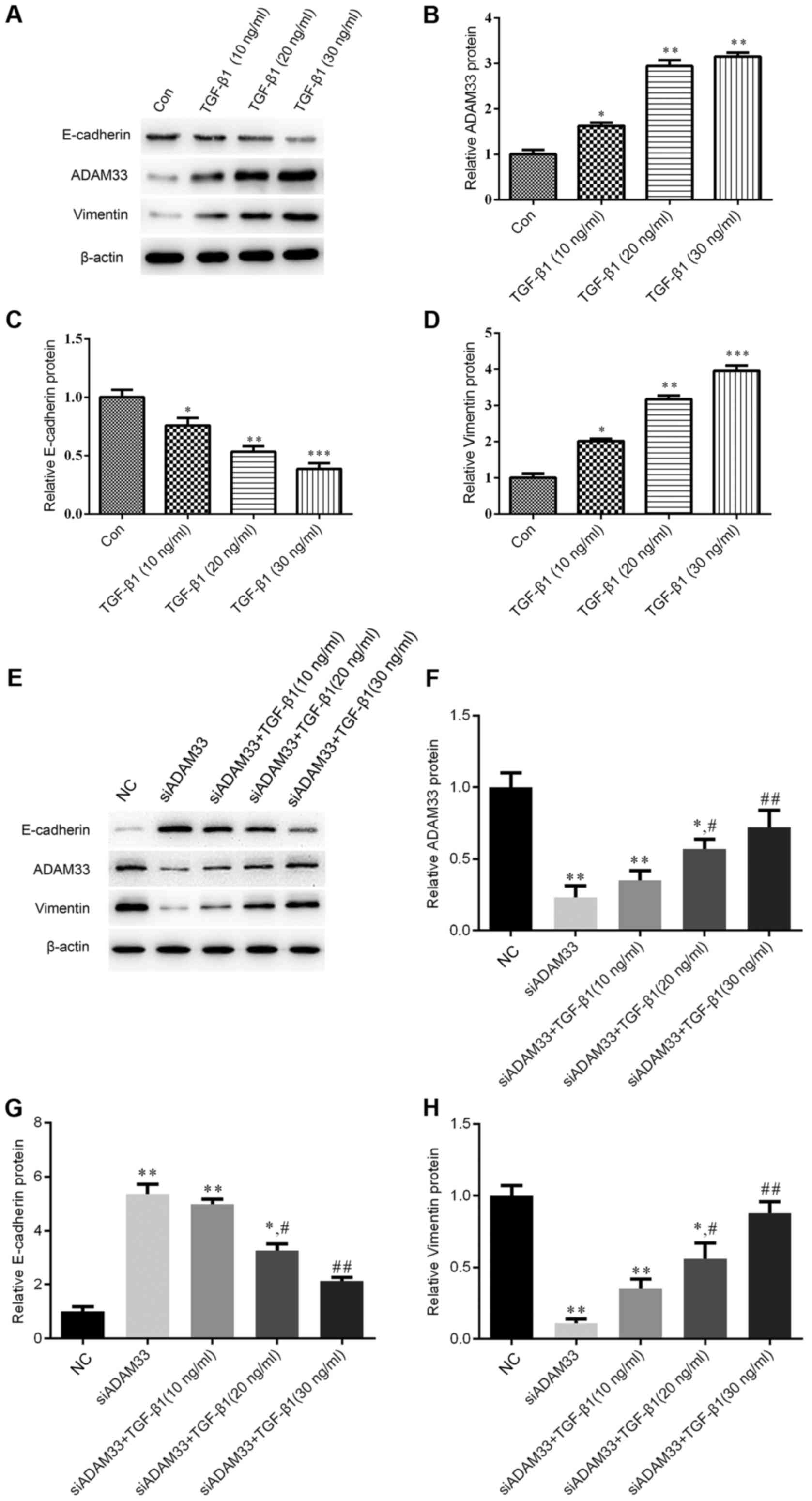 | Figure 2.Effect of TGF-β1 on the protein
expression of ADAM33, E-cadherin and vimentin in the HBE human
bronchial epithelial cell line. (A-D) Cells were treated with
various concentrations of TGF-β1 (10, 20 or 30 ng/ml). (A)
Representative western blot image. Protein expression levels of (B)
ADAM33, (C) E-cadherin and (D) vimentin. (E-G) Cells were
transfected with siADAM33 and treated with various concentrations
of TGF-β1 (10, 20 or 30 ng/ml). (E) Representative western blot
image. Protein expression levels of (F) ADAM33, (G) E-cadherin and
(H) vimentin. *P<0.05, **P<0.01 vs. the control group;
#P<0.05, ##P<0.01 vs. the siADAM33
group. Con, control; TGF, transforming growth factor; NC, negative
control; siADAM33, small interfering RNA targeting disintegrin and
metalloproteinase domain-containing protein 33. |
After transfection of siADAM33 into HBE cells, the
protein expression levels of ADAM33 were significantly decreased
compared with the NC group (Fig. 2E and
F). As presented in Fig. 2E, G and
H, siADAM33 promoted the expression of E-cadherin and repressed
the expression of vimentin compared with the siADAM33 group. These
results indicated that the stimulation of the EMT process by TGF-β1
may be mediated by ADAM33.
Effect of TGF-β1 on the mRNA
expression of ADAM33, E-cadherin and vimentin in HBE cells
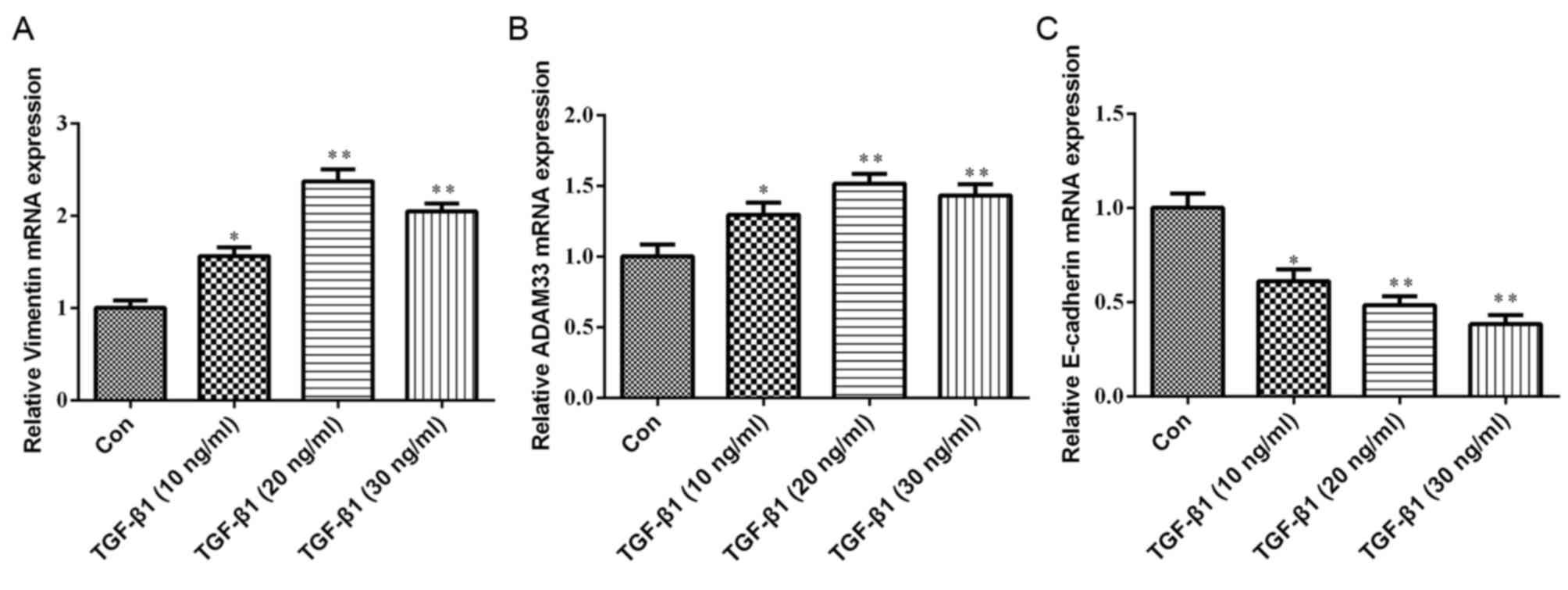
As presented in Fig.
3, compared with the control group, the mRNA expression levels
of ADAM33 and vimentin were significantly increased in HBE cells
treated with various concentrations of TGF-β1 (10, 20 or 30 ng/ml).
Furthermore, TGF-β1 significantly inhibited E-cadherin mRNA
expression in a dose-dependent manner.
Correlation analysis of ADAM33
expression with E-cadherin and vimentin expression
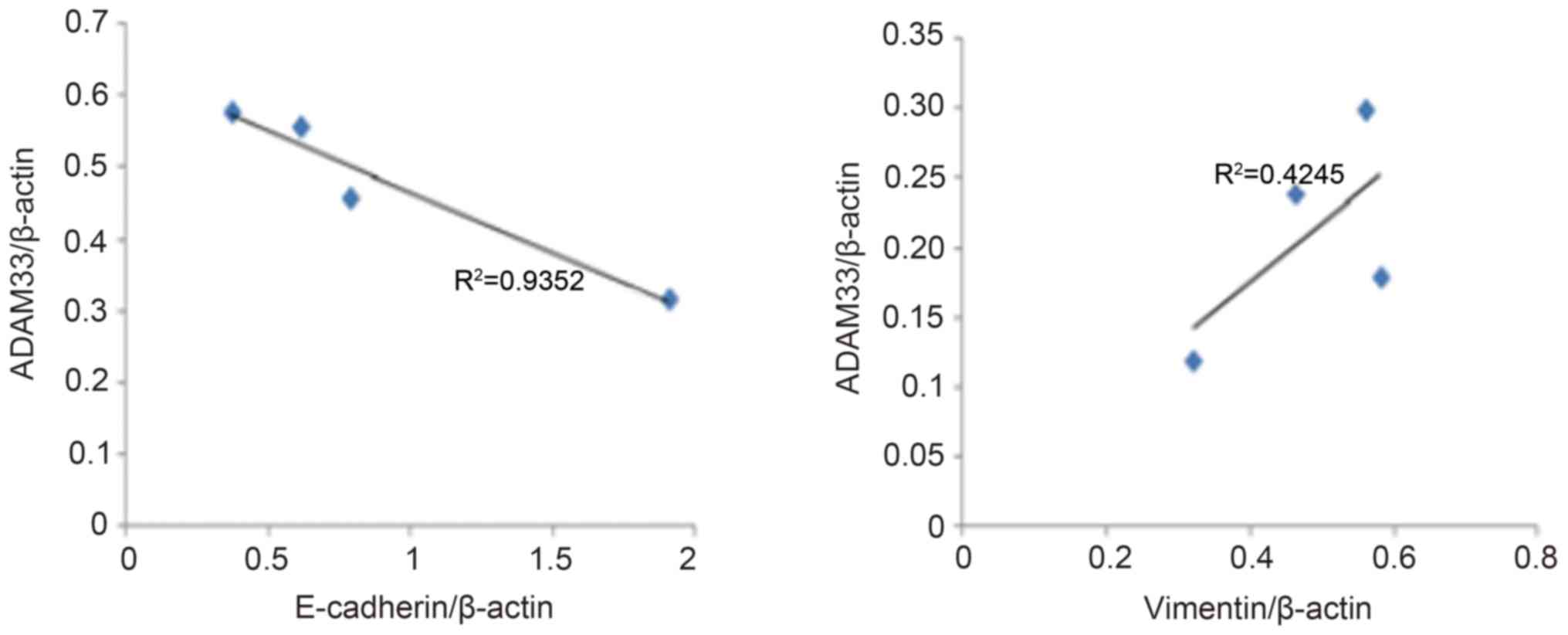
As presented in Fig.
4, the results of the correlation analysis indicated a
significant negative correlation between ADAM33 protein expression
and E-cadherin expression (R2=0.935, P<0.05), whereas
there was a positive correlation between ADAM33 and vimentin
expression (R2=0.425, P<0.05).
Effect of TGF-β1 on the migration and
invasion of HBE cells
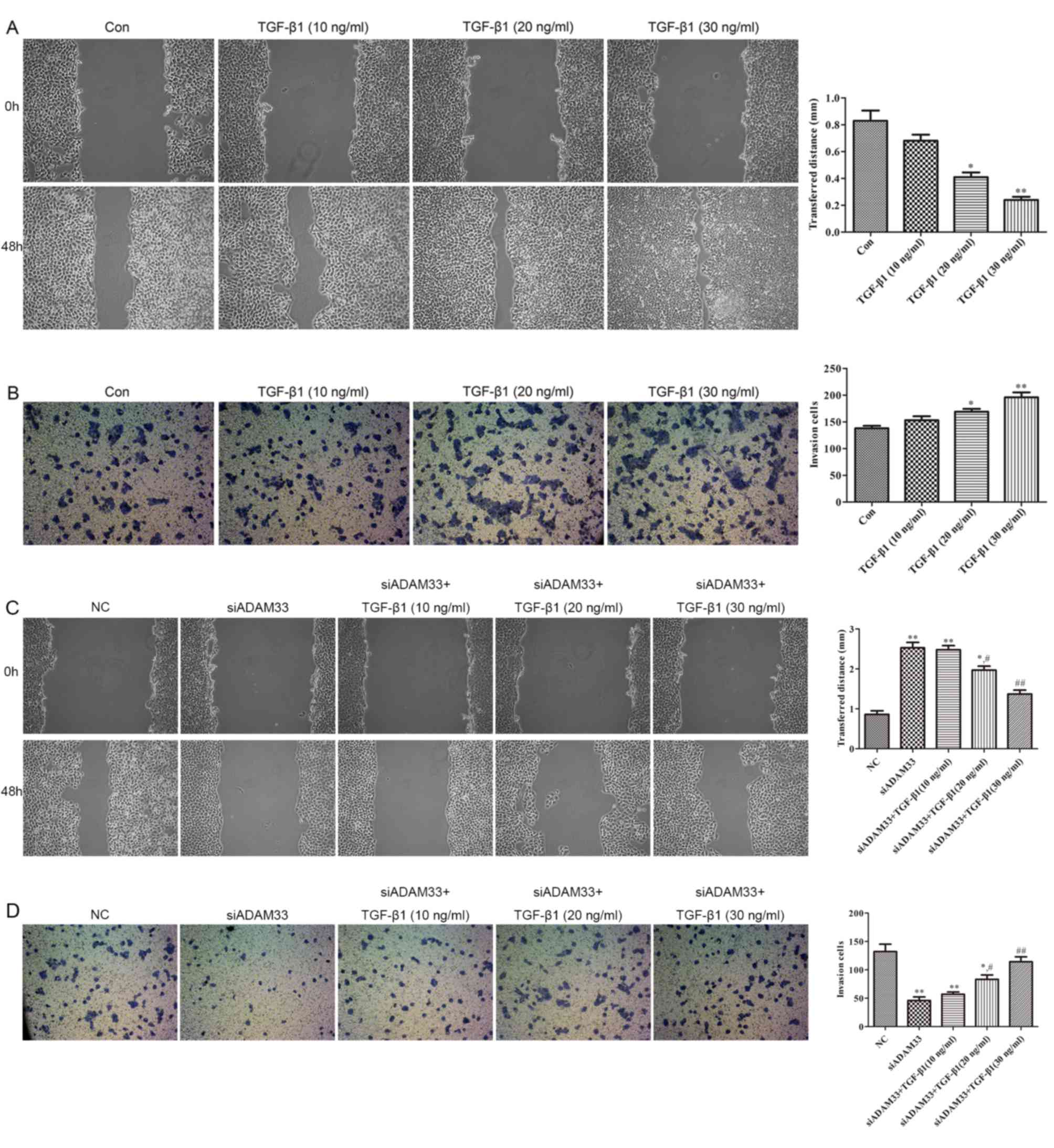
As illustrated in Fig.
5A, compared with the control group, wound healing was promoted
by TGF-β1 in a dose-dependent manner (10, 20 and 30 ng/ml).
Furthermore, the results of the cell invasion assay indicated that
the invasive ability of HBE cells was increased by TGF-β1 in a
dose-dependent manner (10, 20 and 30 ng/ml; Fig. 5B). Following transfection of siADAM33
into HBE cells, wound healing and invasive ability were observed to
be inhibited by siADAM33 compared with the NC group (Fig. 5C and D). These results indicate that
the migration and invasion of HBE cells induced by TGF-β1 may be
mediated by ADAM33.
Discussion
The human ADAM33 gene, which is located on the short
arm of chromosome 20 (20p13), is one of the most significant
susceptibility genes in asthma. The full-length ADAM33 gene is
~2,439 bp, comprising 22 exons and 21 introns, and is divided into
eight domains: A signal peptide domain, a prodomain, a
metalloproteinase domain, a disintegrin domain, a
cysteine-containing domain, an epidermal growth factor domain, a
transmembrane domain and a cytoplasmic domain. The
metalloproteinase domain comprises a zinc finger structure sequence
that encodes a protein with enzymatic activity responsible for
cleaving peptides bound to proteins, and thereby exhibits
proteolytic enzyme action. The presence of the growth factor domain
suggests that ADAM33 may be associated with cell proliferation and
differentiation (7). Due to ADAM33
expression and its molecular structure, as well as functional
characteristics in mesenchymal cells and epithelial cells in human
lung tissue, it is thought that ADAM33 may be involved in the
process of AR in chronic inflammatory airway disease, including
asthma and chronic obstructive pulmonary disease (10,11).
A previous study revealed that the expression levels
of ADAM33 were upregulated in the lung tissues of asthma patients
(12). In another study, in
situ hybridization and immunohistochemistry were used to
determine the mRNA and protein expression levels of ADAM33 in human
bronchial biopsy specimens, and the results indicated that ADAM33
was significantly overexpressed in tissues from asthmatic patients
compared with those from normal subjects (13). In addition, the expression levels of
ADAM33 in human fibroblasts cultured in vitro were reported
to be upregulated by interleukin-4 and −13 stimulation (14) and the same result was obtained in
smooth muscle cells cultured in vitro, but not in cultured
epithelial cells, endothelial cells or neutrophils (15). These studies suggested that the
immune imbalance of T helper type I vs. type II cells in asthma may
contribute to the expression of ADAM33 in interstitial cells. Foley
et al (16) reported that the
expression levels of ADAM33 in patients with severe and moderate
asthma were significantly higher than those in patients with mild
or controlled asthma, and that it was also significantly increased
in epithelial, submucosal and smooth muscle cells. In summary,
these studies indicated that ADAM33 expression was upregulated in
airway epithelial and stromal cells, and that there were
significant differences in ADAM33 expression according to the
severity of asthma, suggesting that ADAM33 may be involved in the
AR process.
The nature of the participation of ADAM33 in AR, and
its associated mechanisms, have remained elusive. A previous study
using a mouse model of asthma demonstrated that the expression
levels of ADAM33 in airway smooth muscle cells were positively
correlated with cytoskeletal protein proliferation, suggesting that
ADAM33 may promote asthma-associated AR by promoting smooth muscle
cell proliferation (17). In
addition, in tissue culture, it was demonstrated that ADAM33 was
involved in the AR process though interacting with α-actin and
other interstitial cell proteins (18). Furthermore, another study indicated
that ADAM33 was activated and overexpressed in response to TGF-β2
stimulation in primary cultured airway fibroblasts (PBF),
suggesting that ADAM33 participates in asthma AR by promoting PBF
activation (19).
Epithelial cells are the primary barrier against
harmful environmental stimuli, including pollutants, harmful gases,
allergens and pathogenic microorganisms. The process of
injury-repair-reinjury of epithelial cells is the primary step in
AR, and EMT is involved and has important roles in this process. It
has been confirmed that epithelial cells participate in
AR-associated processes, such as subepithelial fibrosis, by
undergoing EMT to transform into mesenchymal cells and subsequently
serving the role of fibroblasts. TGF-β1 is the only cytokine that
has been demonstrated to independently induce EMT in airway
epithelial cells (2,20–22). As
a member of the growth factor family, TGF-β1 has a binding site
specific for ADAM33. However, whether TGF-β1 has an effect on
ADAM33 expression in airway epithelial cells, as well as the
association between ADAM33 expression and TGF-β1-induced EMT, have
remained elusive. In the present study, HBE cells were stimulated
with TGF-β1 to observe its effect on ADAM33 expression, cell
migration and invasion in airway epithelial cells, and the
association between ADAM33 expression and TGF-β1-induced
EMT-associated proteins, including E-cadherin, vimentin and α-SMA,
was investigated. The results suggested that stimulation of HBE
cells with various concentrations of TGF-β1 enhanced ADAM33
expression, indicating that TGF-β1 promoted the expression of
ADAM33 in airway epithelial cells in vitro. The results of a
correlation analysis revealed that TGF-β1 promoted the expression
of ADAM33, which was negatively correlated with E-cadherin and
positively correlated with vimentin, suggesting that the
TGF-β1-induced upregulation of ADAM33 is closely associated with
the EMT process induced by TGF-β1, and that it is particularly
strongly associated with the decrease of E-cadherin. In a
loss-of-function experiment, ADAM33 was knocked down in order to
further clarify the association between ADAM33 expression and
EMT-associated proteins in airway epithelial cells stimulated by
TGF-β1. The results of a western blot analysis indicated that
siADAM33 promoted the expression of E-cadherin and repressed the
expression of vimentin, and that TGF-β1 suppressed these changes.
These results confirmed that the stimulation of the EMT process by
TGF-β1 may be mediated via ADAM33.
Acknowledgements
The present study was supported by Jinling Hospital
(grant no. 2015062).
References
|
1
|
Heijink IH, Postma DS, Noordhoek JA,
Broekema M and Kapus A: House dust mite-promoted
epithelial-to-mesenchymal transition in human bronchial epithelium.
Am J Respir Cell Mol Biol. 42:69–79. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Hackett TL, Warner SM, Stefanowicz D,
Shaheen F, Pechkovsky DV, Murray LA, Argentieri R, Kicic A, Stick
SM, Bai TR and Knight DA: Induction of epithelial-mesenchymal
transition in primary airway epithelial cells from patients with
asthma by transforming growth factor-beta1. Am J Respir Crit Care
Med. 180:122–133. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Hackett TL: Epithelial-mesenchymal
transition in the pathophysiology of airway remodelling in asthma.
Curr Opin Allergy Clin Immunol. 12:53–59. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Johnson JR, Roos A, Berg T, Nord M and
Fuxe J: Chronic respiratory aeroallergen exposure in mice induces
epithelial-mesenchymal transition in the large airway. PLoS One.
6:e161752011. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Wang JH, Sun LH, Huang SK and Chen AH:
Effect of co-culturing human primary basic fibroblasts with
respiratory syncytial virus-infected 16-HBE cells. Genet Mol Res.
15:150173392016.
|
|
6
|
Ko H, So Y, Jeon H, Jeong MH, Choi HK, Ryu
SH, Lee SW, Yoon HG and Choi KC: TGF-β1-induced
epithelial-mesenchymal transition and acetylation of Smad2 and
Smad3 are negatively regulated by EGCG in human A549 lung cancer
cells. Cancer Lett. 335:205–213. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Van Eerdewegh P, Little RD, Dupuis J, Del
Mastro RG, Falls K, Simon J, Torrey D, Pandit S, McKenny J,
Braunschweiger K, et al: Association of the ADAM33 gene with asthma
and bronchial hyperresponsiveness. Nature. 418:426–430. 2002.
View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Lee JY, Park SW, Chang HK, Kim HY, Rhim T,
Lee JH, Jang AS, Koh ES and Park CS: A disintegrin and
metalloproteinase 33 protein in patients with asthma: Relevance to
airflow limitation. Am J Respir Crit Care Med. 173:729–735. 2006.
View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Holgate ST: ADAM metallopeptidase domain
33 (ADAM33): Identification and role in airways disease. Drug News
Perspect. 23:381–387. 2010.PubMed/NCBI
|
|
11
|
Xiao J, Han J, Wang X, Hua D, Su D, Bao Y
and Lv F: Association of ADAM33 gene with susceptibility to COPD in
Tibetan population of China. Mol Biol Rep. 38:4941–4945. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Sunadome H, Matsumoto H, Petrova G,
Kanemitsu Y, Tohda Y, Horiguchi T, Kita H, Kuwabara K, Tomii K,
Otsuka K, et al: IL4Rα and ADAM33 as genetic markers in asthma
exacerbations and type-2 inflammatory endotype. Clin Exp Allergy.
47:998–1006. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Ito I, Laporte JD, Fiset PO, Asai K,
Yamauchi Y, Martin JG and Hamid Q: Downregulation of a disintegrin
and metalloproteinase 33 by IFN-gamma in human airway smooth muscle
cells. J Allergy Clin Immunol. 119:89–97. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Jie Z, Jin M, Cai Y, Bai C, Shen Y, Yuan
Z, Hu Y and Holgate S: The effects of Th2 cytokines on the
expression of ADAM33 in allergen-induced chronic airway
inflammation. Respir Physiol Neurobiol. 168:289–294. 2009.
View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Tripathi P, Awasthi S, Husain N, Prasad R
and Mishra V: Increased expression of ADAM33 protein in asthmatic
patients as compared to non-asthmatic controls. Indian J Med Res.
137:507–514. 2013.PubMed/NCBI
|
|
16
|
Foley SC, Mogas AK, Olivenstein R, Fiset
PO, Chakir J, Bourbeau J, Ernst P, Lemière C, Martin JG and Hamid
Q: Increased expression of ADAM33 and ADAM8 with disease
progression in asthma. J Allergy Clin Immunol. 119:863–871. 2007.
View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Lin F, Song A, Wu J, Jiang X, Long J, Chen
J, Duan Y, Shi Y and Deng L: ADAM33 protein expression and the
mechanics of airway smooth muscle cells and highly correlated in
ovalbumin-sensitized rats. Mol Med Rep. 8:1209–1215. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Haitchi HM, Powell RM, Shaw TJ, Howarth
PH, Wilson SJ, Wilson DI, Holgate ST and Davies DE: ADAM33
expression in asthmatic airways and humanembryonic lungs. Am J
Respir Crit Care Med. 171:958–965. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Yang Y, Wicks J, Haitchi HM, Powell RM,
Manuyakorn W, Howarth PH, Holgate ST and Davies DE: Regulation of a
disintegrin and metalloprotease-33 expression by transforming
growth factor-β. Am J Respir Cell Mol Biol. 46:633–640. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Zhang M, Zhang Z, Pan HY, Wang DX, Deng ZT
and Ye XL: TGF-beta1 induces human bronchial epithelial
cell-to-mesenchymal transition in vitro. Lung. 187:187–194. 2009.
View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Kamitani S, Yamauchi Y, Kawasaki S, Takami
K, Takizawa H, Nagase T and Kohyama T: Simultaneous stimulation
with TGF-β1 and TNF-α induces epithelial mesenchymal transition in
bronchial epithelial cells. Int Arch Allergy Immunol. 155:119–128.
2011. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Yang ZC, Yi MJ, Ran N, Wang C, Fu P, Feng
XY, Xu L and Qu ZH: Transforming growth factor-β1 induces bronchial
epithelial cells to mesenchymal transition by activating the Snail
pathway and promotes airway remodeling in asthma. Mol Med Rep.
8:1663–1668. 2013. View Article : Google Scholar : PubMed/NCBI
|