Introduction
Heat stroke (HS) is a hyperthermia-associated
disease that may induce multi-organ dysfunction and death (1). Abnormalities in endothelial cell
(EC) function have been implicated in the pathophysiology of HS
(2). EC dysfunction initiates a
number of events that trigger EC activation and predispose the
vessel wall to increased endothelial permeability, leukocyte
adherence, endothelial apoptosis, pro-oxidation and thrombosis
(3–9). MicroRNAs (miRNAs) participate in the
control of EC-mediated homeostasis (10,11).
miRNAs are small non-coding RNAs of 18–22
nucleotides in length that play an important role in the regulation
of gene expression at the post-transcriptional level. Specific
classes of miRNAs have been demonstrated to affect the function of
ECs. For example, miR-126, miR-128, the miR-17-92 cluster, and the
miR-23-24-27 cluster, have been reported to control blood vessel
development, growth and differentiation; therefore, these miRNAs
and clusters contribute to the regulation of angiogenesis (12–16). Previous studies demonstrated that
certain inducible miRNAs also contribute to the regulation of
inflammation. Inducible miRNAs, including miR-31, miR-17-3p,
miR-126, miR-181b, miR-10a, miR-155, miR-221/222, miR-92,
miR-125a/b, miR-365 and miR-663, may participate in the regulation
of inflammatory cytokine-induced leukocyte adhesion to ECs and
activation of leukocytes (17–27). Recent studies also indicated that
certain miRNAs, including miR-155, miR-320, miR-222, miR-125b,
miR-410 and miR-218, play an important role in controlling the
barrier function of ECs (28–30). These miRNAs reportedly regulate
adherens junction disassembly responses, cell migration and cell
morphology, which contribute to changes in vascular permeability
and integrity. Another set of endothelial miRNAs participate in the
control of vascular tone. Several reports have indeed provided
evidence that miR-155, miR-221, miR-222, miR-125a/b and miR-21 are
associated with the regulation of blood pressure and blood flow by
releasing vasodilators or vasoconstrictors (24,31–33). Upon exposure to certain stimuli,
ECs may secrete miRNAs that mediate cell-to-cell communication. For
example, ECs have been shown to produce microvesicles under shear
stress. These microvesicles are enriched in miRNAs, particularly
miR-143/145 or miR-150, which exert an atheroprotective effect on
smooth muscle cells (34–36). This evidence suggests that miRNAs
are important molecules that modulate the biological processes of
ECs.
Although the involvement of individual miRNAs in
endothelial regulation has been characterized in several diseases,
changes in the general miRNA expression profiles under heat stress
have not been reported. The aim of the present study was to
systematically analyze the profiles of miRNA expression in human
umbilical vein ECs (HUVECs) exposed to acute heat stress using
microarray and bioinformatics analyses, in order to evaluate the
potential role of miRNAs in post-transcriptional regulation.
Materials and methods
HUVEC culture and treatment with
heat
HUVECs (American Type Culture Collection, Manassas,
VA, USA) were cultured in complete extracellular matrix
supplemented with 5% fetal bovine serum (Invitrogen, Thermo Fisher
Scientific, Inc., Waltham, MA, USA), 1% EC growth supplement
(PromoCell, Heidelberg, Germany), 100 U/ml penicillin and 100
µg/ml streptomycin (Invitrogen, Thermo Fisher Scientific,
Inc.), in a humidified atmosphere with 5% CO2 at 37°C.
All the experiments were performed using 4–6 passages of
HUVECs.
In the following experiments, the cells
were randomly divided into two groups
The control cells were always placed in an incubator
at 37°C. The cells in the heat treatment group were cultured in the
incubator at 43°C for 1 h, and then at 37°C for a further 24 h.
RNA extraction
Total RNA from each sample was individually isolated
using TRIzol (Thermo Fisher Scientific, Inc., Carlsbad, CA, USA)
and miRNeasy mini kit (Qiagen, Inc., Valencia, CA, USA) according
to the manufacturers' instructions. This procedure efficiently
recovered all RNA species, including miRNAs. RNA quantity and
quality were measured using a NanoDrop spectrophotometer (ND-1000;
NanoDrop Technologies, Wilmington, DE, USA) and RNA integrity was
detected by gel electrophoresis.
RNA labeling and array hybridization
The isolated RNA was labeled using the miRCURY™
Hy3™/Hy5™ Power Labeling kit (Exiqon A/S, Vedbaek, Denmark)
according to the manufacturer's instructions. Each RNA sample (1
µg) was 3′-end-labeled with Hy3™ fluorescent label, using T4
RNA ligase. Following the array manual, the Hy3™-labeled samples
were then hybridized to the miRCURY™ LNA array (v.18.0) (Exiqon
A/S). The total mixture with hybridization buffer was first
denatured for 2 min at 95°C, incubated on ice for 2 min, and then
hybridized to the microarray for 16–20 h at 56°C in a 12-Bay
Hybridization system (Nimblegen Systems, Inc., Madison, WI, USA).
After the hybridization procedure was terminated, the slides were
washed several times using the wash buffer kit (Exiqon A/S), and
finally dried by centrifugation for 5 min at 340 × g. The slides
were then scanned with the Axon GenePix 4000B microarray scanner
(Axon Instruments, Foster City, CA, USA), which contains 3,100
capture probes annotated in miRBase 18.0, covering all human, mouse
and rat miRNAs, as well as all viral miRNAs related to these
species. In addition, the microarray scanner contains capture
probes for 25 additional new miRPlus™ human miRNAs that are
proprietary and not found in miRBase.
Array data analysis
Scanned images were imported into GenePix Pro 6.0
software (Axon Instruments) for grid alignment and data extraction.
Replicated miRNAs were averaged, and miRNAs with intensities ≥30
were selected for calculation of the normalization factor.
Expressed data were normalized using the median normalization.
Following normalization, significantly differentially expressed
miRNAs were identified through volcano plot change filtering.
Finally, unsupervised hierarchical clustering was performed based
on the normalized data of miRNAs in all samples using Multi
Experiment Viewer software, version 4.6 (The Institute for Genomic
Research, Rockville, MD, USA). A miRNA was defined as
differentially expressed between the compared groups if the P-value
was <0.05 and the fold change was >2.
Reverse transcription-quantitative
polymerase chain reaction (RT-qPCR)
To validate the authenticity of miRNA expression
screened by microarray assay, six differentially expressed miRNAs
were analyzed by RT-qPCR. For this analysis, total RNA (10 ng) was
reversely transcribed into cDNA using the TaqMan®
MicroRNA Reverse Transcription kit (Applied Biosystems, Foster
City, CA, USA) according to the manufacturer's instructions. qPCR
was performed using SYBR-Green (Thermo Fisher Scientific, Inc.)
according to the manufacturer's instructions in the ABI 7300
real-time qPCR system (Bio-Rad Laboratories, Inc., Hercules, CA,
USA). The quantification of each miRNA was performed in relation to
the U6 housekeeping miRNAs that was used as a standard. The
relative abundance of each miRNA was calculated by the comparative
Cq method (2−ΔΔCq), and the results were assessed by
using a t-test.
Bioinformatics analysis
Potential target genes of differentially expressed
miRNAs were predicted using the three most popular databases,
namely, TargetScan, miRanda and miRDB. Only the overlapping genes
that were in all three databases were considered as the target
genes of the differentially expressed miRNAs. The predicted genes
subsequently underwent Gene Ontology (GO) analysis for functional
annotation analysis and Kyoto Encyclopedia of Genes and Genomes
(KEGG) database analysis to identify the involved enriched
pathways. Fisher's exact and Chi-square tests were used to
determine the significance of the GO terms and pathways, and the
false discovery rate (FDR) was calculated to correct the P-value
(37). Only the GO terms and
pathways with an adjusted P-value of <0.05 and an FDR of
<0.05 were selected.
miRNA-gene-network analyses
The target genes identified in the functional and
pathway enrichment analyses were investigated. The 494 overlapping
genes that were in both databases were selected. In addition, to
improve the understanding of the associations between the key
miRNAs and hub genes, networks for miRNA target genes were
constructed according to the regulatory associations between miRNAs
and target genes. The associations of the genes and miRNAs were
evaluated by graph theory. The degree was used to measure the
regulated effect of miRNAs on genes.
miRNA-GO-network analysis
A miRNA-GO-network was constructed according to the
specifically expressed miRNAs and the results of the GO analysis.
The control status of GO and miRNAs in the network was evaluated by
graph theory, and the degree of miRNAs indicated the number of GO
which was regulated by miRNAs in the network. In the same way, a
higher degree of GO indicated that more miRNAs contributed to
GO.
Statistical analysis
All quantitative data are presented as mean ±
standard deviation. Comparisons between the two groups were
performed using the Student's t-test. Comparisons of multiple group
data were performed using one-way analysis of variance followed by
Turkey's post hoc test. P-values <0.05 were considered
statistically significant. Statistical values were calculated using
SPSS software, version 20.0 (IBM Corp., Armonk, NY, USA).
Results
Screening differentially expressed
miRNAs
To identify significantly differentially expressed
miRNAs between heat-treated cells and controls, the screening
threshold was set to a fold change of >2 and a P-value of
<0.05. In total, 31 miRNAs (20 downregulated and 11 upregulated)
were identified (Table I).
 | Table IDifferentially expressed microRNAs
(miRNAs) between heat-treated cells and controls (P<0.05, fold
change >2). |
Table I
Differentially expressed microRNAs
(miRNAs) between heat-treated cells and controls (P<0.05, fold
change >2).
| miRNA | Fold change | Type of
regulation | P-value |
|---|
| hsa-miR-4448 | 0.240462612 | Down | <0.05 |
| hsa-miR-3180 | 0.243442001 | Down | <0.05 |
| hsa-miR-1231 | 0.309725485 | Down | <0.05 |
| hsa-miR-3620 | 0.330975034 | Down | <0.05 |
| hsa-miR-665 | 0.331296255 | Down | <0.05 |
| hsa-miR-4656 | 0.381063941 | Down | <0.05 |
| hsa-miR-4458 | 0.384188702 | Down | <0.05 |
|
hsa-miR-3613-3p | 0.385431663 | Down | <0.05 |
| hsa-miR-486-5p | 0.391376978 | Down | <0.05 |
|
hsa-miR-3180-3p | 0.404438992 | Down | <0.05 |
| hsa-miR-378h | 0.416819304 | Down | <0.05 |
| hsa-miR-1296 | 0.450624607 | Down | <0.05 |
| hsa-miR-616-3p | 0.453329075 | Down | <0.05 |
| hsa-miR-4486 | 0.453329201 | Down | <0.05 |
|
hsa-miR-4749-5p | 0.46624915 | Down | <0.05 |
| hsa-miR-489 | 0.468301966 | Down | <0.05 |
|
hsa-miR-3130-5p | 0.469450744 | Down | <0.05 |
| hsa-miR-411-5p | 0.470553479 | Down | <0.05 |
| hsa-miR-299-3p | 0.476023136 | Down | <0.05 |
|
hsa-miR-4665-5p | 0.492758274 | Down | <0.05 |
| hsa-miR-1290 | 2.07284178 | Up | <0.05 |
| hsa-miR-4430 | 2.135374336 | Up | <0.05 |
|
hsa-miR-4655-5p | 2.191798238 | Up | <0.05 |
| hsa-miR-140-5p | 2.318002303 | Up | <0.05 |
| hsa-miR-4500 | 2.324482718 | Up | <0.05 |
| hsa-miR-485-3p | 2.399120465 | Up | <0.05 |
| hsa-miR-589-5p | 2.399121462 | Up | <0.05 |
| hsa-miR-34c-5p | 2.518895027 | Up | <0.05 |
|
hsa-miR-3158-5p | 3.033915491 | Up | <0.05 |
|
hsa-miR-3613-5p | 3.690735374 | Up | <0.05 |
| hsa-miR-1281 | 4.409294993 | Up | <0.05 |
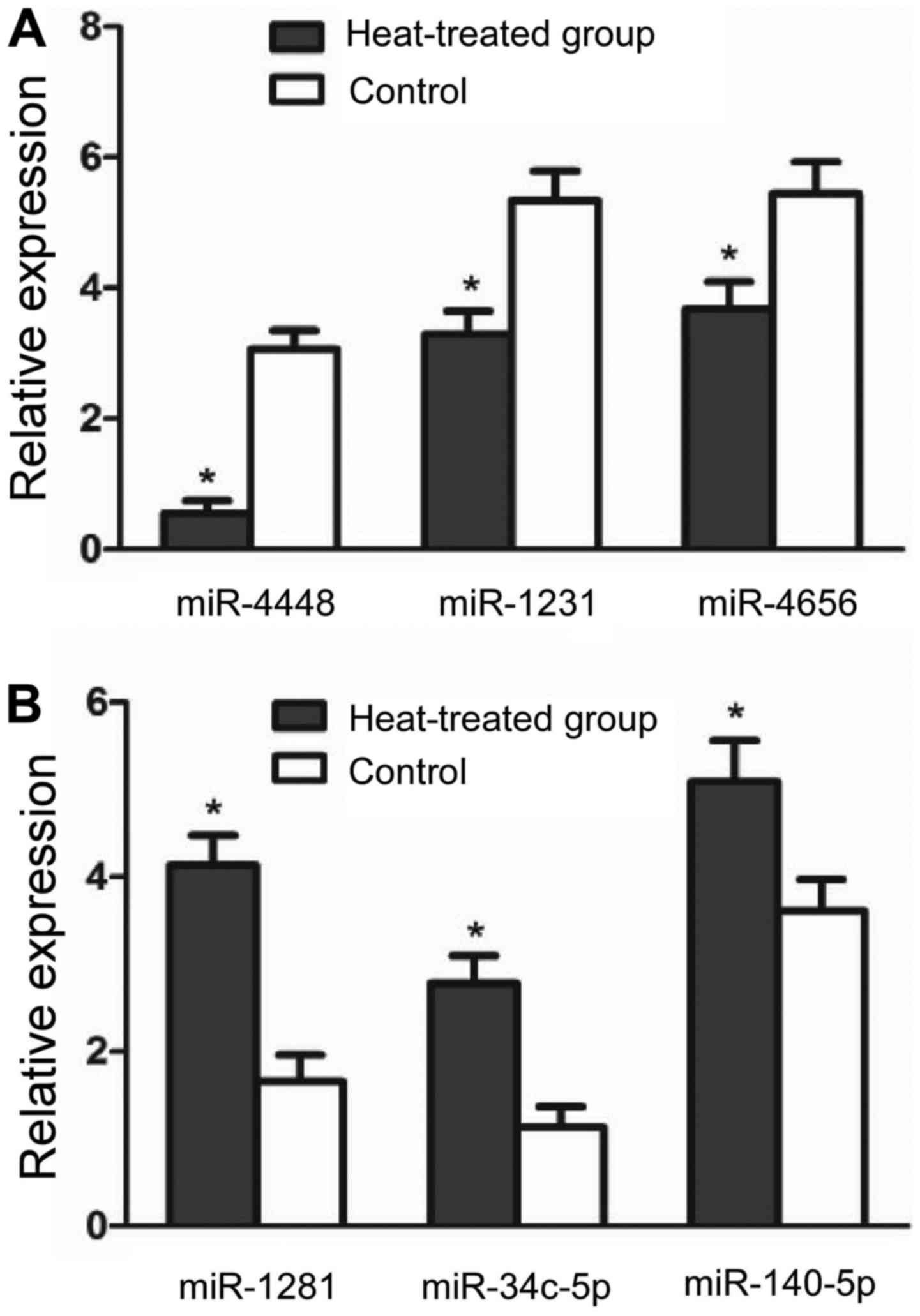
Validation by RT-qPCR
Based on their expression levels and fold change in
expression, 3 downregulated miRNAs (miR-1281, miR-34c-5p and
miR-140-5p) and 3 upregulated miRNAs (miR-4448, miR-1231 and
miR-4656) were selected for validation by RT-qPCR. The results
revealed a significant intra-group difference in the expression of
miRNAs, in a manner consistent with the data obtained from the
microarray analysis (P<0.05) (Fig.
1).
Target gene and bioinformatics
analysis
A total of 2,024 target genes were identified for
the validated miRNAs. To better understand the potential
implications of the differentially expressed miRNAs, these target
genes were subjected to GO analysis to evaluate their potential
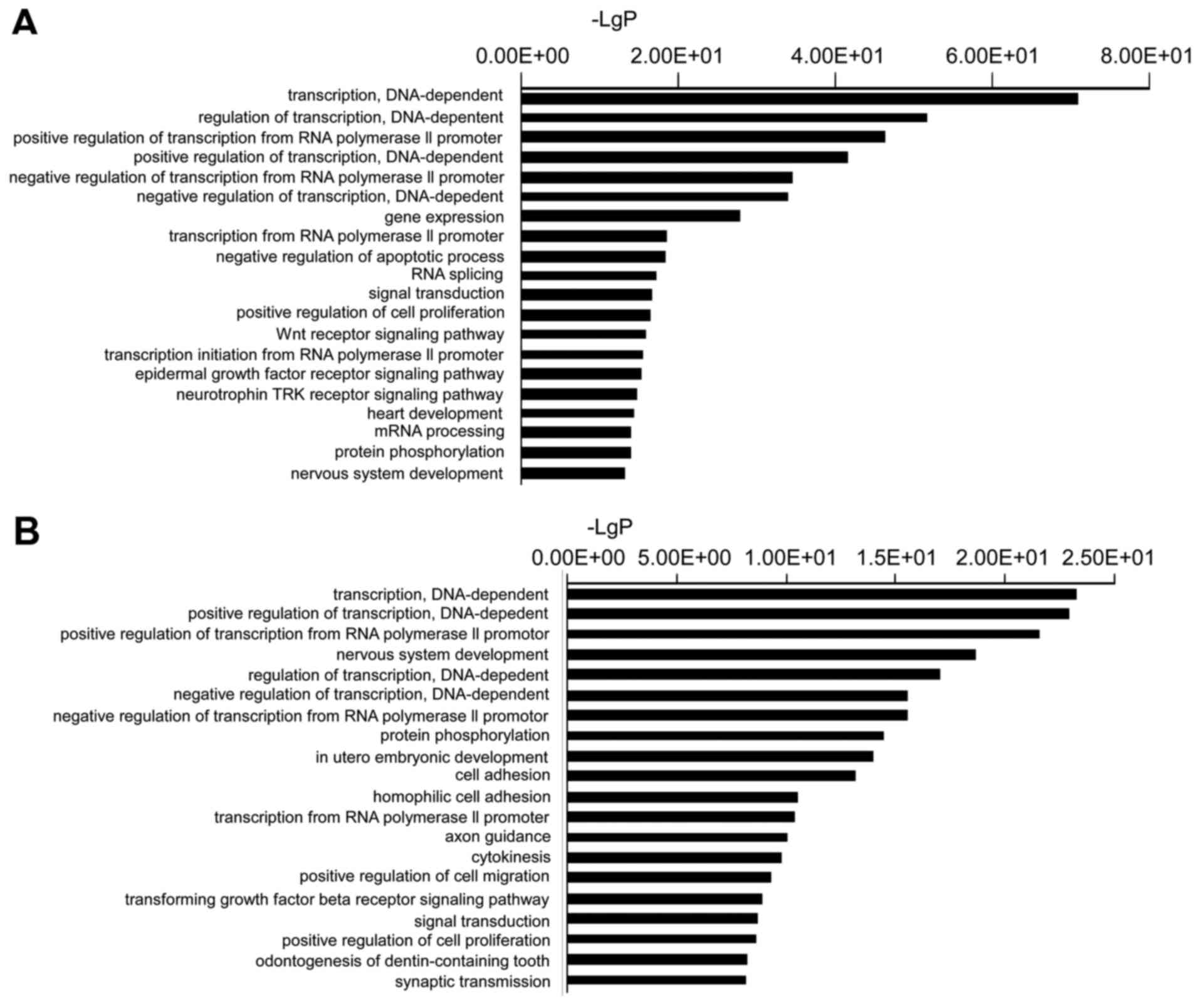
functions. In the present study, the top GO terms of the target
genes of the downregulated miRNAs were transcription
(DNA-dependent), regulation of transcription (DNA-dependent),
positive regulation of transcription from RNA polymerase II
promoter, positive regulation of transcription (DNA-dependent), and
negative regulation of transcription from RNA polymerase II
promoter. The top GO terms of the target genes of the upregulated
miRNAs were transcription (DNA-dependent), positive regulation of
transcription (DNA-dependent), positive regulation of transcription
from RNA polymerase II promoter, nervous system development, and
regulation of transcription (DNA-dependent) (Fig. 2).
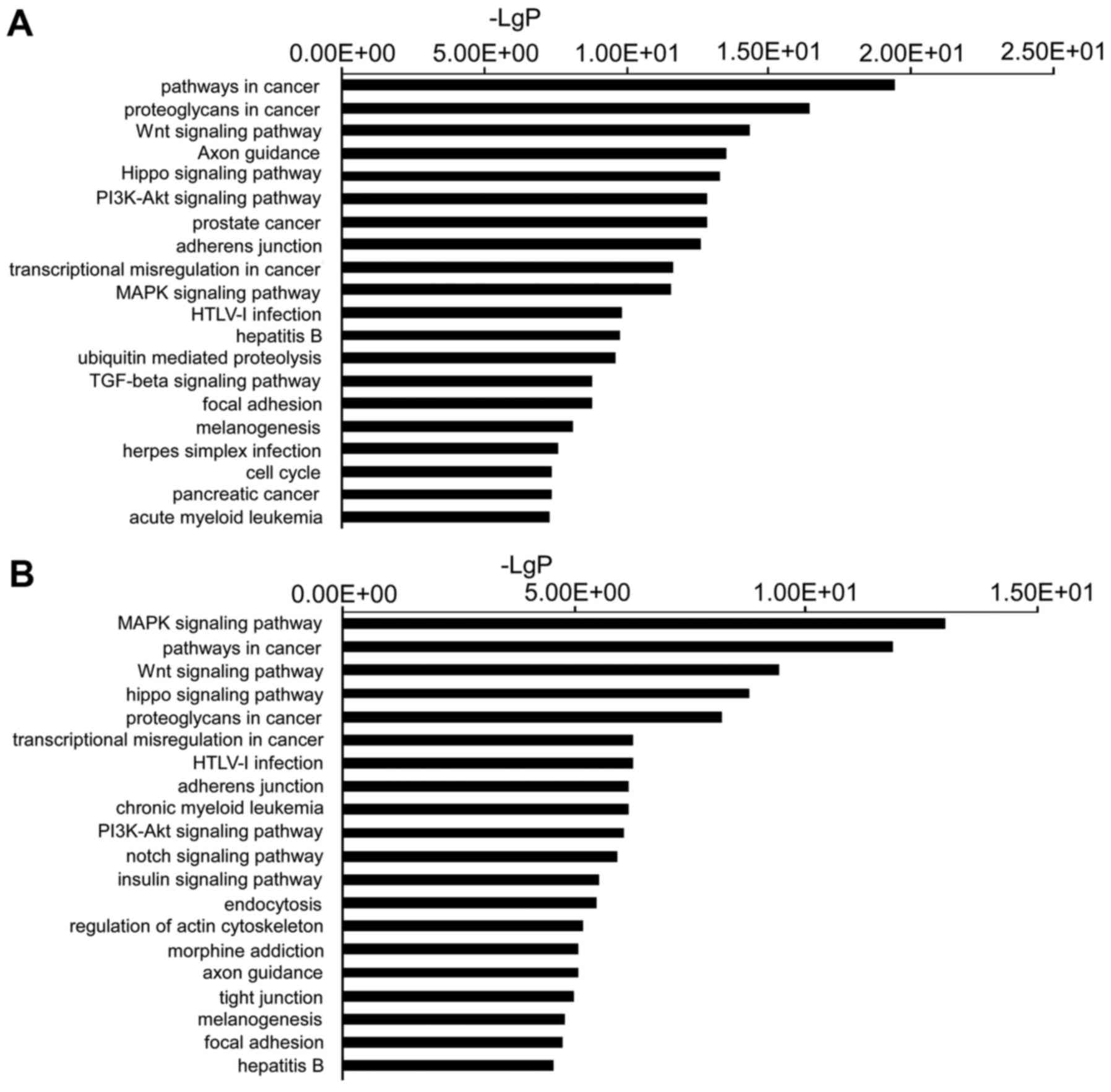
The target genes were then subjected to pathway
enrichment analysis using KEGG pathways to determine the canonical
pathways controlled by the identified miRNAs. Pathways involved in
cancer, proteoglycans involved in cancer, the Wnt signaling
pathway, axon guidance and Hippo signaling pathways, were the most
active pathways regulated by the downregulated miRNAs. The
mitogen-activated protein kinase (MAPK) signaling pathway, pathways
involved in cancer, the Wnt signaling pathway, the Hippo signaling
pathway and proteoglycans involved in cancer, were the top pathways
regulated by the upregulated miRNAs (Fig. 3).
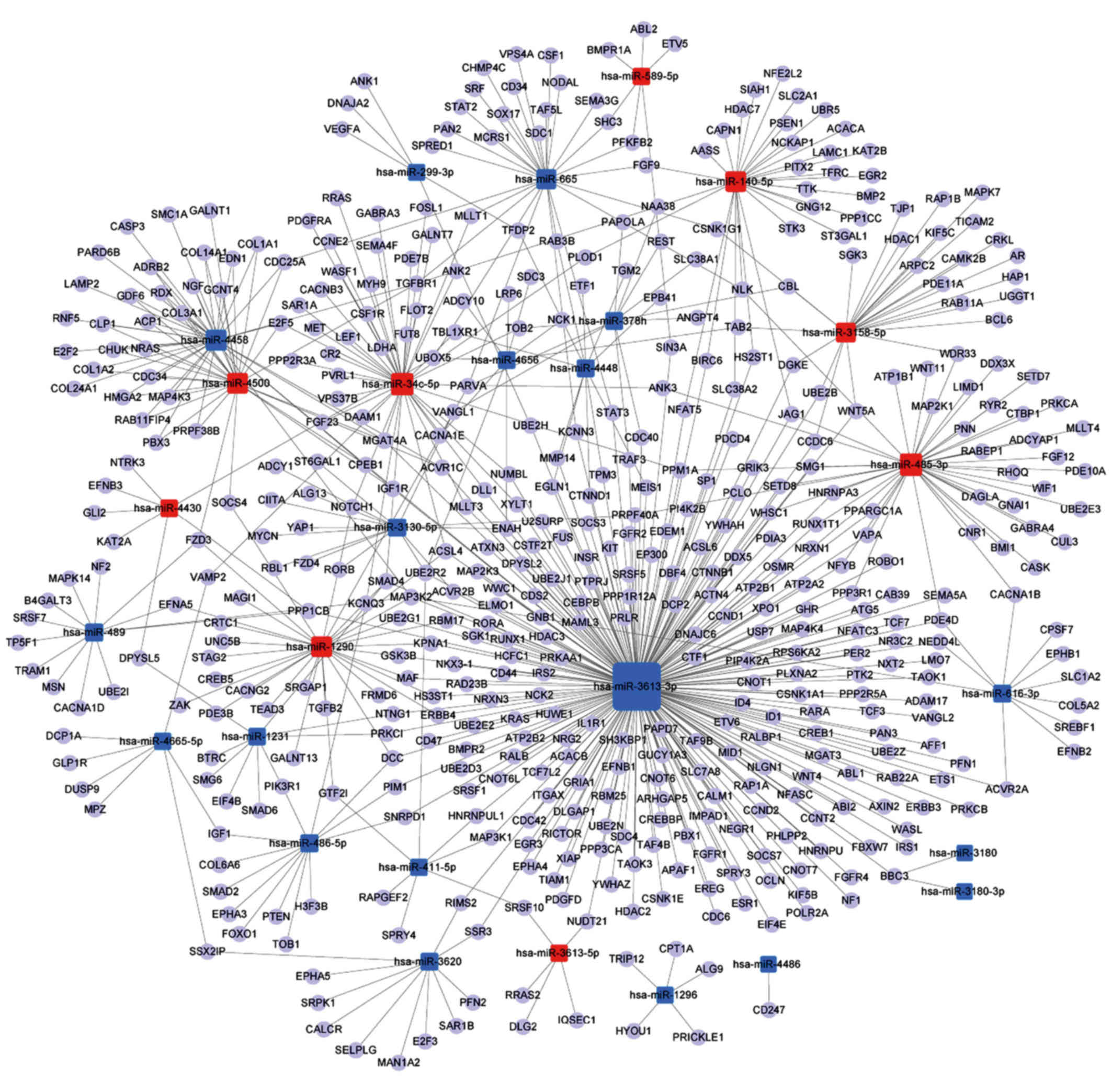
miRNA-gene-network analyses
A miRNA-gene-network was constructed according to
the results of the GO and pathway analyses. The core miRNAs of the
interaction network included miR-3613-3p, miR-34c-5p and miR-485-3p
(Table II). The network also
revealed that MAPK kinase kinase 2 (MAP3K2),
α-1,3-mannosyl-glycoprotein 4-β-N-acetylglu
cosaminyltransferase A (MGAT4A), transforming growth factor β
receptor 1 (TGFBR1), ubiquitin-conjugating enzyme E2 R2 (UBE2R2),
and Sma- and Mad-related family 4 (SMAD4), were the most crucial
target genes (Table III). The
associations of miRNAs with genes are shown in Fig. 4.
 | Table IICrucial microRNAs (miRNAs) in the
miRNA-gene-network (degree >10). |
Table II
Crucial microRNAs (miRNAs) in the
miRNA-gene-network (degree >10).
| miRNA | Degree |
|---|
|
hsa-miR-3613-3p | 237 |
| hsa-miR-34c-5p | 43 |
| hsa-miR-485-3p | 43 |
| hsa-miR-4458 | 34 |
| hsa-miR-4500 | 31 |
| hsa-miR-140-5p | 30 |
| hsa-miR-1290 | 30 |
| hsa-miR-665 | 27 |
|
hsa-miR-3158-5p | 24 |
|
hsa-miR-3130-5p | 14 |
| hsa-miR-486-5p | 13 |
| hsa-miR-489 | 13 |
| hsa-miR-616-3p | 11 |
| hsa-miR-3620 | 11 |
| hsa-miR-4448 | 11 |
| hsa-miR-378h | 11 |
 | Table IIICrucial target genes in the
microRNA-gene-network (degree >3). |
Table III
Crucial target genes in the
microRNA-gene-network (degree >3).
| Gene symbol | Gene
description | Degree |
|---|
| MAP3K2 | Homo sapiens
mitogen-activated protein kinase kinase kinase 2 (MAP3K2),
mRNA | 5 |
| MGAT4A | Homo sapiens
mannosyl (α-1,3-)-glycoprotein
β-1,4-N-acetylglucosaminyltransferase, isozyme A (MGAT4A),
transcript variant 1, mRNA | 4 |
| TGFBR1 | Homo sapiens
transforming growth factor, β receptor 1 (TGFBR1), transcript
variant 1, mRNA | 4 |
| UBE2R2 | Homo sapiens
ubiquitin-conjugating enzyme E2 R2 (UBE2R2), mRNA | 4 |
| SMAD4 | Homo sapiens
SMAD family member 4 (SMAD4), mRNA | 4 |
| PRPF40A | Homo sapiens
PRP40 pre-mRNA processing factor 40 homolog A (S.
cerevisiae) (PRPF40A), mRNA | 3 |
| PARVA | Homo sapiens
parvin, α (PARVA), mRNA | 3 |
| ACSL4 | Homo sapiens
acyl-CoA synthetase long-chain family member 4 (ACSL4), transcript
variant 2, mRNA | 3 |
| ENAH | Homo sapiens
enabled homolog (Drosophila) (ENAH), transcript variant 2,
mRNA | 3 |
| JAG1 | Homo sapiens
jagged 1 (JAG1), mRNA | 3 |
| PDE4D | Homo sapiens
phosphodiesterase 4D, cAMP-specific (PDE4D), transcript variant 2,
mRNA | 3 |
| E2F5 | Homo sapiens
E2F transcription factor 5, p130-binding (E2F5), transcript variant
1, mRNA | 3 |
| DCC | Homo sapiens
deleted in colorectal carcinoma (DCC), mRNA | 3 |
| PPM1A | Homo sapiens
protein phosphatase, Mg2+/Mn2+-dependent, 1A
(PPM1A), transcript variant 3, mRNA | 3 |
| IGF1R | Homo sapiens
insulin-like growth factor 1 receptor (IGF1R), mRNA | 3 |
| CDC25A | Homo sapiens
cell division cycle 25 homolog A (S. pombe) (CDC25A),
transcript variant 2, mRNA | 3 |
| CPEB1 | Homo sapiens
cytoplasmic polyadenylation element binding protein 1 (CPEB1),
transcript variant 1, mRNA | 3 |
| NLK | Homo sapiens
nemo-like kinase (NLK), mRNA | 3 |
| FZD3 | Homo sapiens
frizzled family receptor 3 (FZD3), transcript variant 2, mRNA | 3 |
| PPP1CB | Homo sapiens
protein phosphatase 1, catalytic subunit, β isozyme (PPP1CB),
transcript variant 3, mRNA | 3 |
| UBOX5 | Homo sapiens
U-box domain containing 5 (UBOX5), transcript variant 2, mRNA | 3 |
| BBC3 | Homo sapiens
BCL2-binding component 3 (BBC3), nuclear gene encoding
mitochondrial protein, transcript variant 4, mRNA | 3 |
| ACVR1C | Homo sapiens
activin A receptor, type IC (ACVR1C), transcript variant 1,
mRNA | 3 |
miRNA-GO-network analysis
The network analysis was useful for determining
regulatory associations between the key miRNAs and hub GO. In this
network, miR-3613-3p, miR-4458 and miR-4500, which contributed more
than the other specifically expressed miRNAs, exhibited 479, 217
and 216 GO functions, respectively (Table IV). The most significantly
regulated function-cluster of the total 20 categories was gene
expression, whereas others included positive regulation of
transcription, positive regulation of transcription by RNA
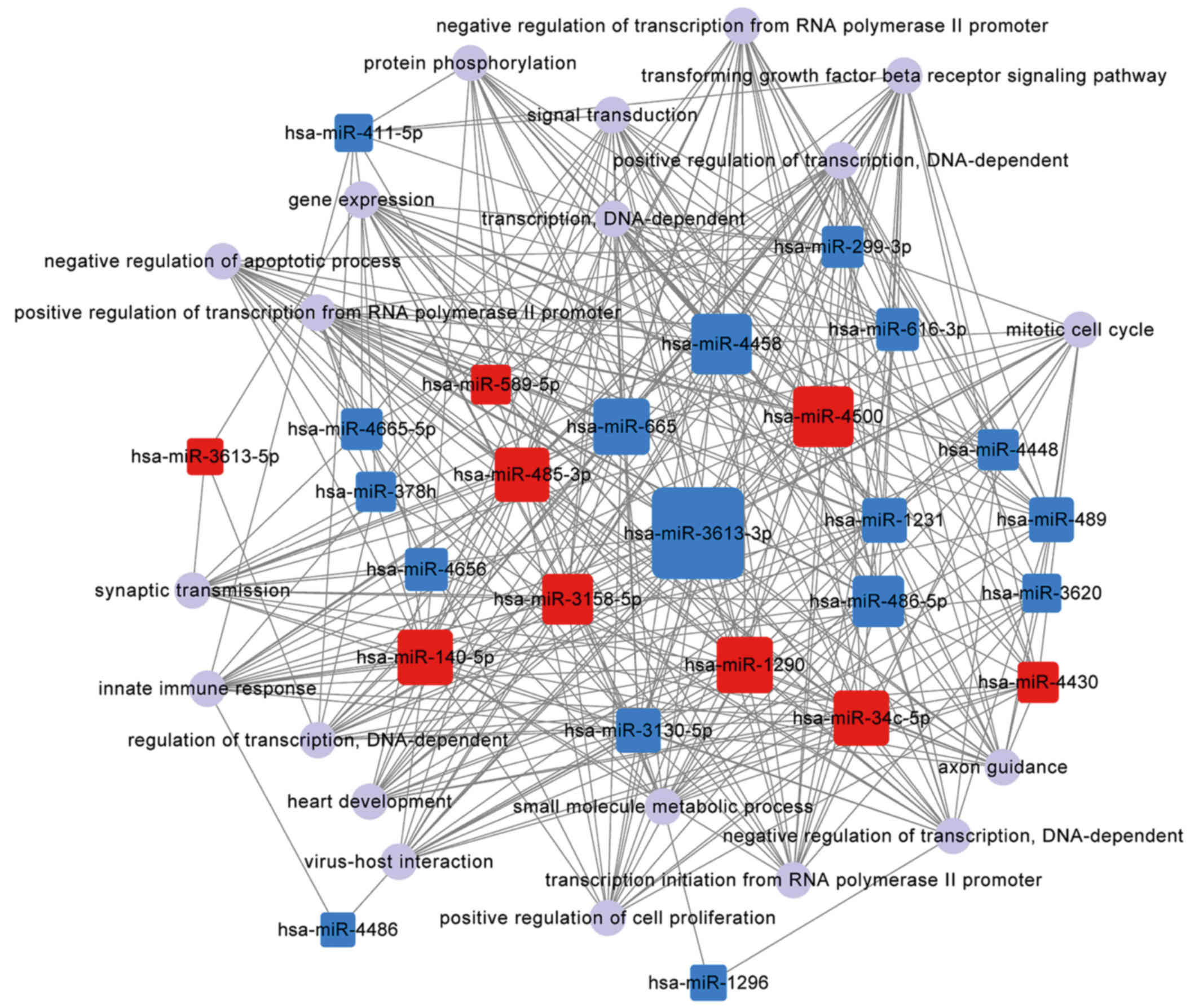
polymerase II promoter and signal transduction (Table V). The significantly complicated
associations of the top 20 functions with miRNAs are shown in
Fig. 5.
 | Table IVCrucial microRNAs (miRNAs) in the
miRNA-GO-network (degree >100). |
Table IV
Crucial microRNAs (miRNAs) in the
miRNA-GO-network (degree >100).
| miRNA | Degree |
|---|
|
hsa-miR-3613-3p | 479 |
| hsa-miR-4458 | 217 |
| hsa-miR-4500 | 216 |
| hsa-miR-665 | 182 |
| hsa-miR-1290 | 180 |
| hsa-miR-140-5p | 173 |
| hsa-miR-34c-5p | 173 |
| hsa-miR-485-3p | 165 |
| hsa-miR-486-5p | 141 |
|
hsa-miR-3158-5p | 135 |
 | Table VCrucial GO functions in the
microRNA-GO-network (degree >15). |
Table V
Crucial GO functions in the
microRNA-GO-network (degree >15).
| GO (name) | Degree |
|---|
| Gene
expression | 21 |
| Positive regulation
of transcription, DNA-dependent | 21 |
| Positive regulation
of transcription from RNA polymerase II promoter | 20 |
| Signal
transduction | 20 |
| Negative regulation
of transcription from RNA polymerase II promoter | 19 |
| Small molecule
metabolic process | 19 |
| Transforming growth
factor-β receptor signaling pathway | 19 |
| Axon guidance | 18 |
| Innate immune
response | 18 |
| Transcription,
DNA-dependent | 18 |
| Virus-host
interaction | 17 |
| Negative regulation
of apoptotic process | 16 |
| Negative regulation
of transcription, DNA-dependent | 16 |
| Regulation of
transcription, DNA-dependent | 16 |
| Synaptic
transmission | 16 |
| Transcription
initiation from RNA polymerase II promoter | 16 |
| Heart
development | 15 |
| Mitotic cell
cycle | 15 |
| Positive regulation
of cell proliferation | 15 |
| Protein
phosphorylation | 15 |
Discussion
In the present study, the miRNA expression profiles
of heat-treated HUVECs were evaluated by microarray analysis. A
number of miRNAs were obtained from both heat-treated cells and
normal controls, and 31 differentially expressed miRNAs were
selected, including 20 downregulated and 11 upregulated miRNAs
(Table I). The target genes of 31
differentially expressed miRNAs were predicted, and 2,024 target
genes were validated. GO and pathway enrichment analysis revealed
the significantly different functions and pathways targeted by the
differentially expressed miRNAs. A total of 755 notable GO terms
and 136 pathways were identified. The most active functions and
pathways are shown in Figs. 2 and
3. The wide target range
indicates that these miRNAs may play important roles in the
biological regulation of HUVECs under heat stress. These results
also demonstrated that the differentially expressed miRNAs were
predicted to control various functions that were most relevant to
the transcription-related process. Moreover, the miRNAs regulated
gene expression in ECs under heat treatment via multiple signaling
pathways, and biological regulatory mechanisms were integrated in
several main pathways. The majority of the pathways were
cancer-related. Following exposure to environmental stressors or
cytokines, these pathways may be activated to regulate cell
proliferation, migration, differentiation, evasion and
angiogenesis.
The network analysis linked the differentially
regulated miRNAs to target genes and the relevant biological
functions, and facilitated a better understanding of the role of
dysregulated miRNAs. In the networks, certain miRNAs functioned as
network hubs, such as miR-3613-3p, miR-34c-5p, miR-485-3p, miR-4458
and miR-4500. Of these miRNAs, miR-3613-3p was the most active
miRNA in the networks, targeting 237 genes and 479 biological
functions (Tables II and
IV and Figs. 4 and 5). Previous studies have not focused on
miR-3613-3p, and it has not been previously described as
participating in EC physiology and pathology. However, plasma
miR-3613-3p has been reported as a specific biomarker of lung
adenocarcinoma (38). Another
study indicated that circulating miR-3613-3p may predict persistent
severe axial pain following exposure to common stressors, and may
be involved in the pathogenesis of post-traumatic musculoskeletal
pain (39). Liu et al
found that miR-3613-3p was upregulated in left atrial appendages in
atrial fibrillation (40).
Moreover, Wang et al reported that urinary expression of
miR-3613-3p was downregulated in patients with immunoglobulin A
nephropathy, and the levels of miR-3613-3p were correlated with
disease severity (41).
Therefore, although the specific mechanism of function of
miR-3613-3 remains unknown, decreased miR-3613-3p expression may
exert a modulatory effect on ECs under heat stress. Additional
studies are required to elucidate the detailed role of miR-3613-3p
in ECs.
The miRNA-gene network analysis revealed a range of
hub genes, such as MAP3K2, MGAT4A, TGFBR1, UBE2R2 and SMAD4
(Table III). MAP3K2 encodes
MAPK kinase kinase 2 (MEKK2), which is a component of a protein
kinase signal transduction cascade that preferentially regulates
the c-Jun N-terminal kinases (JNKs) and extracellular
signal-regulated kinase 5 (ERK5) pathways by phosphorylating and
activating MAP2K5 as well as MAP2K7 in a similar manner (42,43). The MAP2K5/ERK5 pathway is required
for normal cardiovascular development and vascular integrity,
improves EC viability and reduces apoptosis (44,45), whereas JNK MAPKs promote apoptosis
of ECs under most conditions (46). MEKK2 coordinately activates
signaling through MEK5/ERK5 and MEK7/JNK protein kinases to respond
to different stimuli. MEKK2 has also been shown to directly
phosphorylate and activate IκB kinases and, thus, plays a role in
the nuclear factor-κB (NF-κB) signaling pathway (47,48). MEKK2 has also been found to bind
and activate protein kinase C-related kinase 2, which suggests its
involvement in a regulated signaling process (49). MGAT4A encodes the
glycosyltransferase N-acetylglucosaminyltransferase IVa, and
catalyzes the transfer of N-acetylglucosamine (GlcNAc) from
UDP-GlcNAc to the GlcNAcβ1-2Mana1-3 arm of the N-glycan core
via a β1-4 linkage (50,51). As an important post-translational
modification of target proteins, N-glycosylation plays an important
role in cell proliferation, differentiation, apoptosis and
development, during embryogenesis as well as in mature tissues
(52–54). The functions of N-glycans
associated with cell surface receptors are of great importance,
since N-glycans are directly involved in controlling
cellular functions. The target proteins of glycosyltransferases
include epidermal growth factor receptor, human epidermal growth
factor receptor 3, transforming growth factor-β (TGF-β) receptor,
T-cell receptors, β-site APP cleaving enzyme 1, glucose transporter
2, E-cadherin and α5β1 integrin (52). TGFBR1 encodes TGF-β type I
receptor (TβRI), which is the sole known cell surface receptor
serine/threonine kinase in humans and participates in TGF-β
signaling. The binding of TGF-β to TβRII dimers stabilizes the
interaction of TβRII dimers with two TβRI molecules to form a
ligand receptor-complex involving a heterotetrameric receptor and a
ligand dimer (55). Ligand
binding to TβRII causes the phosphorylation of TβRI, which
activates Smad2 and Smad3 (55).
The activated Smad proteins form complexes with Smad4 and are
translocated into the nucleus, where they regulate the
transcription of target genes (56,57). TGF-β1 exerts its effects on a
variety of cells through activation of the Smad proteins and other
signaling pathways, including MAPKs, small GTPases, protein kinase
C and phosphoinositide 3-kinase (58,59). Under most conditions, the activa
tion of the TGF-β1 signaling pathway inhibits EC proliferation and
migration and induces EC apoptosis (60). Smad4 is a central mediator of
TGF-β1/Smad signaling. Smad4 belongs to the evolutionarily
conserved family of Smad proteins that are crucial for transmitting
TGF-β1 superfamily signals from the cell surface to the nucleus
(61). UBE2R is involved in the
ubiquitin cascade. The ubiquitin-proteasome system (UPS) comprises
ubiquitin, ubiquitin-activating enzyme (E1), ubiquitin-binding
enzyme (E2), ubiquitin-ligating enzymes (E3) and the 26S proteasome
(62). Ubiquitin is attached to
substrates by a complex three-step enzymatic cascade, which
involves E1 ubiquitin activation (62,63), E2 ubiquitin conjugation and a
variety of E3 ubiquitin-ligating enzymes (64,65). The UPS plays an important role in
the regulation of cellular processes via degradation of damaged
proteins and the control of protein quantity in cells (66,67). Although the common effect of
ubiquitylation is degradation, the degradation-independent
Ub-signaling system also affects various biological processes,
which include cytokine production, proliferation, apoptosis,
inflammation, canalization, pain sensitization to host defense by
regulation of protein kinase activation, signal transduction,
membrane transport, transcriptional regulation, gene silencing, DNA
injury repair and DNA replication (62,68). All these genes receive
physicochemical stimuli and inputs from a diverse array of
receptors and coordinate the activation of multiple responses.
In addition to the miRNA-gene network analysis, the
miRNA-GO network analysis provides a more detailed view of the
integrated association between the regulation of miRNAs and heat
stress. Among the top 10 functions, gene expression was the most
active and was regulated by 21 differentially expressed miRNAs.
Furthermore, 4 transcription-related functions were regulated by 78
differentially expressed miRNAs (Table V). These findings indicated that
heat stress may affect EC gene expression directly via
differentially expressed miRNAs, or induce alterations in
transcription-related functions indirectly via miRNA profile
changes. This finding was not surprising, as most of the hub genes
participate in gene expression and transcription via certain
downstream molecules. For example, MAP3K2 was one of the top genes
revealed by the miRNA-gene network. The most significant role of
ERK5, which is a downstream target of MAP3K2, appears to be the
regulation of a number of downstream transcription factors,
including the myocyte-enhancer factor (MEF) family, MEF2A, C and D,
which are the best characterized members of this family (69,70). ERK5 also directly enhances the
transcription of c-Myc, cAMP-response element-binding protein, and
Sap1a (71–73). Enhancement of Sap1a occurs via a
serum response element that may also be involved in activating the
c-Fos promoter (74). JNKs,
another target of MAP3K2, translocate from the cytoplasm to the
nucleus once activated, and regulate a series of transcription
factors, such as c-Jun, Ets-like protein-1, activating
transcription factor 2, p53, c-Myc, and nuclear factor of activated
T-cells (75–79).
In the present study, the miRNA-GO network analysis
revealed a significantly complicated association between miRNAs and
different functions (Fig. 5 and
Tables IV and V). First, more functions than genes that
were targeted by miRNAs were found. Second, each differentially
expressed miRNA modulated a large variety of functions, and each
function was controlled by several miRNAs, thus resulting in a
complicated network. For example, in the present study, miR-3613-3p
targeted 479 functions, and the function of gene expression was
regulated by 21 differentially expressed miRNAs (Tables IV and V). Third, one miRNA may simultaneously
regulate opposing functions. For example, certain miRNAs, such as
miR-3613-3p, miR-4458 or miR-4500, were not only involved in the
positive regulation of transcription, but also in the negative
regulation of transcription (data not shown). Finally, certain
miRNAs contributed to the positive regulatory function, and other
miRNAs contributed to the corresponding negative regulatory
function. For example, miR-665 participated in the negative
regulation of the apoptotic process, but miR-4500 not only
participated in the anti-apoptotic process, but also promoted the
cellular apoptosis process (data not shown). Moreover, each
function was affected by both upregulated and downregulated miRNAs
(data not shown). Our results suggested that the functions
regulated by miRNAs were not an isolated event, and this regulation
depends on the balance between the miRNAs that promote the function
and the miRNAs that suppress the function. The miRNA-GO network
provided a global view of the molecular pathophysiology of EC heat
stress and helped identify the key 'players' in the whole
network.
There were several limitations to this study. First,
we were limited by the use of HUVECs. Although the baseline miRNA
expression levels are similar between ECs from different vascular
beds, some unique miRNA phenotypic diversity is present among the
different types of ECs (80). As
only one cell type was assessed, 'cell-specific' miRNAs could not
be identified in our analysis, which could be determined using
other cell types. Second, the number of samples utilized for the
miRNA microarray was relatively small and, therefore, less
predominantly differentiated miRNAs were difficult to detect with
the microarrays due to low power. Third, the miRNA microarray
technique had certain limitations, as some of the microarray
results may have been affected by technical artifacts introduced by
sample selection, the hybridization procedure, or the microarray
platform.
In conclusion, this study demonstrated that 31
miRNAs were differentially expressed in ECs under heat stress. The
bioinformatics analysis predicted the target genes, functions and
signaling pathways of these miRNAs. The miRNA-gene and miRNA-GO
network analyses revealed a range of hub miRNAs, genes and
functions, which may consolidate our understanding of the
involvement of miRNAs in the pathophysiology of heat-treated ECs.
miR-3613-3p may be involved in heat stress by regulating relevant
genes and functions. Although further mechanistic investigation is
required, the findings presented herein provide novel insights into
the molecular mechanisms of heat stress in ECs.
References
|
1
|
Bouchama A and Knochel JP: Heat stroke. N
Engl J Med. 346:1978–1988. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Roberts GT, Ghebeh H, Chishti MA,
Al-Mohanna F, El-Sayed R, Al-Mohanna F and Bouchama A:
Microvascular injury, thrombosis, inflammation, and apoptosis in
the pathogenesis of heatstroke: A study in baboon model.
Arterioscler Thromb Vasc Biol. 28:1130–1136. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Bazzoni G and Dejana E: Endothelial
cell-to-cell junctions: Molecular organization and role in vascular
homeostasis. Physiol Rev. 84:869–901. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Busse R and Fleming I: Vascular
endothelium and blood flow. Handb Exp Pharmacol. 176:43–78. 2006.
View Article : Google Scholar
|
|
5
|
Toda N, Nakanishi S and Tanabe S:
Aldosterone affects blood flow and vascular tone regulated by
endothelium-derived NO: Therapeutic implications. Br J Pharmacol.
168:519–533. 2013. View Article : Google Scholar :
|
|
6
|
Cines DB, Pollak ES, Buck CA, Loscalzo J,
Zimmerman GA, McEver RP, Pober JS, Wick TM, Konkle BA, Schwartz BS,
et al: Endothelial cells in physiology and in the pathophysiology
of vascular disorders. Blood. 91:3527–3561. 1998.PubMed/NCBI
|
|
7
|
Michiels C: Endothelial cell functions. J
Cell Physiol. 196:430–443. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Minshall RD and Malik AB: Transport across
the endothelium: Regulation of endothelial permeability. Handb Exp
Pharmacol. 176:107–144. 2006. View Article : Google Scholar
|
|
9
|
Pober JS and Sessa WC: Evolving functions
of endothelial cells in inflammation. Nat Rev Immunol. 7:803–815.
2007. View
Article : Google Scholar : PubMed/NCBI
|
|
10
|
Santoro MM and Nicoli S: miRNAs in
endothelial cell signaling: The endomiRNAs. Exp Cell Res.
319:1324–1330. 2013. View Article : Google Scholar :
|
|
11
|
Staszel T, Zapała B, Polus A,
Sadakierska-Chudy A, Kieć-Wilk B, Stępień E, Wybrańska I, Chojnacka
M and Dembińska-Kieć A: Role of microRNAs in endothelial cell
pathophysiology. Pol Arch Med Wewn. 121:361–366. 2011.PubMed/NCBI
|
|
12
|
Wang S, Aurora AB, Johnson BA, Qi X,
McAnally J, Hill JA, Richardson JA, Bassel-Duby R and Olson EN: The
endothelial-specific microRNA miR-126 governs vascular integrity
and angiogenesis. Dev Cell. 15:261–271. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Small EM, Sutherland LB, Rajagopalan KN,
Wang S and Olson EN: MicroRNA-218 regulates vascular patterning by
modulation of Slit-Robo signaling. Circ Res. 107:1336–1344. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Doebele C, Bonauer A, Fischer A, Scholz A,
Reiss Y, Urbich C, Hofmann WK, Zeiher AM and Dimmeler S: Members of
the microRNA-17-92 cluster exhibit a cell-intrinsic antiangiogenic
function in endothelial cells. Blood. 115:4944–4950. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Qin X, Wang X, Wang Y, Tang Z, Cui Q, Xi
J, Li YS, Chien S and Wang N: MicroRNA-19a mediates the suppressive
effect of laminar flow on cyclin D1 expression in human umbilical
vein endothelial cells. Proc Natl Acad Sci USA. 107:3240–3244.
2010. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Bang C, Fiedler J and Thum T:
Cardiovascular importance of the microRNA-23/27/24 family.
Microcirculation. 19:208–214. 2012. View Article : Google Scholar
|
|
17
|
O'Connell RM, Taganov KD, Boldin MP, Cheng
G and Baltimore D: MicroRNA-155 is induced during the macrophage
inflammatory response. Proc Natl Acad Sci USA. 104:1604–1609. 2007.
View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Tili E, Michaille JJ, Cimino A, Costinean
S, Dumitru CD, Adair B, Fabbri M, Alder H, Liu CG, Calin GA, et al:
Modulation of miR-155 and miR-125b levels following
lipopolysaccharide/TNF-alpha stimulation and their possible roles
in regulating the response to endotoxin shock. J Immunol.
179:5082–5089. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Sun X, Icli B, Wara AK, Belkin N, He S,
Kobzik L, Hunninghake GM, Vera MP, Blackwell TS, Baron RM, et al:
MICU Registry: MicroRNA-181b regulates NF-κB-mediated vascular
inflammation. J Clin Invest. 122:1973–1990. 2012.PubMed/NCBI
|
|
20
|
Fang Y, Shi C, Manduchi E, Civelek M and
Davies PF: MicroRNA-10a regulation of proinflammatory phenotype in
athero-susceptible endothelium in vivo and in vitro. Proc Natl Acad
Sci USA. 107:13450–13455. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Qin B, Xiao B, Liang D, Xia J, Li Y and
Yang H: MicroRNAs expression in ox-LDL treated HUVECs: MiR-365
modulates apoptosis and Bcl-2 expression. Biochem Biophys Res
Commun. 410:127–133. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Fang Y and Davies PF: Site-specific
microRNA-92a regulation of Kruppel-like factors 4 and 2 in
atherosusceptible endothelium. Arterioscler Thromb Vasc Biol.
32:979–987. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Zhu N, Zhang D, Chen S, Liu X, Lin L,
Huang X, Guo Z, Liu J, Wang Y, Yuan W, et al: Endothelial enriched
microRNAs regulate angiotensin II-induced endothelial inflammation
and migration. Atherosclerosis. 215:286–293. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Li D, Yang P, Xiong Q, Song X, Yang X, Liu
L, Yuan W and Rui YC: MicroRNA-125a/b-5p inhibits endothelin-1
expression in vascular endothelial cells. J Hypertens.
28:1646–1654. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Ni CW, Qiu H and Jo H: MicroRNA-663
upregulated by oscillatory shear stress plays a role in
inflammatory response of endothelial cells. Am J Physiol Heart Circ
Physiol. 300:H1762–H1769. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Suárez Y, Wang C, Manes TD and Pober JS:
Cutting edge: TNF-induced microRNAs regulate TNF-induced expression
of E-selectin and intercellular adhesion molecule-1 on human
endothelial cells: feedback control of inflammation. J Immunol.
184:21–25. 2010. View Article : Google Scholar
|
|
27
|
Harris TA, Yamakuchi M, Ferlito M, Mendell
JT and Lowenstein CJ: MicroRNA-126 regulates endothelial expression
of vascular cell adhesion molecule 1. Proc Natl Acad Sci USA.
105:1516–1521. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Dejana E, Corada M and Lampugnani MG:
Endothelial cell-to-cell junctions. FASEB J. 9:910–918.
1995.PubMed/NCBI
|
|
29
|
Pepini T, Gorbunova EE, Gavrilovskaya IN,
Mackow JE and Mackow ER: Andes virus regulation of cellular
microRNAs contributes to hantavirus-induced endothelial cell
permeability. J Virol. 84:11929–11936. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Nicoloso MS, Spizzo R, Shimizu M, Rossi S
and Calin GA: MicroRNAs - the micro steering wheel of tumour
metastases. Nat Rev Cancer. 9:293–302. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Sun HX, Zeng DY, Li RT, Pang RP, Yang H,
Hu YL, Zhang Q, Jiang Y, Huang LY, Tang YB, et al: Essential role
of microRNA-155 in regulating endothelium-dependent vasorelaxation
by targeting endothelial nitric oxide synthase. Hypertension.
60:1407–1414. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Suárez Y, Fernández-Hernando C, Pober JS
and Sessa WC: Dicer dependent microRNAs regulate gene expression
and functions in human endothelial cells. Circ Res. 100:1164–1173.
2007. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Weber M, Baker MB, Moore JP and Searles
CD: MiR-21 is induced in endothelial cells by shear stress and
modulates apoptosis and eNOS activity. Biochem Biophys Res Commun.
393:643–648. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Hergenreider E, Heydt S, Tréguer K,
Boettger T, Horrevoets AJ, Zeiher AM, Scheffer MP, Frangakis AS,
Yin X, Mayr M, et al: Atheroprotective communication between
endothelial cells and smooth muscle cells through miRNAs. Nat Cell
Biol. 14:249–256. 2012. View
Article : Google Scholar : PubMed/NCBI
|
|
35
|
Zhang Y, Liu D, Chen X, Li J, Li L, Bian
Z, Sun F, Lu J, Yin Y, Cai X, et al: Secreted monocytic miR-150
enhances targeted endothelial cell migration. Mol Cell. 39:133–144.
2010. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Chamorro-Jorganes A, Araldi E and Suárez
Y: MicroRNAs as pharmacological targets in endothelial cell
function and dysfunction. Pharmacol Res. 75:15–27. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Benjamini Y, Drai D, Elmer G, Kafkafi N
and Golani I: Controlling the false discovery rate in behavior
genetics research. Behav Brain Res. 125:279–284. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Pu Q, Huang Y, Lu Y, Peng Y, Zhang J, Feng
G, Wang C, Liu L and Dai Y: Tissue-specific and plasma microRNA
profiles could be promising biomarkers of histological
classification and TNM stage in non-small cell lung cancer. Thorac
Cancer. 7:348–354. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Linnstaedt SD, Walker MG, Parker JS, Yeh
E, Sons RL, Zimny E, Lewandowski C, Hendry PL, Damiron K, Pearson
C, et al: MicroRNA circulating in the early aftermath of motor
vehicle collision predict persistent pain development and suggest a
role for microRNA in sex-specific pain differences. Mol Pain.
11:662015. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Liu H, Chen GX, Liang MY, Qin H, Rong J,
Yao JP and Wu ZK: Atrial fibrillation alters the microRNA
expression profiles of the left atria of patients with mitral
stenosis. BMC Cardiovasc Disord. 14:102014. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Wang N, Bu R, Duan Z, Zhang X, Chen P, Li
Z, Wu J, Cai G and Chen X: Profiling and initial validation of
urinary microRNAs as biomarkers in IgA nephropathy. PeerJ.
3:e9902015. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Dermott JM, Ha JH, Lee CH and Dhanasekaran
N: Differential regulation of Jun N-terminal kinase and p38MAP
kinase by Galpha12. Oncogene. 23:226–232. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Raviv Z, Kalie E and Seger R: MEK5 and
ERK5 are localized in the nuclei of resting as well as stimulated
cells, while MEKK2 translocates from the cytosol to the nucleus
upon stimulation. J Cell Sci. 117:1773–1784. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Nithianandarajah-Jones GN, Wilm B,
Goldring CE, Müller J and Cross MJ: The role of ERK5 in endothelial
cell function. Biochem Soc Trans. 42:1584–1589. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Wu Y and Chakrabarti S: ERK5 mediated
signalling in diabetic retinopathy. Med Hypothesis Discov Innov
Ophthalmol. 4:17–26. 2015.PubMed/NCBI
|
|
46
|
Weston CR and Davis RJ: The JNK signal
transduction pathway. Curr Opin Cell Biol. 19:142–149. 2007.
View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Schmidt C, Peng B, Li Z, Sclabas GM,
Fujioka S, Niu J, Schmidt-Supprian M, Evans DB, Abbruzzese JL and
Chiao PJ: Mechanisms of proinflammatory cytokine-induced biphasic
NF-kappaB activation. Mol Cell. 12:1287–1300. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Winsauer G, Resch U, Hofer-Warbinek R,
Schichl YM and de Martin R: XIAP regulates bi-phasic NF-kappaB
induction involving physical interaction and ubiquitination of
MEKK2. Cell Signal. 20:2107–2112. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
49
|
Sun W, Vincent S, Settleman J and Johnson
GL: MEK kinase 2 binds and activates protein kinase C-related
kinase 2. Bifurcation of kinase regulatory pathways at the level of
an MAPK kinase kinase. J Biol Chem. 275:24421–24428. 2000.
View Article : Google Scholar : PubMed/NCBI
|
|
50
|
Oguri S, Yoshida A, Minowa MT and Takeuchi
M: Kinetic properties and substrate specificities of two
recombinant human N-acetylglucosaminyltransferase-IV isozymes.
Glycoconj J. 23:473–480. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
51
|
Minowa MT, Oguri S, Yoshida A, Hara T,
Iwamatsu A, Ikenaga H and Takeuchi M: cDNA cloning and expression
of bovine UDP-N-acetylglucosamine: alpha1, 3-D-mannoside
beta1,4-N-acetylglucosaminyltransferase IV. J Biol Chem.
273:11556–11562. 1998. View Article : Google Scholar : PubMed/NCBI
|
|
52
|
Takahashi M, Kizuka Y, Ohtsubo K, Gu J and
Taniguchi N: Disease-associated glycans on cell surface proteins.
Mol Aspects Med. 51:56–70. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
53
|
Fan J, Wang S, Yu S, He J, Zheng W and
Zhang J: N-acetylglucosaminyltransferase IVa regulates metastatic
potential of mouse hepatocarcinoma cells through glycosylation of
CD147. Glycoconj J. 29:323–334. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
54
|
Niimi K, Yamamoto E, Fujiwara S, Shinjo K,
Kotani T, Umezu T, Kajiyama H, Shibata K, Ino K and Kikkawa F: High
expression of N-acetylglucosaminyltransferase IVa promotes invasion
of choriocarcinoma. Br J Cancer. 107:1969–1977. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
55
|
López-Casillas F, Wrana JL and Massagué J:
Betaglycan presents ligand to the TGF beta signaling receptor.
Cell. 73:1435–1444. 1993. View Article : Google Scholar : PubMed/NCBI
|
|
56
|
Yin JJ, Selander K, Chirgwin JM, Dallas M,
Grubbs BG, Wieser R, Massagué J, Mundy GR and Guise TA: TGF-beta
signaling blockade inhibits PTHrP secretion by breast cancer cells
and bone metastases development. J Clin Invest. 103:197–206. 1999.
View Article : Google Scholar : PubMed/NCBI
|
|
57
|
Siegel PM and Massagué J: Cytostatic and
apoptotic actions of TGF-beta in homeostasis and cancer. Nat Rev
Cancer. 3:807–821. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
58
|
Smith AL, Iwanaga R, Drasin DJ, Micalizzi
DS, Vartuli RL, Tan AC and Ford HL: The miR-106b-25 cluster targets
Smad7, activates TGF-β signaling, and induces EMT and tumor
initiating cell characteristics downstream of Six1 in human breast
cancer. Oncogene. 31:5162–5171. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
59
|
Chu IM, Lai WC, Aprelikova O, El Touny LH,
Kouros-Mehr H and Green JE: Expression of GATA3 in MDA-MB-231
triple-negative breast cancer cells induces a growth inhibitory
response to TGFβ. PLoS One. 8:e611252013. View Article : Google Scholar
|
|
60
|
Roberts AB and Sporn MB: Regulation of
endothelial cell growth, architecture, and matrix synthesis by
TGF-beta. Am Rev Respir Dis. 140:1126–1128. 1989. View Article : Google Scholar : PubMed/NCBI
|
|
61
|
Zhang Y, Feng XH and Derynck R: Smad3 and
Smad4 cooperate with c-Jun/c-Fos to mediate TGF-beta-induced
transcription. Nature. 394:909–913. 1998. View Article : Google Scholar : PubMed/NCBI
|
|
62
|
Swatek KN and Komander D: Ubiquitin
modifications. Cell Res. 26:399–422. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
63
|
Schulman BA and Harper JW: Ubiquitin-like
protein activation by E1 enzymes: The apex for downstream
signalling pathways. Nat Rev Mol Cell Biol. 10:319–331. 2009.
View Article : Google Scholar : PubMed/NCBI
|
|
64
|
Ye Y and Rape M: Building ubiquitin
chains: E2 enzymes at work. Nat Rev Mol Cell Biol. 10:755–764.
2009. View Article : Google Scholar : PubMed/NCBI
|
|
65
|
Deshaies RJ and Joazeiro CA: RING domain
E3 ubiquitin ligases. Annu Rev Biochem. 78:399–434. 2009.
View Article : Google Scholar : PubMed/NCBI
|
|
66
|
Komander D and Rape M: The ubiquitin code.
Annu Rev Biochem. 81:203–229. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
67
|
Hoeller D, Hecker CM and Dikic I:
Ubiquitin and ubiquitin-like proteins in cancer pathogenesis. Nat
Rev Cancer. 6:776–788. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
68
|
Meier P, Morris O and Broemer M:
Ubiquitin-mediated regulation of cell death, inflammation, and
defense of homeostasis. Curr Top Dev Biol. 114:209–239. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
69
|
Yang CC, Ornatsky OI, McDermott JC, Cruz
TF and Prody CA: Interaction of myocyte enhancer factor 2 (MEF2)
with a mitogen-activated protein kinase, ERK5/BMK1. Nucleic Acids
Res. 26:4771–4777. 1998. View Article : Google Scholar : PubMed/NCBI
|
|
70
|
Kato Y, Zhao M, Morikawa A, Sugiyama T,
Chakravortty D, Koide N, Yoshida T, Tapping RI, Yang Y, Yokochi T,
et al: Big mitogen-activated kinase regulates multiple members of
the MEF2 protein family. J Biol Chem. 275:18534–18540. 2000.
View Article : Google Scholar : PubMed/NCBI
|
|
71
|
Kamakura S, Moriguchi T and Nishida E:
Activation of the protein kinase ERK5/BMK1 by receptor tyrosine
kinases. Identification and characterization of a signaling pathway
to the nucleus. J Biol Chem. 274:26563–26571. 1999. View Article : Google Scholar : PubMed/NCBI
|
|
72
|
English JM, Pearson G, Baer R and Cobb MH:
Identification of substrates and regulators of the
mitogen-activated protein kinase ERK5 using chimeric protein
kinases. J Biol Chem. 273:3854–3860. 1998. View Article : Google Scholar : PubMed/NCBI
|
|
73
|
Watson FL, Heerssen HM, Bhattacharyya A,
Klesse L, Lin MZ and Segal RA: Neurotrophins use the Erk5 pathway
to mediate a retrograde survival response. Nat Neurosci. 4:981–988.
2001. View Article : Google Scholar : PubMed/NCBI
|
|
74
|
Nithianandarajah-Jones GN, Wilm B,
Goldring CE, Müller J and Cross MJ: ERK5: Structure, regulation and
function. Cell Signal. 24:2187–2196. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
75
|
McCubrey JA, Steelman LS, Chappell WH,
Abrams SL, Wong EW, Chang F, Lehmann B, Terrian DM, Milella M,
Tafuri A, et al: Roles of the Raf/MEK/ERK pathway in cell growth,
malignant transformation and drug resistance. Biochim Biophys Acta.
1773:1263–1284. 2007. View Article : Google Scholar
|
|
76
|
Oleinik NV, Krupenko NI and Krupenko SA:
Cooperation between JNK1 and JNK2 in activation of p53 apoptotic
pathway. Oncogene. 26:7222–7230. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
77
|
Iavarone C, Catania A, Marinissen MJ,
Visconti R, Acunzo M, Tarantino C, Carlomagno MS, Bruni CB, Gutkind
JS and Chiariello M: The platelet-derived growth factor controls
c-myc expression through a JNK- and AP-1-dependent signaling
pathway. J Biol Chem. 278:50024–50030. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
78
|
Zhong S, Fromm J and Johnson DL: TBP is
differentially regulated by c-Jun N-terminal kinase 1 (JNK1) and
JNK2 through Elk-1, controlling c-Jun expression and cell
proliferation. Mol Cell Biol. 27:54–64. 2007. View Article : Google Scholar :
|
|
79
|
Chow CW, Dong C, Flavell RA and Davis RJ:
c-Jun NH(2)-terminal kinase inhibits targeting of the protein
phosphatase calcineurin to NFATc1. Mol Cell Biol. 20:5227–5234.
2000. View Article : Google Scholar : PubMed/NCBI
|
|
80
|
McCall MN, Kent OA, Yu J, Fox-Talbot K,
Zaiman AL and Halushka MK: MicroRNA profiling of diverse
endothelial cell types. BMC Med Genomics. 4:782011. View Article : Google Scholar : PubMed/NCBI
|