Introduction
Visualization of alterations in cell-surface
receptor expression in malignant cells may identify molecular
targets enabling specific treatment of an individual tumor. A
spectacular example of such an approach is the clinical use of
imaging of the expression of somatostatin receptors for patient
selection for subsequent therapy using radiolabeled somatostatin
analogues (1). Relatively
recently, in vivo imaging of receptor tyrosine kinase (RTK)
expression has attracted increased attention (2,3).
RTKs normally regulate cellular division, differentiation, motility
and apoptosis, i.e. phenomena that are essential in malignancies.
Aberrant expression of RTKs is often one of the driving forces of a
malignancy, and targeting of overexpressed RTKs is one of the major
directions in development of anticancer drugs (4). The epidermal growth factor receptor
(EGFR) is an RTK that is often overexpressed in a variety of
malignancies (5).
Overexpression/amplification of EGFR is associated with shorter
survival in gastric and esophageal adenocarcinoma (6), pancreatic adenocarcinoma (7), vulvar carcinoma (8), head and neck squamous cell carcinoma
(HNSCC) (9) and glioma (10). EGFR is a well-established target
for monoclonal antibodies and specific tyrosine kinase inhibitors
(11).
The specific character of anti-EGFR therapeutics
necessitates an identification of patients with tumors that will
respond to therapy. The expression level of the receptor is one of
the possible predictors for the response. In some cases,
overexpression of EGFR cannot be a sole predicting biomarker. For
example, presence of specific mutations in the kinase domain of
EGFR is a precondition to response of non-small cell lung cancer
(NSCLC) to the tyrosine kinase inhibitor gefitinib in a number of
settings (12,13). Metastatic colorectal cancer would
not respond to anti-EGFR antibody-treatment in the case of
mutations in the intracellular signaling cascades (14). However, information concerning the
expression level of wild-type EGFR is helpful in selection of the
optimal treatment in many other instances. Non-small cell lung
cancer overexpressing EGFR would be more likely to respond to the
addition of cetuximab to a first-line chemotherapy (15) and to treatment with gefitinib
(16,17) compared to NSCLCs with low EGFR
expression. The addition of cetuximab to chemoradiotherapy of stage
III HNSCC significantly improves survival of patients with tumors
having high EGFR expression (18).
In the case of low EGFR expression, the use of cetuximab shortens
survival. In HNSCC, high expression of EGFR is associated with
relapse after radiotherapy (19).
For such patients, accelerated radiotherapy fractionation would
provide advantages compared to conventional radiation treatment
(20,21). High expression of EGFR in
esophageal squamous cell carcinoma is a precondition for successful
treatment with the TKI icotinib (22). High EGFR expression is a negative
predicting biomarker for response of triple-negative breast cancer
to neoadjuvant therapy using anthracyclines and taxanes (23). The main problem is that the
expression level of EGFR can vary during the metastasis process,
and the discordance rate between biopsy samples from primary NSCLC
and metastases might be up to 50% (24). This necessitates a reliable
methodology for assessment of EGFR expression in disseminated
cancer.
The use of radionuclide molecular imaging has a
potential for non-invasive estimation of EGFR expression in
multiple metastatic sites. Several radiolabeled monoclonal
anti-EGFR antibodies have been evaluated as imaging probes
(25–28). The feasibility of in vivo
imaging of EGFR expression has been demonstrated in these studies.
However, all radiolabeled antibodies clear slowly from blood and
non-specific compartments, which results in moderate contrast and
requires several days between injection of the antibody and
imaging. The use of smaller radiolabeled fragments of cetuximab as
imaging agents increased appreciably the contrast of EGFR imaging
and enabled shortening of the time between injection of the probe
and the imaging session (29,30).
A smaller size of the (Fab′)2-fragment contributed to
both more rapid clearance and better tumor localization, which
demonstrated advantages of a reduction of the imaging probe size
for improved contrast. An alternative to the use of monoclonal
antibodies for imaging of EGFR is the use of affibody
molecules.
Affibody molecules are small affinity proteins that
can be engineered to bind a large repertoire of different target
proteins through creation of combinatorial libraries and a
subsequent selection procedure for the isolation of target-specific
binders (31). Combination of
small size (6–7 kDa) with high affinity makes affibody molecules
attractive as imaging probes, which has been shown in preclinical
(32) and clinical studies
(33). Several anti-EGFR affibody
molecules have been developed and labeled with 111In
(34,35) for SPECT and 18F
(36,37) or 89Zr (38) for PET imaging. The present study
was focused on the ZEGFR:2377 affibody molecule (35,39),
which has equal affinity to human and murine EGFR. This feature is
essential since EGFR is expressed in a number of normal tissues.
The cross-reactivity with murine receptor makes mouse models
adequate to study all aspects of in vivo targeting.
The rapid progress in the design and performance of
SPECT cameras during the last years (40) suggests that the development of
molecular imaging probes labeled with single-photon emitters
remains to be of interest. The use of the generator-produced
radionuclide 99mTc as a label would be preferable for
this purpose, as this nuclide offers a number of advantages
compared to alternative labels, such as 111In or
123I which includes low cost, excellent availability and
favorable dosimetry (41).
We have reported earlier labeling of anti-HER2
affibody molecule ZHER2:2395 with 99mTc using
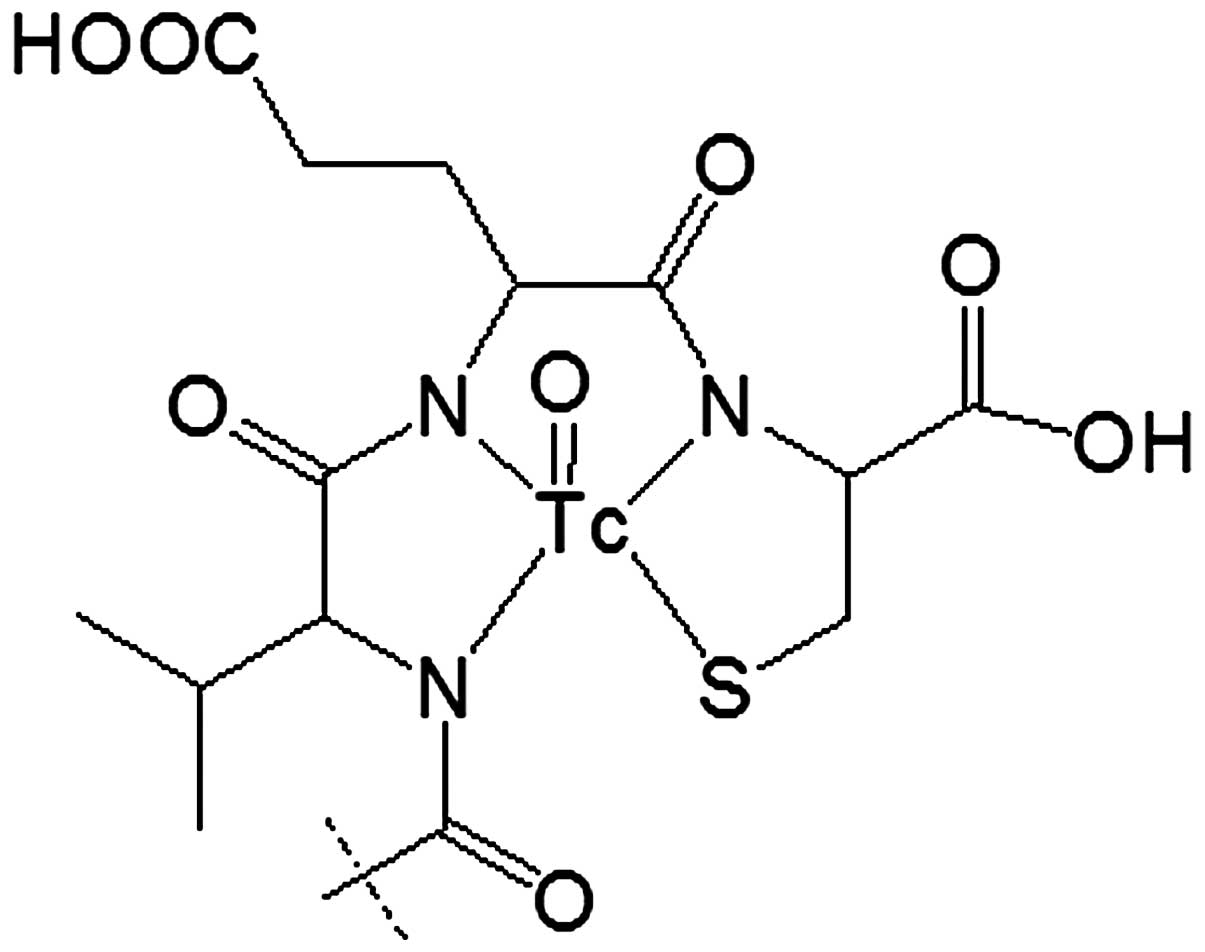
incorporation of cysteine at C-terminus (42). The thiol group of cysteine forms
together with amide nitrogen of adjacent amino acids a
N3S chelator providing a stable complex with
99mTc (42) (Fig. 1). Such approach might be considered
for anti-EGFR affibody ZEGFR:2377 as well. However, our studies
have demonstrated that modification of the amino acid sequence of
affibody molecule might create an alternative binding site for
technetium-99m, providing lower stability of the complex (43). Since the amino acid composition of
ZEGFR:2377 differs profoundly from the composition of ZHER2:2395 we
had to consider formation of an alternative chelating site as
well.
The aim of the present study was to evaluate the
feasibility of imaging of EGFR expression with the ZEGFR:2377
affibody molecule labeled with 99mTc using peptide-based
cysteine-containing chelator.
Materials and methods
99mTc-pertechnetate was obtained by
elution of a generator (Mallinckrodt-Tyco, Petten, The Netherlands)
with sterile 0.9% sodium chloride. Radioactivity in cell and
biodistribution studies was measured by an automated
gamma-spectrometer (1480 Wizard; Wallac Oy, Turku, Finland). The
Cyclone Storage Phosphor system (Perkin-Elmer, Wellesley, MA, USA)
was used for quantitative measurement of radioactivity distribution
in instant thin-layer chromatography strips and electrophoresis
gels.
An unpaired t-test was used to determine if the
difference between the measured values in the biological
experiments was significant (P<0.05).
Preparation of
99mTc-ZEGFR:2377
The EGFR specific affibody molecule ZEGFR:2377 was
produced as previously described (45). Briefly, the affibody molecule was
recombinantly produced overnight at 25°C in E. coli BL21
Star (DE3) (Thermo Fisher Scientific, Inc., Stockholm, Sweden) by
induction with 100 μM IPTG at an OD600 of 0.6. The lysed cells were
heat-shocked at 90°C for 10 min and the affibody molecule was
recovered using a Q-Sepharose column (GE-Healthcare, Uppsala,
Sweden). A polishing step was done using RP-HPLC (Zorbax 300SB-C18;
Agilent Technologies, Inc., Palo Alto, CA, USA). The purity was
measured using RP-HPLC using a gradient from 20–60 for 25 min at a
flow rate of 1 ml/min (A: 0.1% trifluoroacetic acid in water; B:
0.1% trifluoroacetic acid in acetonitrile). Determination of mass
was done using electrospray on-line mass spectrometry (Agilent
Technologies).
Before labeling, the ZEGFR:2377 affibody molecule
was treated with dithiothreitol (DTT; Merck, Darmstadt, Germany) to
reduce the disulphide bonds formed by cysteines. For this purpose,
a solution of DTT (3.5 μl, 1 M in degassed 0.2 M sodium phosphate
buffer, pH 8.0) was mixed with affibody molecules (100 μl, 5 mg/ml
in 0.2 M sodium phosphate buffer, pH 8.0) to obtain a final DTT
concentration 30 mM. The mixture was incubated at 40°C for 90 min
under argon atmosphere. Purification of reduced affibody molecules
was performed using NAP-5 column equilibrated and eluted with PBS.
The solution of the reduced affibody molecules was divided in
aliquots, 100 μg in 180 μl PBS each and stored at −80°C.
A freeze-dried labeling kit containing 75 mg of tin
(II) chloride dihydrate (Fluka Chemika, Buchs, Switzerland), 5 mg
of gluconic acid sodium salt (Celsus Laboratories, Geel, Belgium)
and 100 μg of EDTA (Sigma-Aldrich, Munich, Germany) was prepared
for labeling of affibody molecules with 99mTc as
describe earlier (44).
For a typical labeling, 50 μg of the reduced
ZEGFR:2377 in PBS (90 μl), was mixed with one freeze-dried kit and
100 μl of 99mTc-pertechnetate generator eluate
(typically, 500–800 MBq) was added. The mixture was incubated at
90°C for 60 min. To remove loosly affibody-bound 99mTc,
a treatment with excess of cysteine was used. A fresh solution of
cysteine (1 mg/ml in PBS, 300-fold molar excess to affibody
molecules, was added to the labeled conjugate and incubated at 90°C
for 15 min. After challenge, 99mTc-ZEGFR:2377 was
isolated using size-exclusion chromatography on disposable NAP-5
columns, pre-equilibrated and eluted with PBS.
Labeling yield and purity of affibody molecules were
analyzed using ITLC-SG (Agilent Technologies) developed with PBS
(affibody: Rf=0.0; other forms of 99mTc: Rf=1.0). The
reduced hydrolyzed technetium colloid (RHT) level in the product
was measured using pyridine:acetic acid:water (5:3:1.5) as the
mobile phase (99mTc colloid: Rf=0.0, other forms of
99mTc and radiolabeled affibody molecule: Rf=1.0). The
results of the ITLC measurement were validated by a sodium dodecyl
sulfate polyacrylamide gel electrophoresis (SDS-PAGE). This was
performed using NuPAGE 4–12% Bis-Tris Gel in MES buffer (both from
Invitrogen AB, Stockholm, Sweden) at 200 V constant during 30
min.
In vitro stability was evaluated using
cysteine challenge (45). Samples
of 99mTc-ZEGFR:2377 (5.6 μg, 50 μl) after purification
were mixed with cysteine (5.2 μg, 1 mg/ml in PBS) to obtain a
300-fold molar excess of cysteine. Control samples were mixed with
equal volume of PBS. In addition, a separate set of affibody
samples was mixed with sodium ascorbate (13 μg, 1 mg/ml in PBS).
The samples were incubated at 37°C. At 1, 2 and 4 h after mixing,
the radiochemical purity was determined by radio-ITLC as describe
above. The experiment was performed in triplicate.
In vitro studies
The three different EGFR-expressing cell lines A431,
MDA468 and PC3 (ATCC; purchased via LGC Promochem, Borås, Sweden),
were used in the present study. The cell lines were cultured in
McCoy's medium, supplemented with 10% fetal bovine serum
(Sigma-Aldrich), 1% L-glutamine, and PEST (penicillin 100 U/ml and
100 μg/ml streptomycin), all from Bookroom AG (Berlin, Germany).
The cells were cultured at 37°C in a humidified incubator with 5%
CO2.
Binding specificity of the labeled affibody
molecules was tested using cell lines with three different levels
of EGFR expression (A431, MDA468 and PC3). Cells were seeded to
three sets of dishes, ~106 cells/dish, and incubated for
1 h at 37°C with 10 nM 99mTc-ZEGFR:2377. To two sets of
control dishes, a 50-fold molar excess of either non-labeled
ZEGFR:2377 or cetuximab was added before adding
99mTc-ZEGFR:2377. After incubation, the medium was
aspired; the cells were washed with medium, detached using trypsin
and collected. Radioactivity of cells was measured, and percentage
of cell-bound radioactivity was calculated. The experiments were
performed in triplicate.
To measure the affinity of the conjugate to EGFR,
kinetics of binding of 99mTc-ZEGFR:2377 to and its
dissociation from A431 cells were measured using a LigandTracer
Yellow instrument (Ridgeview Instruments AB, Vänge, Sweden). The
measurements were performed at room temperature to prevent
internalization. Uptake curves were recorded at 0.33, 1 and 3 nM of
99mTc-ZEGFR:2377, thereafter the radioactive medium was
withdrawn, fresh non-radioactive medium was added and the
dissociation curve was recorded. The data were analyzed using the
Interaction Map software (Ridgeview Diagnostics AB, Uppsala,
Sweden) to calculate association rate, dissociation rate and
dissociation constant at equilibrium (KD).
Cellular processing of bound
99mTc-ZEGFR:2377 was evaluated using MDA486 and A432
cells. The cells were incubated with 10 nM
99mTc-ZEGFR:2377 at 37°C. At 1, 2, 4, 6 and 24 h after
incubation start, the internalized fraction was determined by an
acid wash method adapted and validated for affibody molecules by
Wållberg and Orlova (46). The
membrane-bound affibody molecules were removed from cells by
treatment with 4 M urea solution in a 0.1 M glycine buffer, pH 2.5,
for 5 min on ice. The cell debris containing the internalized
conjugates was detached by treatment with 1 M NaOH. Radioactivity
of cells was measured, and percentage of membrane-bound and
internalized radioactivity was calculated. The experiments were
performed in triplicate.
In vivo studies
All applicable international, Swedish national and
Uppsala University guidelines for the care and the use of the
animals were followed. All procedures performed in studies
involving animals were in accordance with the ethical standards of
the Uppsala University. The studies were approved by the Ethics
Committee for Animal Research in Uppsala. Female BALB/C nu/nu mice
were purchased from Taconic M&B a/S (Ry, Denmark).
EGFR-expressing xenografts were established by subcutaneous
injection of 107 A431 cells in the right hind legs of
mice. The experiments were performed 12–14 days after tumor
implantation. The average animal weight at the time of experiment
was 19±2 g. The average tumor weight was 550±240 mg.
For biodistribution measurements, mice were randomly
divided into groups of four animals each. To investigate the
specificity of conjugate in vivo, EGF receptors were
saturated in a control group of mice by subcutaneous injection of
2.5 mg of cetuximab 48 and 24 h before labeled conjugate injection.
Two groups of mice (including control group) were intravenously
injected with 99mTc-ZEGFR:2377 (60 kBq in 100 μl PBS per
mouse) for biodistribution measurement at 3 h after injection. One
group of mice was injected with 480 kBq for measurement of
biodistribution at 24 h after injection. The injected protein dose
was adjusted to 38 μg per mouse. This protein dose has been found
to be optimal for imaging using 111In-DOTA-ZEGFR:2377 in
earlier studies (35).
The mice were euthanized at 3 and 24 h pi by
overdosing of anesthesia. Blood, salivary glands, thyroid, lung,
liver, spleen, colon, kidney, tumor, muscle and bone were collected
in weighed plastic bottles. The radioactivity was measured and
uptake was calculated as percent injected dose per gram tissue
(%ID/g).
microSPECT/CT imaging of EGFR-expressing
xenografts using 99mTc-ZEGFR:2377
BALB/C nu/nu mice bearing subcutaneous A431
xenografts were intravenously injected with 38 μg (30 MBq)
99mTc-ZEGFR:2377. Whole body scans were acquired using
nanoScan SC (Mediso Medical Imaging Systems, Budapest, Hungary) at
3 and 24 h after injections. The mice were euthanized by
CO2 asphyxiation immediately before being placed on
camera. The computed tomography (CT) acquisition was carried out at
the following parameters: energy peak of 50 kV, 670 μA, 480
projections, 2.29-min acquisition time. SPECT acquisition was
performed at the following parameters: 99mTc energy peak
of 140 keV, window width of 20%, matrix of 256×256, acquisition
time: 1 h. CT images were reconstructed in real-time using Nucline
2.03 Software (Mediso Medical Imaging Systems). SPECT raw data were
reconstructed using Tera-Tomo™ 3D SPECT reconstruction technology.
Coronal SPECT-CT images of the scans are presented as maximum
intensity projections (MIP) in RGB color scale.
Results
Preparation of
99mTc-ZEGFR:2377
The anti-EGFR binding ZEGFR:2377 affibody molecule
was produced and subsequently purified using heat-treatment of
cells, anion exchange for product recovery and a reverse phase-HPLC
polishing step. The purity of the protein was determined to 97.2%
and the mass was resolved using mass-spectrometry to 14,781 Da
(theoretical mass of dimer 14,782 Da and monomer 7,391 Da).
Initially, the labeling was performed according to
protocol of Ahlgren and co-workers (44), i.e. without intermediate challenge
before purification. The radiochemical yield was 99.6% and, after
purification using NAP-5 column, radiochemical purity was 100%.
However, the results of the in vitro stability test
(Table I) demonstrated rapid
release of 99mTc not only under cysteine challenge but
also during storage in PBS.
 | Table IIn vitro stability of
99mTc-ZEGFR:2377. |
Table I
In vitro stability of
99mTc-ZEGFR:2377.
| Affibody-bound
radioactivity, % |
|---|
|
|---|
| No cysteine
challenge before purification | Cysteine challenge
before purification |
|---|
|
|
|---|
| PBS | 300-fold cysteine
excess | PBS | 300-fold cysteine
excess | Sodium
ascorbate |
|---|
| 1 h | 60.0±0.4 | 66±2 | 96±3 | 98±1 | 99.6±0.1 |
| 2 h | 60.7±0.4 | 65.3±0.1 | 95±4 | 96±2 | 99.4±0.2 |
| 4 h | 62.6±0.8 | 63±4 | 92±3 | 95.3±0.7 | 99±1 |
There was a possibility that part of
99mTc was bound not to a cysteine-containing chelator,
but to a much weaker chelating site. Therefore, we have introduced
a cysteine challenge between labeling and chromatographic
purification. The isolated yield of such procedure was 69.2±1.2%,
and the purity of the conjugate after NAP-5 purification was
100±0%. The presence of RHT was 0.2±0.2% in all experiments. The
maximum specific activity of 9 MBq/μg (66.5 GBq/μmol) was
obtained.
The release of 99mTc under in
vitro stability test was appreciably reduced, and >95% of
radioactivity was associated with the affibody molecule after the
4-h cysteine challenge. Notably, there was higher release in the
control than in cysteine challenge group. This indicated that a
re-oxidation of 99mTc might play a role in its release.
To test the hypothesis, we performed the stability test using
sodium ascorbate as an antioxidant. The release of 99mTc
has been appreciable reduced in this case (Table I). Sodium ascorbate has hence been
included in formulation of 99mTc-ZEGFR:2377 for animal
studies.
In vitro studies
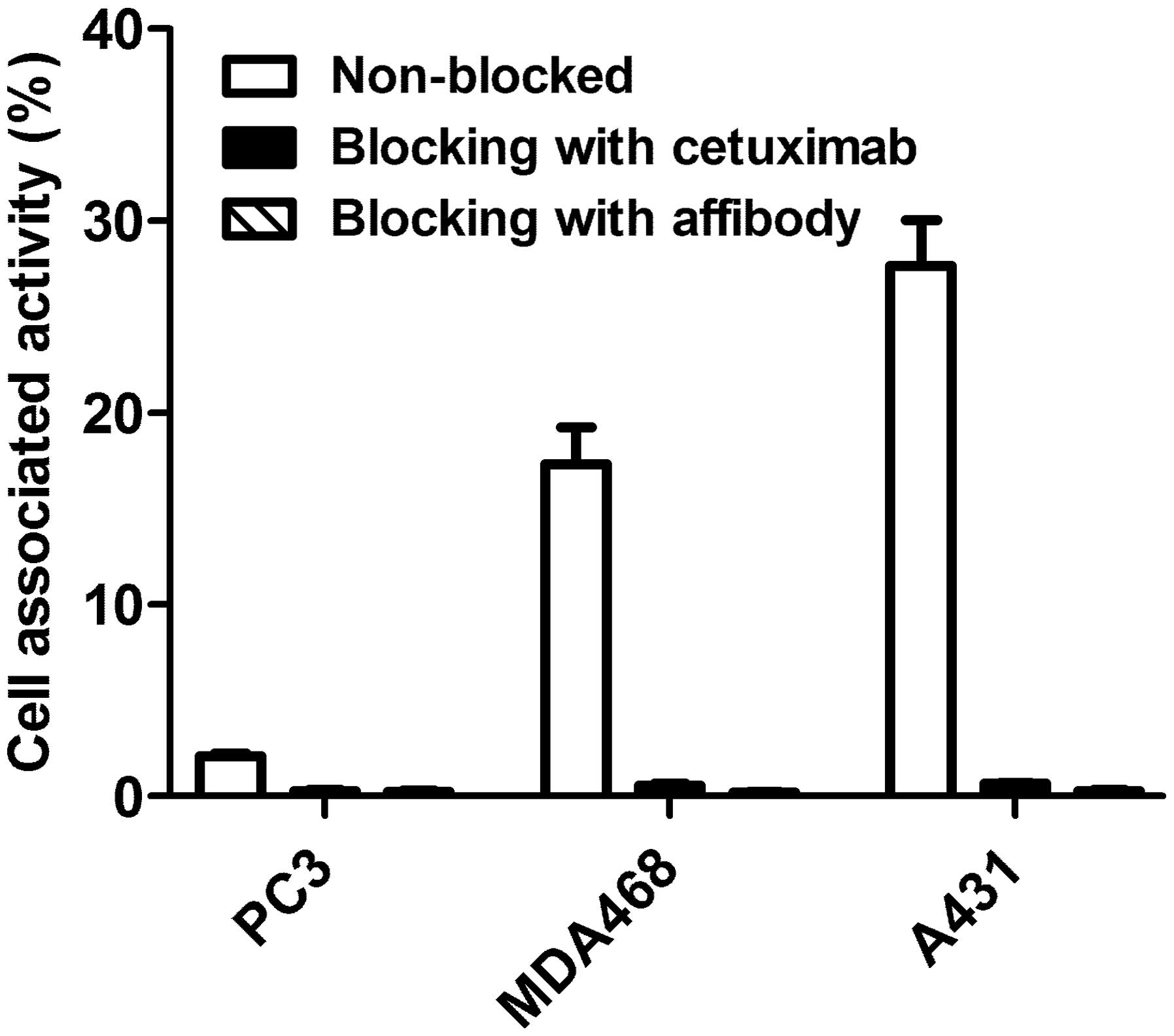
The results of the in vitro specificity test
of 99mTc-ZEGFR:2377 are presented in Fig. 2. Pre-saturation of EGFR with a
large excess of both non-labeled ZEGFR:2377 and anti-EGFR antibody
cetuximab reduced significantly (P<0.0001) binding of
99mTc-ZEGFR:2377 to EGFR-expressing cells. The saturable
character of the binding suggests its specificity.
The LigandTracer measurements of the
99mTc-ZEGFR:2377 interaction with A431 cells showed a
rapid binding (association rate of 2.77×105 1/M × sec)
and slow dissociation (dissociation rate of 7.73×10−5
1/sec). The dissociation constant (KD) of
99mTc-ZEGFR:2377 interaction with A431 cells was 274
pM.
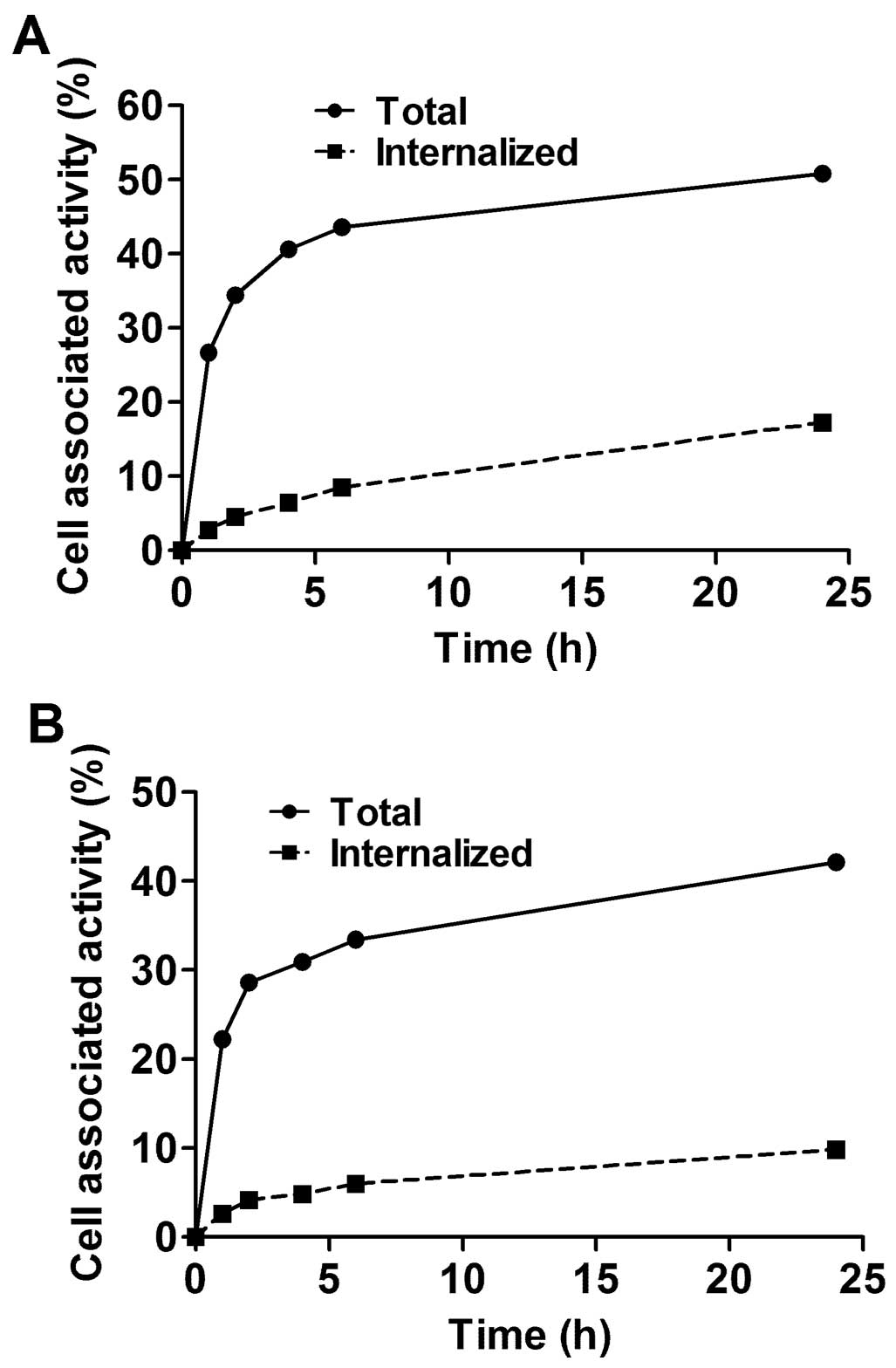
The processing of bound 99mTc-ZEGFR:2377
by MDA468 and A431 cancer cell lines is presented in Fig. 3. The common feature for both cell
lines was rapid binding and relatively slow internalization of
99mTc-ZEGFR:2377. Although the pattern of
internalization was similar, the internalization by A431 cells was
slower than by MDA468. The internalized fractions after 24-h
incubation were 23.3±0.3 and 34.9±0.3% of total cell-bound
radioactivity, for A431 and MD468, respectively.
In vivo studies
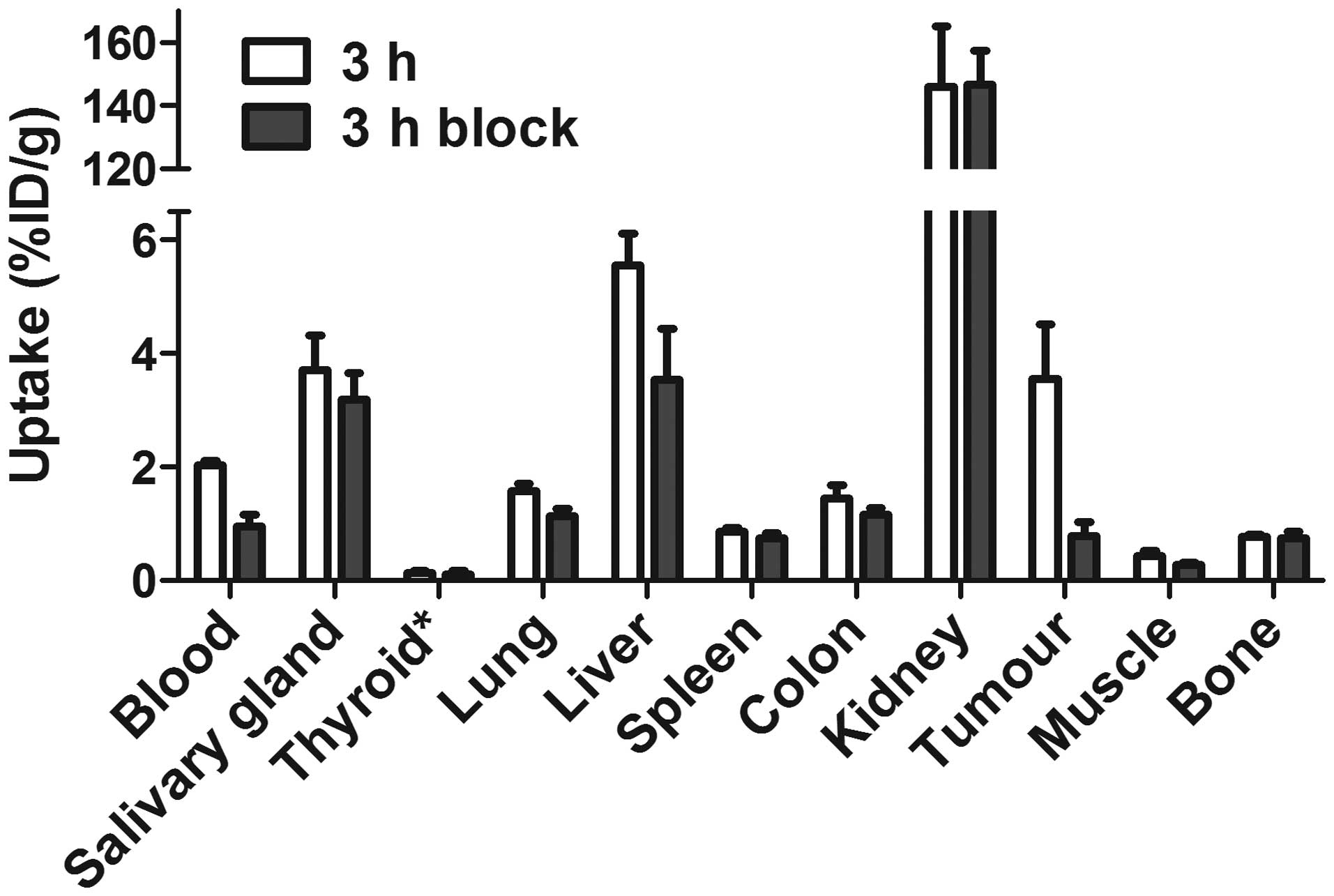
The specificity of 99mTc-ZEGFR:2377
binding to EGFR in vivo was demonstrated by in vivo
saturation of receptors by injection of the monoclonal antibody
cetuximab (Fig. 4). The tumor
uptake in the control group was reduced 4.6-fold (P<0.005).
Significant (P<0.05) reduction of radioactivity uptake in blood,
liver, lung and muscle was also observed.
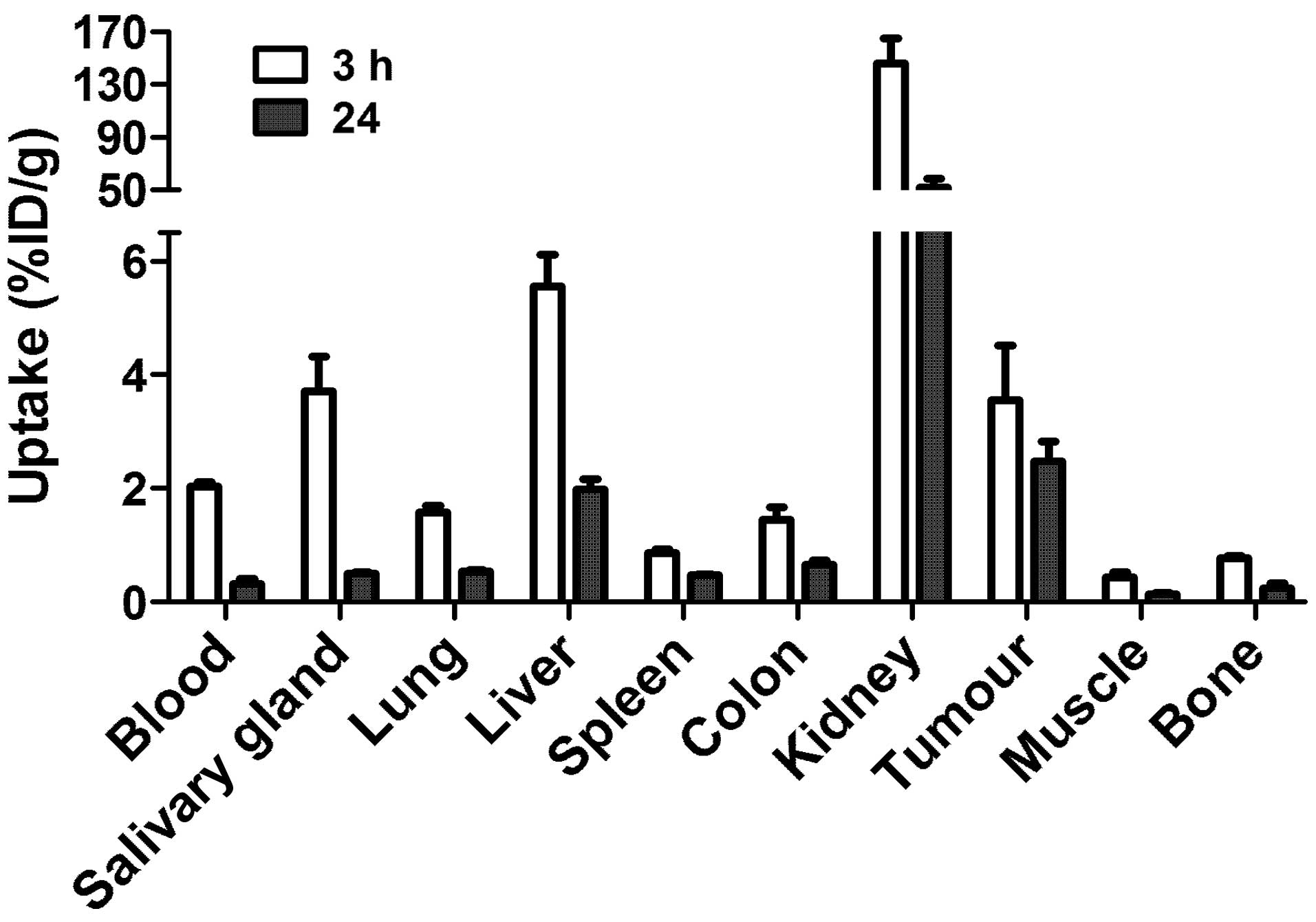
Fig. 5 shows the
biodistribution of the 99mTc-ZEGFR:2377 conjugate at 3
and 24 h after injection in BALB/C nu/nu mice bearing
EGFR-expressing A431 xenografts. The blood concentration was
2.03±0.08% ID/g at 3 h after injection, which suggests rather rapid
blood clearance. The high uptake in kidneys (146±19% ID/g)
indicates that the predominant excretion was via glomerular
filtration followed by re-absorption which is typical for affibody
molecules. Liver was the other organ with prominent uptake
(5.6±0.6% ID/g), but excretion via the bile was minor. The
gastrointestinal tracts with content contained only 4.0±0.5% of
injected radioactivity. The tumor uptake (3.6±1% ID/g) exceeded the
uptake in other organs and tissues except from liver, salivary
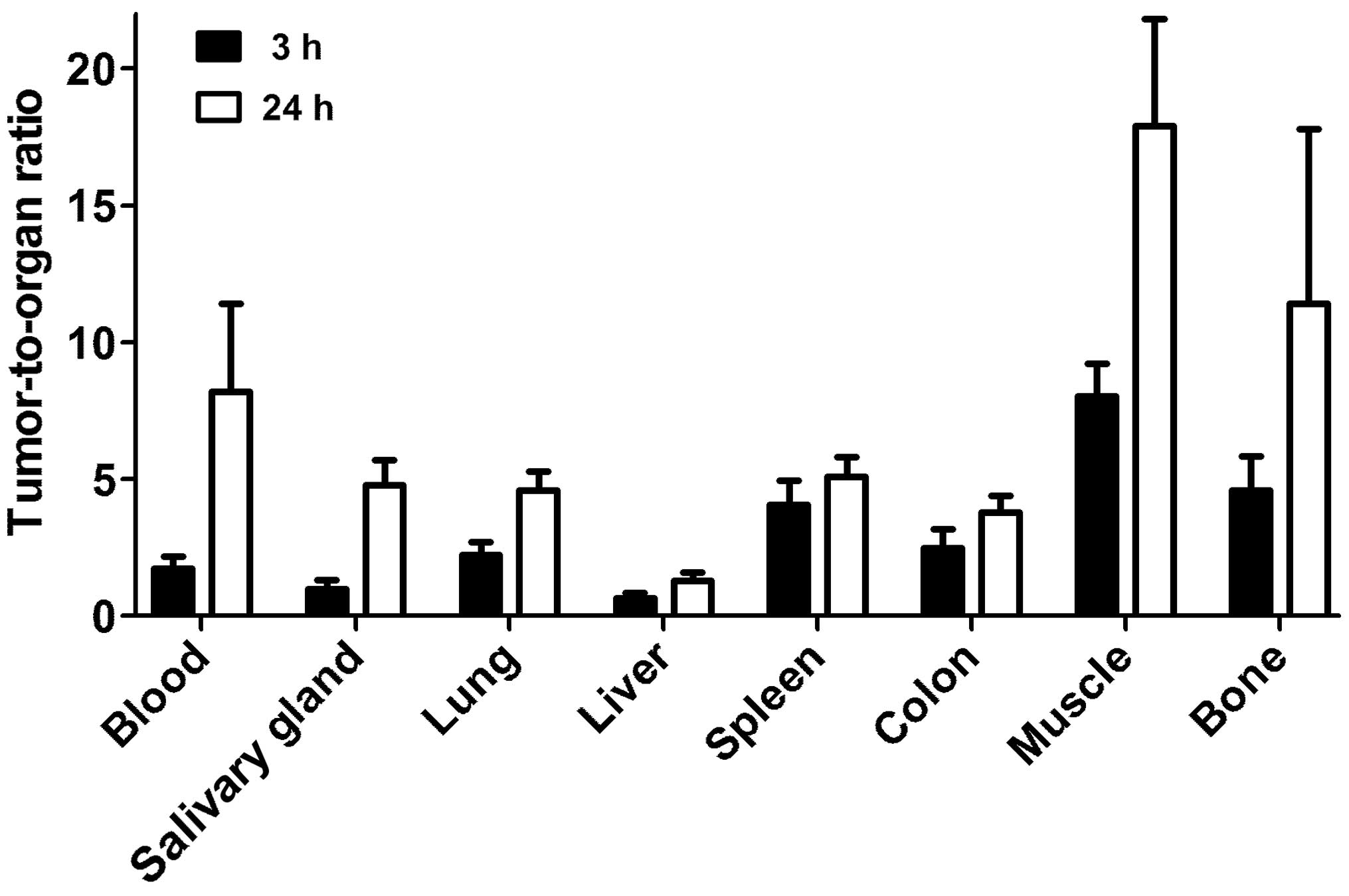
glands and kidneys. However, the tumor-to-organ ratios (Fig. 6) were modest at this time-point.
The tumor uptake had a tendency to decrease at 24 h, but the
difference between tumor uptake values at 3 and 24 h was not
significant (P=0.12). There was a highly significant (P<0.01)
decrease of uptake in all other organs and tissues. This resulted
in an appreciable increase of tumor-to-organ ratios at 24 h after
injection (Fig. 6). This suggests
that the preferable time-point for imaging would be at 24 h after
injection.
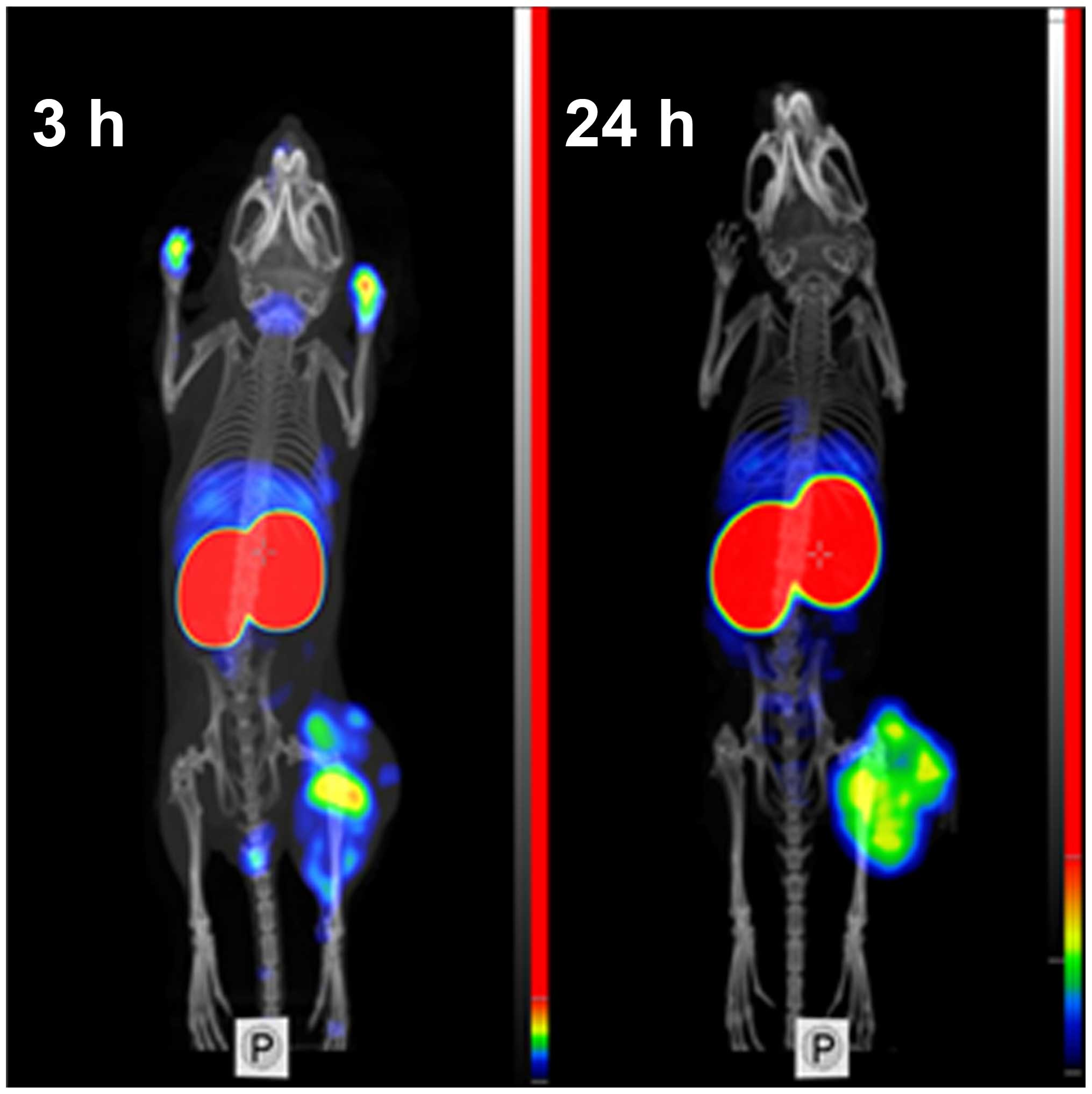
Feasibility of imaging of EGFR-expressing tumors
using 99mTc-ZEGFR:2377 was demonstrated using
microSPECT/CT (Fig. 7). Tumors on
hind legs were clearly visualized. The irregular pattern of uptake
reflected necrotic areas in the tumors. Radioactivity uptake in
kidneys was considerably higher than in tumors. Other places with
high radioactivity accumulation were liver and salivary gland (at 3
h after injection). At 3 h after injection, urinary bladder and,
occasionally, urine contamination on front legs were also
visualized. There was a reasonable contrast between tumors and
other organs and tissues.
Discussion
Clinical studies have demonstrated that affibody
molecules provide excellent sensitivity and specificity in imaging
of HER2-expressing tumors using SPECT (47) and PET (33). Moreover, preclinical studies
demonstrated feasibility of application of affibody molecules for
imaging several molecular targets in cancer xenografts, e.g. EGFR
(33), HER3 (48) IGF-1R (49), PDGFRβ (50) and CAIX (51). It would be attractive to apply our
experience in development of HER2-imaging affibody molecule probes
for development of probes to new targets. However, there are
multiple amino-acid-alterations in the positions forming the target
binding sites of new probes, which might influence both labeling
chemistry and biodistribution pattern. The present study evaluated
preconditions to labeling of affibody molecule-derived imaging
probes using peptide-based cysteine-containing chelators. Chelators
of that type provided a stable labeling of anti-HER2 affibody
molecules with 99mTc and 188Re (42,52).
An apparent advantage of such chelators is that they are ‘built in’
in the recombinantly produced affibody molecule, and there is no
need in additional steps of coupling of the chelator and
purification of the conjugate. In addition, selection of amino
acids in the chelator may enable a fine-tuning of residualizing
properties of the label (52–54).
However, there was a risk of losing site-specificity of the label
when amino acid composition of an affibody molecules was modified
(43). Indeed, the first
experiments demonstrated release of over 30% of the
99mTc after dilution or under cysteine challenge
(Table I). This might be an
indication that some amino acids in ZEGFR:2377 form a chelating
site, which competes with the cysteine-containing chelator, but has
appreciably weaker complexing of the nuclide. To minimize the
influence of such site, we introduced a cysteine challenge of the
labeled conjugate before the final purification. We assumed that
weakly bound nuclide would be stripped from the conjugate. The
overall isolated yield has decreased to 69±1% in this case, but the
conjugate could withstand the cysteine challenge (Table I). Adding an anti-oxidizing agent,
sodium ascorbate, reduced the releases further, which indicates
that re-oxidation of technetium is another, although minor, reason
of release.
The 99mTc-ZEGFR:2377 retained capacity of
specific binding to EGFR-expressing cells as it has been
demonstrated by saturation of binding sites with both non-labeled
ZEGFR:2377 and anti-EGFR antibody cetuximab (Fig. 2). Affinity of
99mTc-ZEGFR:2377 was 274±16 pM, in the same picomolar
range as affinity of 89Zr-labeled DFO-ZEGFR:2377 (160±60
pM) (38). The internalization by
two EGFR-expressing cell lines was relatively slow (Fig. 3), which is typical for EGFR-binding
affibody molecules (35,38). Blocking of EGFR receptors in A431
xenografts with the antibody cetuximab reduced significantly
(P<0.005) uptake of 99mTc-ZEGFR:2377, which
unambiguously demonstrated specificity of in vivo targeting
(Fig. 4).
Recently, several approaches for radiofluorination
of anti-EGFR affibody molecules have been reported (36,37).
In similar tumor models (A431 xenografts), these tracers
demonstrated tumor-to-blood ratios in the range from 1.5 to 2.4 at
3 h after injection. These values are similar to values provided by
99mTc-ZEGFR:2377 at this time-point (1.8±0.4). For
comparison, the anti-HER2 99mTc-ZHER2:2395 affibody
molecule provided the tumor-to-blood ratio of 9±2 already 1 h after
injection (42). The moderate
values of tumor-to-blood ratios at early time-points (a few hours
after injection) for anti-EGFR affibody molecules might be
determined by two factors. First, there is an expression of EGFR in
normal tissues, particularly in liver, and these tissues sequester
part of the radiolabeled affibody molecules. Second, the affibody
molecules are internalized quite slowly after binding to EGFR, i.e.
remain reversibly bound to a cell surface. Thus, EGFR expressing
tissues act as depots. Decrease of blood concentration of affibody
molecules due to renal clearance shifts the equilibrium towards
dissociation, and affibody molecules start to return to blood
stream. Due to slower blood clearance, a good contrast might be
obtained only several hours or at the next day after injection
(35). This suggests that
short-lived radionuclides, such as 18F
(T1/2=109.8 min) or 68Ga
(T1/2=67.6 min) are suboptimal for labeling of anti-EGFR
affibody molecules. The longer half-life of 99mTc, 6.01
h, permits clinical imaging at several hours or even at the next
day after injection (55,56). Due to absence of corpuscular
radiation, more than 1100 MBq of 99mTc-labeled compounds
might be injected resulting in low dose burden to the patient
(57). This study demonstrated
that tumor-to-organ ratios for 99mTc-ZEGFR:2377
increased appreciably at 24 h after injection compared to 3 h after
injection. For example, the tumor-to-blood ratio increased
4.7-fold, and tumor-to-liver, tumor-to-lung, tumor-to-muscle and
tumor-to-bone ratios increased 2–2.5-fold. This determines better
contrast of imaging at later time-points and should provide better
sensitivity of clinical imaging.
It has to be noted that the physical half-life of
the label nuclide is only one of several factors determining
contrast and feasibility of imaging. Chemical properties of a
radionuclide and linker or chelators for its attachment to a
tumor-targeting protein might influence the affinity, the cellular
processing and retention of a radionuclide in tumor and normal
tissues, its off-target interactions, and predominant excretion
routes of a tracer and radiocatabolites (39). This might be exemplified by
comparison of the biodistribution of ZEGFR:2377 labeled with
99mTc and with a long-lived (T1/2=78.4 h)
positron emitting radionuclide 89Zr. We have recently
studied ZEGFR:2377 labeled with 89Zr using deferoxamine
(DFO) chelator (38). Both studies
used the same experimental methods and the same animal models,
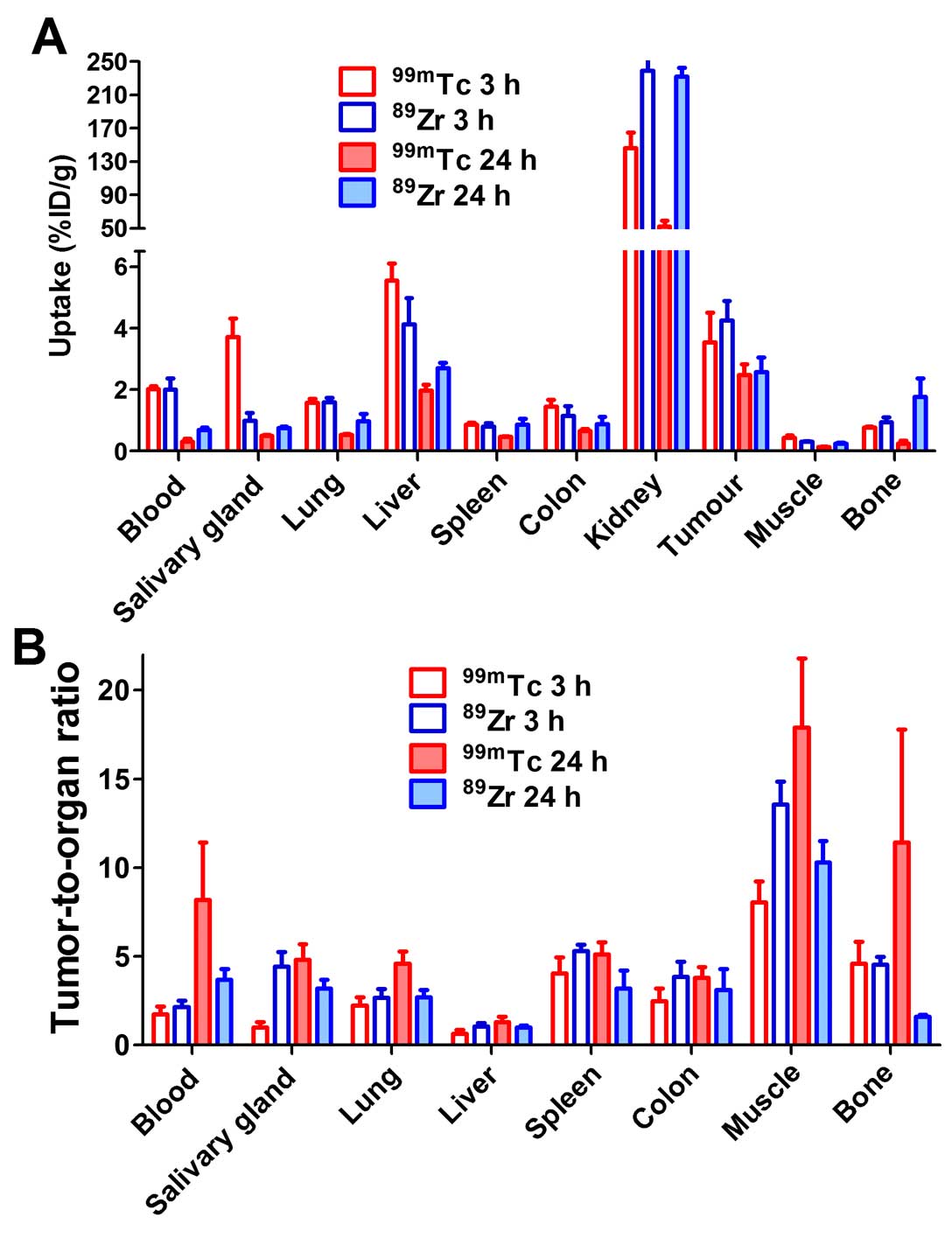
which facilitates comparison. The tumor uptake of both probes was
similar at 3 and 24 h after injection (Fig. 8). At 3 h after injection, the
uptake in normal organs and tissues was also quite similar; with
exception that uptake of 99mTc is appreciably higher in
salivary gland but lower in kidneys. At 24 h after injection,
uptake of 99mTc is lower in all normal organ and
tissues, which results in higher tumor-to-organ ratios compared to
89Zr-ZEGFR:2377 (Fig.
8). Insight into why the differences occur might be gained by
considering kidney retention. Affibody molecules are re-absorbed in
proximal tubuli of kidneys and are rapidly internalized. Thus, the
renal retention of a radionuclide depends on its residualizing
properties. The renal uptake of 89Zr is equal at 3 and
24 h, which suggests that this nuclide has strong residualizing
properties. The renal uptake of 99mTc decreased ~3-fold
at 24 h. This indicates that the residualizing properties of
99mTc label are moderate, and radiometabolites ‘leak’
from cells after internalization. It is likely that this is the
reason why 99mTc clears more rapidly from other normal
tissues.
99mTc-ZEGFR:2377 enabled good-contrast
imaging of EGFR-expressing xenografts both at 3 and 24 h after
injection (Fig. 7). The above
suggests that the use of peptide-based cysteine-containing chelator
at the C-terminus of ZEGFR:2377 affibody molecules for labeling
with 99mTc is an adequate approach, providing a
conjugate capable of visualization of EGFR expression in tumors. It
would be attractive, of course, to find a labelling approach
permitting a single-vial kit formulation, as for HER2-targeting
affibody molecule (44). For
ZEGFR:2377, a two-vial kit is possible, and a purification of the
labeled is required. This complicates somewhat the labelling
procedure. An evaluation of alternative approaches for
99mTc-labeling might be required before decision for
clinical translation could be made.
In conclusion, the use of a peptide-based
cysteine-containing chelator at the C-terminus provides labeling of
the ZEGFR:2377 affibody molecule with 99mTc. However, an
intermediate cysteine challenge is required to ensure that the
nuclide is not complexed by a weak chelating site. Translation of
previously gained knowledge to new affibody molecules is possible,
but requires re-optimization of labeling chemistry since changes in
amino acid composition might influence the site-specificity of
labeling as well as the stability of the label.
Acknowledgements
The present study was supported by grants from the
Swedish Research Council (Vetenskapsrådet) and the Swedish Cancer
Society (Cancerfonden). The molecular imaging work in this
publication has been supported by the Wallenberg infrastructure for
PET-MRI (WIPPET), a Swedish nationally available imaging platform
at Uppsala University, Sweden.
Abbreviations:
|
EGFR
|
epidermal growth factor receptor
|
|
RTK
|
receptor tyrosine kinase
|
|
HNSCC
|
head and neck squamous cell
carcinoma
|
|
NSCLC
|
non-small cell lung cancer
|
|
TKI
|
tyrosine kinase inhibitor
|
|
SPECT
|
single photon emission computed
tomography
|
|
PBS
|
phosphate-buffered saline
|
|
DTT
|
dithiothreitol
|
|
SDS-PAGE
|
sodium dodecyl sulfate polyacrylamide
gel electrophoresis
|
|
HPLC
|
high performance liquid
chromatography
|
|
RHT
|
reduced hydrolyzed technetium
|
|
KD
|
equilibrium dissociation constant
|
|
PET
|
positron emission tomography
|
References
|
1
|
Kulkarni HR and Baum RP: Patient selection
for personalized peptide receptor radionuclide therapy using Ga-68
somatostatin receptor PET/CT. PET Clin. 9:83–90. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Slobbe P, Poot AJ, Windhorst AD and van
Dongen GA: PET imaging with small-molecule tyrosine kinase
inhibitors: TKI-PET. Drug Discov Today. 17:1175–1187. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Altai M, Orlova A and Tolmachev V:
Radiolabeled probes targeting tyrosine-kinase receptors for
personalized medicine. Curr Pharm Des. 20:2275–2292. 2014.
View Article : Google Scholar
|
|
4
|
Krause DS and Van Etten RA: Tyrosine
kinases as targets for cancer therapy. N Engl J Med. 353:172–187.
2005. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Salomon DS, Brandt R, Ciardiello F and
Normanno N: Epidermal growth factor-related peptides and their
receptors in human malignancies. Crit Rev Oncol Hematol.
19:183–232. 1995. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Hedner C, Borg D, Nodin B, Karnevi E,
Jirström K and Eberhard J: Expression and prognostic significance
of human epidermal growth factor receptors 1 and 3 in gastric and
esophageal adenocarcinoma. PLoS One. 11:e01481012016. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Ueda S, Ogata S, Tsuda H, Kawarabayashi N,
Kimura M, Sugiura Y, Tamai S, Matsubara O, Hatsuse K and Mochizuki
H: The correlation between cytoplasmic overexpression of epidermal
growth factor receptor and tumor aggressiveness: Poor prognosis in
patients with pancreatic ductal adenocarcinoma. Pancreas. 29:e1–e8.
2004. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Growdon WB, Boisvert SL, Akhavanfard S,
Oliva E, Dias-Santagata DC, Kojiro S, Horowitz NS, Iafrate AJ,
Borger DR and Rueda BR: Decreased survival in EGFR gene amplified
vulvar carcinoma. Gynecol Oncol. 111:289–297. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Young RJ, Rischin D, Fisher R, McArthur
GA, Fox SB, Peters LJ, Corry J, Lim A, Waldeck K and Solomon B:
Relationship between epidermal growth factor receptor status,
p16INK4A, and outcome in head and neck squamous cell
carcinoma. Cancer Epidemiol Biomarkers Prev. 20:1230–1237. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Kros JM, Huizer K, Hernández-Laín A,
Marucci G, Michotte A, Pollo B, Rushing EJ, Ribalta T, French P,
Jaminé D, et al: Evidence-based diagnostic algorithm for glioma:
Analysis of the results of Pathology Panel Review and Molecular
Parameters of EORTC 26951 and 26882 Trials. J Clin Oncol.
33:1943–1950. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Roskoski R Jr: The ErbB/HER family of
protein-tyrosine kinases and cancer. Pharmacol Res. 79:34–74. 2014.
View Article : Google Scholar
|
|
12
|
Lynch TJ, Bell DW, Sordella R,
Gurubhagavatula S, Okimoto RA, Brannigan BW, Harris PL, Haserlat
SM, Supko JG, Haluska FG, et al: Activating mutations in the
epidermal growth factor receptor underlying responsiveness of
non-small-cell lung cancer to gefitinib. N Engl J Med.
350:2129–2139. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Paez JG, Jänne PA, Lee JC, Tracy S,
Greulich H, Gabriel S, Herman P, Kaye FJ, Lindeman N, Boggon TJ, et
al: EGFR mutations in lung cancer: Correlation with clinical
response to gefitinib therapy. Science. 304:1497–1500. 2004.
View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Therkildsen C, Bergmann TK,
Henrichsen-Schnack T, Ladelund S and Nilbert M: The predictive
value of KRAS, NRAS, BRAF, PIK3CA and PTEN for anti-EGFR treatment
in metastatic colorectal cancer: A systematic review and
meta-analysis. Acta Oncol. 53:852–864. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Pirker R, Pereira JR, von Pawel J,
Krzakowski M, Ramlau R, Park K, de Marinis F, Eberhardt WE,
Paz-Ares L, Störkel S, et al: EGFR expression as a predictor of
survival for first-line chemotherapy plus cetuximab in patients
with advanced non-small-cell lung cancer: Analysis of data from the
phase 3 FLEX study. Lancet Oncol. 13:33–42. 2012. View Article : Google Scholar
|
|
16
|
Cappuzzo F, Hirsch FR, Rossi E, Bartolini
S, Ceresoli GL, Bemis L, Haney J, Witta S, Danenberg K, Domenichini
I, et al: Epidermal growth factor receptor gene and protein and
gefitinib sensitivity in non-small-cell lung cancer. J Natl Cancer
Inst. 97:643–655. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Hirsch FR, Varella-Garcia M, Bunn PA Jr,
Franklin WA, Dziadziuszko R, Thatcher N, Chang A, Parikh P, Pereira
JR, Ciuleanu T, et al: Molecular predictors of outcome with
gefitinib in a phase III placebo-controlled study in advanced
non-small-cell lung cancer. J Clin Oncol. 24:5034–5042. 2006.
View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Bradley JD, Paulus R, Komaki R, Masters G,
Blumenschein G, Schild S, Bogart J, Hu C, Forster K, Magliocco A,
et al: Standard-dose versus high-dose conformal radiotherapy with
concurrent and consolidation carboplatin plus paclitaxel with or
without cetuximab for patients with stage IIIA or IIIB
non-small-cell lung cancer (RTOG 0617): A randomised, two-by-two
factorial phase 3 study. Lancet Oncol. 16:187–199. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Ang KK, Berkey BA, Tu X, Zhang HZ, Katz R,
Hammond EH, Fu KK and Milas L: Impact of epidermal growth factor
receptor expression on survival and pattern of relapse in patients
with advanced head and neck carcinoma. Cancer Res. 62:7350–7356.
2002.PubMed/NCBI
|
|
20
|
Bentzen SM, Atasoy BM, Daley FM, Dische S,
Richman PI, Saunders MI, Trott KR and Wilson GD: Epidermal growth
factor receptor expression in pretreatment biopsies from head and
neck squamous cell carcinoma as a predictive factor for a benefit
from accelerated radiation therapy in a randomized controlled
trial. J Clin Oncol. 23:5560–5567. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Eriksen JG, Steiniche T and Overgaard J;
Danish Head and Neck Cancer study group (DAHANCA). The influence of
epidermal growth factor receptor and tumor differentiation on the
response to accelerated radiotherapy of squamous cell carcinomas of
the head and neck in the randomized DAHANCA 6 and 7 study.
Radiother Oncol. 74:93–100. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Wang X, Niu H, Fan Q, Lu P, Ma C, Liu W,
Liu Y, Li W, Hu S, Ling Y, et al: Predictive value of EGFR
overexpression and gene amplification on icotinib efficacy in
patients with advanced esophageal squamous cell carcinoma.
Oncotarget. 7:24744–24751. 2016.PubMed/NCBI
|
|
23
|
Humbert O, Riedinger JM, Charon-Barra C,
Berriolo-Riedinger A, Desmoulins I, Lorgis V, Kanoun S, Coutant C,
Fumoleau P, Cochet A, et al: Identification of biomarkers including
18FDG-PET/CT for early prediction of response to
neoadjuvant chemotherapy in triple negative breast cancer. Clin
Cancer Res. 21:5460–5468. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Vignot S, Besse B, André F, Spano JP and
Soria JC: Discrepancies between primary tumor and metastasis: A
literature review on clinically established biomarkers. Crit Rev
Oncol Hematol. 84:301–313. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Divgi CR, Welt S, Kris M, Real FX, Yeh SD,
Gralla R, Merchant B, Schweighart S, Unger M, Larson SM, et al:
Phase I and imaging trial of indium 111-labeled anti-epidermal
growth factor receptor monoclonal antibody 225 in patients with
squamous cell lung carcinoma. J Natl Cancer Inst. 83:97–104. 1991.
View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Cai W, Chen K, He L, Cao Q, Koong A and
Chen X: Quantitative PET of EGFR expression in xenograft-bearing
mice using 64Cu-labeled cetuximab, a chimeric anti-EGFR
monoclonal antibody. Eur J Nucl Med Mol Imaging. 34:850–858. 2007.
View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Nayak TK, Garmestani K, Milenic DE and
Brechbiel MW: PET and MRI of metastatic peritoneal and pulmonary
colorectal cancer in mice with human epidermal growth factor
receptor 1-targeted 89Zr-labeled panitumumab. J Nucl
Med. 53:113–120. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Chang AJ, De Silva RA and Lapi SE:
Development and characterization of 89Zr-labeled
panitumumab for immuno-positron emission tomographic imaging of the
epidermal growth factor receptor. Mol Imaging. 12:17–27.
2013.PubMed/NCBI
|
|
29
|
van Dijk LK, Hoeben BA, Kaanders JH,
Franssen GM, Boerman OC and Bussink J: Imaging of epidermal growth
factor receptor expression in head and neck cancer with SPECT/CT
and 111In-labeled cetuximab-F(ab′)2. J Nucl
Med. 54:2118–2124. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
van Dijk LK, Yim CB, Franssen GM, Kaanders
JH, Rajander J, Solin O, Grönroos TJ, Boerman OC and Bussink J: PET
of EGFR with 64Cu-cetuximab-F(ab′)2 in mice
with head and neck squamous cell carcinoma xenografts. Contrast
Media Mol Imaging. 11:65–70. 2016. View Article : Google Scholar
|
|
31
|
Nygren PA and Skerra A: Binding proteins
from alternative scaffolds. J Immunol Methods. 290:3–28. 2004.
View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Ahlgren S and Tolmachev V: Radionuclide
molecular imaging using Affibody molecules. Curr Pharm Biotechnol.
11:581–589. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Sörensen J, Velikyan I, Sandberg D,
Wennborg A, Feldwisch J, Tolmachev V, Orlova A, Sandström M,
Lubberink M, Olofsson H, et al: Measuring HER2-receptor expression
in metastatic breast cancer using [68Ga]ABY-025 affibody
PET/CT. Theranostics. 6:262–271. 2016. View Article : Google Scholar
|
|
34
|
Tolmachev V, Friedman M, Sandström M,
Eriksson TL, Rosik D, Hodik M, Ståhl S, Frejd FY and Orlova A:
Affibody molecules for epidermal growth factor receptor targeting
in vivo: Aspects of dimerization and labeling chemistry. J Nucl
Med. 50:274–283. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Tolmachev V, Rosik D, Wållberg H, Sjöberg
A, Sandström M, Hansson M, Wennborg A and Orlova A: Imaging of EGFR
expression in murine xenografts using site-specifically labelled
anti-EGFR 111In-DOTA-ZEGFR:2377 affibody
molecule: aspect of the injected tracer amount. Eur J Nucl Med Mol
Imaging. 37:613–622. 2010. View Article : Google Scholar
|
|
36
|
Miao Z, Ren G, Liu H, Qi S, Wu S and Cheng
Z: PET of EGFR expression with an 18F-labeled affibody
molecule. J Nucl Med. 53:1110–1118. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Su X, Cheng K, Jeon J, Shen B, Venturin
GT, Hu X, Rao J, Chin FT, Wu H and Cheng Z: Comparison of two
site-specifically 18F-labeled affibodies for PET imaging
of EGFR positive tumors. Mol Pharm. 11:3947–3956. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Garousi J, Andersson KG, Mitran B, Pichl
ML, Ståhl S, Orlova A, Löfblom J and Tolmachev V: PET imaging of
epidermal growth factor receptor expression in tumours using
89Zr-labelled ZEGFR:2377 affibody molecules. Int J
Oncol. 48:1325–1332. 2016.PubMed/NCBI
|
|
39
|
Tolmachev V and Orlova A: Influence of
labelling methods on biodistribution and imaging properties of
radiolabelled peptides for visualisation of molecular therapeutic
targets. Curr Med Chem. 17:2636–2655. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Slomka PJ, Pan T, Berman DS and Germano G:
Advances in SPECT and PET Hardware. Prog Cardiovasc Dis.
57:566–578. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Banerjee SR, Maresca KP, Francesconi L,
Valliant J, Babich JW and Zubieta J: New directions in the
coordination chemistry of 99mTc: A reflection on
technetium core structures and a strategy for new chelate design.
Nucl Med Biol. 32:1–20. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Ahlgren S, Wållberg H, Tran TA, Widström
C, Hjertman M, Abrahmsén L, Berndorff D, Dinkelborg LM, Cyr JE,
Feldwisch J, et al: Targeting of HER2-expressing tumors with a
site-specifically 99mTc-labeled recombinant affibody
molecule, ZHER2:2395, with C-terminally engineered
cysteine. J Nucl Med. 50:781–789. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Lindberg H, Hofström C, Altai M, Honorvar
H, Wållberg H, Orlova A, Ståhl S, Gräslund T and Tolmachev V:
Evaluation of a HER2-targeting affibody molecule combining an
N-terminal HEHEHE-tag with a GGGC chelator for
99mTc-labelling at the C terminus. Tumour Biol.
33:641–651. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Ahlgren S, Andersson K and Tolmachev V:
Kit formulation for 99mTc-labeling of recombinant
anti-HER2 affibody molecules with a C-terminally engineered
cysteine. Nucl Med Biol. 37:539–546. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Hnatowich DJ, Virzi F, Fogarasi M,
Rusckowski M and Winnard P Jr: Can a cysteine challenge assay
predict the in vivo behavior of 99mTc-labeled
antibodies? Nucl Med Biol. 21:1035–1044. 1994. View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Wållberg H and Orlova A: Slow
internalization of anti-HER2 synthetic affibody monomer
111In-DOTA-ZHER2:342-pep2: implications for
development of labeled tracers. Cancer Biother Radiopharm.
23:435–442. 2008. View Article : Google Scholar
|
|
47
|
Sörensen J, Sandberg D, Sandström M,
Wennborg A, Feldwisch J, Tolmachev V, Åström G, Lubberink M,
Garske-Román U, Carlsson J, et al: First-in-human molecular imaging
of HER2 expression in breast cancer metastases using the
111In-ABY-025 affibody molecule. J Nucl Med. 55:730–735.
2014. View Article : Google Scholar
|
|
48
|
Andersson KG, Rosestedt M, Varasteh Z,
Malm M, Sandström M, Tolmachev V, Löfblom J, Ståhl S and Orlova A:
Comparative evaluation of 111In-labeled NOTA-conjugated
affibody molecules for visualization of HER3 expression in
malignant tumors. Oncol Rep. 34:1042–1048. 2015.PubMed/NCBI
|
|
49
|
Mitran B, Altai M, Hofström C, Honarvar H,
Sandström M, Orlova A, Tolmachev V and Gräslund T: Evaluation of
99mTc-Z IGF1R:4551-GGGC affibody molecule, a
new probe for imaging of insulin-like growth factor type 1 receptor
expression. Amino Acids. 47:303–315. 2015. View Article : Google Scholar
|
|
50
|
Tolmachev V, Varasteh Z, Honarvar H,
Hosseinimehr SJ, Eriksson O, Jonasson P, Frejd FY, Abrahmsen L and
Orlova A: Imaging of platelet-derived growth factor receptor β
expression in glioblastoma xenografts using affibody molecule
111In-DOTA-Z09591. J Nucl Med. 55:294–300. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
51
|
Honarvar H, Garousi J, Gunneriusson E,
Höidén-Guthenberg I, Altai M, Widström C, Tolmachev V and Frejd FY:
Imaging of CAIX-expressing xenografts in vivo using
99mTc-HEHEHE-ZCAIX:1 affibody molecule. Int J Oncol.
46:513–520. 2015.
|
|
52
|
Altai M, Honarvar H, Wållberg H, Strand J,
Varasteh Z, Rosestedt M, Orlova A, Dunås F, Sandström M, Löfblom J,
et al: Selection of an optimal cysteine-containing peptide-based
chelator for labeling of affibody molecules with 188Re.
Eur J Med Chem. 87:519–528. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
53
|
Wållberg H, Orlova A, Altai M,
Hosseinimehr SJ, Widström C, Malmberg J, Ståhl S and Tolmachev V:
Molecular design and optimization of 99mTc-labeled
recombinant affibody molecules improves their biodistribution and
imaging properties. J Nucl Med. 52:461–469. 2011. View Article : Google Scholar
|
|
54
|
Altai M, Wållberg H, Orlova A, Rosestedt
M, Hosseinimehr SJ, Tolmachev V and Ståhl S: Order of amino acids
in C-terminal cysteine-containing peptide-based chelators
influences cellular processing and biodistribution of
99mTc-labeled recombinant Affibody molecules. Amino
Acid. 42:1975–1985. 2012. View Article : Google Scholar
|
|
55
|
Moffat FL Jr, Pinsky CM, Hammershaimb L,
Petrelli NJ, Patt YZ, Whaley FS and Goldenberg DM; The Immunomedics
Study Group. Clinical utility of external immunoscintigraphy with
the IMMU-4 technetium-99m Fab′ antibody fragment in patients
undergoing surgery for carcinoma of the colon and rectum: Results
of a pivotal, phase III trial. J Clin Oncol. 14:2295–2305.
1996.PubMed/NCBI
|
|
56
|
Serafini AN, Klein JL, Wolff BG, Baum R,
Chetanneau A, Pecking A, Fischman AJ, Hoover HC Jr, Wynant GE,
Subramanian R, et al: Radioimmunoscintigraphy of recurrent,
metastatic, or occult colorectal cancer with technetium 99m-labeled
totally human monoclonal antibody 88BV59: Results of pivotal, phase
III multicenter studies. J Clin Oncol. 16:1777–1787.
1998.PubMed/NCBI
|
|
57
|
Schwochau K: Technetium
radiopharmaceuticals: Fundamentals, synthesis, structure and
development. Angew Chem Int Ed Engl. 33:2258–2267. 1994. View Article : Google Scholar
|