Introduction
Carcinoma of the bladder is the most common
urothelial malignancy, and is the 6th commonest cancer in the world
by incidence (1). Despite
significant improvements in management, bladder cancer remains one
of the commonest causes of cancer related mortality, ranked 6th and
11th in the UK in 2008 in men and women, respectively (2). Additionally, curative surgery or
radiotherapy commonly results in significant long-term
morbidity.
Localised bladder cancer may be divided into either
non-muscle invasive bladder cancer (NMIBC) or into muscle invasive
bladder cancer (MIBC). While the former may be cured by
transurethral resection, treatment of the latter requires radical
local management using either cystectomy or radical radiotherapy
(with or without concurrent chemotherapy). Approximately a third of
NMIBC tumours will recur and progress to muscle invasive over time
(3). Several clinicopathological
factors including tumour grade, stage, size, presence of carcinoma
in situ, previous recurrence (4), and p53 status are associated with
increased likelihood of progression. However, these factors are not
currently clearly sufficient to identify patients who need more
radical therapy at the outset. Similarly, stratification of
patients with muscle invasive cancer into those most likely to
recur or progress following local therapy would enable a more
personalised approach to treatment.
Previous studies have employed microarrays to
investigate gene expression profiles in bladder cancer (5–8). The
identification of differentially regulated genes and gene networks
allows the dissection of pathways and processes that are
dysregulated in bladder cancer. This, in turn, provides invaluable
clues relating to pathogenesis and moreover, may provide novel
targets for drug development.
Here, we describe a study in which we profiled gene
expression in 20 bladder cancer samples and compared this to 11
healthy urothelial samples. In addition to defining gene expression
signatures for different subgroups, the study identified several
genes and gene networks which are significantly deregulated in
bladder cancer. These include osteopontin and the classical
complement pathway. In this study we used the Affymetrix Human Gene
1.0 ST array which is a whole transcript based array compared to
many other studies which used arrays querying only the 3′ end of
transcripts.
Materials and methods
Patients and tumour samples
Cystoscopic tissue biopsies were collected between
May 2004 and November 2005. Prior to undergoing cystoscopy,
participants gave written informed consent for additional biopsies
to be taken for this study during their medical procedure. Patients
eligible for inclusion were those aged 18–80 years with a diagnosis
of histologically confirmed superficial (NMIBC) or muscle invasive
(MIBC) transitional cell carcinoma (n=14 and n=6, respectively),
and who had not received any prior therapy. Control (C) samples
were obtained from uninvolved tissues, for which no abnormalities
were observed upon histological examination (n=11). Ten of these
were paired with bladder cancer biopsies (4 nonmuscle invasive and
5 muscle invasive), while the last was from a patient without a
diagnosis of malignancy. Demographic characteristics of the patient
cohort are presented in Table I.
MIBC samples displayed a basal immuno-phenotype.
 | Table IPatient demographics. |
Table I
Patient demographics.
| Demographics | Controls
(n=11) | NMIBC (n=14) | MIBC (n=6) |
|---|
| Age in years |
| Median, range | 79.2 (64–85.8) | 70.8
(47.0–81.9) | 80.14
(65.3–85.8) |
| Gender |
| Male | 8 (72.2) | 11 (78.6) | 3 (60.0) |
| Female | 3 (27.2) | 3 (21.4) | 2 (40.0) |
| Grade |
| G1 | N/A | 6 (42.8) | 0 (0) |
| G2 | N/A | 1 (7.1) | 0 (0) |
| G3 | N/A | 7 (50.0) | 6 (100) |
| T stage |
| PT1 | N/A | 5 (35.7) | 0 (0) |
| PT1B | N/A | 1 (7.1) | 0 (0) |
| Pta | N/A | 8 (57.1) | 0 (0) |
| PT2 | N/A | 0 (0) | 6 (100) |
| Recurrence |
| No | N/A | 12 (92.3) | 0 (0) |
| Yes | N/A | 1 (7.7) | 5 (100) |
| Survival in months
(95% CI) | N/A | 81.2 (73.3-not
estimatable) | 12.26 (0–35.6) |
Tumour samples for IHC analysis were obtained from
the Liverpool Bioinnovation Hub Biobank (Liverpool, UK).
Pathologically assessed samples used healthy bladder tissue (n=3)
and MIBC tumour tissue (n=3).
Ethical considerations
Approval for the present study was obtained from the
South Birmingham Local Research Ethics Committee. All participants
gave written informed consent for their respective biopsies to be
utilized in this study. Approval for further anaylsis was granted
by the Liverpool Tissue Bank Ethics Committee (application no.
12-09). All participants gave written informed consent for their
respective biopsies to be used in biomedical analysis.
Tissue sample preparation
Tissue biopsy samples were snap-frozen and stored in
liquid nitrogen prior to RNA extraction.
RNA isolation
Total RNA was extracted from each tissue using
TRIzol (Invitrogen) prior to purification on silica (RNeasy Mini
columns; Qiagen, Hilden, Germany) according to the manufacturer's
recommendation. An Agilent Bioanalyser was used to confirm RNA
integrity and only samples having a RIN of 6.0 or more were
used.
cDNA probe synthesis
A total of 200 ng of total RNA was converted to cDNA
including targeted depletion of rRNA transcripts using a WT
Expression kit (Ambion, Cambridge, MA, USA) according to the
manufacturer's procedures. Resultant cDNA was fragmented and biotin
end labelled using an Affymetrix WT Terminal Labelling kit
(Affymetrix, Santa Clara, CA, USA).
Microarray hybridisation analysis
Labelled cDNA was hybridised to Affymetrix Human
Gene 1.0 ST whole transcript-based arrays followed by washing and
staining on an Affymetrix FS450 fluidics station. The Human Gene
1.0 ST Array (Affymetrix) comprised 764,885 distinct 25-nt probes
interrogating 28,869 annotated genes based on the March 2006 (UCSC
hg18, NCBI Build 36) human genome sequence assembly with
comprehensive coverage of RefSeq, Ensembl and putative complete CDS
GenBank transcripts.
Signal collection and analysis
Arrays were scanned using an Affymetrix Scanner 3000
7G. Affymetrix Command Console was used for instrument control and
data acquisition. All procedures were carried out according to the
standard Affymetrix protocols. Raw microarray data were submitted
to ArrayExpress (Accession: E-MTAB-1560).
Statistical analysis of microarray
data
Data transformation
Robust multi-array averaging, adjusted for
non-specific binding, (GCRMA) was used for normalisation and
summarisation of the probe intensity signals (9,10).
Data were subject to RMA background correction and quantile
normalisation across all chips (to ensure comparable mean and
standard deviation of the probe intensities), prior to log2
transformation and median polish probe set summarisation.
Array data quality estimation
The normalised unscaled standard error (NUSE) of
each array was calculated to compare data variability between and
within arrays (11). A NUSE value
of 1.05 was set as the upper limit of acceptable quality.
Data analysis
Principal components analysis using a covariance
dispersion matrix was performed to map high dimensional data of
each sample to 3 dimensions for estimation of clustering according
to classification and identification of outlying samples.
A mixed model analysis of variance (ANOVA) including
methods of moments estimation was used to identify differential
expression between non-invasive tumours, invasive tumours and
healthy bladder tissue. Batch effects were taken into consideration
within the model. ANOVA was conducted including 'patient' as a
factor, taking the paired samples into consideration. Differences
in gene expression between tumours of differing grade, and also
differential expression between tumour samples were assessed using
the t-test. To correct for multiple testing, genes were considered
to be significantly differentially expressed if a false discovery
rate (FDR) adjusted P<0.05 was obtained. The false discovery
rate was calculated as previously described (12). Those genes whose fold change
expression level was found to significantly differ by ±2× or
greater were used for further analysis.
Unsupervised hierarchical agglomerative clustering
(Euclidean distance with average linkage clustering) was used to
identify samples with similar expression profiles.
MetaCore version 6.7 (GeneGo, Inc., St. Joseph, MI,
USA), IPA (Ingenuity Systems, Redwood City, CA, USA) and DAVID
Bioinformatics resource v 6.7 (13) were employed to identify canonical
pathways and curated networks having statistically significant
enrichment of differentially expressed genes. Pathways from
MetaCore analysis filtered on FDR <0.05 are presented in
Results, unless otherwise stated. Univariate Cox proportional
hazards regression analysis was performed to evaluate the
association between disease progression and differential gene
expression.
Meta-analysis
Meta-analysis was conducted using publicly available
gene expression data from Gene Expression Omnibus (14). Out of the possible 12 datasets
identified (GSE88, GSE89, GSE7476, GSE30522, GSE24152, GSE3167,
GSE5287, GSE12630, GSE27448, GSE19915, GSE13507 and GSE5479), we
selected two datasets using the same microarray platform
(Affymetrix Human Genome U133A array) with a large number of
samples for further analysis. Raw CEL files from GSE3167 (15) with 60 samples and GSE5287 (16) with 30 samples were imported into
Partek Genomic Suite (Partek, Inc., Chesterfield, MO, USA) and data
were processed and differential genes identified following the same
protocol described earlier for our own data. Clinico-pathological
data for the datasets were kindly provided by the authors. The
GSE3167 dataset contains superficial transitional cell carcinoma
(NMIBC) with surrounding carcinoma in situ (CIS) (13
patients), without surrounding CIS lesions (15 patients), muscle
invasive carcinoma (MIBC, 13 patients), CIS only (n=5) and healthy
bladder (n=14). The GSE2587 dataset contains 30 samples from
patients with muscle invasive bladder cancer.
Immunostaining
Western blotting
Western blotting was performed to validate the
specificity of anti-OPN antibody (AF1433; R&D Systems, Inc.,
Minneapolis, MN, USA) for subsequent IHC analysis. Protein lysates
were prepared in RIPA buffer from a MIBC cell line, 253-J and a
NMIBC cell line, RT112. Bradford was used for protein
quantification. Proteins were separated by SDS-PAGE prior to
transferring onto a PDVF membrane. Blocking was performed for 1 h
with 10% milk, prior to overnight incubation in 1:1,000 dilution of
AF1433. Secondary antibody was applied for 1 h before visualising
using ECL. β-actin (ab8227; Abcam) was used to demonstrate equal
protein loading.
Immunohistochemistry
Conditions for IHC with AF1433 were optimised in
clear cell renal carcinoma and kidney FFPE sections, using a no
primary antibody control, a goat IgG isotype control (AB-108-C;
R&D Systems) and a negative tissue control-breast skin. Antigen
retreval was performed using the PT-link at pH 9.0, prior to
peroxidase block (Dako), 1-h primary antibody (1:50 AF1433)
incubation at RT, 1-h secondary antibody incubation at RT (1:100)
prior to visualisation with DAB Chromagen (Dako). Staining
intensity was scored by a consultant pathologist. Briefly, staining
intensity was graded as 0 (negative), + (weak), ++ (moderate) and
+++ (strong).
Results
Bladder cancer tissues exhibit specific
gene signatures
A total of 418 genes were differentially expressed
between cancer and control samples, using a fold change of at least
2.5, and a false discovery rate (FDR) of 0.05. Of these, 368 genes
showed reduced expression in tumours, while 50 genes showed
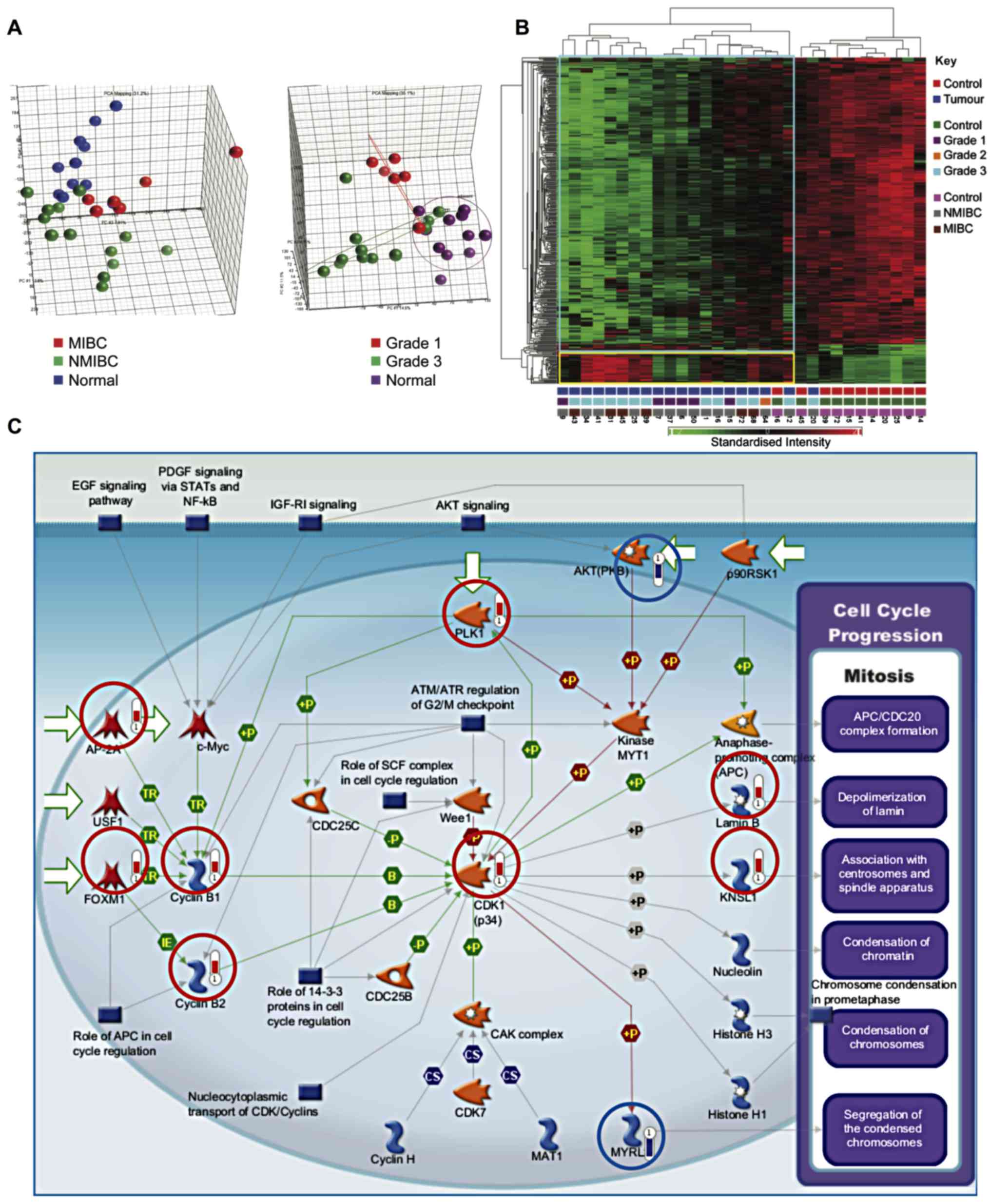
increased expression in tumours. Principal components analysis
demonstrated segregation of samples by tissue type and tumour grade
(Fig. 1A). Similarly, hierarchical
clustering segregated healthy bladder tissue and bladder cancer
into two groups (Fig. 1B).
Multiple genes involved in proliferation were
upregulated in bladder cancer samples as compared to healthy
samples (P=2.59×10−9) (Fig.
1C). These included components of the cyclin-dependent kinase 1
(CDK1)/cydin B complex, activation of which is required to drive
progression from G2- to M-phase. Initiation of cyclin B
transcription is carried out by FOXM1. Moreover, the equilibrium
between nuclear import and export of cyclin B1 is influenced by its
phosphorylation status by PLK1, which was also upregulated. The
substrates of CDKl/cyclin B include Lamin B, leading to
depolymerization of the lamin nuclear-cytoskeleton, and
Kinesin-related motor protein Eg5 (KNSL1), which associates with
centrosomes and spindle apparatus.
Genes involved in myogenesis and
epithelial-to-mesenchymal transition (EMT) are downregulated in
cancer samples
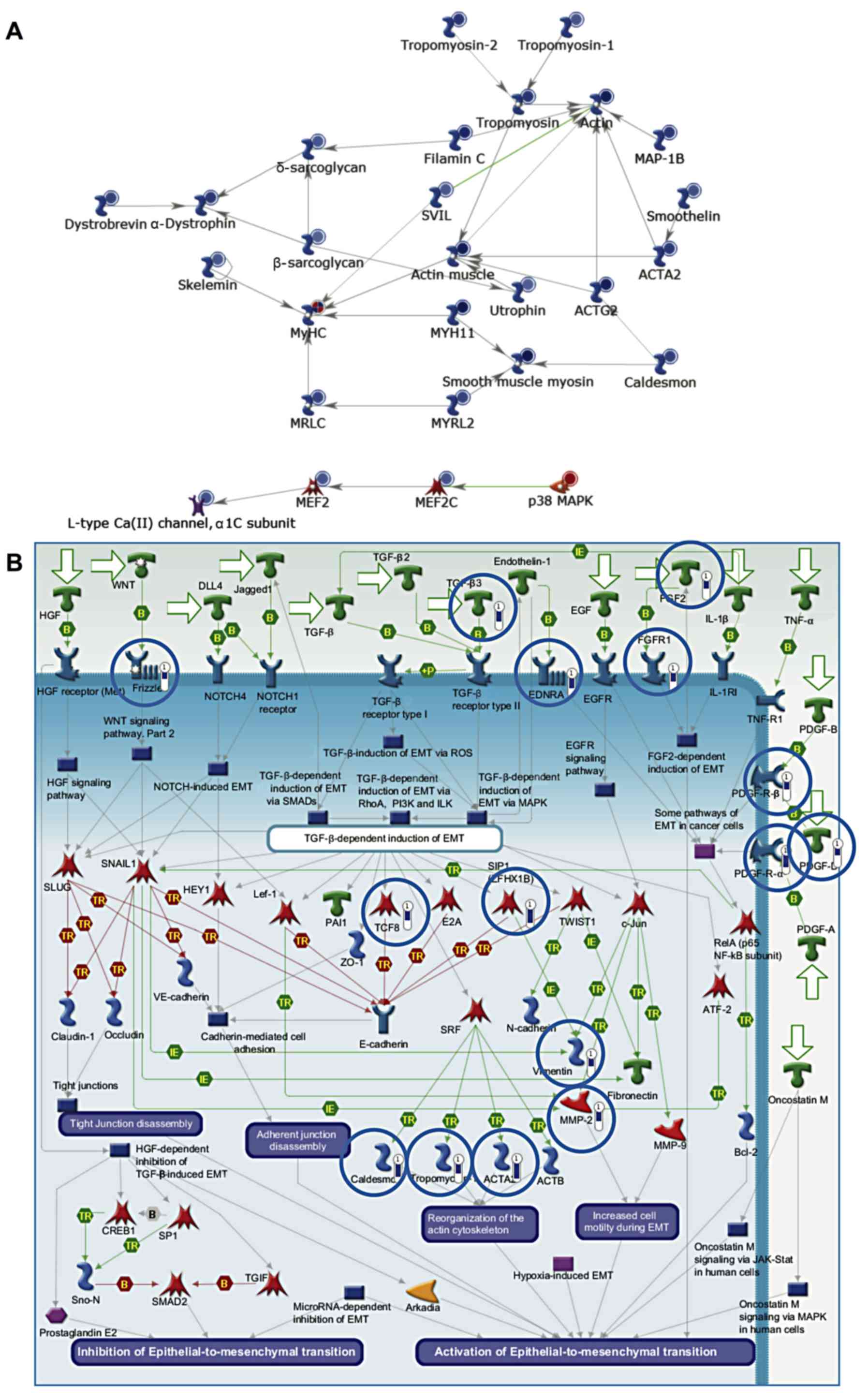
Genes downregulated in cancer (compared to healthy
tissue) included several which are involved in myogenesis
(P=1.22×10−10) (Fig.
2A). These included the myocyte enhancer factor, Mef2, which is
expressed during myogenesis. Mef2 and MyoD interact leading to MyoD
activation and hence the expression of myosin heavy and light
chains. In addition, genes involved in EMT were also downregulated
(P=1.41×10−9) (Fig.
2B). EMT is critical during myogenesis, particularly as the
muscle precursor cells delaminate from the dermomyotome. Muscle
precursors invade the developing tissue and differentiate into
mature muscle cells.
Comparison of NMIBC and healthy bladder
tissue
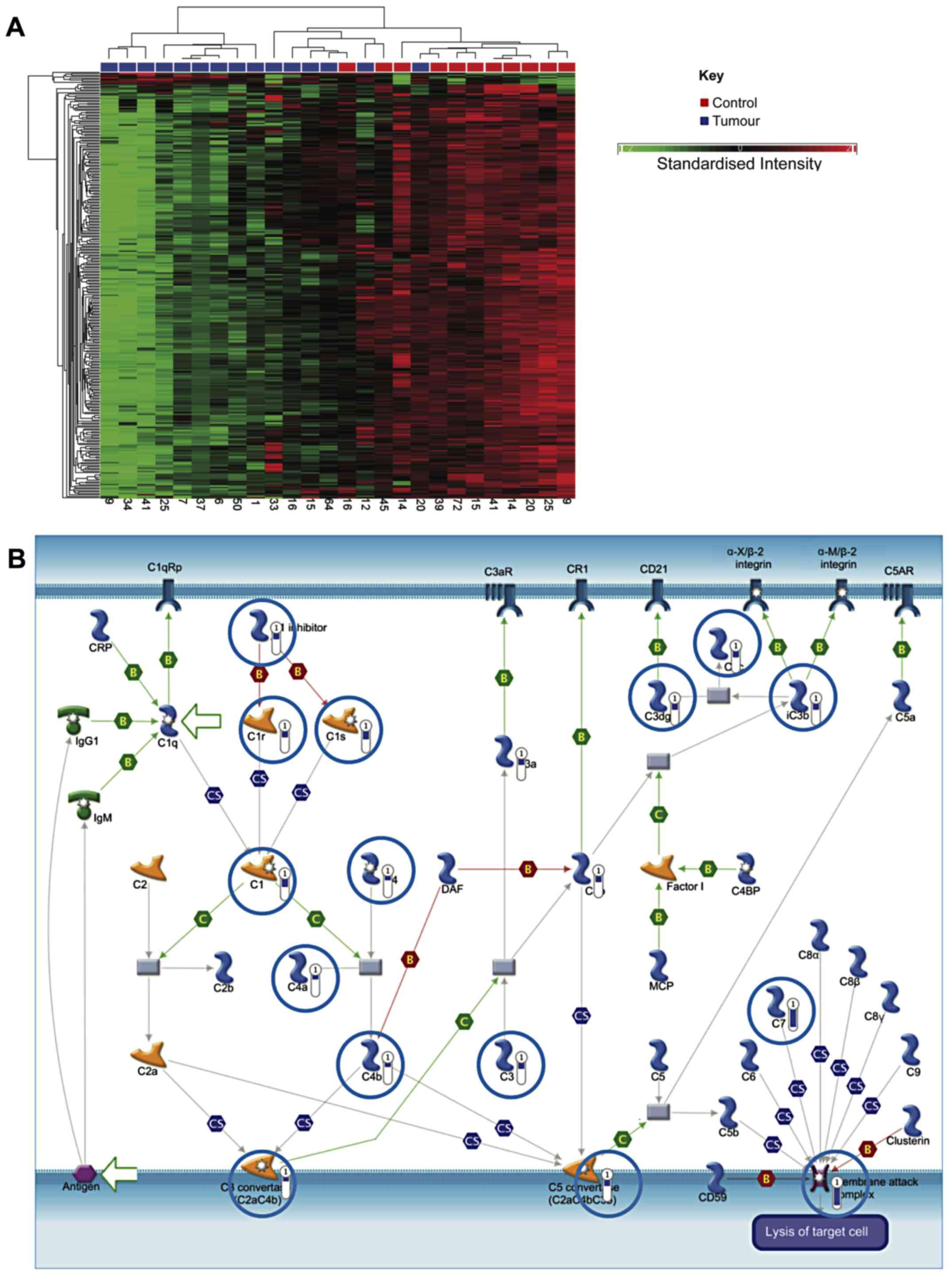
A specific comparison was also made between NMIBC
and healthy tissue. This identified 410 genes, which were
differentially expressed by ≥2.5-fold (P<0.05, false discovery
rate). Of these, 393 were downregulated in cancer, while increased
expression was observed in 17 (data not shown). Hierarchical
clustering generally segregated into two groups, but with
additional branching in one cluster as shown in Fig. 3A.
Similar to the analysis comparing all bladder cancer
samples to healthy samples, genes involved in smooth muscle
development and EMT were downregulated (P=4.8×10−12).
Notably, genes associated with the classical complement pathway
were strongly downregulated in NMIBC samples, which may indicate a
mechanism for immune evasion (P=5.12×10−10) (Fig. 3B).
MIBC vs. healthy tissue
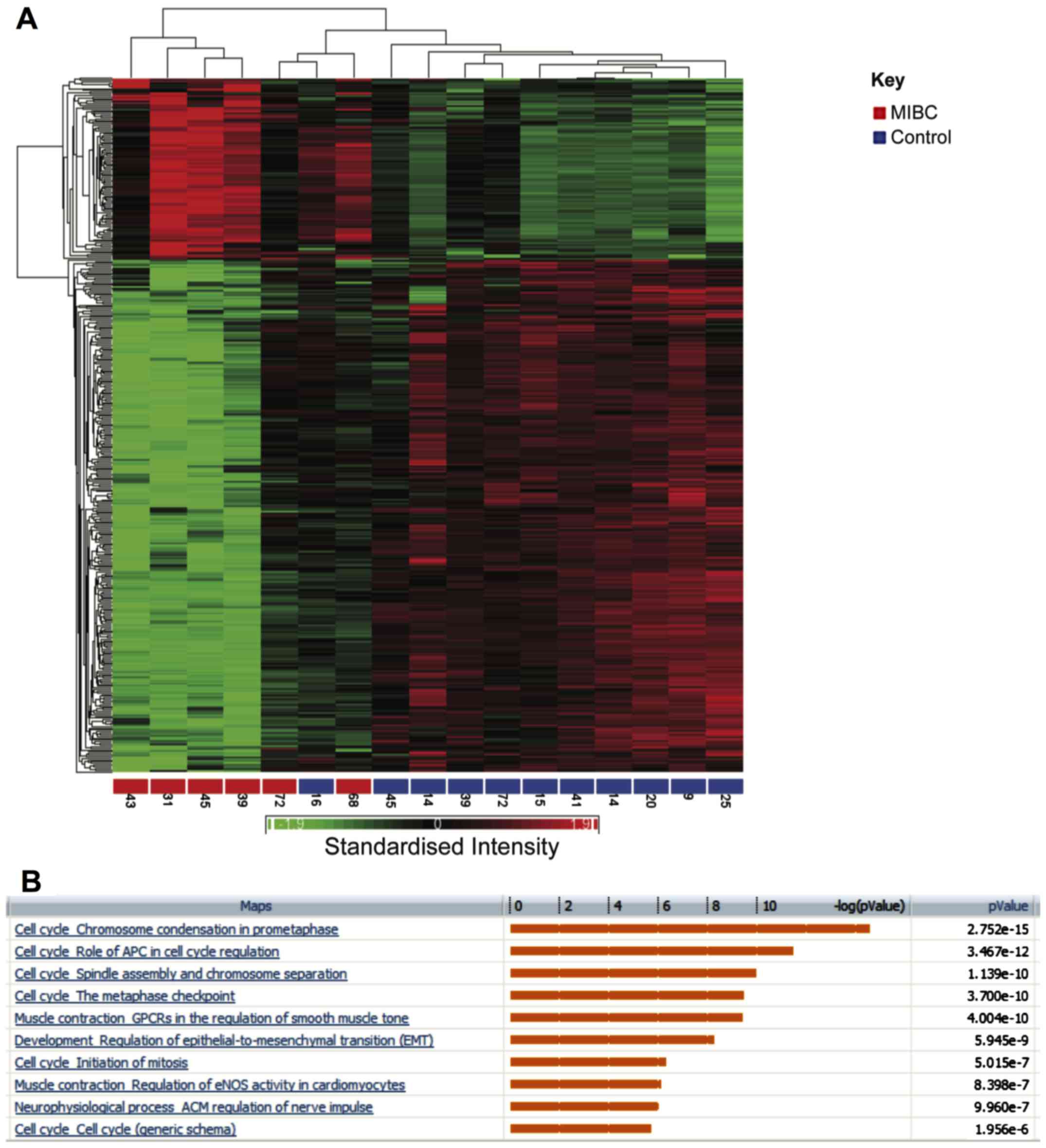
Hierarchical clustering showed segregation of MIBC
and healthy samples into two groups (Fig. 4A). There were 404 genes that were
altered by at least 3-fold (P<0.05 false discovery rate). Out of
these, 107 showed increased expression between invasive tumours and
healthy samples, while 297 were decreased in expression. Pathways
enriched for these differentially regulated genes are shown in
Fig. 4B. Genes involved in
chromosome condensation in prometaphase were upregulated in
invasive bladder cancer (P=2.75×10−15). Similarly,
several regulators of the anaphase promoting complex (APC) were
also upregulated (P=3.47×10−12). These included the
kinases, PLK1 and CDK1.
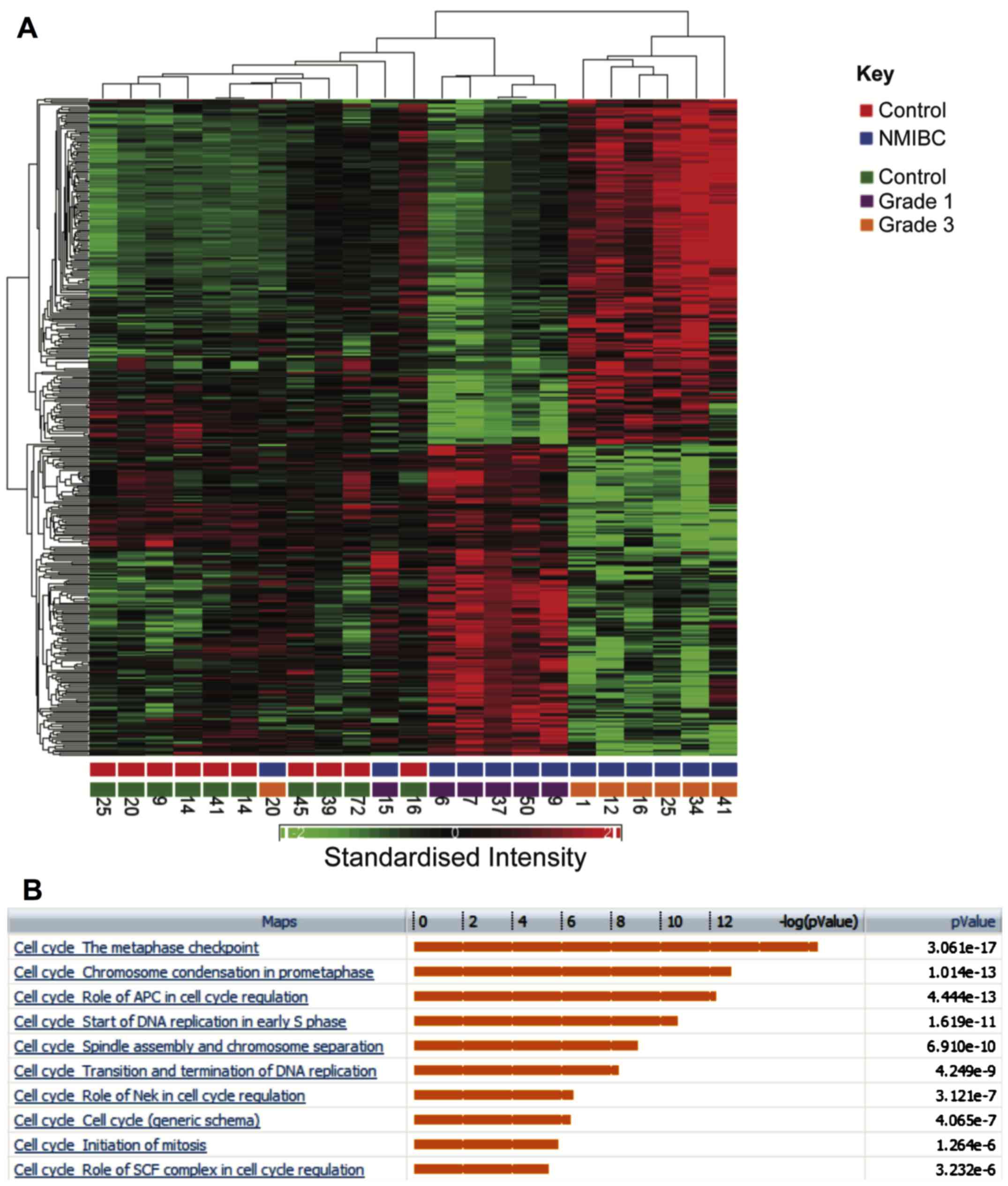
Comparison of grade 1 and 3 NMIBC
A further comparison was made between gene
expression profiles in grade 1 and 3 bladder cancers, which
identified 341 genes that were differentially regulated by at least
2-fold (FDR P<0.05). Hierarchical clustering clearly segregates
between grade 1 and 3 (Fig. 5A).
Enriched pathways are shown in Fig.
5B. Notably multiple pathways associated with cell cycle
progression were upregulated in grade 3 tumours and multiple core
cell cycle components were upregulated in grade 3 tumours. Genes
involved in the metaphase checkpoint were particularly upregulated
(P=3.06×10−17). A comparison was also made of genes that
were upregulated both in grade 3 compared to grade 1 NMIBC and in
MIBC compared to healthy samples.
Comparing grade 3 NMIBC and MIBC
A total of 292 genes were differentially expressed 2
or more fold (P<0.05). Of these, 144 showed increased expression
in MIBC while the remaining 148 exhibited decreased expression in
MIBC. Hierarchical clustering resulted in clear segregation between
the two groups. Table II lists
genes that were differentially expressed by >5-fold in grade 3
NMIBC compared to MIBC. Among the genes most upregulated in MIBC
were the proteinases. Overexpression and activation of proteinases,
that destroy the connective tissue, are involved in various
pathological processes including tumour progression. The most
proteolytic of enzymes are the metalloproteinases. The matrix
metalloproteinases (MMPs) comprise a family of enzymes that
collectively can degrade all components of the extracellular matrix
(ECM), these were upregulated in this study. For example, the
matrix metalloproteinases MMP1 and MMP16 were upregulated 4.8- and
2.6-fold, respectively in MIBC compared to NMIBC.
 | Table IIGenes exhibiting >5 fold
differential expression between grade 3 and MIBC (FDR
P<0.05). |
Table II
Genes exhibiting >5 fold
differential expression between grade 3 and MIBC (FDR
P<0.05).
| Gene | ID | RefSeq | P-value | Fold change
(MIBC/NMIBC |
|---|
| Family with
sequence similarity 83, member D | FAM83D | NM_030919 | 5.50E-06 | 7.87015 |
| Retinoic acid
receptor responder (tazarotene induced) 1 | RARRES1 | NM_206963 | 0.001291 | 5.96921 |
| Centromere protein
I | CENPI | NM 006733 | 1.35E-05 | 5.92042 |
| Fibronectin 1 | FN1 | NM_212482 | 0.000579 | 5.86325 |
| Centromere protein
F, 350/400 ka (mitosin) | CENPF | NM_016343 | 0.000505 | 5.7622 |
| N mannosidase, α,
class 1C, member 1 | MANICI | NM_020379 | 0.000401 | 5.52916 |
| Topoisomerase (DNA)
II α 170 kDa | TOP2A | NM_001067 | 0.00091 | 5.32363 |
| TPX2,
microtubule-associated, homolog (Xenopus laevis) | TPX2 | NM_012112 | 0.000892 | 5.27537 |
| Ras association
(RalGDS/AF-6) domain family member 2 | RASSF2 | NM_014737 | 0.000644 | 5.04613 |
| Anillin, actin
binding protein | ANLN | NM_018685 | 0.000489 | 5.01334 |
Further functional analysis using
Database for Annotation, Visualization and Integrated Discovery
(DAVID)
A further analysis of 143 genes, which were
upregulated in MIBC compared to NMIBC was performed using the DAVID
database. A summary of annotation clusters is given in Table III.
 | Table IIIDAVID functional annotation
clusters. |
Table III
DAVID functional annotation
clusters.
| Annotation
cluster | Enrichment
score | Count | P-value |
|---|
| Extracellular
glycoproteins | 6.47 | 37 | 2.7E-07 |
| Serine-type
endopeptidase inhibitor activity | 3.29 | 7 | 7.90E-05 |
| Negative regulation
of immune system processes | 2.21 | 6 | 4.60E-04 |
| Zymogen
(proteolytic enzyme precursors) | 1.85 | 14 | 5.10E-04 |
| Cell migration | 1.33 | 9 | 1.40E-03 |
| EGF | 1.18 | 5 | 1.50E-02 |
| Response to
wounding | 0.95 | 13 | 8.20E-04 |
Meta-analysis of gene expression
data
In order to validate our results, we used a
meta-analysis of a larger dataset consisting of 90 samples,
generated by the combination of two publicly available bladder
cancer gene expression datasets (15,16).
When we followed the same criteria used with our dataset for
classifying MIBC and NMIBC, there were 29 NMIBC samples and 42 MIBC
samples in the meta-analysis dataset. There were also 14 healthy
bladder tissue samples, while the rest of the samples (carcinoma
in situ) were not used in analysis.
A total of 4513 genes were differentially expressed
in NMIBC compared to healthy tissue, when a cut-off of FDR <0.05
and fold-change >2.5 were applied. When compared with
differential genes from our data, there were 118 genes common in
both analyses. Surprisingly except for one gene, PELO, all the
other genes showed concordant expression. This included all the 8
genes upregulated in NMIBC and 109 genes downregulated in NMIBC
compared to healthy bladder tissue (data not shown).
When MIBC samples were compared with healthy bladder
samples, with a filter of FDR <0.05 and fold-change >3, 3905
genes were differentially expressed in the meta-analysis. A
comparison with differential genes from our data under similar
conditions and filtrations revealed 44 common genes. Similarly, the
expressions of all except 2 genes (SERPINEB13 and CASC5) were
concordant in both the analyses. All the 30 genes downregulated in
MIBC compared to healthy tissue showed concordant expression, while
12 out of 14 genes highly expressed in MIBC in this study showed
similar expression in the meta-analysis (data not shown).
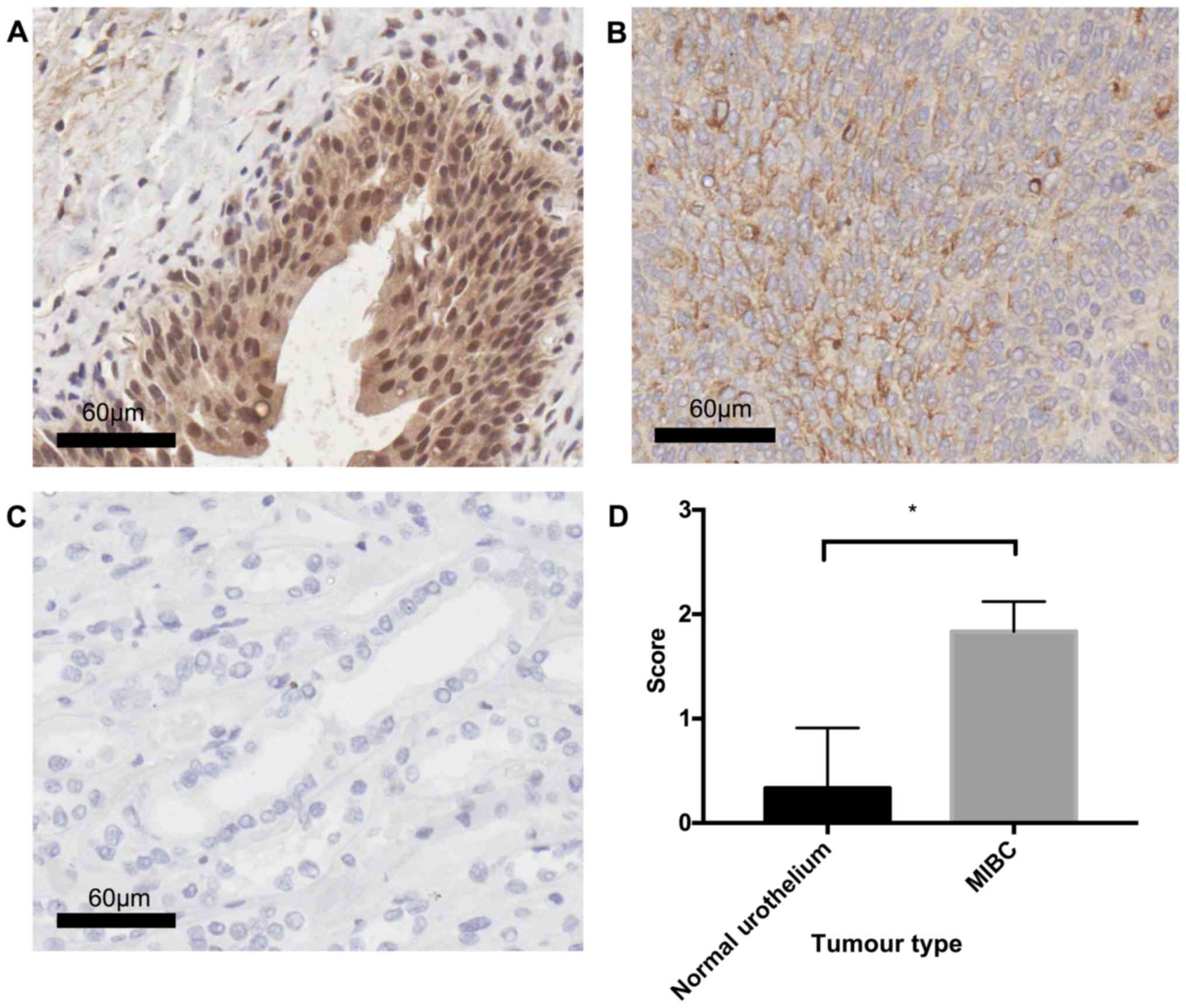
Osteopontin protein is overexpressed in
MIBC compared to healthy urothelium
In order to determine the validity of the microarray
data, OPN expression was examined in MIBC (n=3) and healthy
urothelial tissue (n=3) using the IHC technique. Statistical
analysis using Welch's unpaired t-test revealed a significant
difference (P=0.0286) between MIBC and healthy tissue, with MIBC
displaying diffuse cytoplasmic OPN staining of moderate (++)
intensity, whilst healthy urothelium was typically OPN negative (−)
(Fig. 6).
Discussion
The present study contributes to a body of work that
uses microarray data to profile bladder cancer, and provides fresh
insights into pathways involved in its pathogenesis. Distinctive
patterns of gene expression differentiating between NMIBC and MIBC,
and between high and low grade cancers were identified.
A striking upregulation in pathways involved in cell
cycle control was observed in comparisons of MIBC and NMIBC, and
between grade 3 and grade 1 NMIBC tumours. A significant overlap
was observed between genes upregulated in grade 3 NMIBC tumours and
those upregulated in MIBC compared to healthy tissue and these
genes were predominantly mapped to cell cycle control pathways.
This is likely to reflect the increased aggressiveness and
likelihood of progression of grade 3 NMIBC. This panel of genes may
therefore be useful in predicting progression of NMIBC.
Cell cycle genes which were upregulated included
anillin (>8-fold upregulation), a substrate of APC, which is
involved in cytokinesis (17).
Anillin is upregulated in various cancers including pancreatic
(18) and renal carcinoma
(19). TPX2, which is involved in
spindle assembly (20), was also
upregulated.
This upregulation of cell cycle related genes in
bladder cancer is not unexpected, and was similarly shown in a
study by Zaravinos et al (21). Notably, several of the upregulated
genes have been identified as potential therapeutic targets for
bladder cancers. Inhibitors of TOP2A include anthracyclines and
etoposide, two well-established chemotherapy agents. The
anthracycline doxorubicin, for example, exhibits antibladder cancer
effects and is a component of the commonly employed MVAC regimen.
PLK1 inhibitors are currently in phase 1 and 2 trials, and
preliminary studies suggest that PLK1 inhibitors may reduce
proliferation of bladder cancer cell lines (22). However, results obtained from a
recent phase 2 study evaluating the efficacy of the PLK1 inhbitor
volasertib as a second-line treatment in patients with locally
advanced or metastatic urothelial cancer, suggests that although
volasertib was well-tolerated with an acceptable safety profile the
antitumour activity of volasertib was not suffecient for further
evaluation as a monotherapy (23).
Several CDK inhibitors are also currently in phase 1 and 2 clinical
trials.
The expression of osteopontin (OPN) 'secreted
phosphoprotein 1', a matricellular protein, is strongly associated
with malignancies (reviewed in ref. 24). Furthermore, OPN RNA-silencing in BC
significantly attenuates tumour cell invasion (25). The finding that OPN expression was
strikingly upregulated (>10-fold) in invasive bladder cancers
compared with the healthy tissue is consistent with previous
studies demonstrating upregulation of OPN in urothelial malignancy
(22,26–28).
Whilst OPN RNA expression data is well represented in BC tissue
(28,29), protein expression data are limited
(26,27). Therefore, OPN expression was
examined between MIBC and healthy urothelium to determine whether
commonality between RNA and protein expression exists in BC. OPN
was found to be signifcantly (P=0.0286) overexpressed in MIBC
compared to healthy urothelium. These data are consistent with
obeservations in gastric (30),
oesophageal (31), cervical
(32) and prostate (33) cancers (26,34).
Whilst a significant relationship between OPN and poor prognosis
was noted in upper urinary tract urothelial cell carcinoma no
correlation between stage and grades was found (27). Conversely, OPN expression is
associated with improved survival in pancreatic adenocarcinoma
(35).
Elevated levels of osteopontin have also been
detected in the plasma of patients with advanced transitional cell
carcinoma (TCC), suggesting that OPN could be a novel biomarker of
disease stage (36,37). The utility of OPN expression as a
prognostic marker (and/or predictor of treatment response),
however, remains unproven (26,27,38).
The differences in OPN expression (in blood and tissue) observed in
studies could be explained, at least in part, by the expression of
OPN isoforms. Specifically, OPN splice variants which exhibit
divergent biological functions are differentially expressed in
cancers. Intriguingly, CD44, the hyaluronic acid receptor, for
which osteopontin is a key ligand (37), was found to be upregulated among
invasive bladder cancers, further supporting a potential role for
OPN in pathogenesis of TCC.
When comparing grade 3 NMIBC with MIBC, we observed
an overexpression of the cell-matrix interaction network. In
particular, there was upregulation in proteolytic enzymes such as
the matrix metalloproteases (MMPs). This is consistent with the
increased ability of MIBC to invade local tissues. It would be
important to evaluate if overexpression of MMPs predicts NMIBC
progression.
Genes involved in myogenesis and
epithelial-to-mesenchymal transition (EMT) were downregulated in
tumours, and this was particularly noted in NMIBC samples compared
with the healthy tissue. This is intriguing, and could represent an
inhibitory effect of bladder cancer on normal bladder muscle
growth. EMT describes a process by which epithelial cells lose
their characteristic apical-basal polarity, intercellular junctions
and epithelial markers, while upregulating mesenchymal markers,
acquiring front-rear polarity, and undergoing cytoskeleton
reorganization into mesenchymal cells (39). EMT is associated with invasive
cancers and metastases, portends a poor prognosis in patients with
bladder cancer (40). In the
present study, tumour samples were not micro-dissected, and the
possible 'admixture' with normal muscle tissue within samples could
have confounded the analysis of myogenesis and EMT genes.
One novel finding in this study is the
downregulation of genes involved in the classical complement
pathway within the NMIBC group. Previously, others have shown that
complement inhibitors such as CD46 and complement factor H (CFH)
are upregulated in bladder cancers (41). The aggregate data suggest that
repression of complement activity is an important pathogenic
mechanism in bladder cancer development. Intriguingly, CFH binds to
the OPN protein and masks putative enzymatic-cleavage sites, and
could thereby, alter OPN activity. Urinary levels of CFH were also
investigated as markers of bladder malignancy (42,43),
but were found to exhibit low sensitivity and specificity. This
could be explained by changes in CFH levels during disease
progression, and supports our findings that downregulation of
complement genes occurred in NMIBC rather than MIBC samples.
ESRP1 and ESRP2 are upregulated in NMIBC (compared
to healthy tissue), and are involved in the regulation of FGFR2
splicing (44). While decreased
expression of FGFR2b (one of the FGFR2 splice variants) was
observed in a subset of bladder cancers (45), upregulation of FGFR2c (another
splice variant) was detected in a bladder cancer metastasis model
(46). Consistent with reports on
differential protein isoforms during cancer development, our
findings support the hypothesis that differential expression of
FGFR2 splice variants is an important pathogenic mechanism in
bladder cancer. In agreement with data published from the Cancer
Genome Atlas Reseach Network in 2013 (47) which comprehensively evaluated the
molecular signature of 131 urothelial carcinomas and revealed that
mutational spectrum was involved in cell cycle regulation,
chromatin regulation, and kinase signalling pathways with potential
therapeutic application, the present study highlights several
dysregulated transcripts in BC that potentially could be targeted
in BC treatment.
Whilst the long-term clinical outcome data would
have allowed more robust conclusions to be drawn from the small
cohort, meta-analysis of the microarray data together with the
protein expression experiment highlight the importance of the
presented data, in addition to the role of OPN in invasion
(25). Furthermore, this study
supplements a growing body of molecular data in bladder cancer
allowing further understanding of this complex and heterogeneous
disease.
In conclusion, this study complements the published
literature, and gives credence to observed associations between
patterns of gene signatures with the stage and grade of bladder
cancers. Ultimately such signatures would be most clinically useful
if they improve prognostication of disease, particularly among
NMIBC, where a more aggressive therapeutic approach could be
employed in cancers that were more likely to progress. The present
study identified expression patterns, which may help this
prediction (such as proteolytic enzymes), and these may be
incorporated into future prospective studies. Significantly, we
also identified several genes and pathways (such as OPN and the
classical complement pathway) which were strikingly deregulated in
bladder cancer, and which may be valuable drug targets.
Abbreviations:
|
NMIBC
|
non-muscle invasive bladder cancer
|
|
MIBC
|
muscle invasive bladder cancer
|
|
CDK1
|
cyclin-dependent kinase 1
|
|
FOXM1
|
forkhead box protein M1
|
|
PLK1
|
polo-like kinase 1
|
|
KNSL1
|
kinesin family member 11
|
|
Mef2
|
myocyte enhancer factor-2
|
|
MyoD
|
myogenic differentiation 1
|
|
EMT
|
epithelial to mesenchymal
transition
|
|
MMPs
|
matrix metalloproteinases
|
|
MMP1
|
matrix metalloproteinase 1
|
|
MMP16
|
matrix metalloproteinase 16
|
|
PELO
|
protein pelota homolog
|
|
SERPINB13
|
Serpin peptidase inhibitor, clade B
(ovalbumin), member 13
|
|
CASC5
|
cancer susceptiility candidate 5
|
|
APC
|
anaphase promoting complex
|
|
TPX2
|
targeting protein for Xklp2
|
|
TOP2A
|
topoisomerase 2-alpha
|
|
MVAC
|
methotrexate, vinblastine, adriamycin
and cisplatin
|
|
OPN
|
osteopontin
|
|
TCC
|
transitional cell carcinoma
|
|
CFH
|
compliment factor H
|
|
ESRP1
|
epithelial splicing regulatory protein
1
|
|
ESRP2
|
epithelial splicing regularoty protein
2
|
|
FGFR2(b,c)
|
fibroblast growth factor receptor 2 (b
isoform, c isoform)
|
|
IHC
|
immunohistochemistry
|
Acknowledgments
We thank Sim Sihota for her skilled technical
assistance in the extraction and microarray analysis of RNA.
References
|
1
|
IARC: Cancer incidence and mortality
Worldwide. IARC CancerBase No. 10. Journal. 2015, http://globocan.iarc.fr.
|
|
2
|
Cancer Research UK: Bladder cancer
statistics. 2015, http://www.cancerresearchuk.org/health-professional/cancer-statistics/statistics-by-cancer-type/bladder-cancer.
|
|
3
|
Kaufman DS, Shipley WU and Feldman AS:
Bladder cancer. Lancet. 374:239–249. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Sylvester RJ, van der Meijden AP,
Oosterlinck W, Witjes JA, Bouffioux C, Denis L, Newling DW and
Kurth K: Predicting recurrence and progression in individual
patients with stage Ta T1 bladder cancer using EORTC risk tables: a
combined analysis of 2596 patients from seven EORTC trials. Eur
Urol. 475–467. 2006.
|
|
5
|
Blaveri E, Simko JP, Korkola JE, Brewer
JL, Baehner F, Mehta K, Devries S, Koppie T, Pejavar S, Carroll P,
et al: Bladder cancer outcome and subtype classification by gene
expression. Clin Cancer Res. 11:4044–4055. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Elsamman E, Fukumori T, Ewis AA, Ali N,
Kajimoto K, Shinohara Y, Ishikawa M, Takahashi M, Nishitani MA,
Baba Y, et al: Differences in gene expression between noninvasive
and invasive transitional cell carcinoma of the human bladder using
complementary deoxyribonucleic acid microarray: Preliminary
results. Urol Oncol. 24:109–115. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Riester M, Taylor JM, Feifer A, Koppie T,
Rosenberg JE, Downey RJ, Bochner BH and Michor F: Combination of a
novel gene expression signature with a clinical nomogram improves
the prediction of survival in high-risk bladder cancer. Clin Cancer
Res. 18:1323–1333. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Sanchez-Carbayo M, Socci ND, Lozano J,
Saint F and Cordon-Cardo C: Defining molecular profiles of poor
outcome in patients with invasive bladder cancer using
oligonucleotide microarrays. J Clin Oncol. 24:778–789. 2006.
View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Irizarry RA, Hobbs B, Collin F,
Beazer-Barclay YD, Antonellis KJ, Scherf U and Speed TP:
Exploration, normalization, and summaries of high density
oligonucleotide array probe level data. Biostatistics. 4:249–264.
2003. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Wu Z, Irizarry RA, Gentleman R,
Martinez-Murillo F and Spencer F: A Model-based background
adjustment for oligonucleotide expression arrays. J Am Stat Assoc.
99:909–917. 2004. View Article : Google Scholar
|
|
11
|
Bolstad BM, Collin F, Simpson KM, Irizarry
RA and Speed TP: Experimental design and low-level analysis of
microarray data. Int Rev Neurobiol. 60:25–58. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Pawitan Y, Michiels S, Koscielny S,
Gusnanto A and Ploner A: False discovery rate, sensitivity and
sample size for microarray studies. Bioinformatics. 21:3017–3024.
2005. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Huang W, Sherman BT and Lempicki RA:
Systematic and integrative analysis of large gene lists using DAVID
bioinformatics resources. Nat Protoc. 4:44–57. 2009. View Article : Google Scholar
|
|
14
|
Edgar R, Domrachev M and Lash AE: Gene
Expression Omnibus: NCBI gene expression and hybridization array
data repository. Nucleic Acids Res. 30:207–210. 2002. View Article : Google Scholar :
|
|
15
|
Dyrskjøt L, Kruhøffer M, Thykjaer T,
Marcussen N, Jensen JL, Møller K and Ørntoft TF: Gene expression in
the urinary bladder: A common carcinoma in situ gene expression
signature exists disregarding histopathological classification.
Cancer Res. 64:4040–4048. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Als AB, Dyrskjøt L, von der Maase H, Koed
K, Mansilla F, Toldbod HE, Jensen JL, Ulhøi BP, Sengeløv L, Jensen
KM, et al: Emmprin and survivin predict response and survival
following cisplatin-containing chemotherapy in patients with
advanced bladder cancer. Clin Cancer Res. 13:4407–4414. 2007.
View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Zhao WM and Fang G: Anillin is a substrate
of anaphase-promoting complex/cyclosome (APC/C) that controls
spatial contractility of myosin during late cytokinesis. J Biol
Chem. 280:33516–33524. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Olakowski M, Tyszkiewicz T, Jarzab M, Krol
R, Oczko-Wojciechowska M, Kowalska M, Kowal M, Gala GM, Kajor M,
Lange D, et al: NBL1 and anillin (ANLN) genes over-expression in
pancreatic carcinoma. Folia Histochem Cytobiol. 47:249–255. 2009.
View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Gruss OJ and Vernos I: The mechanism of
spindle assembly: Functions of Ran and its target TPX2. J Cell
Biol. 166:949–955. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Ronkainen H, Hirvikoski P, Kauppila S and
Vaarala MH: Anillin expression is a marker of favourable prognosis
in patients with renal cell carcinoma. Oncol Rep. 25:129–133.
2011.
|
|
21
|
Zaravinos A, Lambrou GI, Volanis D,
Delakas D and Spandidos DA: Spotlight on differentially expressed
genes in urinary bladder cancer. PLoS One. 6:e182552011. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Zhang Z, Zhang G and Kong C: High
expression of polo-like kinase 1 is associated with the metastasis
and recurrence in urothelial carcinoma of bladder. Urol Oncol.
31:1222–1230. 2013. View Article : Google Scholar
|
|
23
|
Stadler WM, Vaughn DJ, Sonpavde G,
Vogelzang NJ, Tagawa ST, Petrylak DP, Rosen P, Lin CC, Mahoney J,
Modi S, et al: An open-label, single-arm, phase 2 trial of the
Polo-like kinase inhibitor volasertib (BI 6727) in patients with
locally advanced or metastatic urothelial cancer. Cancer.
120:976–982. 2014. View Article : Google Scholar
|
|
24
|
Ahmed M, Behera R, Chakraborty G, Jain S,
Kumar V, Sharma P, Bulbule A, Kale S, Kumar S, Mishra R, et al:
Osteopontin: A potentially important therapeutic target in cancer.
Expert. Opin Ther Targets. 15:1113–1126. 2011. View Article : Google Scholar
|
|
25
|
Xu ST, Guo C, Ding X, Fan WJ, Zhang FH, Xu
WL and Ma YC: Role of osteopontin in the regulation of human
bladder cancer proliferation and migration in T24 cells. Mol Med
Rep. 11:3701–3707. 2015.PubMed/NCBI
|
|
26
|
Coppola D, Szabo M, Boulware D, Muraca P,
Alsarraj M, Chambers AF and Yeatman TJ: Correlation of osteopontin
protein expression and pathological stage across a wide variety of
tumor histologies. Clin Cancer Res. 10:184–190. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Ke HL, Chang LL, Yang SF, Lin HH, Li CC,
Wu DC and Wu WJ: Osteopontin overexpression predicts poor prognosis
of upper urinary tract urothelial carcinoma. Urol Oncol.
29:703–709. 2011. View Article : Google Scholar
|
|
28
|
Zaravinos A, Volanis D, Lambrou GI,
Delakas D and Spandidos DA: Role of the angiogenic components,
VEGFA, FGF2, OPN and RHOC, in urothelial cell carcinoma of the
urinary bladder. Oncol Rep. 28:1159–1166. 2012.PubMed/NCBI
|
|
29
|
Fang ZQ, Zang WD, Chen R, Ye BW, Wang XW,
Yi SH, Chen W, He F and Ye G: Gene expression profile and
enrichment pathways in different stages of bladder cancer. Genet
Mol Res. 12:1479–1489. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Imano M, Satou T, Itoh T, Sakai K,
Ishimaru E, Yasuda A, Peng YF, Shinkai M, Akai F, Yasuda T, et al:
Immunohistochemical expression of osteopontin in gastric cancer. JJ
Gastrointest Surg. 13:1577–1582. 2009. View Article : Google Scholar
|
|
31
|
Kita Y, Natsugoe S, Okumura H, Matsumoto
M, Uchikado Y, Setoyama T, Owaki T, Ishigami S and Aikou T:
Expression of osteopontin in oesophageal squamous cell carcinoma.
Br J Cancer. 95:634–638. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Song JY, Lee JK, Lee NW, Yeom BW, Kim SH
and Lee KW: Osteopontin expression correlates with invasiveness in
cervical cancer. Aust N Z J Obstet Gynaecol. 49:434–438. 2009.
View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Forootan SS, Foster CS, Aachi VR, Adamson
J, Smith PH, Lin K and Ke Y: Prognostic significance of osteopontin
expression in human prostate cancer. Int J Cancer. 118:2255–2261.
2006. View Article : Google Scholar
|
|
34
|
Weber GF, Lett GS and Haubein NC:
Osteopontin is a marker for cancer aggressiveness and patient
survival. Br J Cancer. 103:861–869. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Collins AL, Rock J, Malhotra L, Frankel WL
and Bloomston M: Osteopontin expression is associated with improved
survival in patients with pancreatic adenocarcinoma. Ann Surg
Oncol. 19:2673–2678. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Zhao L, Wang Y, Qu N, Huang C and Chen L:
Significance of plasma osteopontin levels in patients with bladder
urothelial carcinomas. Mol Diagn Ther. 16:311–316. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Naor D, Sionov RV and Ish-Shalom D: CD44:
Structure, function, and association with the malignant process.
Adv Cancer Res. 71:241–319. 1997. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Ang C, Chambers AF, Tuck AB, Winquist E
and Izawa JI: Plasma osteopontin levels are predictive of disease
stage in patients with transitional cell carcinoma of the bladder.
BJU Int. 96:803–805. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Lim J and Thiery JP:
Epithelial-mesenchymal transitions: Insights from development.
Development. 139:3471–3486. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Baumgart E, Cohen MS, Silva Neto B, Jacobs
MA, Wotkowicz C, Rieger-Christ KM, Biolo A, Zeheb R, Loda M,
Libertino JA, et al: Identification and prognostic significance of
an epithelialmesenchymal transition expression profile in human
bladder tumors. Clin Cancer Res. 13:1685–1694. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Varela JC, Atkinson C, Woolson R, Keane TE
and Tomlinson S: Upregulated expression of complement inhibitory
proteins on bladder cancer cells and anti-MUC1 antibody immune
selection. Int J Cancer. 123:1357–1363. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Babjuk M, Soukup V, Pesl M, Kostirova M,
Drncova E, Smolova H, Szakacsova M, Getzenberg R, Pavlik I and
Dvoracek J: Urinary cytology and quantitative BTA and UBC tests in
surveillance of patients with pTapT1 bladder urothelial carcinoma.
Urology. 71:718–722. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Cheng ZZ, Corey MJ, Pärepalo M, Majno S,
Hellwage J, Zipfel PF, Kinders RJ, Raitanen M, Meri S and Jokiranta
TS: Complement factor H as a marker for detection of bladder
cancer. Clin Chem. 51:856–863. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Warzecha CC, Sato TK, Nabet B, Hogenesch
JB and Carstens RP: ESRP1 and ESRP2 are epithelial
cell-type-specific regulators of FGFR2 splicing. Mol Cell.
33:591–601. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Diez de Medina SG, Chopin D, El Marjou A,
Delouvee A, LaRochelle WJ, Hoznek A, Abbou C, Aaronson SA, Thiery
JP and Radvanyi F: Decreased expression of keratinocyte growth
factor receptor in a subset of human transitional cell bladder
carcinomas. Oncogene. 14:323–330. 1997. View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Chaffer CL, Brennan JP, Slavin JL, Blick
T, Thompson EW and Williams ED: Mesenchymal-to-epithelial
transition facilitates bladder cancer metastasis: Role of
fibroblast growth factor receptor-2. Cancer Res. 66:11271–11278.
2006. View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Cancer Genome Atlas Research Network:
Comprehensive molecular characterization of urothelial bladder
carcinoma. Nature. 507:315–322. 2014. View Article : Google Scholar : PubMed/NCBI
|