Introduction
Colorectal cancer (CRC) is steadily increasing
worldwide, but has a particularly high incidence in developed
countries. Approxiamtely 20% of CRC patients already have
metastases at diagnosis (1). It is
well known that metastasis spreads cancer to other organs such as
the liver, lungs and lymph nodes, and it accounts for over 90% of
cancer-related mortality (2).
Mitochondria play an important role in the
production of ATP via oxidative phosphorylation (OXPHOS).
Mitochondria are also involved in biosynthetic metabolic processes
involving cholesterol, heme, lipids and nucleotides. Moreover,
mitochondria are considered the main sources of reactive oxygen
species and nitrogen species (3).
Mitochondrial biogenesis is a complex process that occurs by the
cooperation of numerous genes between nuclear and mitochondrial
genomes. It is regulated by a complex transcriptional network of
key factors including peroxisome proliferator-activated receptor
co-activator 1 alpha (PGC-1α), the nuclear respiratory factors
(NRFs), estrogen receptor-related receptor alpha (ERRα) and
mitochondrial transcription factor A (mtTFA) (4).
Research has shown an association between
mitochondrial biogenesis and metastasis. Cells with depleted
mitochondrial DNA (mtDNA) exhibit increased invasiveness through
the upregulation of cathepsin L expression (5). Depletion of mtDNA also promotes
invasion by modulating the expression of matrix metalloproteinases,
tissue inhibitors of the MMP family and other invasion-related
genes (6). However, increased
mitochondrial biogenesis is induced by treatment with bezafibrate,
a pan agonist of peroxisome proliferator-activated receptors
(PPARs), in cancer cells such as 143B, HeLa and MDA-MB-231.
Bezafibrate treatment reduces cell growth, glucose utilization and
invasion in these cells (7). When
wild-type KISS1, a metastasis suppressor, is upregulated, aerobic
glycolysis decreases and OXPHOS predominates. KISS1-expressing
cells show that increased mitochondrial biogenesis is accompanied
by increased mitochondrial gene expression and higher expression of
PGC-1α, a key co-activator that regulates mitochondrial mass and
metabolism (8).
Proton beam therapy (PBT), an alternative to gamma
or X-ray irradiation therapy, has produced promising clinical
results worldwide (9). In
particular, PBT has recently been used to treat hepatocellular
carcinoma (10), non-small cell
lung (11), prostate (12) and head and neck tumors (13). Some in vitro studies have
shown that PBT inhibits metastatic characteristics such as the
increase of adhesion and migration (14,15).
In previous studies, we also found that proton beam irradiation
inhibits metastatic potentials such as migration, invasion and the
expression of MMP-2 and MMP-9 in HT-29 cells (16,17).
Furthermore, proton beam irradiation inhibits gene expression and
the activities of molecules involved in integrin trafficking and
integrin-mediated signaling, which are essential processes in tumor
progression (18). However, the
contribution of abnormal mitochondrial biogenesis to colon cancer
metastasis is unknown. In the present study, we investigated the
effects of proton beam irradiation on mitochondrial biogenesis by
determining: mtDNA mass; the expression of mitochondria-specific
transcription factors, dynamic regulators, and functional
molecules; and the activities of signaling molecules in
12-O-tetradecanoyl phorbol-13-acetate (TPA)-induced
aggressive HT-29 cells.
Materials and methods
Materials
The following items were purchased from the stated
commercial sources: TPA (P1585) and 4′,6-diamidino-2-phenylindole
(DAPI, D9542) were from Sigma-Aldrich Co. (St. Louis, MO, USA);
rabbit anti-human phospho AMPK (#2535), mouse anti-human AMPK
(#2793), and rabbit anti-human phospho SIRT1 (#2314) antibodies
were from Cell Signaling Technology (Danvers, MA, USA); rabbit
anti-human SIRT1 (sc-15404), rabbit anti-human PGC-1α (sc-13067),
mouse anti-human β-actin (sc-47778), goat anti-mouse
IgG-horseradish peroxidase (HRP) (sc-2005), and goat anti-rabbit
IgG-HRP (sc-2004) secondary antibodies were from Santa Cruz
Biotechnology Inc., (Santa Cruz, CA, USA); ECL Plus Western
Blotting Substrate (34580) was from Pierce Biotechnology (Rockford,
IL, USA); TRIzol (15596026) and MitoTracker Red CMXRos (M7512) were
from Invitrogen-Life Technologies (Carlsbad, CA, USA); the
PrimeScript First Strand cDNA Synthesis kit (#6110A) was from
Takara Bio Inc. (Shiga, Japan); the FastStart Universal SYBR-Green
Master Mix (04 913 850 001) was from Roche Applied Science
(Mannheim, Germany); and the Phosphatase Inhibitor Cocktail (P3200)
and Protease Inhibitor Cocktail (P3100) solutions were from
GenDEPOT (Barker, TX, USA).
Cell culture
As previously described in detail (18), the HT-29 cell line was purchased
from the Korean Cell Line Bank (KCLB no. 30038, Seoul, Republic of
Korea). Cells were maintained in 5% CO2 at 37°C in
Roswell Park Memorial Institute (RPMI-1640) medium including 10%
fetal bovine serum (FBS; SH30084.03; HyClone Laboratories-GE
Healthcare Life Science, Victoria, Australia) and 100 U/ml
penicillin-streptomycin (15140122; Invitrogen). For induction of
metastatic capacity, cells were incubated with 150 nM TPA for 1 h
and then RPMI-1640 medium was added before proton beam
irradiation.
Proton beam irradiation
According to previously described methods (19), cell irradiation was performed with
a 100-MeV proton beam using the 100 MeV linac equipment
(Scanditronix, Uppsala, Sweden) at the Korea Multi-purpose
Accelerator Complex (KOMAC, Gyeongju, Republic of Korea). Briefly,
cells in a 12.5-cm2 flask filled with RPMI-1640 medium
were placed on a beam stage and irradiated (1, 4 and 8 Gy) at the
center of the Bragg peaks modulated to 6-cm widths. The dose during
each beam irradiation was measured using radiochromic film
(GAF-MD55; GAF Corp., Wayne, NJ, USA) as an in situ
measuring tool.
Mitochondrial biogenesis analysis
We cultured the irradiated cells for 48 and 72 h to
investigate the effect of proton beam irradiation on mitochondrial
biogenesis. As previously described in detail (20), DNA from cells was extracted using a
genomic DNA purification kit (A1120; Promega Corp., Madison, WI,
USA), according to the manufacturer's instructions. Mitochondrial
biogenesis was determined by the ratio of cytochrome c
oxidase subunit I (COX I) for mtDNA to ribosomal protein large p0
(RPLP0) for nuclear DNA (nDNA), quantified by quantitative reverse
transcription polymerase chain reaction (RT-qPCR). Data were
normalized against the expression of RPLP0.
Mitochondrial staining and
quantification
As previously described in detail (20), the irradiated cells were cultured
in 8-well chamber slides (12-565-8; Thermo Fisher Scientific,
Rochester, NY, USA) for 48 and 72 h. After washing with
phosphate-buffered saline (PBS), the cells were incubated with
RPMI-1640 medium containing 20 nM MitoTracker Red CMXRos at 37°C
for 30 min. After washing with PBS, the cells were fixed with 4%
paraformaldehyde for 10 min, and stained with 1 µg/ml DAPI
for 10 min. Fluorescent images were obtained using a digital
fluorescence microscope (Nikon Eclipse 80i; Nikon, Tokyo, Japan) at
×400 magnification. For quantification of the fluorescent signal,
the cells were dissolved with a lysis buffer (0.1 M potassium
phosphate, 1% Triton X-100, pH 10.0) for 10 min. Dimethyl sulfoxide
(DMSO) was added to the lysates, which were then incubated for 10
min. The fluorescent signal intensity from the lysates was measured
using a microplate reader (FLUOstar OPTIMA; BMG Labtech, Ortenberg,
Germany) at 466 nm excitation and 540 nm emission.
RT-qPCR analysis
As previously described in detail (20), total RNA from cells was obtained
using TRIzol (Invitrogen), and cDNA was produced using a
PrimeScript First strand cDNA Synthesis kit (Takara Bio Inc.)
according to the manufacturer's instructions. To analyze gene
expression, RT-qPCR was conducted using a FastStart SYBR-Green
Master mix (Roche Diagnostics) in an ABI Prism 7300 Sequence
Detection System (Applied Biosystems, Foster City, CA, USA). Levels
of target gene expression were calculated relative to the
expression level of the endogenous reference gene, β-actin, using
the delta cycle threshold method (Table I).
 | Table IPrimers for quantitative RT-PCR
analysis. |
Table I
Primers for quantitative RT-PCR
analysis.
| Forward | Reverse |
|---|
| PGC-1α |
GTTGCCTGCATGAGTGTGTG |
TCACTGCACCACTTGAGTCC |
| mtTFA |
ATGCTTATAGGGCGGAGTGG |
CTTCAGCTTTTCCTGCGGTG |
| NRF1 |
AGAACTCTCCCTGCTGGACT |
AGCACACTTACACGACGACT |
| ERRα |
GCGATGTCCTTTTGTGTCCT |
TCCGAGGAACCCTTTGGACT |
| TOMM40 |
CTCTGACCTCTCCCCTAGCAG |
CGATTGTGCTGAGGGCTACT |
| TIMM44 |
TGCTACGGAAGAAGCTTGGG |
CTTCTTCACGGACTCCACCC |
| CPT1α |
TTCAGTTCACGGTCACTCCG |
TGACCACGTTCTTCGTCTGG |
| CytC |
CAGTGCCACACCGTTGAAAA |
TGCATCGGTTATTTCACACTCC |
| ATP5B |
TGCCCCTGCTACTACGTTTG |
TGGCTGAGACAAGAAACGCT |
| RPLP0 |
CCTTCTCCTTTGGGCTGGTCATCCA |
CAGACACTGGCAACATTGCGGACAC |
| COX1 |
TCACCCACACTGTGCCCATCTACGA |
CAGCGGAACCGCTCATTGCCAATGG |
| β-actin |
TGACAGGATGCAGAAGGAGAT |
GCGCTCAGGAGGAGCAAT |
Western blot analysis
As previously described in detail (20), the irradiated cells were cultured
for 48 and 72 h, and lysed in radio-immunoprecipitation assay
(RIPA) buffer (89900; Thermo Fisher Scientific, Waltham, MA, USA).
Total protein lysates were separated by 8% SDS-polyacrylamide gel
electrophoresis and transferred to nitrocellulose membranes (10 401
396; Whatman GmbH, Dassel, Germany). The membranes were incubated
with 5% non-fat milk for 1 h at 25°C, and incubated with the
primary antibody (1:1,000) overnight at 4°C. After washing with
Tris-buffered saline (TBS) containing 0.1% Tween-20 for 30 min, the
membranes were incubated with the anti-mouse IgG, or anti-rabbit
IgG, HRP-conjugated secondary antibodies (1:3,000) for 1 h at 25°C.
After washing with TBS containing 0.1% Tween-20 for 30 min, the
western blotting images were obtained with an ECL Plus Western
blotting substrate (Pierce Biotechnology), using a Lumino image
analyzer (LAS 4000 mini; Fujifilm, Tokyo, Japan), and quantified
with ImageJ software (ver. 1.51j; National Institutes of Health,
Bethesda, MD, USA).
Statistical analysis
All data are shown as the mean ± SEM. The data were
analyzed by the one-way analysis of variance (ANOVA). Differences
between mean values were assessed using the Dunnett's multiple
comparisons test. Statistical significance was defined as
P<0.05.
Results
Proton beam irradiation stimulates
mitochondrial biogenesis in TPA-induced aggressive HT-29 cells
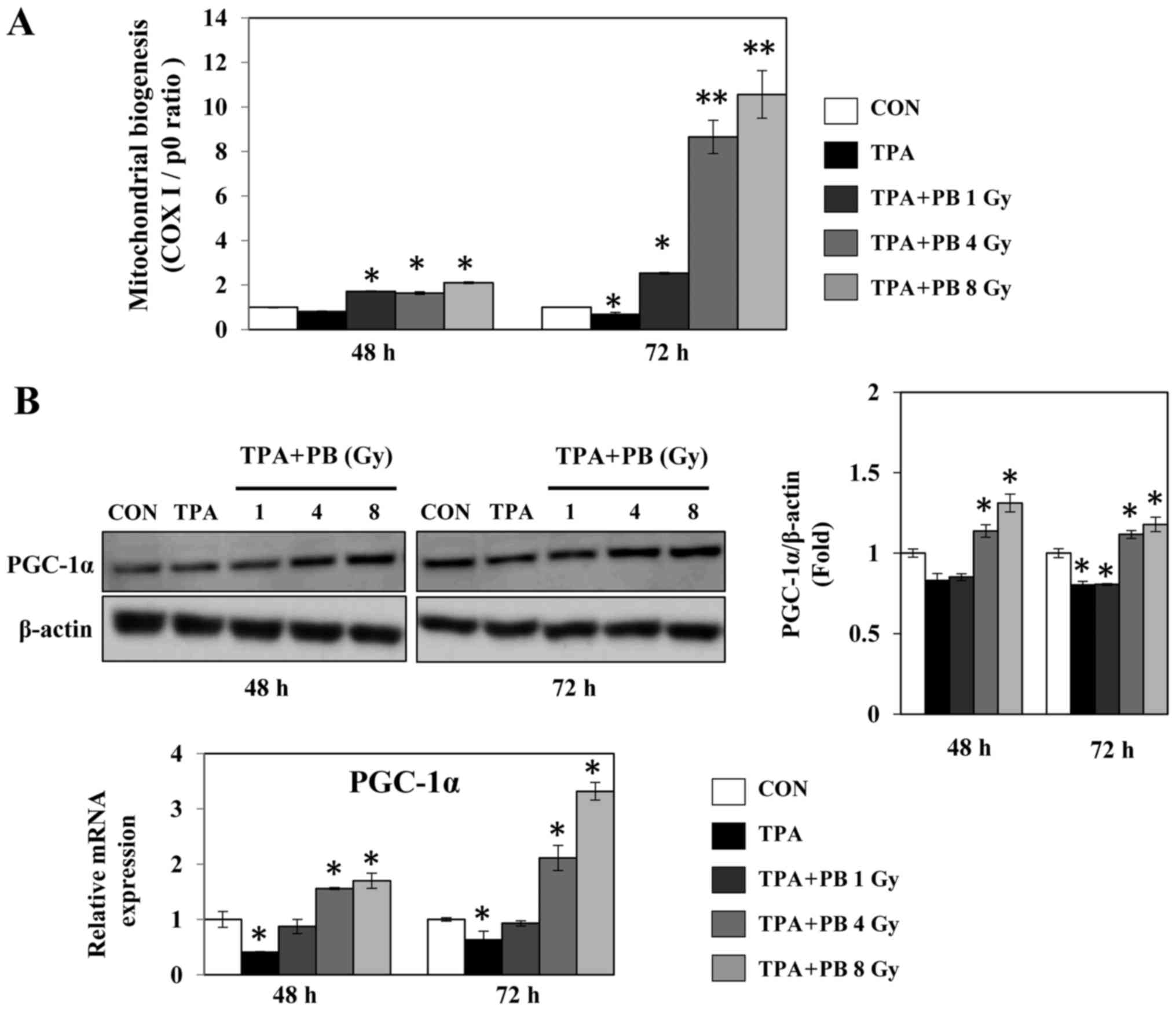
To determine whether proton beam irradiation
regulates mitochondrial biogenesis, we analyzed the gene expression
ratio of COX I and RPLP0, representing mtDNA and nDNA,
respectively, in TPA-induced aggressive HT-29 cells after
irradiation with a proton beam at 1, 4 or 8 Gy. Proton beam
irradiation increased mitochondrial biogenesis in a dose- and
time-dependent manner (Fig. 1A).
At the same time, proton beam irradiation increased the expression
of PGC-1α, a key transcription factor for mitochondrial biogenesis,
in a dose-dependent manner (Fig.
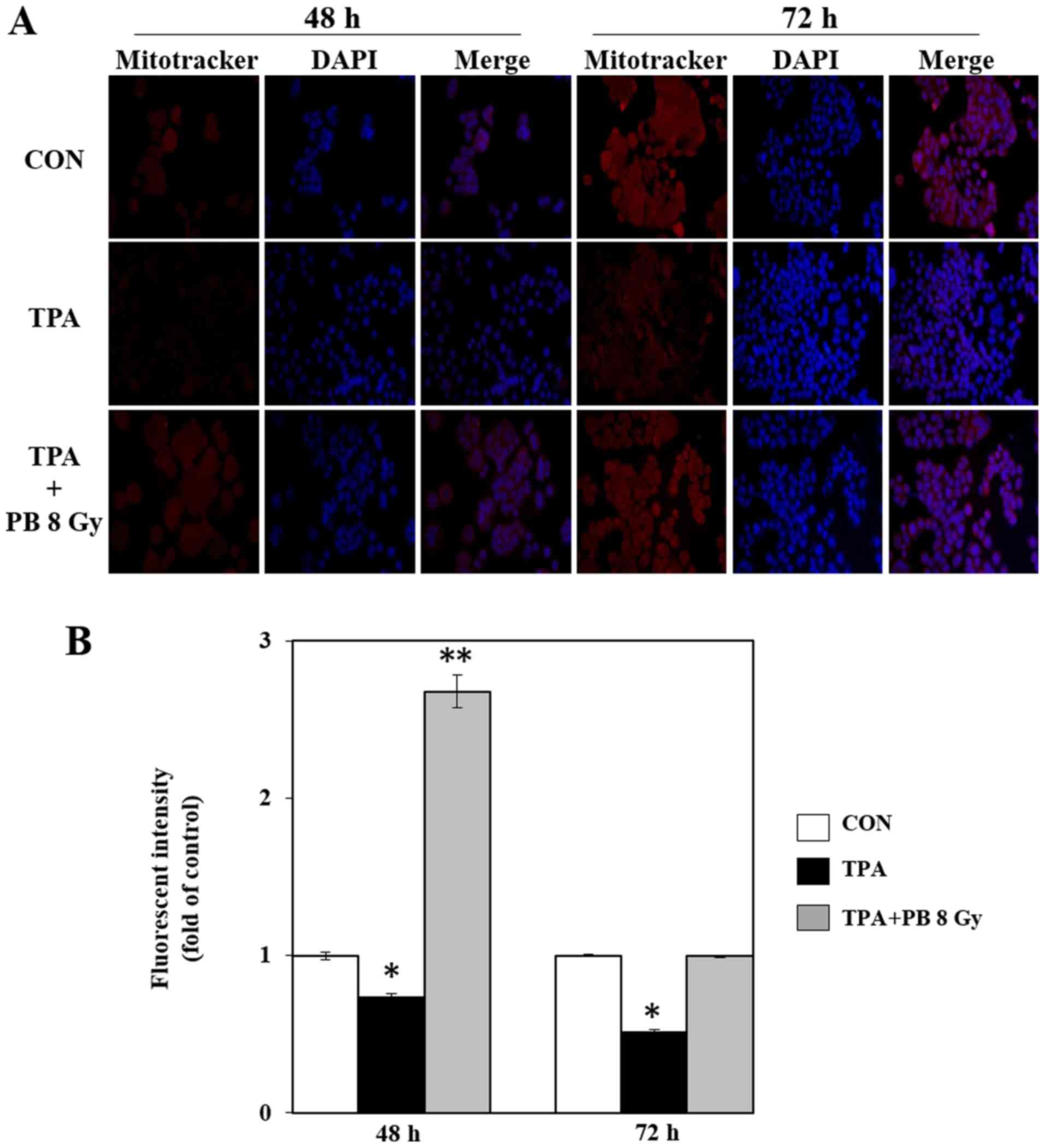
1B). To determine whether proton beam irradiation-induced
mitochondrial biogenesis is involved in mitochondrial function, we
next stained mitochondria in TPA-induced aggressive HT-29 cells
using MitoTracker Red CMXRos, a mitochondrial membrane potential
(ΔΨm)-sensitive fluorochrome that accumulates membrane
potential-dependently in live cells. Proton beam irradiation
increased mitochondrial staining compared with staining in the
group treated with TPA alone (Fig.
2). Therefore, these results indicate that proton beam
irradiation-induced mitochondrial biogenesis is associated with the
inhibitory effects of proton beam irradiation on metastatic
potential in TPA-induced aggressive HT-29 cells.
Proton beam irradiation stimulates
mitochondrial biogenesis by AMPK signaling pathways in TPA-induced
aggressive HT-29 cells
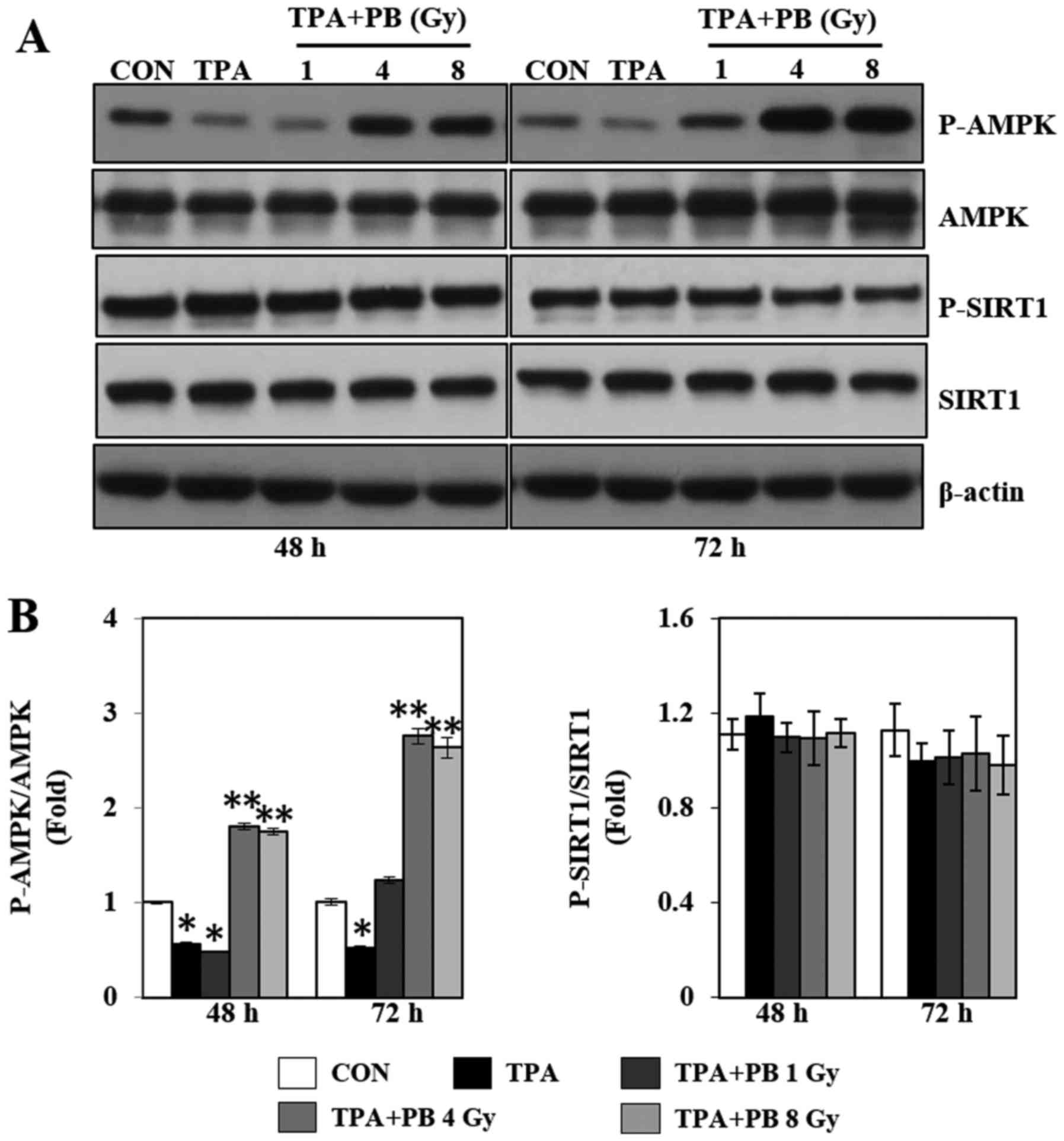
To further study the mechanism by which proton beam
irradiation affects mitochondrial biogenesis and function, we next
investigated the phosphorylation of AMPK and SIRT1, which are
upstream molecules associated with PGC-1α and regulators of
mitochondrial biogenesis. As shown in Fig. 3, TPA treatment decreased AMPK
phosphorylation, whereas proton beam irradiation increased AMPK
phosphorylation in a dose- and time-dependent manner. However,
proton beam irradiation did not affect SIRT1 phosphorylation.
Therefore, these results indicate that the proton beam
irradiation-induced increase in mitochondrial biogenesis might be
regulated by the AMPK signaling pathways.
Proton beam irradiation stimulates the
expression of mitochondrial transcription factors and mitochondria
functional genes in TPA-induced aggressive HT-29 cells
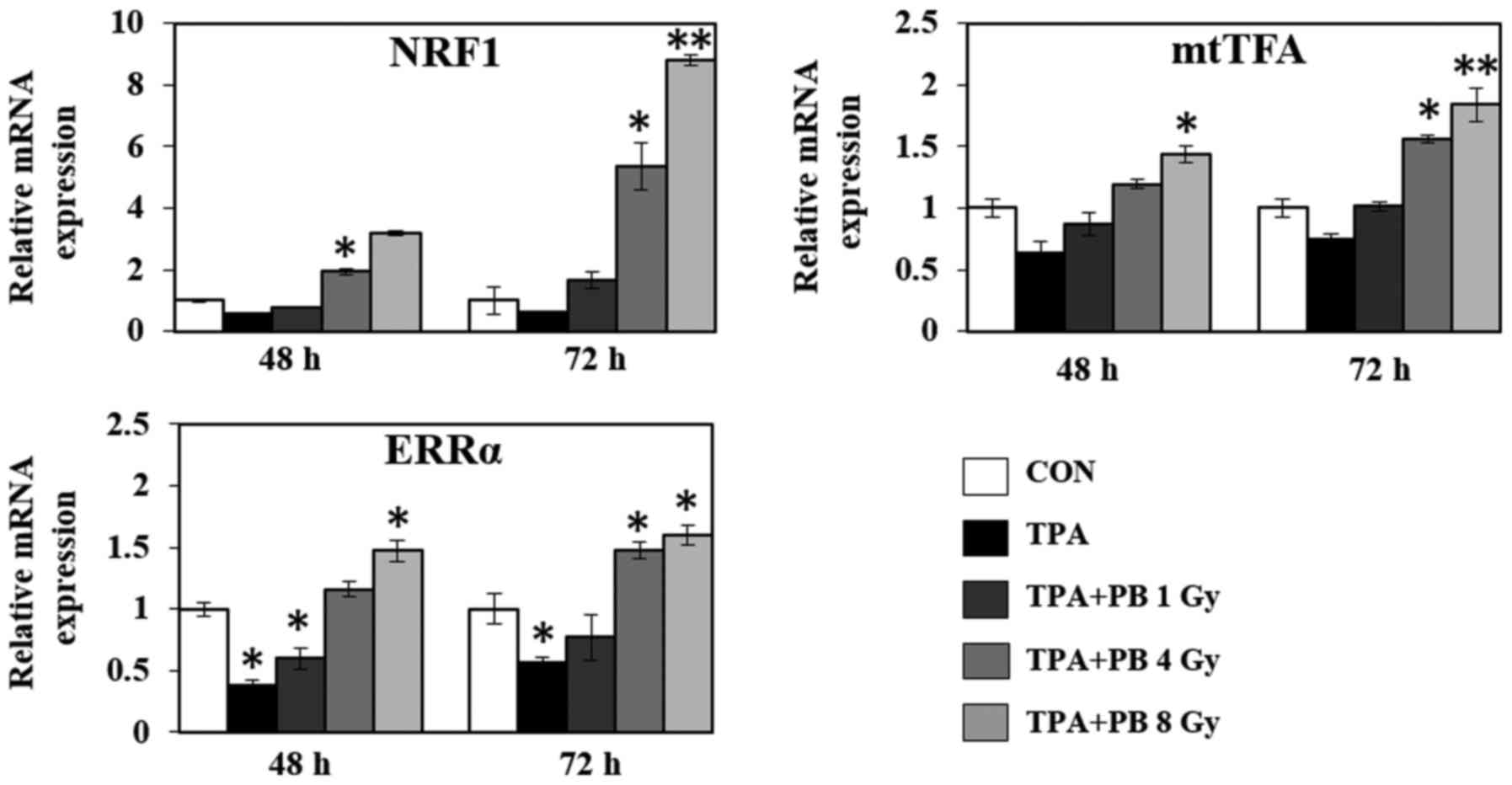
To further investigate the underlying regulatory
mechanism associated with mitochondrial biogenesis, we investigated
the mRNA expression of various genes involved in mitochondrial
transcription, mitochondrial dynamics, the electron transport
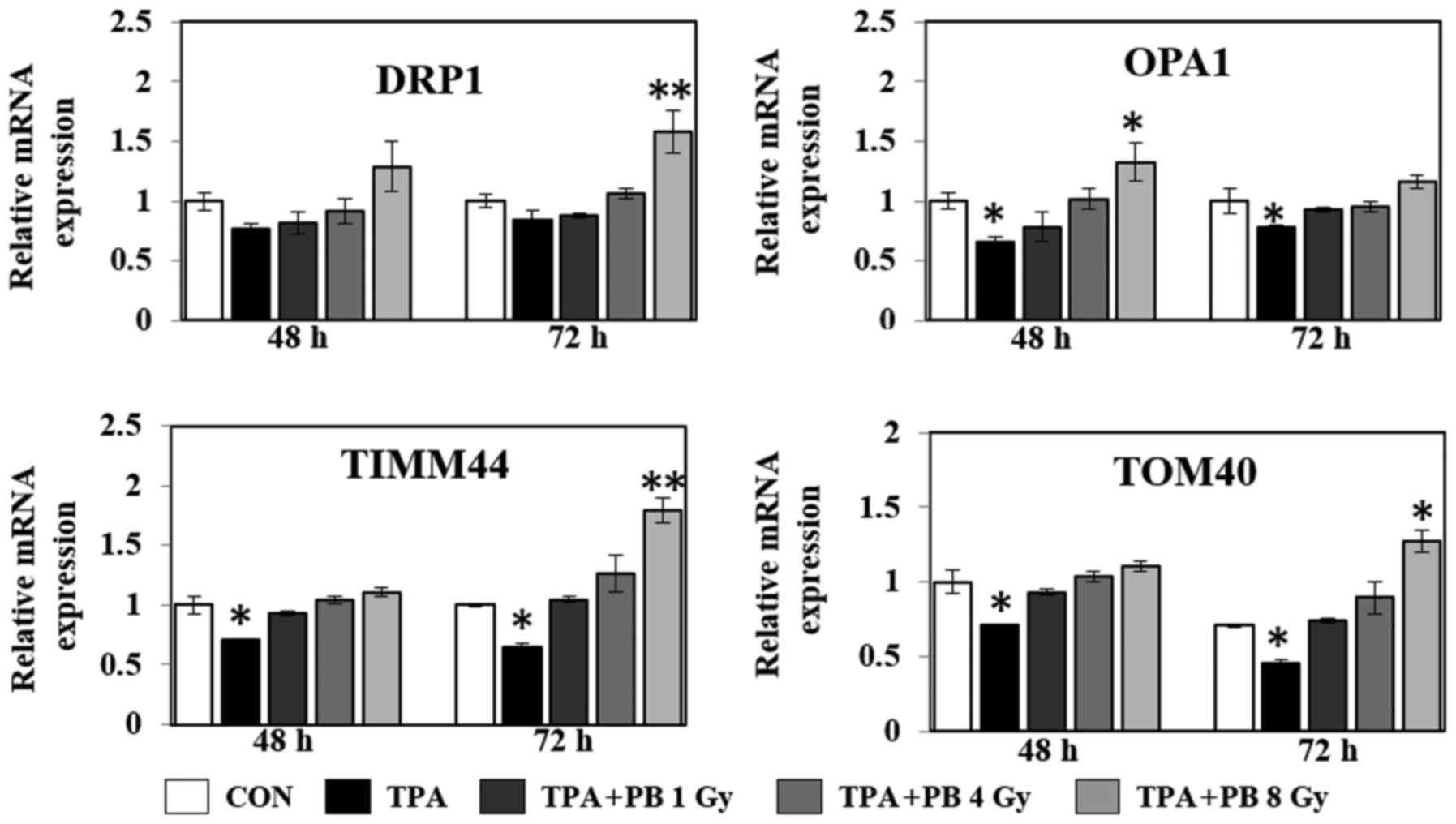
chain, ATP synthesis and fatty acid oxidation. As shown in Fig. 4, proton beam irradiation increased
the gene expression of the transcriptional co-activators NRF1,
mtTFA and ERRα in a dose- and time-dependent manner. At the same
time, proton beam irradiation increased the gene expression of
dynamin-related protein 1 (DRP1; a GTPase that regulates
mitochondrial fission), optic atrophy 1 (OPA1; a dynamin-related
GTPase, which participates in mitochondrial fusion and
mitochondrial cristae remodeling), translocase of outer
mitochondrial membrane 40 (TOMM40), and translocase of inner
mitochondrial membrane 44 (TIMM44), which play a role in
mitochondrial protein import (Fig.
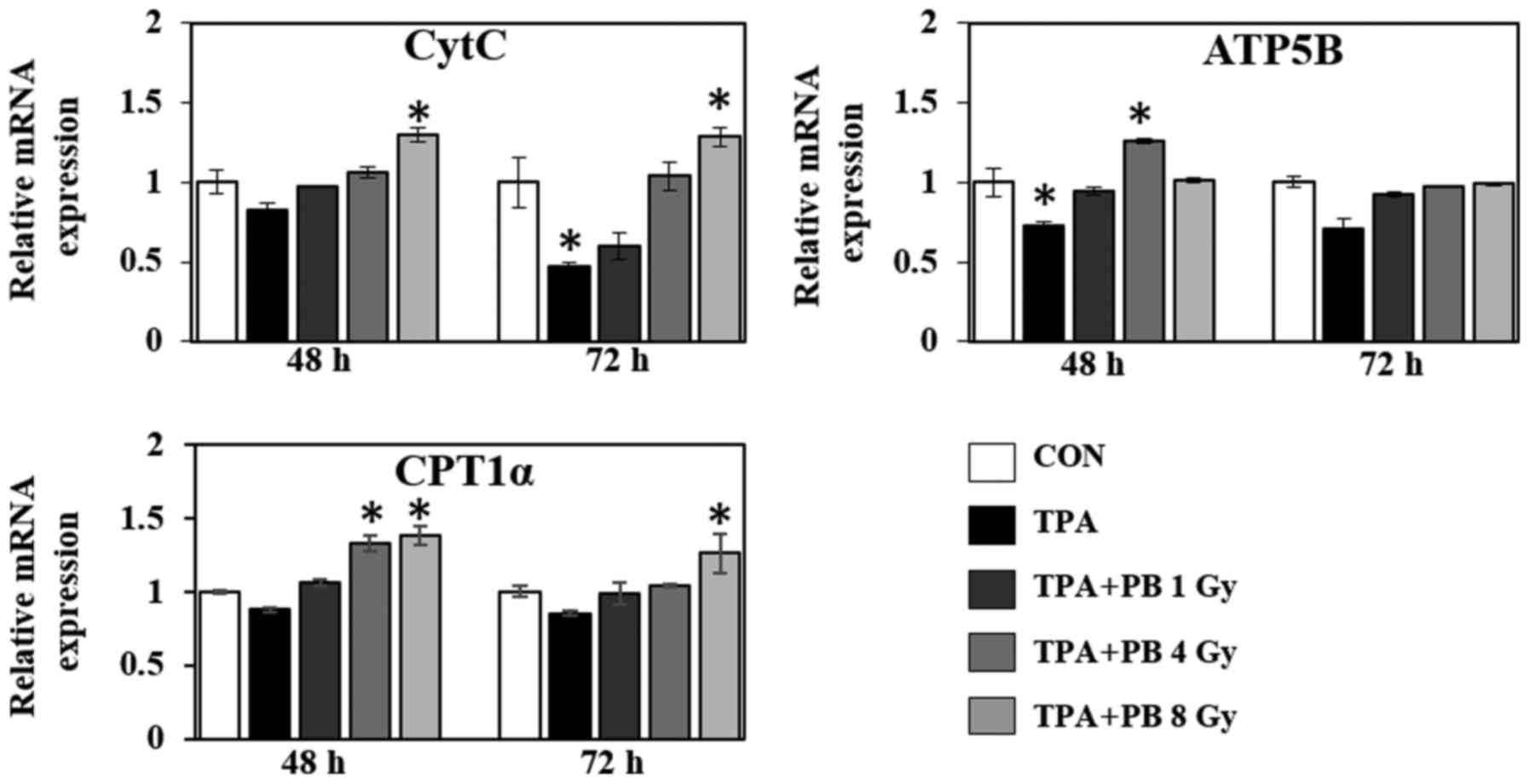
5). Moreover, in the context of mitochondrial function, proton
beam irradiation increased the gene expression of cytochrome
c (CytC), an essential component of the electron transport
chain, ATP synthase subunit β (ATP5B), a subunit of mitochondrial
ATP synthase, and carnitine palmitoyltransferase 1-α (CPT1-α), a
mitochondrial enzyme responsible for the β-oxidation of long-chain
fatty acids, in a dose- and time-dependent manner (Fig. 6). Therefore, these results indicate
that proton beam irradiation is an effective stimulator of
mitochondrial biogenesis and function in TPA-induced aggressive
HT-29 cells.
Discussion
Proton beam therapy (PBT) is a new form of
radiotherapy used to treat cancer. Compared with other
radiotherapies such as gamma and X-ray, PBT has better dose
distribution owing to its unique absorption spectrum in tissues,
known as the Bragg's peak, which allows the deposition of
devastating energy at the tumor while limiting the damage to normal
tissues (21). However, the
biological effects and detailed mechanism of proton beam
irradiation on cancer metabolism are still unclear, and further
studies are needed to determine the biological role of proton beam
irradiation for efficient application. In the present study, we
proposed that proton beam irradiation suppresses colon cancer
metastasis by stimulating mitochondrial biogenesis and inducing the
expression of mitochondrial transcription factors, functional
regulators, and dynamic-related molecules, and by upregulating AMPK
activity in TPA-induced aggressive HT-29 cells.
PGC-1α is the main regulator of mitochondrial
biogenesis, and acts through interactions with transcription
factors such as NRF1 and ERRα (4).
It is sensitive to cellular energy status, which is influenced by
cold, exercise and fasting and its activity is closely controlled
by several post-translational modifications such as phosphorylation
and acetylation. Both AMPK-mediated phosphorylation and
SIRT1-mediated deacetylation activate PGC-1α under energy
deprivation conditions (21).
NRF-1 regulates mtTFA expression, thereby mediating the increased
expression of nuclear mitochondrial genes with increased mtDNA
replication and expression (22,23).
ERRα acts as a co-transcription factor of PGC-1α, and regulates the
expression of genes involved in fatty acid oxidation, the TCA
cycle, mitochondrial biogenesis and dynamics and OXPHOS. ERRα also
acts both directly on genes involved in mitochondrial structural
components and enzymes (24).
Recent studies have revealed that the PGC-1α-mediated
transcriptional cascade suppresses invasion and migration in
different types of cancer cells. Integrative metabolomics and
transcriptomics using genetically engineered mouse models and
xenografts have shown that PGC-1α activates an ERRα-dependent
transcriptional network to induce a catabolic state and suppress
metastasis in prostate cancer (25). In melanoma, PGC-1α directly
increases the transcription of DNA-binding protein inhibitor (ID2),
which in turn binds to and inactivates transcription factor 4
(TCF4). TCF4 inactivation downregulates metastasis-related genes
including integrins and integrin-mediated signaling pathways that
play a role in invasion and migration (26). In the present study, we also found
that proton beam irradiation effectively increased mitochondrial
biogenesis by upregulating PGC-1α expression and its
co-transcriptional factors such as NRF1 and ERRα, as well as the
mitochondrial transcription factor mtTFA. Therefore, our findings
demonstrate that increased mitochondrial biogenesis, dynamics, and
function by a PGC-1α-mediated transcriptional cascade might be a
novel mechanism underlying the inhibitory effects of proton beam
irradiation on metastatic potential in TPA-induced aggressive HT-29
cells.
AMPK is a hetero-trimeric protein complex composed
of a catalytic subunit and two regulatory subunits; it plays an
essential role in regulating metabolic and energy homeostasis
(27). In particular, AMPK is well
known as the main regulator for lipid, cholesterol and glucose
metabolism in various tissues, such as the liver, muscles and
adipose tissues. Owing to this physiological role, AMPK has been
considered a key therapeutic target in patients with metabolic
diseases such as obesity and diabetes (28). Moreover, therapeutic manipulation
for targeting AMPK signal pathways using some diabetes drugs in
cancer patients shows the association between AMPK and several
tumor suppressors (29). Recent
studies have indicated that AMPK not only inhibits cancer cell
proliferation but also invasion. The siRNA-mediated knockdown and
pharmacologic activation of AMPK by OSU-53, a novel allosteric AMPK
activator, suppresses the epithelial-mesenchymal transition (EMT)
by modulating Akt-MDM2-Foxo3 signaling in breast and prostate
cancer cells (30). Treatment with
metformin, an AMPK activator, inhibits EMT in an AMPK/p53-dependent
manner in metastatic melanoma cells (31) and inhibits both angiogenesis and
metastatic spread in ovarian cancer (32). Similarly, Kim et al
(33) reported that AMPK
activation by berberine, another AMPK activator, suppresses human
melanoma cell migration by decreasing Erk activity and cytochrome
c oxidase subunit II expression. Notably, our findings
showed that proton beam irradiation increased TPA-inhibited AMPK
phosphorylation, which supports the increased expression of PGC1-α
and its co-transcription factors including NRF1 and ERRα, as well
as target genes including CytC, ATP5B and CPT-1α, which are
associated with OXPHOS components and mitochondrial functional
enzymes. Collectively, AMPK activation by proton beam irradiation
is decisive for mitochondrial biogenesis, and may exert a tumor
suppressor function and enhance mitochondrial function. However,
the specific role of AMPK activation by proton beam irradiation on
metastasis and mitochondrial function requires further study.
Mitochondria are highly dynamic organelles that are
continuously undergoing fusion and fission; they are necessary for
maintaining cellular physiological functions such as survival,
death and metabolic homeostasis (34). Although mitochondrial dysfunction
has been associated with tumorigenesis, the roles of mitochondrial
dynamics in metastasis are poorly understood. Altered mitochondrial
dynamics have been associated with changed mitochondrial physiology
and cell dysfunction (35), which
are involved in many human diseases such as cancer,
neurodegenerative diseases including Alzheimer's, Huntington's and
Parkinson's diseases, and cardiometabolic diseases including
diabetes and cardiomyopathy (36).
A recent study showed that human lung cancer cells have excess
mitochondrial fission and impaired mitochondrial fusion due to
imbalanced DRP1/MFN expression (37). Our findings suggest that proton
beam irradiation effectively regulates dysfunctional mitochondrial
dynamics in metastatic cancer by increasing the expression levels
of DRP1 and OPA1.
Over 99% of proteins involved in mitochondrial
biogenesis are imported from the cytoplasm. This mitochondrial
protein import process occurs through channel protein complexes
such as the translocase of the outer membrane (TOM) and the
translocases of the inner membrane (TIM) (38). Mitochondrial membrane channels such
as ion and protein channels are considered promising therapeutic
targets for cancer transformation owing to their correlation in
metabolic and apoptotic functions (39). In particular, many subunits of the
mitochondrial protein import complexes are dysregulated in cancer
cell mitochondria (40). The
mitochondrial protein import system subunits TIMM44 and TOM40 play
an important role in mitochondrial functions such as the import of
OXPHOS-related proteins and mitochondrial redox balance (41). Our findings show that proton beam
irradiation might be an effective regulator of TIMM44 and TOM40
expression.
In conclusion, our findings show that proton beam
irradiation increases mitochondrial biogenesis by modulating the
expression of the main mitochondrial transcription factors,
functional regulators and dynamic-related molecules by inducing
AMPK activity in TPA-induced aggressive HT-29 cells. We propose
that targeting mitochondrial biogenesis may be a novel inhibitory
mechanism for metastatic cancer metabolism.
Abbreviations:
|
TPA
|
12-O-tetradecanoylphorbol-13-acetate
|
|
PGC-1α
|
peroxisome proliferator-activated
receptor co-activator 1α
|
|
NRF
|
nuclear respiratory factor
|
|
mtTFA
|
mitochondrial transcription factor
A
|
|
ERRα
|
estrogen receptor-related receptor
alpha
|
|
AMPK
|
AMP-activated protein kinase
|
|
PBT
|
proton beam therapy
|
Acknowledgments
The present study was supported by the National
Nuclear R&D Program through the National Research Foundation
(NRF) of Korea funded by the Ministry of Science, ICT and Future
Planning (NRF- 2016M2B2A4910909).
References
|
1
|
Favoriti P, Carbone G, Greco M, Pirozzi F,
Pirozzi RE and Corcione F: Worldwide burden of colorectal cancer: A
review. Updates Surg. 68:7–11. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Steeg PS: Targeting metastasis. Nat Rev
Cancer. 16:201–218. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Suliman HB and Piantadosi CA:
Mitochondrial quality control as a therapeutic target. Pharmacol
Rev. 68:20–48. 2016. View Article : Google Scholar
|
|
4
|
Scarpulla RC, Vega RB and Kelly DP:
Transcriptional integration of mitochondrial biogenesis. Trends
Endocrinol Metab. 23:459–466. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Biswas G, Guha M and Avadhani NG:
Mitochondria-to-nucleus stress signaling in mammalian cells: Nature
of nuclear gene targets, transcription regulation, and induced
resistance to apoptosis. Gene. 354:132–139. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
van Waveren C, Sun Y, Cheung HS and Moraes
CT: Oxidative phosphorylation dysfunction modulates expression of
extracellular matrix–remodeling genes and invasion. Carcinogenesis.
27:409–418. 2006. View Article : Google Scholar
|
|
7
|
Wang X and Moraes CT: Increases in
mitochondrial biogenesis impair carcinogenesis at multiple levels.
Mol Oncol. 5:399–409. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Liu W, Beck BH, Vaidya KS, Nash KT, Feeley
KP, Ballinger SW, Pounds KM, Denning WL, Diers AR, Landar A, et al:
Metastasis suppressor KISS1 seems to reverse the Warburg effect by
enhancing mitochondrial biogenesis. Cancer Res. 74:954–963. 2014.
View Article : Google Scholar
|
|
9
|
Mohan R and Grosshans D: Proton therapy -
Present and future. Adv Drug Deliv Rev. 109:26–44. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Fukuda K, Okumura T, Abei M, Fukumitsu N,
Ishige K, Mizumoto M, Hasegawa N, Numajiri H, Ohnishi K, Ishikawa
H, et al: Long-term outcomes of proton beam therapy in patients
with previously untreated hepatocellular carcinoma. Cancer Sci.
108:497–503. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Hatayama Y, Nakamura T, Suzuki M, Azami Y,
Ono T, Yabuuchi T, Hayashi Y, Kimura K, Hirose K, Wada H, et al:
Clinical outcomes and prognostic factors of high-dose proton beam
therapy for peripheral stage I non-Small-cell lung cancer. Clin
Lung Cancer. 17:427–432. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Yamoah K and Johnstone PA: Proton beam
therapy: Clinical utility and current status in prostate cancer.
Onco Targets Ther. 9:5721–5727. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Romesser PB, Cahlon O, Scher E, Zhou Y,
Berry SL, Rybkin A, Sine KM, Tang S, Sherman EJ, Wong R, et al:
Proton beam radiation therapy results in significantly reduced
toxicity compared with intensity-modulated radiation therapy for
head and neck tumors that require ipsilateral radiation. Radiother
Oncol. 118:286–292. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Ogata T, Teshima T, Kagawa K, Hishikawa Y,
Takahashi Y, Kawaguchi A, Suzumoto Y, Nojima K, Furusawa Y and
Matsuura N: Particle irradiation suppresses metastatic potential of
cancer cells. Cancer Res. 65:113–120. 2005.PubMed/NCBI
|
|
15
|
Narang H, Kumar A, Bhat N, Pandey BN and
Ghosh A: Effect of proton and gamma irradiation on human lung
carcinoma cells: Gene expression, cell cycle, cell death,
epithelial-mesenchymal transition and cancer-stem cell trait as
biological end points. Mutat Res. 780:35–46. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Nam KS, Kim MK and Shon YH: Cancer
chemopreventive enzymes of human colorectal adenocarcinoma cells
irradiated with proton beams. J Korean Phys Soc. 52:945–948. 2008.
View Article : Google Scholar
|
|
17
|
Nam KS and Shon YH: Suppression of
metastatic potential in human colorectal adenocarcinoma cells
irradiated with proton beams. J Korean Phys Soc. 59:709–712. 2011.
View Article : Google Scholar
|
|
18
|
Ha BG, Park JE, Cho HJ, Lim YB and Shon
YH: Inhibitory effects of proton beam irradiation on integrin
expression and signaling pathway in human colon carcinoma HT29
cells. Int J Oncol. 46:2621–2628. 2015.PubMed/NCBI
|
|
19
|
Kim B, Bae H, Lee H, Lee S, Park JC, Kim
KR and Kim SJ: Proton beams inhibit proliferation of breast cancer
cells by altering DNA methylation status. J Cancer. 7:344–352.
2016. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Ha BG, Park JE, Cho HJ and Shon YH:
Stimulatory effects of balanced deep sea water on mitochondrial
biogenesis and function. PLoS One. 10:e01299722015. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
MacDonald SM, DeLaney TF and Loeffler JS:
Proton beam radiation therapy. Cancer Invest. 24:199–208. 2006.
View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Wenz T: Regulation of mitochondrial
biogenesis and PGC-1α under cellular stress. Mitochondrion.
13:134–142. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Zhang Y and Xiang Y: Molecular and
cellular basis for the unique functioning of Nrf1, an indispensable
transcription factor for maintaining cell homoeostasis and organ
integrity. Biochem J. 473:961–1000. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Villena JA and Kralli A: ERRalpha: A
metabolic function for the oldest orphan. Trends Endocrinol Metab.
19:269–276. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Torrano V, Valcarcel-Jimenez L, Cortazar
AR, Liu X, Urosevic J, Castillo-Martin M, Fernández-Ruiz S,
Morciano G, Caro-Maldonado A, Guiu M, et al: The metabolic
co-regulator PGC1α suppresses prostate cancer metastasis. Nat Cell
Biol. 18:645–656. 2016. View
Article : Google Scholar : PubMed/NCBI
|
|
26
|
Luo C, Lim JH, Lee Y, Granter SR, Thomas
A, Vazquez F, Widlund HR and Puigserver P: A PGC1α-mediated
transcriptional axis suppresses melanoma metastasis. Nature.
537:422–426. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Hardie DG, Schaffer BE and Brunet A: AMPK:
An energy-sensing pathway with multiple inputs and outputs. Trends
Cell Biol. 26:190–201. 2016. View Article : Google Scholar
|
|
28
|
Gasparrini M, Giampieri F, Alvarez Suarez
JM, Mazzoni L, Y Forbes Hernandez T, Quiles JL, Bullon P and
Battino M: AMPK as a new attractive therapeutic target for disease
prevention: The role of dietary compounds AMPK and disease
prevention. Curr Drug Targets. 17:865–889. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Shackelford DB and Shaw RJ: The LKB1-AMPK
pathway: Metabolism and growth control in tumour suppression. Nat
Rev Cancer. 9:563–575. 2009. View
Article : Google Scholar : PubMed/NCBI
|
|
30
|
Chou CC, Lee KH, Lai IL, Wang D, Mo X,
Kulp SK, Shapiro CL and Chen CS: AMPK reverses the mesenchymal
phenotype of cancer cells by targeting the Akt-MDM2-Foxo3a
signaling axis. Cancer Res. 74:4783–4795. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Cerezo M, Tichet M, Abbe P, Ohanna M,
Lehraiki A, Rouaud F, Allegra M, Giacchero D, Bahadoran P,
Bertolotto C, et al: Metformin blocks melanoma invasion and
metastasis development in AMPK/p53-dependent manner. Mol Cancer
Ther. 12:1605–1615. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Rattan R, Graham RP, Maguire JL, Giri S
and Shridhar V: Metformin suppresses ovarian cancer growth and
metastasis with enhancement of cisplatin cytotoxicity in vivo.
Neoplasia. 13:483–491. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Kim HS, Kim MJ, Kim EJ, Yang Y, Lee MS and
Lim JS: Berberine-induced AMPK activation inhibits the metastatic
potential of melanoma cells via reduction of ERK activity and COX-2
protein expression. Biochem Pharmacol. 83:385–394. 2012. View Article : Google Scholar
|
|
34
|
Westermann B: Mitochondrial fusion and
fission in cell life and death. Nat Rev Mol Cell Biol. 11:872–884.
2010. View
Article : Google Scholar : PubMed/NCBI
|
|
35
|
Detmer SA and Chan DC: Functions and
dysfunctions of mitochondrial dynamics. Nat Rev Mol Cell Biol.
8:870–879. 2007. View
Article : Google Scholar : PubMed/NCBI
|
|
36
|
Mishra P: Interfaces between mitochondrial
dynamics and disease. Cell Calcium. 60:190–198. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Zhao J, Zhang J, Yu M, Xie Y, Huang Y,
Wolff DW, Abel PW and Tu Y: Mitochondrial dynamics regulates
migration and invasion of breast cancer cells. Oncogene.
32:4814–4824. 2013. View Article : Google Scholar
|
|
38
|
Chacinska A, Koehler CM, Milenkovic D,
Lithgow T and Pfanner N: Importing mitochondrial proteins:
Machineries and mechanisms. Cell. 138:628–644. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Madamba SM, Damri KN, Dejean LM and
Peixoto PM: Mitochondrial ion channels in cancer transformation.
Front Oncol. 5:1202015. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Sotgia F, Whitaker-Menezes D,
Martinez-Outschoorn UE, Salem AF, Tsirigos A, Lamb R, Sneddon S,
Hulit J, Howell A and Lisanti MP: Mitochondria 'fuel' breast cancer
metabolism: Fifteen markers of mitochondrial biogenesis label
epithelial cancer cells, but are excluded from adjacent stromal
cells. Cell Cycle. 11:4390–4401. 2012. View
Article : Google Scholar : PubMed/NCBI
|
|
41
|
Baker MJ, Frazier AE, Gulbis JM and Ryan
MT: Mitochondrial protein-import machinery: Correlating structure
with function. Trends Cell Biol. 17:456–464. 2007. View Article : Google Scholar : PubMed/NCBI
|