|
1
|
Ilic M and Ilic I: Epidemiology of
pancreatic cancer. World J Gastroenterol. 22:9694–9705. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Morris JP IV, Wang SC and Hebrok M: KRAS,
Hedgehog, Wnt and the twisted developmental biology of pancreatic
ductal adenocarcinoma. Nat Rev Cancer. 10:683–695. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Ying H, Dey P, Yao W, Kimmelman AC,
Draetta GF, Maitra A and DePinho RA: Genetics and biology of
pancreatic ductal adenocarcinoma. Genes Dev. 30:355–385. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Bartel DP: MicroRNAs: Genomics,
biogenesis, mechanism, and function. Cell. 116:281–297. 2004.
View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Bloomston M, Frankel WL, Petrocca F,
Volinia S, Alder H, Hagan JP, Liu CG, Bhatt D, Taccioli C and Croce
CM: MicroRNA expression patterns to differentiate pancreatic
adenocarcinoma from normal pancreas and chronic pancreatitis. JAMA.
297:1901–1908. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Rachagani S, Macha MA, Menning MS, Dey P,
Pai P, Smith LM, Mo YY and Batra SK: Changes in microRNA (miRNA)
expression during pancreatic cancer development and progression in
a genetically engineered KrasG12D;Pdx1-Cre mouse (KC) model.
Oncotarget. 6:40295–40309. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Jin X, Sun Y, Yang H, Li J, Yu S, Chang X,
Lu Z and Chen J: Deregulation of the miR-193b-KRAS axis contributes
to impaired cell growth in pancreatic cancer. PLoS One.
10:e01255152015. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Keklikoglou I, Hosaka K, Bender C, Bott A,
Koerner C, Mitra D, Will R, Woerner A, Muenstermann E, Wilhelm H,
et al: MicroRNA-206 functions as a pleiotropic modulator of cell
proliferation, invasion and lymphangiogenesis in pancreatic
adenocarcinoma by targeting ANXA2 and KRAS genes. Oncogene.
34:4867–4878. 2015. View Article : Google Scholar :
|
|
9
|
Hu Y, Ou Y, Wu K, Chen Y and Sun W:
miR-143 inhibits the metastasis of pancreatic cancer and an
associated signaling pathway. Tumour Biol. 33:1863–1870. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Zhao WG, Yu SN, Lu ZH, Ma YH, Gu YM and
Chen J: The miR-217 microRNA functions as a potential tumor
suppressor in pancreatic ductal adenocarcinoma by targeting KRAS.
Carcinogenesis. 31:1726–1733. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Yu S, Lu Z, Liu C, Meng Y, Ma Y, Zhao W,
Liu J, Yu J and Chen J: miRNA-96 suppresses KRAS and functions as a
tumor suppressor gene in pancreatic cancer. Cancer Res.
70:6015–6025. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Wapinski O and Chang HY: Long noncoding
RNAs and human disease. Trends Cell Biol. 21:354–361. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Novikova IV, Hennelly SP, Tung CS and
Sanbonmatsu KY: Rise of the RNA machines: Exploring the structure
of long non-coding RNAs. J Mol Biol. 425:3731–3746. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Tripathi V, Ellis JD, Shen Z, Song DY, Pan
Q, Watt AT, Freier SM, Bennett CF, Sharma A, Bubulya PA, et al: The
nuclear-retained noncoding RNA MALAT1 regulates alternative
splicing by modulating SR splicing factor phosphorylation. Mol
Cell. 39:925–938. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Wang KC, Yang YW, Liu B, Sanyal A,
Corces-Zimmerman R, Chen Y, Lajoie BR, Protacio A, Flynn RA, Gupta
RA, et al: A long noncoding RNA maintains active chromatin to
coordinate homeotic gene expression. Nature. 472:120–124. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Muers M: RNA: Genome-wide views of long
non-coding RNAs. Nat Rev Genet. 12:742–743. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Chen X and Yan GY: Novel human
lncRNA-disease association inference based on lncRNA expression
profiles. Bioinformatics. 29:2617–2624. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Harries LW: Long non-coding RNAs and human
disease. Biochem Soc Trans. 40:902–906. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Tahira AC, Kubrusly MS, Faria MF, Dazzani
B, Fonseca RS, Maracaja-Coutinho V, Verjovski-Almeida S, Machado MC
and Reis EM: Long noncoding intronic RNAs are differentially
expressed in primary and metastatic pancreatic cancer. Mol Cancer.
10:1412011. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Kim K, Jutooru I, Chadalapaka G, Johnson
G, Frank J, Burghardt R, Kim S and Safe S: HOTAIR is a negative
prognostic factor and exhibits pro-oncogenic activity in pancreatic
cancer. Oncogene. 32:1616–1625. 2013. View Article : Google Scholar
|
|
21
|
Li Z, Zhao X, Zhou Y, Liu Y, Zhou Q, Ye H,
Wang Y, Zeng J, Song Y, Gao W, et al: The long non-coding RNA
HOTTIP promotes progression and gemcitabine resistance by
regulating HOXA13 in pancreatic cancer. J Transl Med. 13:842015.
View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Pang EJ, Yang R, Fu XB and Liu YF:
Overexpression of long non-coding RNA MALAT1 is correlated with
clinical progression and unfavorable prognosis in pancreatic
cancer. Tumour Biol. 36:2403–2407. 2015. View Article : Google Scholar
|
|
23
|
Peng W, Gao W and Feng J: Long noncoding
RNA HULC is a novel biomarker of poor prognosis in patients with
pancreatic cancer. Med Oncol. 31:3462014. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Liu JH, Chen G, Dang YW, Li CJ and Luo DZ:
Expression and prognostic significance of lncRNA MALAT1 in
pancreatic cancer tissues. Asian Pac J Cancer Prev. 15:2971–2977.
2014. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
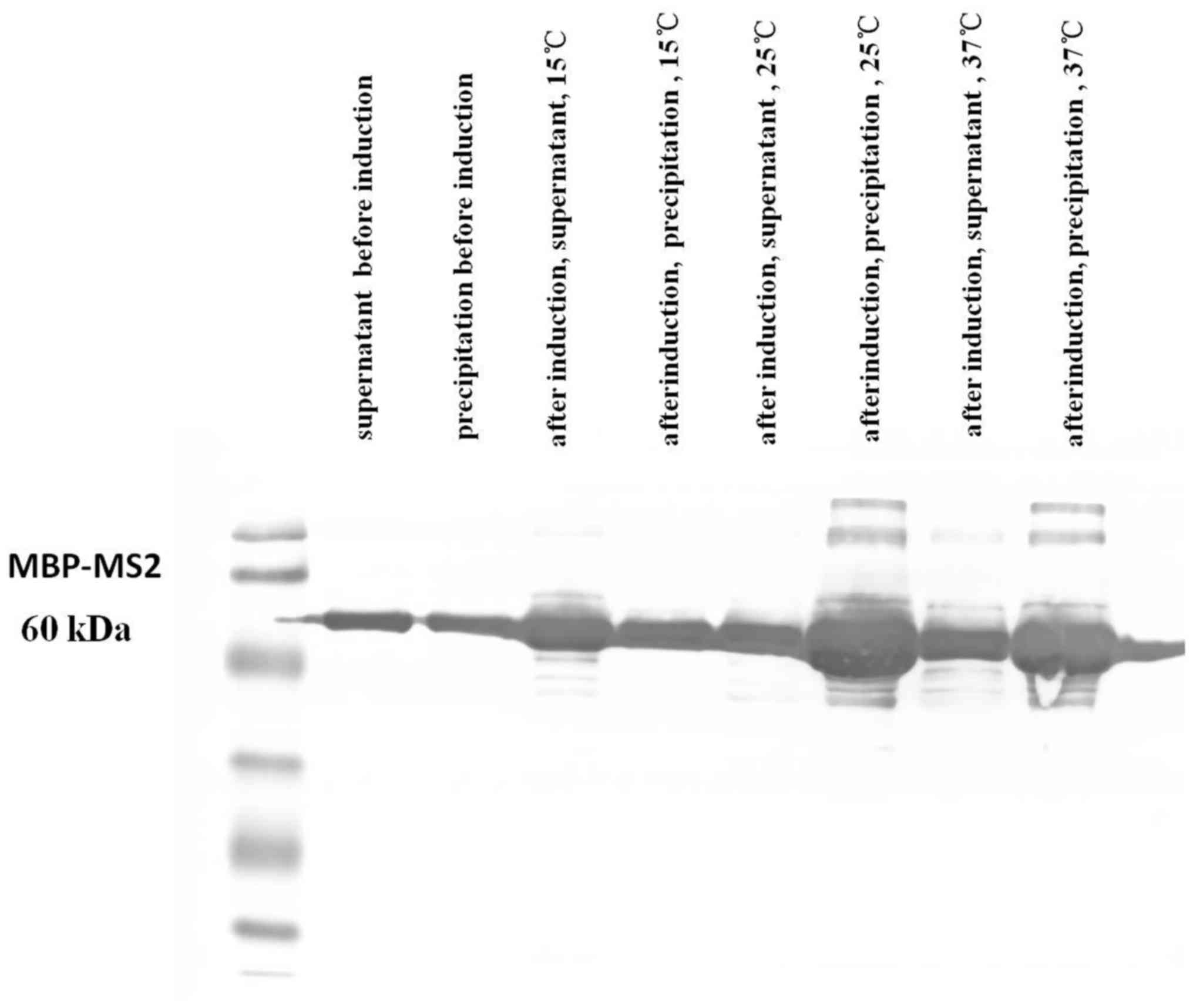
Kuzmanovic DA, Elashvili I, Wick C,
O’Connell C and Krueger S: The MS2 coat protein shell is likely
assembled under tension: A novel role for the MS2 bacteriophage A
protein as revealed by small-angle neutron scattering. J Mol Biol.
355:1095–1111. 2006. View Article : Google Scholar
|
|
26
|
Batey RT and Kieft JS: Improved native
affinity purification of RNA. RNA. 13:1384–1389. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Gong C, Popp MW and Maquat LE: Biochemical
analysis of long non-coding RNA-containing ribonucleoprotein
complexes. Methods. 58:88–93. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Zhou Z and Reed R: Purification of
functional RNA-protein complexes using MS2-MBP. Curr Protoc Mol
Biol Chapter. 27:32003.
|
|
29
|
Corcoran CP, Rieder R, Podkaminski D,
Hofmann B and Vogel J: Use of aptamer tagging to identify in vivo
protein binding partners of small regulatory RNAs. Methods Mol
Biol. 905:177–200. 2012.PubMed/NCBI
|
|
30
|
Said N, Rieder R, Hurwitz R, Deckert J,
Urlaub H and Vogel J: In vivo expression and purification of
aptamer-tagged small RNA regulators. Nucleic Acids Res.
37:e1332009. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Bardwell VJ and Wickens M: Purification of
RNA and RNA-protein complexes by an R17 coat protein affinity
method. Nucleic Acids Res. 18:6587–6594. 1990. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
SenGupta DJ, Zhang B, Kraemer B, Pochart
P, Fields S and Wickens M: A three-hybrid system to detect
RNA-protein interactions in vivo. Proc Natl Acad Sci USA.
93:8496–8501. 1996. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(−ΔΔC(T)) Method. Methods. 25:402–408. 2001. View Article : Google Scholar
|
|
34
|
Cai X, Hagedorn CH and Cullen BR: Human
microRNAs are processed from capped, polyadenylated transcripts
that can also function as mRNAs. RNA. 10:1957–1966. 2004.
View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Lee Y, Kim M, Han J, Yeom KH, Lee S, Baek
SH and Kim VN: MicroRNA genes are transcribed by RNA polymerase II.
EMBO J. 23:4051–4060. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Wu Y, Wei B, Liu H, Li T and Rayner S:
MiRPara: A SVM-based software tool for prediction of most probable
microRNA coding regions in genome scale sequences. BMC
Bioinformatics. 12:1072011. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Pylayeva-Gupta Y, Grabocka E and Bar-Sagi
D: RAS oncogenes: Weaving a tumorigenic web. Nat Rev Cancer.
11:761–774. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Rozenblum E, Schutte M, Goggins M, Hahn
SA, Panzer S, Zahurak M, Goodman SN, Sohn TA, Hruban RH, Yeo CJ, et
al: Tumor-suppressive pathways in pancreatic carcinoma. Cancer Res.
57:1731–1734. 1997.PubMed/NCBI
|
|
39
|
Jiao LR, Frampton AE, Jacob J, Pellegrino
L, Krell J, Giamas G, Tsim N, Vlavianos P, Cohen P, Ahmad R, et al:
MicroRNAs targeting oncogenes are down-regulated in pancreatic
malignant transformation from benign tumors. PLoS One.
7:e320682012. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Xu B, Niu X, Zhang X, Tao J, Wu D, Wang Z,
Li P, Zhang W, Wu H, Feng N, et al: miR-143 decreases prostate
cancer cells proliferation and migration and enhances their
sensitivity to docetaxel through suppression of KRAS. Mol Cell
Biochem. 350:207–213. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Johnson SM, Grosshans H, Shingara J, Byrom
M, Jarvis R, Cheng A, Labourier E, Reinert KL, Brown D and Slack
FJ: RAS is regulated by the let-7 microRNA family. Cell.
120:635–647. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Kumar MS, Erkeland SJ, Pester RE, Chen CY,
Ebert MS, Sharp PA and Jacks T: Suppression of non-small cell lung
tumor development by the let-7 microRNA family. Proc Natl Acad Sci
USA. 105:3903–3908. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Boyerinas B, Park SM, Hau A, Murmann AE
and Peter ME: The role of let-7 in cell differentiation and cancer.
Endocr Relat Cancer. 17:F19–F36. 2010. View Article : Google Scholar
|
|
44
|
Torrisani J, Bournet B, du Rieu MC,
Bouisson M, Souque A, Escourrou J, Buscail L and Cordelier P: let-7
MicroRNA transfer in pancreatic cancer-derived cells inhibits in
vitro cell proliferation but fails to alter tumor progression. Hum
Gene Ther. 20:831–844. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Kent OA, Chivukula RR, Mullendore M,
Wentzel EA, Feldmann G, Lee KH, Liu S, Leach SD, Maitra A and
Mendell JT: Repression of the miR-143/145 cluster by oncogenic Ras
initiates a tumor-promoting feed-forward pathway. Genes Dev.
24:2754–2759. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Tsang WP and Kwok TT: The
miR-18a* microRNA functions as a potential tumor
suppressor by targeting on K-Ras. Carcinogenesis. 30:953–959. 2009.
View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Shin KH, Bae SD, Hong HS, Kim RH, Kang MK
and Park NH: miR-181a shows tumor suppressive effect against oral
squamous cell carcinoma cells by downregulating K-ras. Biochem
Biophys Res Commun. 404:896–902. 2011. View Article : Google Scholar
|
|
48
|
Wang XR, Luo H, Li HL, Cao L, Wang XF, Yan
W, Wang YY, Zhang JX, Jiang T, Kang CS, et al Chinese Glioma
Cooperative Group (CGCG): Overexpressed let-7a inhibits glioma cell
malignancy by directly targeting K-ras, independently of PTEN.
Neuro-oncol. 15:1491–1501. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
49
|
Chen KJ, Hou Y, Wang K, Li J, Xia Y, Yang
XY, Lv G, Xing XL and Shen F: Reexpression of Let-7g microRNA
inhibits the proliferation and migration via K-Ras/HMGA2/snail axis
in hepatocellular carcinoma. Biomed Res Int.
2014:7424172014.PubMed/NCBI
|
|
50
|
Zhu L, Wang Z, Fan Q, Wang R and Sun Y:
microRNA-27a functions as a tumor suppressor in esophageal squamous
cell carcinoma by targeting KRAS. Oncol Rep. 31:280–286. 2014.
View Article : Google Scholar
|
|
51
|
Liao WT, Ye YP, Zhang NJ, Li TT, Wang SY,
Cui YM, Qi L, Wu P, Jiao HL, Xie YJ, et al: MicroRNA-30b functions
as a tumour suppressor in human colorectal cancer by targeting
KRAS, PIK3CD and BCL2. J Pathol. 232:415–427. 2014. View Article : Google Scholar
|
|
52
|
Chen X, Guo X, Zhang H, Xiang Y, Chen J,
Yin Y, Cai X, Wang K, Wang G, Ba Y, et al: Role of miR-143
targeting KRAS in colorectal tumorigenesis. Oncogene. 28:1385–1392.
2009. View Article : Google Scholar : PubMed/NCBI
|
|
53
|
Tanic M, Yanowsky K, Rodriguez-Antona C,
Andrés R, Márquez-Rodas I, Osorio A, Benitez J and Martinez-Delgado
B: Deregulated miRNAs in hereditary breast cancer revealed a role
for miR-30c in regulating KRAS oncogene. PLoS One. 7:e388472012.
View Article : Google Scholar : PubMed/NCBI
|
|
54
|
Yang F, Zhang L, Huo XS, Yuan JH, Xu D,
Yuan SX, Zhu N, Zhou WP, Yang GS, Wang YZ, et al: Long noncoding
RNA high expression in hepatocellular carcinoma facilitates tumor
growth through enhancer of zeste homolog 2 in humans. Hepatology.
54:1679–1689. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
55
|
Prensner JR, Iyer MK, Balbin OA,
Dhanasekaran SM, Cao Q, Brenner JC, Laxman B, Asangani IA, Grasso
CS, Kominsky HD, et al: Transcriptome sequencing across a prostate
cancer cohort identifies PCAT-1, an unannotated lincRNA implicated
in disease progression. Nat Biotechnol. 29:742–749. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
56
|
Prensner JR, Chen W, Iyer MK, Cao Q, Ma T,
Han S, Sahu A, Malik R, Wilder-Romans K, Navone N, et al: PCAT-1, a
long noncoding RNA, regulates BRCA2 and controls homologous
recombination in cancer. Cancer Res. 74:1651–1660. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
57
|
Mourtada-Maarabouni M, Pickard MR, Hedge
VL, Farzaneh F and Williams GT: GAS5, a non-protein-coding RNA,
controls apoptosis and is downregulated in breast cancer. Oncogene.
28:195–208. 2009. View Article : Google Scholar
|
|
58
|
Lu KH, Li W, Liu XH, Sun M, Zhang ML, Wu
WQ, Xie WP and Hou YY: Long non-coding RNA MEG3 inhibits NSCLC
cells proliferation and induces apoptosis by affecting p53
expression. BMC Cancer. 13:4612013. View Article : Google Scholar : PubMed/NCBI
|
|
59
|
Qin R, Chen Z, Ding Y, Hao J, Hu J and Guo
F: Long non-coding RNA MEG3 inhibits the proliferation of cervical
carcinoma cells through the induction of cell cycle arrest and
apoptosis. Neoplasma. 60:486–492. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
60
|
Jiao F, Hu H, Yuan C and Wang L, Jiang W,
Jin Z, Guo Z and Wang L: Elevated expression level of long
noncoding RNA MALAT-1 facilitates cell growth, migration and
invasion in pancreatic cancer. Oncol Rep. 32:2485–2492. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
61
|
Lu X, Fang Y, Wang Z, Xie J, Zhan Q, Deng
X, Chen H, Jin J, Peng C, Li H, et al: Downregulation of gas5
increases pancreatic cancer cell proliferation by regulating CDK6.
Cell Tissue Res. 354:891–896. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
62
|
Sun YW, Chen YF, Li J, Huo YM, Liu DJ, Hua
R, Zhang JF, Liu W, Yang JY, Fu XL, et al: A novel long non-coding
RNA ENST00000480739 suppresses tumour cell invasion by regulating
OS-9 and HIF-1α in pancreatic ductal adenocarcinoma. Br J Cancer.
111:2131–2141. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
63
|
Ma C, Nong K, Zhu H, Wang W, Huang X, Yuan
Z and Ai K: H19 promotes pancreatic cancer metastasis by
derepressing let-7′s suppression on its target HMGA2-mediated EMT.
Tumour Biol. 35:9163–9169. 2014. View Article : Google Scholar : PubMed/NCBI
|