Introduction
Bone cancer is one of the most lethal malignancies
and the prognosis remains poor due to a lack of effective
therapeutics that significantly improve the quality of life or
survival rate of the patients (1,2).
Bone cancer occurs in the skeleton and its members, but currently
the mechanism of its initiation remains unknown (3). Malignant bone cancer is a typical
systemic malignant disease that commonly causes symptoms including
bone and joint pain or swelling and fatigue (4,5). In
recent years, new strategies have been proposed; however, the
overall prognosis for patients with osteosarcoma has not markedly
improved (6,7). Additionally, it has been previously
reported that osteosarcoma cells are resistant to apoptosis
(8–10). Furthermore, apoptotic resistance
has become a major challenge in cancer therapy due to resistance of
tumor cells through various molecular mechanisms (11,12).
Therefore, development of more effective target therapies is
required to improve for patient outcomes.
B-cell-specific Moloney murine leukemia virus
integration site 1 protein (Bmi-1) represses tumor suppressor gene
expression by forming complexes with multiple other polycomb group
(PcG) family members. Elevated Bmi-1 expression is associated with
dysplastic cell transformation during carcinogenesis and is
required for cancer cell replication and survival (13). Bmi-1 has also been identified as a
transcription factor with a prognostic role in several malignancies
(14,15). Additionally, previous reports have
indicated that Bmi-1 has an important role in the development and
progression of cancer, and essentially functions as an oncogene
(16). Furthermore, overexpression
of Bmi-1 promoted apoptosis resistance of cancer cells via
activation of nuclear factor-κB (NF-κB) signaling (17). By contrast, Bmi-1 knockdown induced
cell-cycle arrest and upregulated the expression tumor-suppressive
genes, including homeobox C13 (HOXC13), cyclin dependent kinase
inhibitor 2A (p16INK4a), and HOXA9 (18).
Notably, previous reports have suggested that Bmi-1
promotes the aggressiveness of glioma via activation of the
NF-κB/matrix metalloproteinase 9 (MMP-9) signaling pathway
(19). In addition, another report
demonstrated that NF-κB promotes the generation of
CD133+, Bmi-1 keratinocytes and the growth of xenograft
tumors in mice (20). Furthermore,
a novel NF-κB/MMP-3 signal pathway involved in the aggressiveness
of glioma promoted by Bmi-1 has been investigated in a previous
report, and the results demonstrated that Bmi-1 promotes glioma
cell migration and invasion via NF-κB-mediated upregulation of
MMP-3 (21). Therefore, these
reports indicated that Bmi-1 upregulation may be associated with
tumorigenesis through regulation of the NF-κB signaling
pathway.
Aberrant activation of NF-κB is observed in the
majority of human cancers (22,23).
Evidence has suggested that poor survival rate and insufficient
outcomes of patients with bone cancer are associated with
aberrantly activated NF-κB signaling (24,25).
The indicator of NF-κB activation, p65, has also been demonstrated
to be highly active in clinical specimens of bone cancer (25). The NF-κB signaling pathway is
involved in apoptosis resistance induced by chemotherapy, and
enhances tumors cell survival, proliferation, survival, invasion
and angiogenesis (26). Developing
novel molecules that regulate aberrant activation of NF-κB
signaling pathway may be beneficial for clinical osteosarcoma as
targeted therapeutics.
In this study, it was observed that knockdown of
Bmi-1 inhibited the migration and invasion of osteosarcoma cells.
The data demonstrated that Bim-1 stimulation of the invasive
phenotype was mechanistically associated with activation of NF-κB
and subsequent upregulation and activation of MMP-9. A full-length
antibody targeting Bmi-1 (AbBmi-1) was produced, which inhibited
migration and invasion of osteosarcoma in vitro and in
vivo through inactivation of the NF-κB signaling pathway. In
conclusion, the findings provided novel evidence that targeting
Bmi-1 inhibited the progression of bone cancer and that Bmi-1 may
represent a novel therapeutic target for bone cancer treatment.
Materials and methods
Ethics statement
This study was performed in strict accordance with
the recommendations in the Guide for the Care and Use of Laboratory
Animals of Taihe Hospital Affiliated to Hubei University of
Medicine (Shiyan, China). All experimental protocols and animal
procedures were performed in accordance with National Institutes of
Health guidelines and approved by the Committee on the Ethics of
Animal Experiments Defence Research of Taihe Hospital. All surgery
and euthanasia were performed with efforts to minimize
suffering.
Cells and reagents
The MG-63 bone tumor cell line and MC3T3-E1 normal
human osteoblast cells were purchased from American Type Culture
Collection (Manassas, VA, USA). MG-63 cells were cultured in 1640
medium (Gibco; Thermo Fisher Scientific, Inc., Waltham, MA, USA)
supplemented with 10% fetal bovine serum (Invitrogen; Thermo Fisher
Scientific, Inc.). MC3T3-E1 cells were cultured in Dulbecco's
modified Eagle's medium (DMEM; Sigma-Aldrich; Merck KGaA,
Darmstadt, Germany) medium supplemented with 10% fetal calf serum
(Gibco; Thermo Fisher Scientific, Inc.). All cells were cultured in
a 37°C humidified atmosphere of 5% CO2 and treated with
cisplatin (25 µM) or MMP-9 inhibitor (20 nM; sc-311437; Santa Cruz
Biotechnology, Inc., Dallas, TX, USA) for 3 days.
Tissue specimens and patient
information
Paraffin-embedded, archived normal samples obtained
from donors who did not suffer from bone-related disease, and
osteosarcoma specimens histopathologically diagnosed at Taihe
Hospital affiliated to Hubei University of Medicine between June
2007 and July 2013 were obtained. The use of the clinical specimens
was approved by the local Institutional Review Board of Taihe
Hospital affiliated to Hubei University of Medicine.
Bmi-1 overexpression
The Bmi-1 overexpression experiment was performed on
pCDH-EF1-MCS-T2A-copGFP vector (CD521A-1; System Biosciences, Inc.,
Palo Alto, CA, USA). First PCR was used to synthesize the full
length of Bmi-1 CDS region, with 293 cell cDNA as a template. The
primer sequence was: Bmi-1 sense: 5′-GAGGGTACTTCATTGATGCCAC-3′
Bmi-1 antisense: 5′-CCAGTTCTCCAGCATTTGTCAG-3′), in the meantime,
the restriction enzyme site was also cloned, and the Bmi-1 CDS
region ligated to the PCDH vector, with Sanger sequencing used to
confirm the fragment in the plasmid. Following the acquisition of
the PCDH-Bmi-1 expression vector, the vector was transfected into
MG-63 using Lipofectamine 2000 (Invitrogen, Thermo Fisher
Scientific, Inc.) according to the manufacturer's protocol.
Briefly, MG-63 cells were seeded onto 24-well plates
(2×105 per well) and, following attachment, mixed with 5
µg plasmid with 50 µl medium without serum and 2 µl Lipofectamine
2000 with 50 µl medium without serum. They were then incubated for
5 min at room temperature and, after 48 h, the cells were used for
the next experiment.
Production of full-length antibody
AbBmi-1
The single chain variable fragments of the mouse
anti-human Bmi-1 antibody were cloned and ligated into the pET-27b
vector (pET-27bBmi-1), purchased from Addgene, Inc. (Cambridge, MA,
USA). First the cDNA clone primers were designed by Takara
Biotechnology Co., Ltd. (Dalian, China), then 293 cell cDNA was
used as a template and PCR used to synthesize the Bmi-1. In the
meantime, the restriction enzyme site was cloned: Primer sequences
were sense, 5-GCTGTACAAGTCCGGACTCAGAT-3; antisense,
5-CCGGATCTAGATAACTGATCATAA-3. Then the PCR fragments were cloned to
the pET-27b vector and Sanger sequencing used to confirm the
fragment. The constant domain heavy chain linked Fc and light chain
were inserted into the pET-27bBmi-1 vector. Subsequently,
full-length antibody targeting of Bmi-1 was termed AbBmi-1. The
pET-27bBmi-1 vector was transfected into the E. coli
Rossetta (DE3) using electroporation. IPTG was added to a final
concentration of 0.5 mM, when the absorbance reached 0.6 at 600 nm
wavelength as measured by a microplate reader
(Varioskan® Flash Spectral Scanning Multimode Readers,
Thermo Fisher Scientific, Inc.). Finally, cells were spun down,
disrupted and dissolved in 400 W Ultrasonic for 45 times, 8 sec
each time with intervals of 15 sec. The supernatant and precipitate
were collected. The supernatant was filtered (45 µm) and purified
using the ÄKTAprime plus kit (GE Healthcare Bio-Sciences,
Pittsburgh, PA, USA). The obtained AbBmi-1 was further purified by
gel filtration chromatography, then the concentration of antibody
is quantified by Pierce BCA Protein Assay kit (Thermo Fisher
Scientific, Inc.). The synthesis of the fusion protein, Bmi-1- CPPs
(cell penetrating peptides) antibody was performed by LifeTein
Company (Beijing, China).
MTT cytotoxicity
MG-63 cells (3,000 per well) were incubated with 0.5
µg/ml Bmi-1 antibody (produced by our laboratory), PBS or AbBmi-1
in 96-well plates for 72 h in triplicate for each condition. Then,
20 µl MTT (5 mg/ml) in PBS was added to each well and the cells
were incubated for a further 4 h. The entire medium was removed and
100 µl dimethyl sulfoxide was added into the wells to solubilize
the crystals. The optical density was measured by a Bio-Rad reader
at wavelength of 450 nm (Bio-Rad Laboratories, Inc., Hercules, CA,
USA).
Cell migration, wound and invasion
assays
For the migration assay, MG-63 cells
(2×106) were treated with Bim-1, PBS or AbBim-1 and
incubated for 72 h using a control insert (BD Biosciences), then
the cells were seeded onto a 24-well plate, and 10 µl pipette tips
used to scratch a line. Then, 12 h afterwards, cell migration
distance and velocity were calculated. For the invasion assay,
Bim-1, PBS or AbBim-1-treated cells were suspended at a density of
1×105 in 200 µl in serum-free DMEM and 500 µl 10% FBS +
DMEM medium added to the lower chamber of BD BioCoat Matrigel
Invasion Chambers (BD Biosciences). The MG-63 cells in serum-free
medium were added to the upper chamber according to the
manufacturer's protocols. After 12 h, cells were stained with 0.1%
crystal violet for 30 min in room temperature. The tumor cells
invasion and migration were counted in at least three random
stained fields under the light microscope (Olympus 1X71; Olympus
Corporation, Tokyo, Japan).
Reverse transcription-quantitative
polymerase chain reaction (RT-qPCR) analysis
Total RNA was obtained from MG-63 and MC3T3-E1 cells
by using RNAeasy Mini kit (Qiagen, Inc., Valencia, CA, USA). Bmi-1
expression levels in MG-63 and MC3T3-E1cells were determined by
applying RT-qPCR. RT-qPCR was performed using PrimeScript RT
reagent kit with gDNA Eraser (Takara Biotechnology Co., Ltd.),
according to the manufacturer's protocol. First 5X gDNA Eraser
Buffer, gDNA Eraser and total RNA were mixed together, at 42°C for
2 min, then mixed with 5X PrimeScript Buffer 2, PrimeScript RT
Enzyme Mix 1 and RT Primer Mix at 37°C for 15 min giving a reaction
volume of 15 µl in each well. All the forward and reverse primers
were synthesized by Invitrogen (Thermo Fisher Scientific, Inc.).
mRNA was detected by SYBR Premix Ex Taq TM II (Takara Biotechnology
Co., Ltd.); primer sequences: Bmi-1 sense:
5′-TCATCCTTCTGCTGATGCTG-3′, Bmi-1 antisense:
5′-CCGATCCAATCTGTTCTGGT-3′; GAPDH sense:
5′-TATGCTCTCCTCATGCATTG-3′, GAPDH antisense
5′-GGGACGACCTTCGATCTACC-3′. The thermocycling conditions were in
accordance with the manufacturer's protocols (7500 fast instrument:
Applied Biosystems; Thermo Fisher Scientific, Inc.): 95°C for 30
sec, 95°C for 3 sec and 60°C for 30 sec for 40 cycles. Relative
mRNA expression level changes were calculated by 2−ΔΔCq,
RT-qPCR was performed as described previously (27). The results are expressed as the
n-fold change compared with control. Every experiment was
replicated 3 times.
Western blot analysis
MG-63 cells were treated with 30 µM NF-κB inhibitor
(JSH-23, 20 µM) and AbBim-1 (0.5 µg/ml) for 72 h, PBS or AbBim-1
and homogenized in lysate buffer (RIPA Beyotime Institute of
Biotechnology, Suzhou, China) containing protease-inhibitor and
were centrifuged at 7,168 × g and 4°C for 10 min. The supernatant
was used for analysis of proteins. For detection of target
proteins, transmembrane proteins were extracted by using
Transmembrane Protein Extraction kit (Qiagen, Inc.) according to
the manufacturer's protocols. Protein concentration was calculated
using a BCA kit (Thermo Fisher Scientific, Inc.) and SDS-PAGE
assays were performed as previously described (28). For western blotting, the primary
antibodies were Bmi-1 (6964), p65 (8242), p84 (9172), IKK-β (8943),
IkBα (9242) and GAPDH (5174), all at 1:1,000, and the secondary
antibodies anti mouse IgG, HRP-linked antibody (7076) and
anti-rabbit IgG, HRP-linked antibody (5127, all from Cell Signaling
Technology, Inc., Danvers, MA, USA), all at 1:5,000. Incubation
with primary antibodies was performed at 4°C overnight. Membranes
were washed with TBST (NaCl 137 mM, KCl 2.7 mM, Tris base 19 mM) 3
times, 15 mins each wash, and then incubated with secondary
antibodies for 30 min in room temperature. The results were
visualized using a Pierce ECL Western Blotting Substrate (Thermo
Fisher Scientific, Inc.).
Immunohistochemical staining
Immunohistochemical staining was performed by an
avidin-biotin-peroxidase technique on the patient samples and mouse
specimens. Paraffin-embedded tumor tissue sections (formalin fixed
and 4–5 µm thick) were prepared and epitope retrieval was performed
for further analysis. The slides were incubated in 10X antigen
retrieval solution (Antigen Retrieval Reagent-Basic; R&D
Systems, Inc., Minneapolis, MN, USA) at 92–95°C for 2–10 min, then
cooled to room temperature, rinsed in ddH2O, and then
washed by PBS. The paraffin sections were subjected to hydrogen
peroxide (3%) for 10–15 min and subsequently blocked by a regular
blocking solution (StartingBlock Blocking Buffer; Thermo Fisher
Scientific, Inc.) for 10–15 min at 37°C. Finally, the sections were
incubated in anti-CD31 (3528) and anti-Ki67 (9449; both Cell
Signaling Technology, Inc., Danvers, MA, USA), primary antibodies
diluted by 1:200, and terminal deoxynucleotidyl transferase dUTP
nick-end labeling (TUNEL) reagent/DAPI at room temperature for 30
min after blocking. All sections were washed 3 times and incubated
with secondary antibodies Anti-mouse IgG (H+L), F(ab')2 Fragment
(Alexa Fluor® 488 Conjugate; 4408; Cell Signaling
Technology, Inc.) for 1 h at 37°C, and then 6 random fields were
observed under a fluorescence inverted phase microscope (1X71;
Olympus Corporation).
In vivo experiments
Specific pathogen-free (SPF) female BALB/c nude
(n=90; 4–6 weeks old) mice were purchased from Harbin Veterinary
Research Institute (Harbin, China). All the mice were kept in a SPF
room, with a 14-h light/10-h dark cycle at a temperature of
18–23°C, humidity of 40–60% and with food and water ad
libitum. The fat content of the diet ranged from 4–11%. MG-63
(5×106) cells were diluted in PBS, then injected
subcutaneously into the dorsum of the mice at a total volume of 100
µl. The treatments for tumor-bearing mice were initiated when tumor
diameters reached 5–8 mm on day 5 after tumor inoculation. Mice
were randomly divided into three groups (n=30 in each experimental
group) and injected intratumorally with 100 mg/kg Bim-1 or AbBim-1,
or the same volume of PBS. The detailed procedures are described in
a previous report (29). The
treatments were performed 7 times at intervals of every 2 days.
Tumor diameters were recorded once every 2 days and tumor volumes
were calculated using the formula, 0.52 × smallest
diameter2 × largest diameter. After 5 weeks, the mice
were sacrificed by cervical vertebra dislocation and dissected.
Dual luciferase reporter assays
The assay was performed using a Dual Luciferase
Assay kit (Promega Corporation, Madison, WI, USA). First the
promoter region of NF-κB was cloned to a luciferase vector, MG-63
cells were seeded onto a 24-well plate, 30,000 cells in each well,
and then the cells were transfected with luciferase and Renilla.
After 24 h, luciferase and Renilla activity was detected by a
Thermo Multiskan plate reader (Thermo Fisher Scientific, Inc.)
according to the manufacturer's protocol. Activity of NF-κB was
defined as the ratio of firefly luciferase activity compared with
corresponding Renilla luciferase activity. Each experiment was
repeated by 3 times.
ELISA
The Bmi-1 kit used for ELISA was purchased from
Biomatik (EKC35069) and the MMP9 kit purchased from Thermo Fisher
Scientific, Inc. (KHC3061). Briefly, 50 µl of standard and sample
was added to the plate, cultured for 2 h at room temperature, and
washed 4 times using the wash buffer from the kit. Then 100 µl of
antibody was added and incubated for 1 h in room temperature,
followed by washing 4 times. HRP conjugate was added and cultured
for 30 min at room temperature and the wells washed 4 times and
treated with chromogenic substrate. Finally, the results were
defined by absorbance at 450 and 55 nm using a Thermo Multiskan
plate reader (Thermo Fisher Scientific, Inc.); each experiment was
replicated 3 times.
Immunofluorescence
The supernatant of culture cells was aspirated and
the cells fixed in 4% formaldehyde diluted in warm PBS, for 15 min
at room temperature, then washed in PBS for 3 times. The blocking
buffer (1X PBS/5% normal serum/0.3% Triton™ X-100, serum purchased
from Gibco; Thermo Fisher Scientific, Inc.) was added and blocked
for 60 min. Next, the diluted primary antibody (Bmi-1; 6964; 1:200;
Cell Signaling Technology, Inc.) was added and incubated overnight
at 4°C. The cells were then incubated in fluorochrome-conjugated
secondary antibody (Anti-rabbit IgG (H+L), F(ab')2 Fragment (Alexa
Fluor® 555 Conjugate) 4413; 1:500; Cell Signaling
Technology, Inc.). The cells were then washed and stained with DAPI
(8961; Cell Signaling Technology, Inc.), and examined under the
microscope (1X71; Olympus Corporation), which was used to capture
images.
Apoptosis detection
Cell apoptosis was detected using an Annexin V
APC/PI double staining kit from Sungene Biotech Company (Tianjing,
China). Briefly, the cells were first digested and then washed by
PBS for 3 times. The cells were diluted using the binding buffer in
the kit, and stained for Annexin V APC antibody for 10 min at room
temperature, followed by staining with PI, then immediately using
FACS (BD FACSCalibur, BD Biosciences, Franklin Lakes, NJ, USA) to
detect the apoptosis, using CellQuest Pro software version 5.1 (BD
Biosciences).
Statistical analysis
All data are represented as the mean + standard
error. Unpaired data were analyzed by Student's t test. Comparisons
of data between multiple groups were analyzed by one-way analysis
of variance and post hoc tests performed using Duncan's new
multiple range test. Statistical analyses were performed using SPSS
19.0 (IBM Corp., Armonk, NY, USA). P<0.05 was considered to
indicate a statistically significant difference.
Results
Bmi-1 expression in osteosarcoma cells
and clinical tissues and the characteristics of AbBmi-1
In order to investigate the effect of Bmi-1 on the
aggressiveness of osteosarcoma cells, the Bmi-1 expression levels
in osteosarcoma cells and clinical tissues were determined. As
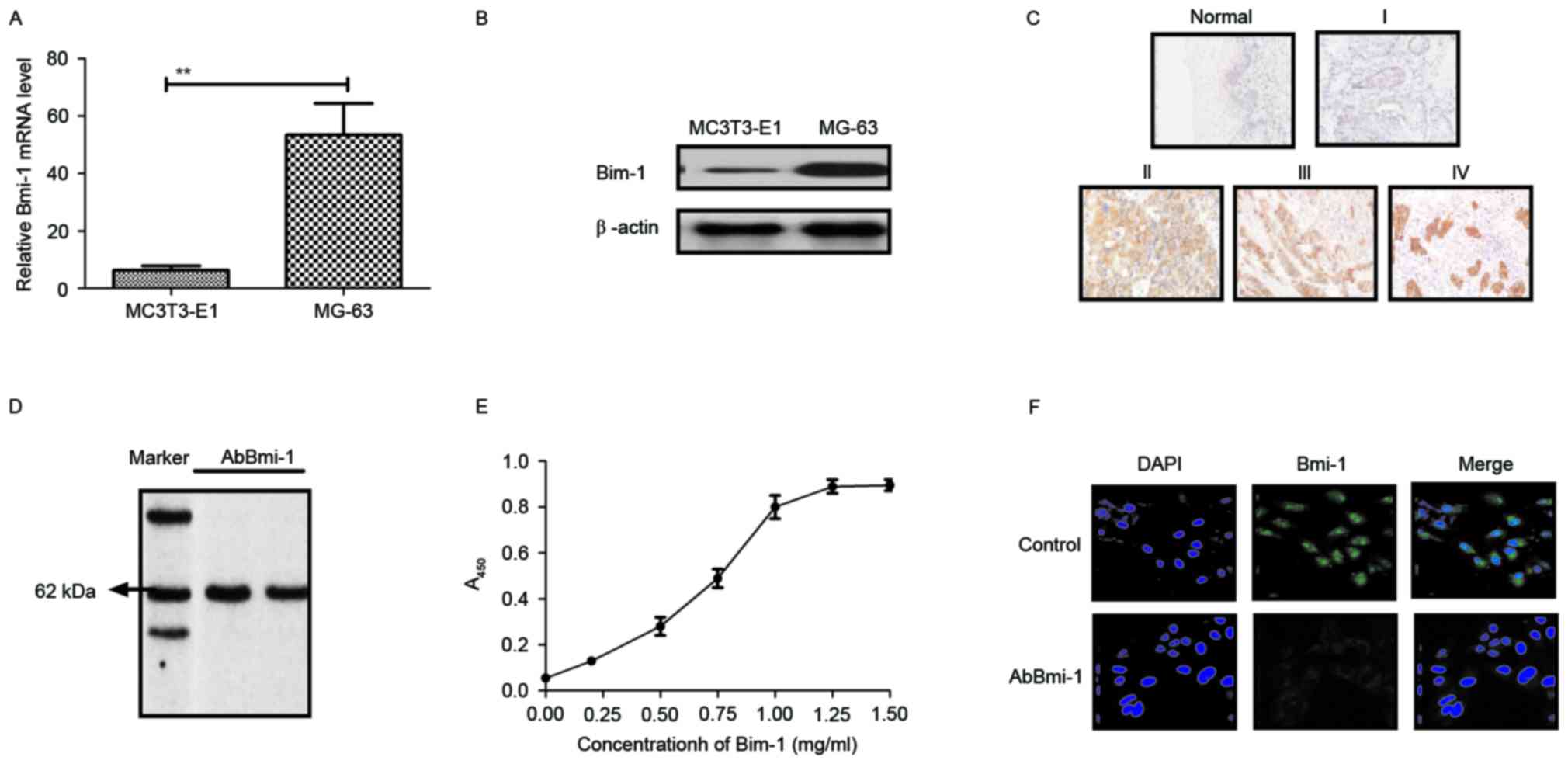
presented in Fig. 1A and B, Bmi-1
expression was higher in MG-63 osteosarcoma cells than in MC3T3-E1
human normal osteoblast cells. The results in Fig. 1C demonstrated that Bmi-1 expression
was also increased in diseased tissues compared with normal
adjacent tissues. Additionally, the affinity of AbBmi-1 for Bmi-1
was determined using ELISA and western blot analysis. AbBmi-1
detected a band ~65 kDa under constant denaturing gel
electrophoresis and specific binding to Bmi-1 was confirmed by
ELISA assay (Fig. 1D and E).
Immunofluorescence also demonstrated that AbBmi-1 efficiently
decreased the Bmi-1 fluorescence signal in MG-63 cells (Fig. 1F). These data suggest that Bmi-1
may be a potential target for the treatment of osteosarcoma and
AbBmi-1 can efficiently bind to Bmi-1.
Efficacy of AbBmi-1 for growth and
invasion of osteosarcoma cells
In order to investigate the role of Bmi-1 in
osteosarcoma cells, bone carcinoma MG-63 cells were treated with
PBS, Bmi-1 and AbBmi-1. The growth, migration and invasion of
osteosarcoma cells were then analyzed in vitro. The data in
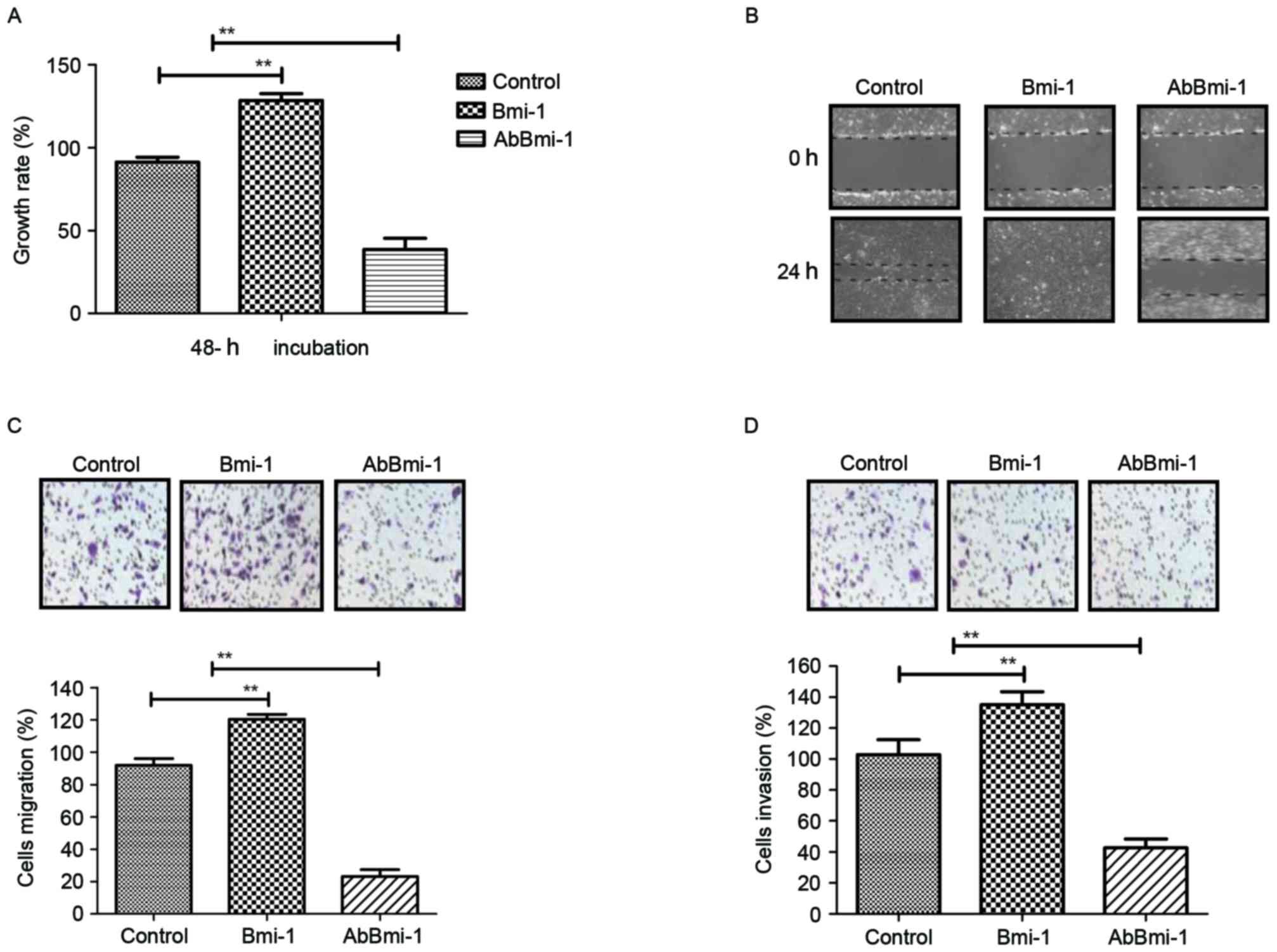
Fig. 2A demonstrated that Bmi-1
treatment significantly increased the growth of osteosarcoma cells,
while AbBmi-1 markedly inhibited osteosarcoma cell growth.
AbBmi-1-treated cells exhibited cellular morphologies typical of a
lower invasive phenotype compared with the control cells and
presented decreased numbers of outward projections as analyzed by a
wound healing assay (Fig. 2B).
Notably, migration and invasion assays revealed that Bmi-1
treatment markedly increased migration of MG63 cells compared with
control cells and AbBmi-1 induced the opposite effect (Fig. 2C and D). Additionally, AbBmi-1
treatment increased apoptosis of MG-63 cells compared with control
cells, while Bmi-1 treatment increased the apoptotic resistance of
MG-63 cells induced by cisplatin (Fig.
2E). Furthermore, the results also that Bmi-1 treatment
increased the number of tumor clones compared with control, whereas
AbBmi-1 reduced clone formation compared with control (Fig. 2F). Taken together, the data
indicate that AbBmi-1 can inhibit the growth and aggressiveness of
osteosarcoma cells and increase apoptosis.
AbBmi-1 inhibits osteosarcoma cells
growth through MMP-9-mediated the NF-κB signaling pathway
To understand the mechanism of how AbBmi-1
suppressed the migration and invasion of osteosarcoma cells, the
expression and activity of MMP-9 in MG-63 cells was determined. The
results in Fig. 3A demonstrated
that MMP-9 expression was reduced in AbBmi-1-treated cells compared
with that in control groups. Additionally, AbBmi-1 treatment
markedly reduced MMP-9 activity in osteosarcoma compared to control
cells (Fig. 3B). Furthermore,
NF-κB target genes, including cyclin D1, B-cell lymphoma-extra
large, tumor necrosis factor-α, vascular endothelial growth
factor-C and MYC were decreased following AbBmi-1 treatment in
MG-63 cells compared with control cells (Fig. 3C). Furthermore, when osteosarcoma
cells were transfected with Bmi-1 overexpression vector, the
increased migratory and invasive ability of Bmi-1-overexpression
osteosarcoma cells were dramatically reversed by treatment with
NF-κB inhibitor (JSH-23) and these effects were accompanied by a
reduction in MMP-9 activity (Fig.
3D-F). Western blotting assays demonstrated that the protein
levels of p65, p84, inhibitor of NF-κB kinase-β (IKK-β) and NF-κB
inhibitor α (IκBα) were obviously reduced by treatment with AbBmi-1
compared with control cells (Fig.
3G). Notably, AbBmi-1 treatment reversed the stimulatory effect
of Bmi-1 on NF-κB phosphorylation in bone carcinoma cells (Fig. 3H). Collectively, these results
indicated that the AbBmi-1 reduced the aggressive phenotype of
osteosarcoma cells, leading to the downregulation of the NF-κB
target gene MMP-9 through reduced activation of the NF-κB signaling
pathway.
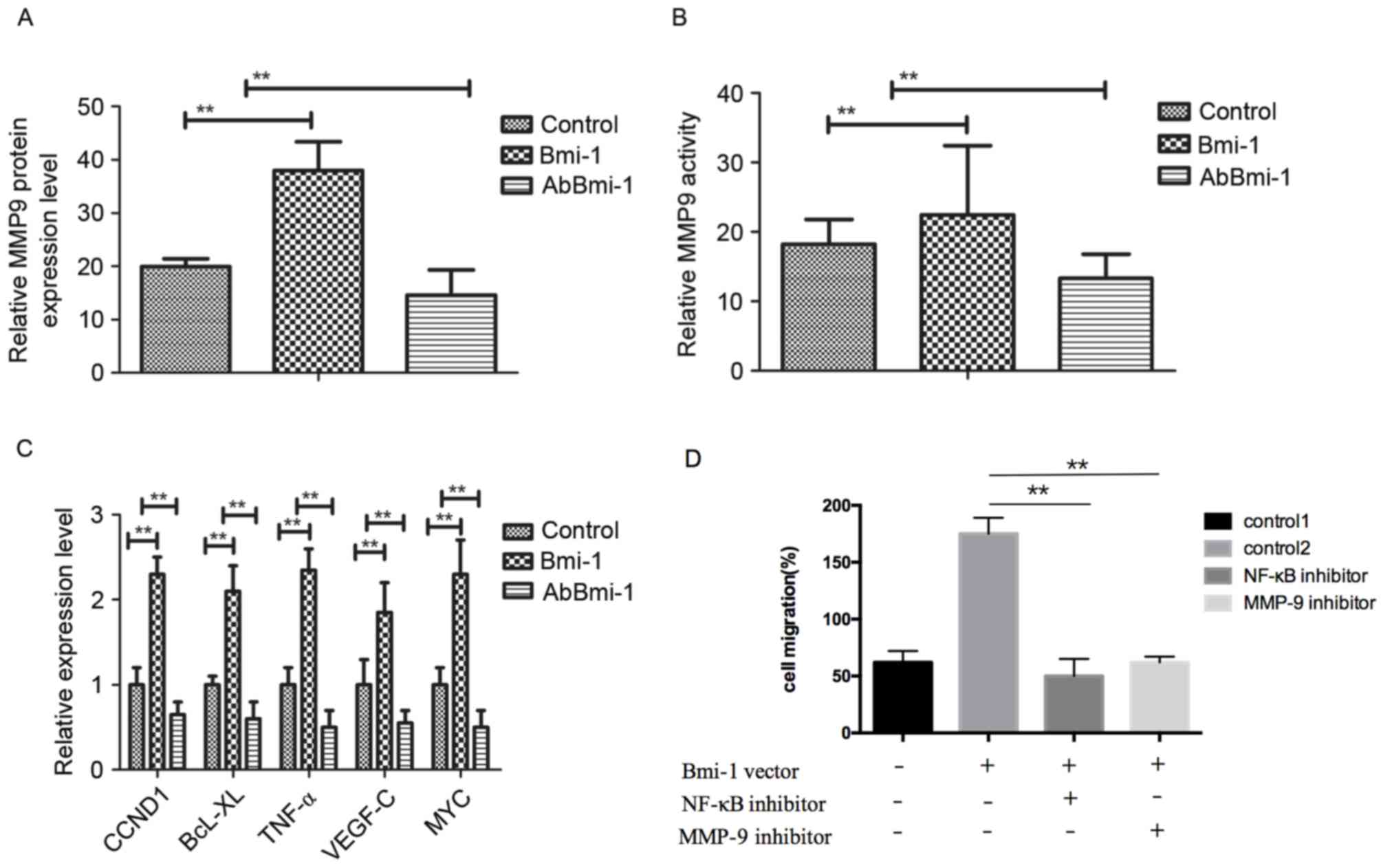 | Figure 3.AbBmi-1 inhibits MG-63 osteosarcoma
cell growth via MMP-9 and NF-κB signaling pathway. (A) MMP-9 mRNA
expression level was suppressed by AbBmi-1. (B) MMP-9 protein
activity was inhibited by AbBmi-1; **P<0.01, Bmi-1 and AbBmi-1
vs. Control group. (C) Analysis of NF-κB target genes expression
after treatment with AbBmi-1. (D) Cells were transfected with Bmi-1
overexpression vector for 48 h, then cell migration, (E) cell
invasion and (F) MMP-9 activity were determined following treatment
with NF-κB inhibitor (JSH-23) and MMP-9 inhibitor; **P<0.01,
NF-κB inhibitor and MMP-9 inhibitor vs. control 2 group. (G) The
protein expression levels of p65, p84, IKK-β and IκBα were
determined following treatment with Bmi-1, JSH-23 and AbBmi-1. (H)
Analysis of NF-κB phosphorylation activation following treatment
with PBS, Bmi-1 and AbBmi-1; **P<0.01, Bmi-1 and AbBmi-1 vs.
Control group. The data are presented as the mean + standard error.
MMP-9, matrix metalloproteinase-9; Bmi-1, B-cell-specific Moloney
murine leukemia virus integration site 1 protein; AbBmi-1,
Bmi-1-targeting antibody; CCND1, cyclin D1; BcL-XL, B-cell
lymphoma-extra large; TNF-α, tumor necrosis factor-α; VEGF-C,
vascular endothelial growth factor C; NF-κB, nuclear factor-κB;
IKK-β, inhibitor of NF-κB kinase-β; IκBα, NF-κB inhibitor α. |
AbBmi-1 targets Bim-1 and shows
benefits for bone cancer therapy in vivo
To further evaluate the effects of AbBmi-1 on bone
carcinoma cell growth, bone carcinoma growth and metastasis were
further analyzed by establishing an osteosarcoma xenograft mice
tumor model. As presented in Fig.
4A, the data revealed that Bmi-1 promoted tumor growth and
AbBmi-1 significantly inhibited tumor growth compared with
PBS-treated mice. The tumor weight was also reduced in the AbBmi-1
group compared with Bmi-1 and PBS groups (Fig. 4B). Representative tumors from each
group are presented in Fig. 4C,
indicating the AbBmi-1 significantly inhibited tumor growth in
vivo. Histological analysis demonstrated that AbBmi-1 treatment
decreased the number of Ki67-positive cells, microvascular density
(CD31 staining) and TUNEL-positive cells (Fig. 4D-F). Notably, the results also
demonstrated that NF-κB luciferase activity was upregulated by
Bmi-1 and downregulated by AbBmi-1 in MG-63 cells, (Fig. 4G). Additionally, it was observed
that MMP-9 and NF-κB protein expression levels were by after
treatment with the AbBmi-1 target-therapy agents (Fig. 4H).
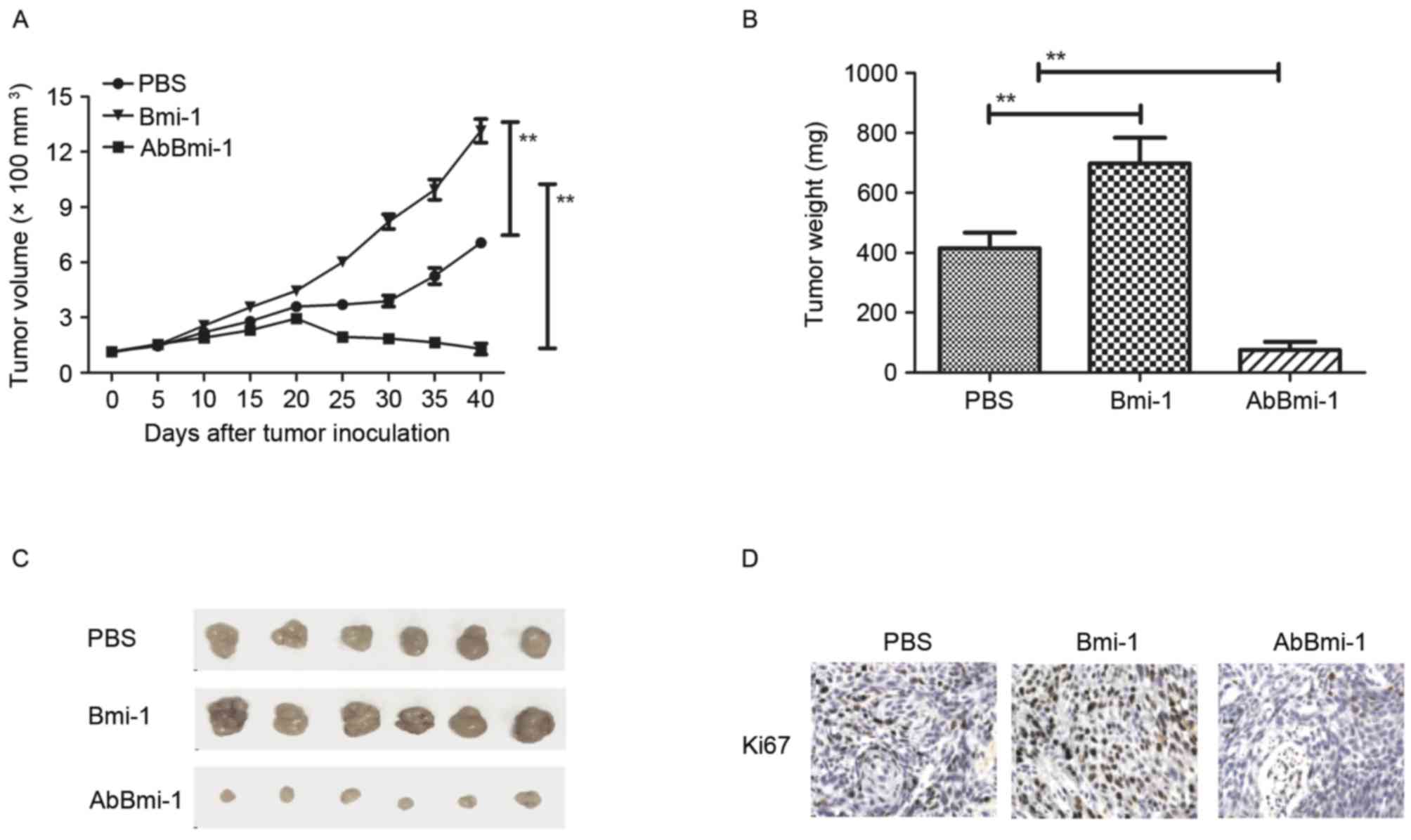 | Figure 4.In vivo action of AbBmi-1 in
osteosarcoma-bearing mice. (A) Mean tumor volumes were measured
following treatment with PBS, Bmi-1 and AbBmi-1; **P<0.01. (B)
Mean tumor weight following treatment with PBS, Bmi-1 and AbBmi-1;
**P<0.01, Bmi-1 and AbBmi-1 vs. PBS group. (C) Representative
images of tumors from experimental mice in each group. Expression
of (D) Ki67 and (E) CD31 in tumors from mice treated with PBS,
Bmi-1 and AbBmi-1. determined by IHC staining. (F) TUNEL-positive
cells in tumors treatment with PBS, Bmi-1 and AbBmi-1. (G) NF-κB
luciferase activity in tumors treated with PBS, Bmi-1 and AbBmi-1;
**P<0.01, Bmi-1 and AbBmi-1 vs. Control group. (H) MMP-9 and
NF-κB expression in tumors following treatment with PBS, Bmi-1 and
AbBmi-1. The data are presented as the mean + standard error.
Bmi-1, B-cell-specific Moloney murine leukemia virus integration
site 1 protein; AbBmi-1, Bmi-1-targeting antibody; TUNEL, terminal
deoxynucleotidyl transferase dUTP nick-end labeling; NF-κB, nuclear
factor-κB; MMP-9, matrix metalloproteinase-9. |
Discussion
The aim of this current study was to examine the
tumorigenic function of Bmi-1 and the therapeutic effects of
AbBmi-1 in bone carcinoma in vitro and in vivo.
Although previous studies have reported that Bmi-1 induced an
aggressive cancer phenotype through modulation of the NF-κB
signaling pathway, the role of the signaling pathway in bone
carcinoma has not been reported and remains unclear (30,31).
Therefore, understanding the role of Bmi-1 may be essential for the
development of osteosarcoma treatments (32). The data of the present study
demonstrated that Bmi-1 increases the migration and invasion of
osteosarcoma cells by activating NF-κB signaling pathway and
subsequent upregulation of MMP-9 expression. Overexpression of
Bmi-1 promoted angiogenesis, tumorigenicity, and increased
apoptosis resistance induced by cisplatin through the NF-κB signal
pathway. Notably, AbBmi-1-treated tumors in xenograft mice were
significantly smaller and had reduced tumor weights compared with
control tumors. These data suggested that Bmi-1 may act as a
potential molecular targets and AbBmi-1 may be a potential
anti-cancer agent acting through inhibition of the NF-κB signaling
pathway for osteosarcoma therapy.
Bmi-1 is a member of the PcG family and is
frequently overexpressed in human tumor cells, suggesting that
Bmi-1 is a potential oncogene involved in the initiation of cancer
tumorigenesis (33). In addition,
Bmi-1 suppress its targets, cyclin dependent kinase inhibitor 2A
(p14ARF) and p16INK4a, and previous reports demonstrated that Bmi-1
promotes tumor cell migration by suppressing the p14 ARF/MDM2/p53
and/or p16/RB transcriptional corepressor 1 signaling pathways
(34). Furthermore, Bmi-1
upregulation was demonstrated to enhance the aggressiveness of
human carcinoma and regulate epithelial-mesenchymal transition
through modulation of the phosphoinositide 3-kinase/Akt/glycogen
synthase kinase-3β pathway (35).
This current study suggested that Bmi-1 promoted an aggressive
phenotype in human osteosarcoma by regulating the NF-κB/MMP-9
signaling pathway, indicating that Bmi-1 may be a potential
therapeutic target for osteosarcoma therapy.
Apoptosis-resistance is a major obstacle in cancer
clinical treatment (33,34). A previous study demonstrated that
overexpression of Bmi-1 in EC9706 esophageal carcinoma cells
promoted cell cycle progression, migration and enhanced the
resistance to apoptosis (35).
Downregulation of Bmi-1 is reported to be associated with
suppressed tumorigenesis and induced apoptosis in CD44+
nasopharyngeal carcinoma cancer stem-like cells (36). Therefore, reducing the
apoptosis-resistance of cancer cells and tumors tissues may enhance
the outcomes of patients undergoing oncotherapy in the clinic.
Bmi-1 is reported as an oncogene that promotes tumor growth,
aggressiveness and tumor angiogenesis; knockdown of Bmi-1
expression exhibits tumor growth, migration, invasion and tumor
angiogenesis in human colorectal cancer cells (15,37).
In addition, Bmi-1 overexpression promotes cancer cell
proliferation and is a predictor of poor survival in patients with
colorectal cancer (38,39). In the current study, Bmi-1
regulated the proliferation and tumorigenicity of bone carcinoma,
whereas neutralizing Bmi-1 induced opposite outcomes through
inactivation of the NF-κB signaling pathway. The results suggest
that therapy targeting Bmi-1 may be an efficient and promising
molecular therapy for the treatment of osteosarcoma in the
clinic.
Notably, different signaling pathways that promote
the aggressiveness of osteosarcoma are involved in the modulation
of MMP-9 transcription (40,41).
Bmi-1 has been recently reported to be critical in the maintenance
of genome integrity, and the p21/cyclin E pathway modulates the
anticlastogenic activity of Bmi-1 in cancer cells (42). A previous study reported that NF-κB
induces the expression and activation of MMP-9 by interacting with
promoter sites and consequently promoting tumor progression
(21). The current study indicated
that Bmi-1 induces MMP-9 expression and activity through a
mechanism associated with NF-κB activation, whereas AbBmi-1 blocked
the activity of NF-κB, downregulated the pro-invasive effect of
Bmi-1 and prevented MMP-9 activity.
In conclusion, this study provided evidence that
Bmi-1 is overexpressed in bone cancer cells and clinical bone
cancer tissues. Reduced NF-activation caused by AbBmi-1 inhibited
growth, aggressiveness and migration, and increased apoptosis in
bone cancer in vitro and in vivo. According to this
molecular analysis, Bmi-1 is a potential target in osteosarcoma and
AbBmi-1 may be useful as a therapeutic agent for the treatment of
human bone cancer.
References
|
1
|
Błogowski W, Bodnarczuk T and Starzyńska
T: Concise review: Pancreatic cancer and bone marrow-derived stem
cells. Stem Cells Transl Med. 5:938–945. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Assi R, Mukherji D, Haydar A, Saroufim M,
Temraz S and Shamseddine A: Metastatic colorectal cancer presenting
with bone marrow metastasis: A case series and review of
literature. J Gastrointest Oncol. 7:284–297. 2016.PubMed/NCBI
|
|
3
|
Pazionis TJ, Alradwan H, Deheshi BM,
Turcotte R, Farrokhyar F and Ghert M: A systematic review and
meta-analysis of En-Bloc vs. intralesional resection for giant cell
tumor of bone of the distal radius. Open Orthop J. 7:103–108. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Maeyama I: Review of bone tumor. Iryo.
24(Suppl): S2271970.(In Japanese).
|
|
5
|
Sanchez-Pareja A, Larousserie F,
Boudabbous S, Beaulieu JY, Mach N, Saiji E and Rougemont AL: Giant
cell tumor of bone with pseudosarcomatous changes leading to
premature denosumab therapy interruption: A case report with review
of the literature. Int J Surg Pathol. 24:366–372. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Mohammadi M, Goudarzi PK, Rahmani O,
Kaghazian P, Yahaghi E, Taheriazam A and Ahmadi K: Evaluation of
gene expression level of CDC5L and MACC1 in poor prognosis and
progression of osteosarcoma. Tumour Biol. 37:8153–8157. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Bahador R, Taheriazam A, Mirghasemi A,
Torkaman A, Shakeri M, Yahaghi E and Goudarzi PK: Tissue expression
levels of miR-29b and miR-422a in children, adolescents, and young
adults' age groups and their association with prediction of poor
prognosis in human osteosarcoma. Tumour Biol. 37:3091–3095. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Zhou Y, Zhao RH, Tseng KF, Li KP, Lu ZG,
Liu Y, Han K, Gan ZH, Lin SC, Hu HY and Min DL: Sirolimus induces
apoptosis and reverses multidrug resistance in human osteosarcoma
cells in vitro via increasing microRNA-34b expression. Acta
Pharmacol Sin. 37:519–529. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Zhao H, Peng C, Ruan G, Zhou J, Li Y and
Hai Y: Adenovirus-delivered PDCD5 counteracts adriamycin resistance
of osteosarcoma cells through enhancing apoptosis and inhibiting
Pgp. Int J Clin Exp Med. 7:5429–5436. 2014.PubMed/NCBI
|
|
10
|
Tsai HC, Huang CY, Su HL and Tang CH: CCN2
enhances resistance to cisplatin-mediating cell apoptosis in human
osteosarcoma. PLoS One. 9:e901592014. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Locklin RM, Federici E, Espina B, Hulley
PA, Russell RG and Edwards CM: Selective targeting of death
receptor 5 circumvents resistance of MG-63 osteosarcoma cells to
TRAIL-induced apoptosis. Mol Cancer Ther. 6:3219–3228. 2007.
View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Vourvouhaki E, Carvalho C and Aguiar P:
Model for Osteosarcoma-9 as a potent factor in cell survival and
resistance to apoptosis. Phys Rev E Stat Nonlin Soft Matter Phys.
76:0119262007. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Kang MK, Kim RH, Kim SJ, Yip FK, Shin KH,
Dimri GP, Christensen R, Han T and Park NH: Elevated Bmi-1
expression is associated with dysplastic cell transformation during
oral carcinogenesis and is required for cancer cell replication and
survival. Br J Cancer. 96:126–133. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Lin MX, Wen ZF, Feng ZY and Li ZK:
Association of Bmi-1 expression with clinicopathological features
and prognosis of colorectal cancer. Nan Fang Yi Ke Da Xue Xue Bao.
29:1816–1819. 2009.(In Chinese). PubMed/NCBI
|
|
15
|
Choi YJ, Choi YL, Cho EY, Shin YK, Sung
KW, Hwang YK, Lee SJ, Kong G, Lee JE, Kim JS, et al: Expression of
Bmi-1 protein in tumor tissues is associated with favorable
prognosis in breast cancer patients. Breast Cancer Res Treat.
113:83–93. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Raaphorst FM, Meijer CJ and Otte AP:
Correspondence re: S. Beá et al., BMI-1 gene amplification and
overexpression in hematological malignancies occur mainly in mantle
cell lymphomas. Cancer Res. 61:2409–2412, 2001. Cancer Res 62:
618–619. 2002.
|
|
17
|
Li J, Gong LY, Song LB, Jiang LL, Liu LP,
Wu J, Yuan J, Cai JC, He M, Wang L, et al: Oncoprotein Bmi-1
renders apoptotic resistance to glioma cells through activation of
the IKK-nuclear factor-kappaB pathway. Am J Pathol. 176:699–709.
2010. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Wu X, Liu X, Sengupta J, Bu Y, Yi F, Wang
C, Shi Y, Zhu Y, Jiao Q and Song F: Silencing of Bmi-1 gene by RNA
interference enhances sensitivity to doxorubicin in breast cancer
cells. Indian J Exp Biol. 49:105–112. 2011.PubMed/NCBI
|
|
19
|
Jiang L, Wu J, Yang Y, Liu L, Song L, Li J
and Li M: Bmi-1 promotes the aggressiveness of glioma via
activating the NF-kappaB/MMP-9 signaling pathway. BMC Cancer.
12:4062012. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Lai J, Cai Q, Biel MA, Wang C, Hu X, Wang
S and Lin J: Id1 and NF-κB promote the generation of CD133+ and
BMI-1+ keratinocytes and the growth of xenograft tumors in mice.
Int J Oncol. 44:1481–1489. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Sun P, Mu Y and Zhang S: A novel
NF-κB/MMP-3 signal pathway involves in the aggressivity of glioma
promoted by Bmi-1. Tumour Biol. 35:12721–12727. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Hassanzadeh P: Colorectal cancer and NF-κB
signaling pathway. Gastroenterol Hepatol Bed Bench. 4:127–132.
2011.PubMed/NCBI
|
|
23
|
Wang Y, Zhou Y, Jia G, Han B, Liu J, Teng
Y, Lv J, Song Z, Li Y, Ji L, et al: Shikonin suppresses tumor
growth and synergizes with gemcitabine in a pancreatic cancer
xenograft model: Involvement of NF-κB signaling pathway. Biochem
Pharmacol. 88:322–333. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Wang Z, Cao CJ, Huang LL, Ke ZF, Luo CJ,
Lin ZW, Wang F, Zhang YQ and Wang LT: EFEMP1 promotes the migration
and invasion of osteosarcoma via MMP-2 with induction by AEG-1 via
NF-κB signaling pathway. Oncotarget. 6:14191–14208. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Zhu LB, Jiang J, Zhu XP, Wang TF, Chen XY,
Luo QF, Shu Y, Liu ZL and Huang SH: Knockdown of Aurora-B inhibits
osteosarcoma cell invasion and migration via modulating
PI3K/Akt/NF-κB signaling pathway. Int J Clin Exp Pathol.
7:3984–3991. 2014.PubMed/NCBI
|
|
26
|
Jamshidi M, Fagerholm R, Khan S, Aittomäki
K, Czene K, Darabi H, Li J, Andrulis IL, Chang-Claude J, Devilee P,
et al: SNP-SNP interaction analysis of NF-κB signaling pathway on
breast cancer survival. Oncotarget. 6:37979–37994. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Wai-Hoe L, Wing-Seng L, Ismail Z and
Lay-Harn G: SDS-PAGE-based quantitative assay for screening of
kidney stone disease. Biol Proced Online. 11:145–160. 2009.
View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Bai FL, Yu YH, Tian H, Ren GP, Wang H,
Zhou B, Han XH, Yu QZ and Li DS: Genetically engineered Newcastle
disease virus expressing interleukin-2 and TNF-related
apoptosis-inducing ligand for cancer therapy. Cancer Biol Ther.
15:1226–1238. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Bansal N, Bartucci M, Yusuff S, Davis S,
Flaherty K, Huselid E, Patrizii M, Jones D, Cao L, Sydorenko N, et
al: BMI-1 targeting interferes with patient-derived
tumor-initiating cell survival and tumor growth in prostate cancer.
Clin Cancer Res. 22:6176–6191. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Chen F, Chen L, He H, Huang W, Zhang R, Li
P, Meng Y and Jiang X: Up-regulation of microRNA-16 in glioblastoma
inhibits the function of endothelial cells and tumor angiogenesis
by targeting Bmi-1. Anticancer Agents Med Chem. 16:609–620. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Song LB, Li J, Liao WT, Feng Y, Yu CP, Hu
LJ, Kong QL, Xu LH, Zhang X, Liu WL, et al: The polycomb group
protein Bmi-1 represses the tumor suppressor PTEN and induces
epithelial-mesenchymal transition in human nasopharyngeal
epithelial cells. J Clin Invest. 119:3626–3636. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Wang MC, Li CL, Cui J, Jiao M, Wu T, Jing
LI and Nan KJ: BMI-1, a promising therapeutic target for human
cancer. Oncol Lett. 10:583–588. 2015.PubMed/NCBI
|
|
34
|
Junan Li, Jye Poi Ming and Ming-Daw Tsai:
The regulatory mechanisms of tumor suppressor P16INK4A
and relevance to cancer. Biochemistry. 50:5566–5582. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Xu W, Yang Z and Lu N: A new role for the
PI3K/Akt signaling pathway in the epithelial-mesenchymal
transition. Cell Adh Migr. 9:317–324. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Chinchar E, Makey KL, Gibson J, Chen F,
Cole SA, Megason GC, Vijayakumar S, Miele L and Gu JW: Sunitinib
significantly suppresses the proliferation, migration, apoptosis
resistance, tumor angiogenesis and growth of triple-negative breast
cancers but increases breast cancer stem cells. Vasc Cell.
6:122014. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Guidicelli G, Chaigne-Delalande B,
Dilhuydy MS, Pinson B, Mahfouf W, Pasquet JM, Mahon FX, Pourquier
P, Moreau JF and Legembre P: The necrotic signal induced by
mycophenolic acid overcomes apoptosis-resistance in tumor cells.
PLoS One. 4:e54932009. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Wang JF, Liu Y, Liu WJ and He SY:
Expression of Bmi-1 gene in esophageal carcinoma cell EC9706 and
its effect on cell cycle, apoptosis and migration. Chin J Cancer.
29:689–696. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Xu X, Liu Y, Su J, Li D, Hu J, Huang Q, Lu
M, Liu X, Ren J, Chen W and Sun L: Downregulation of Bmi-1 is
associated with suppressed tumorigenesis and induced apoptosis in
CD44 (+) nasopharyngeal carcinoma cancer stem-like cells. Oncol
Rep. 35:923–931. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Guo WJ, Zeng MS, Yadav A, Song LB, Guo BH,
Band V and Dimri GP: Mel-18 acts as a tumor suppressor by
repressing Bmi-1 expression and down-regulating Akt activity in
breast cancer cells. Cancer Res. 67:5083–5089. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Taran K, Wysocka A, Sitkiewicz A, Kobos J
and Andrzejewska E: Evaluation of potential prognostic value of
Bmi-1 gene product and selected markers of proliferation (Ki-67)
and apoptosis (p53) in the neuroblastoma group of tumors. Postepy
Hig Med Dosw (Online). 70:110–116. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Jacobs JJ, Scheijen B, Voncken JW, Kieboom
K, Berns A and van Lohuizen M: Bmi-1 collaborates with c-Myc in
tumorigenesis by inhibiting c-Myc-induced apoptosis via INK4a/ARF.
Genes Dev. 13:2678–2690. 1999. View Article : Google Scholar : PubMed/NCBI
|